INTRODUCTION
MicroRNAs (miRNAs) are small non-coding RNAs in 18–25 nucleotides binding to the 3′-untranslated region of massager RNA, implying a crucial important role in regulating gene expression [1, 2]. So far, there are 2588 validated human mature miRNAs according to miRBase release 21 at the website ‘http://www.mirbase.org/’. MiRNAs have been shown to play essential roles in various biological processes, and implicated as potential diagnostic and/or prognostic biomarkers in many human diseases including cancers [3]. Interestingly, a large amount of miRNA can be detected in human body fluids, including plasma, serum, urine, cerebrospinal fluid (CSF) and saliva, and they are termed as extracellular miRNAs [4]. Accumulating evidence suggests that extracellular miRNAs are exported from cells. Recently, the interest in exploring extracellular miRNAs as potential biomarkers for early cancer diagnosis and prognosis prediction is proliferating. The detection of cancer-specific extracellular miRNAs has been convincingly reported in many cancer types, including lung cancer, breast cancer, brain cancer and digestive cancers [5–7].
Recently, miR-21, as one of the most studied miRNAs, is observed to be significantly over-expressed in a broad range of cancers [8, 9] and it can serve as a potential diagnostic biomarker for cancer patients [10–14]. Besides in tissues, recent evidence indicates the presence of miR-21 in various types of extracellular fluid, such as plasma [10, 11, 15–33], serum [17, 29, 34–50], CSF [51–56], saliva [32, 57, 58], gastric juice [59], pancreatic juice [60], sputum [61], and pancreatic cyst fluid [62]. Although the diagnostic efficiency of extracellular miR-21 in cancers has been proposed by many researchers, the results are conflicting and inconclusive. To fill this gap in knowledge, we therefore designed a comprehensive meta-analysis to evaluate the diagnostic efficiency of extracellular miR-21 in various cancer types and different sample types.
In this two-phase study, we systematically investigated the expression of extracellular miR-21. In the first phase, via a comprehensive meta-analysis, we discussed the diagnostic efficiency of extracellular miR-21 in various cancer types and different sample types. Additionally, by using a validation study comprised of matched tissue and CSF samples, we evaluated the clinical significance of miR-21 as a potential diagnostic and prognostic biomarker for glioma patients. In the second phase, we examined whether cultured glioma cells can secrete miR-21 into the culture medium, aiming to establish the secretory mechanisms of extracellular miR-21 in glioma cells.
RESULTS
Literature search and a meta-analysis based on 81 studies
In the first phase, we conducted a meta-analysis according to the guidelines set forth by the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement. A flow diagram schematizing the process of article exclusion with specific reasons is presented in Figure S1. In brief, 1352 potentially relevant articles were obtained after initial search, and 1163 of them were excluded after applying further inclusion/exclusion criteria. Finally, 81 studies from 59 articles were included in this meta-analysis [10, 11, 13, 15–70]. All qualified studies recruiting 4428 corresponding cancer patients and 3066 controls were published between the year 2009 and 2015. The basic characteristics of all qualified studies are summarized in Table S1. The mostly investigated cancers included brain cancer (n = 14), lung cancer (n = 11), colorectal cancer (n = 11), pancreatic cancer (n = 9), breast cancer (n = 8), gastric cancer (n = 7), esophageal cancer (n = 6) and hepatocellular carcinoma (n = 4). Sample sources are consisted of plasma (n = 34), serum (n = 25), CSF (n = 12), and digestive juice (n = 5). Out of 81 studies, 55 were conducted in Asian populations, 20 in Caucasian populations, 2 in African populations, 1 in Caucasian & African populations and 1 in Latinos population. The meta-analysis on diagnostic accuracy of extracellular miR-21 are shown in Figure 1. After excluding outliers, overall sensitivity, specificity and area under the summary receiver operating characteristic (SROC) curve (AUC) of extracellular miR-21 for diagnosing cancers were 0.77 (0.73–0.80), 0.81 (0.79–0.84) and 0.86 (0.83–0.89) followed by their corresponding 95% confidence intervals (95%CI), respectively (Table 1).
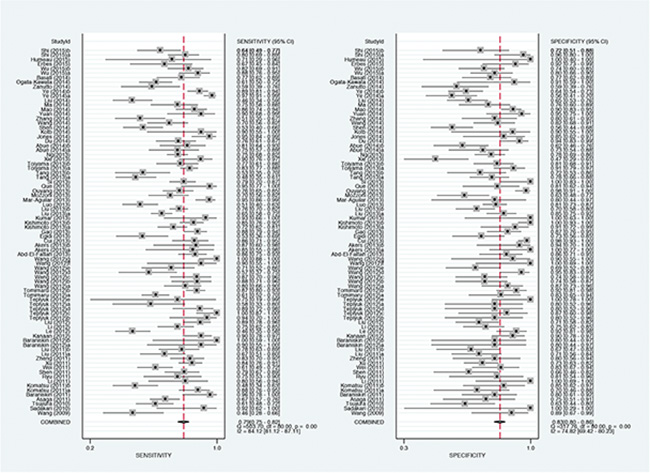
Figure 1: Forest plots of sensitivities and specificities for extracellular miR-21 test accuracy in cancer.
Table 1: Summary estimates of diagnostic criteria and their 95% confidence intervals (95%CI) for extracellular miR-21 in cancer detection
Analysis |
No. of studies |
Sensitivity (95% CI) |
Specitivity(95% CI) |
PLR (95% CI) |
NLR (95% CI) |
DOR (95% CI) |
AUC (95% CI) |
|---|---|---|---|---|---|---|---|
Ethnicity |
|
|
|
|
|
|
|
Asian |
55 |
0.78 (0.74–0.81) |
0.83 (0.79–0.86) |
4.66 (3.76–5.78) |
0.26 (0.22–0.31) |
18 (13–25) |
0.88 (0.84–0.90) |
Caucasian |
20 |
0.80 (0.71–0.87) |
0.81 (0.75–0.86) |
4.29 (3.04–6.07) |
0.25 (0.16–0.38) |
17 (8–35) |
0.87 (0.84–0.90) |
Cancer sites |
|
|
|
|
|
|
|
Brain cancer |
14 |
0.89 (0.80–0.94) |
0.89 (0.83–0.93) |
8.17 (4.95–13.47) |
0.13 (0.07–0.24) |
65 (25–166) |
0.94 (0.92–0.96) |
Breast cancer |
8 |
0.82 (0.74–0.88) |
0.86 (0.78–0.91) |
5.84 (3.58–9.51) |
0.21 (0.14–0.32) |
27 (12–61) |
0.91 (0.88–0.93) |
Lung cancer |
11 |
0.68 (0.58–0.77) |
0.77 (0.69–0.84) |
2.98 (2.11–4.22) |
0.41 (0.30–0.56) |
7 (4–13) |
0.80 (0.76–0.83) |
Esophageal cancer |
6 |
0.86 (0.70–0.94) |
0.65 (0.55–0.74) |
2.48 (2.02–3.04) |
0.22 (0.11–0.44) |
11 (6–22) |
0.77 (0.73–0.80) |
Gastric cancer |
7 |
0.77 (0.68–0.84) |
0.85 (0.72–0.92) |
5.00 (2.53–9.87) |
0.27 (0.18–0.40) |
19 (7–49) |
0.86 (0.83–0.89) |
Hepatocellular carcinoma |
4 |
0.82 (0.70–0.90) |
0.80 (0.70–0.88) |
4.16 (2.63–6.59) |
2.22 (0.13–0.39) |
19 (8–42) |
0.88 (0.85–0.90) |
Pancreatic cancer |
9 |
0.76 (0.66–0.83) |
0.74 (0.67–0.80) |
2.89 (2.28–3.68) |
0.33 (0.23–0.47) |
9 (5–14) |
0.79 (0.75–0.82) |
Colorectal cancer |
11 |
0.72 (0.63–0.79) |
0.83 (0.78–0.87) |
4.22 (3.90–20.78) |
0.34 (0.25–0.47) |
12 (7–23) |
0.85 (0.82–0.88) |
Sample sources |
|
|
|
|
|
|
|
Cerebrospinal fluid |
12 |
0.89 (0.81–0.94) |
0.88 (0.82–0.92) |
7.61 (5.07–11.41) |
0.12 (0.07–0.22) |
61 (30–126) |
0.93 (0.90–0.95) |
Digestive juice |
5 |
0.80 (0.65–0.90) |
0.84 (0.51–0.96) |
4.97 (1.23–20.00) |
0.23 (0.11–0.51) |
21 (3–160) |
0.88 (0.84–0.90) |
Serum |
25 |
0.79 (0.73–0.83) |
0.83 (0.79–0.86) |
4.55 (3.75–5.52) |
0.26 (0.20–0.33) |
18 (12–25) |
0.88 (0.84–0.90) |
Plasma |
34 |
0.75 (0.69–0.80) |
0.81 (0.76–0.85) |
3.91 (2.99–5.12) |
0.31 (0.24–0.39) |
13 (8–20) |
0.85 (0.82–0.88) |
Overall |
81 |
0.79 (0.75–0.82) |
0.83 (0.80–0.86) |
4.59 (3.83–5.49) |
0.26 (0.22–0.30) |
18 (13–24) |
0.88 (0.85–0.90) |
Outliers excluded |
74 |
0.77 (0.73–0.80) |
0.81 (0.79–0.84) |
4.12 (3.53–4.80) |
0.28 (0.24–0.33) |
15 (11–19) |
0.86 (0.83–0.89) |
CI: confidence interval, PLR: positive likelihood ratio, NLR: negative likelihood ratio, DOR: diagnostic odds ratio, AUC: area under the curve.
Finally, Goodness of fit and bivariate normality analysis revealed that the bivariate random-effects model was robust for this meta-analysis (Figure S2A and S2B). Furthermore, we conducted an outlier detection to account for potential sources of heterogeneity. There were seven studies, 23, 35, 39, 43, 50, 57 and 71, as sources of heterogeneity for this meta-analysis. After excluding the deviated studies, there was no significant difference relative to the analysis with deviated studies (Figure S2C and S2D). We used Deek’s funnel plot to evaluate publication bias of included studies. The shape of the funnel plot revealed between-study heterogeneity (P = 0.08, Figure S3).
Subgroup analysis: Extracellular miR-21 as a potential biomarker in glioma
To account for the potential sources of between-study heterogeneity, subgroup analyses were further conducted based on ethnicity, cancer sites, and sample sources, respectively (Table 1). We found that ethnicity exerted on impact on the AUC of extracellular miR-21 (Figure S4). In contrast, the diagnostic accuracies of extracellular miR-21 varied in detecting different cancer types (Figure 2 and Table 1). Our results revealed that extracellular miR-21 had a relatively high diagnostic accuracy in detecting brain cancer, especially in detecting glioma, with a pooled AUC of 0.95 (95% CI: 0.92–0.96) (Table 2 and Figure S5). Additionally, we also found that diagnostic efficiency of extracellular miR-21for cancer differed across different sample types (Table 2 and Figure 3). Compared with other three sample types, CSF-based miR-21 detection had the highest diagnostic efficiency (sensitivity: 0.88; specificity: 0.89 and AUC = 0.94), suggesting a potential clinical role of CSF-based miR-21 in detecting patients with glioma (Figure 3). P values of the Deek’s funnel plot for glioma and CSF subgroups were 0.41 and 0.47, respectively, indicating less likelihood of publication bias (Figure S6 and S7).
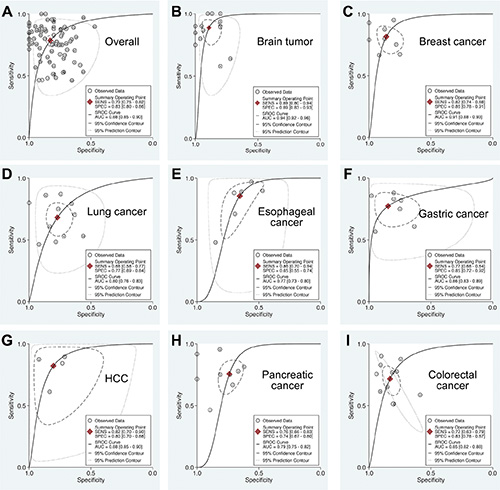
Figure 2: Summary ROC curve of extracellular miR-21 diagnostic values in different cancer types. (A) Overall; (B) Brain tumor; (C) Breast cancer; (D) Lung cancer; (E) Esophageal cancer; (F) Gastric cancer; (G) Hepatocellular carcinoma; (H) Pancreatic cancer; (I) Colorectal cancer.
Table 2: Summary estimates of diagnostic criteria and their 95% confidence intervals (95%CI) for extracellular miR-21 in detection of different types of brain cancer
Variable |
Subtypes of brain cancer |
||
|---|---|---|---|
Overall |
Glioma |
Other brain cancers |
|
No. of studies |
14 |
9 |
5 |
AUC (95% CI) |
0.94 (0.92–0.96) |
0.95 (0.92–0.96) |
0.92 (0.89–0.94) |
Sensitivity (95% CI) |
0.89 (0.80–0.94) |
0.84 (0.73–0.91) |
0.94 (0.85–0.98) |
Specitivity (95% CI) |
0.89 (0.83–0.93) |
0.92 (0.83–0.96) |
0.86 (0.75–0.92) |
PLR (95% CI) |
8.17 (4.95–13.47) |
10.4 (4.4–24.4) |
6.6 (3.7–11.8) |
NLR (95% CI) |
0.13 (0.07–0.24) |
0.17 (0.09–0.31) |
0.07 (0.03–0.19) |
DOR (95% CI) |
65 (25–166) |
61 (16–230) |
94 (27–320) |
CI: confidence interval, PLR: positive likelihood ratio, NLR: negative likelihood ratio, DOR: diagnostic odds ratio, AUC: area under the curve.
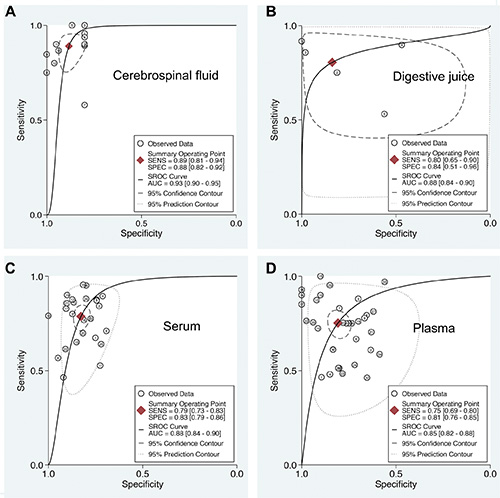
Figure 3: Summary ROC curve of extracellular miR-21 diagnostic values in different sample types. (A) Cerebrospinal fluid; (B) Digestive juice; (C) Serum; (D) Plasma.
Clinical evaluation of CSF-based extracellular miR-21 level in glioma
To further evaluate the clinical potentials of miR-21 detection in glioma, we conducted a validation study by comprehensively collecting brain tissues, and paired CSF samples from 35 glioma patients and 10 non-cancer patients. The clinicopathological characteristics of 35 glioma patients are shown in Table S3. We firstly screened the expression of 15 cancer-related miRNAs in 35 glioma cancer patients (miR-125, miR-126, miR-141, miR-155, miR-17–3p, miR-182, miR-184, miR-195, miR-200, miR-21, miR-223, miR-25, miR-503, miR-92 and miR-98), which were previously reported to exist in human body fluid samples [71]. High miR-21 expression (change in expression of at least 1.5-fold when comparing the means of non-cancer tissues) was detected in 23 out of 35 (65.7%) glioma tissues (Figure 4A). Consistent with the trend in tissue samples, CSF levels of miR-21 in glioma were also significantly higher than that of non-cancer group (P = 0.004, Figure 4B). Moreover, we also found a strong correlation between expression levels of miR-21 in CSF samples and cancer tissues (r = 0.506, P = 0.002), indicating a close relationship between CSF and tissues expressing miR-21 (Figure 4C). Considering the high CSF-based miR-21 levels in glioma patients, we next evaluated the diagnostic accuracy of CSF-based miR-21 in glioma diagnosis. Our results showed that CSF-based miR-21 level had a high diagnostic potential in glioma diagnosis (AUC = 0.81; 95% CI: 0.68–0.93) (Figure 4D), consistent with the meta-analytical results in this study. Moreover, we found CSF-based miR-21 level also exhibited a better prognostic accuracy for glioma (Log Rank test P = 0.004) (Figure 4E), compared with tissue-based miR-21 level, which was previously shown to be a candidate prognostic biomarker for glioma (Figure S8, data from SurvMicro website [72]). Taken together, our data provided robust evidence for clinical implication of CSF-based miR-21 level for the diagnosis and prognosis in glioma.
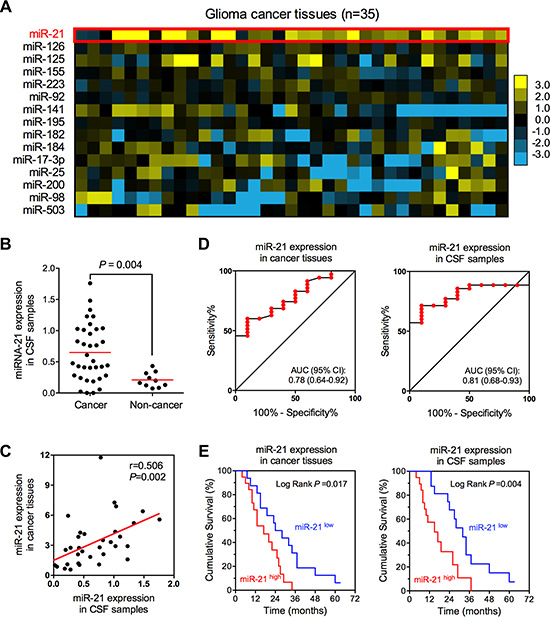
Figure 4: The expression of miR-21 in glioma tissue and CSF samples. (A) Expression profile of 15 cancer-related miRNAs in glioma tissues. (B) CSF-based miR-21 expression in glioma patients and healthy volunteers. (C) Expression correlation between tissue- and CSF-based miR-21 in patients with glioma. (D) Diagnostic efficiencies of tissue- and CSF-basedmiR-21 in glioma. (E) Prognostic efficiencies of tissue- and CSF-based miR-21 in glioma.
MiR-21 in extracellular environment is secreted by glioma cells
Given the strong expression correlation between intracellular (cancer tissues) and extracellular (CSF samples) miR-21 in glioma, we speculated that extracellular miR-21 might be secreted by glioma cells which had high intracellular miR-21 expression. To test this hypothesis, we conducted a high pri-miR-21 expressing glioma cells and collected their conditioned medium for detection of extracellular miR-21 levels (Figure 5A). As shown in Figure 5B, we found that both intracellular and extracellular miR-21 were increased in pri-miR-21 vector transfected glioma cells. Moreover, we also found that the extracellular miR-21 levels in culture medium followed a time-dependent manner (Figure 5C), further suggesting the secretory process of miR-21 in glioma cells.
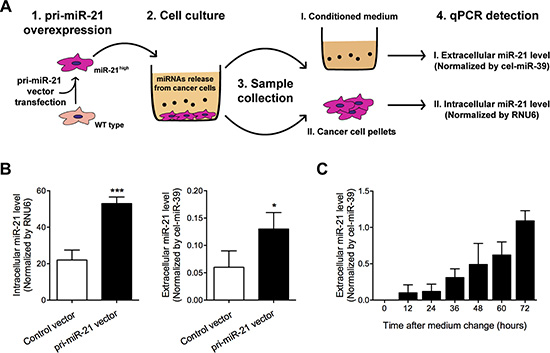
Figure 5: Extracellular and intracellular miR-21 levels were detected in pri-miR-21 overexpressed glioma cells after culture. (A) Study design; (B) Extracellular and intracellular miR-21 levels were detected using qRT-PCR; (C) Extracellular miR-21 levels were detected in medium after cell culture for 0–72 hours. *P < 0.05 and ***P < 0.001.
TGF-β signaling induced miR-21 expression in glioma cells
Previous studies suggested that TGF-β pathway involved in intracellular regulation of miR-21 [73, 74]. However, the corresponding molecular mechanisms remains unclear. In this study, we found that addition of extracellular TGF-β1, the most powerful activator of TGF-β signaling, can significantly induce intracellular miR-21 expression (Figure 6A). Additionally, the fold change of intracellular miR-21 was shown to be much higher (approximately three-fold increase) in U251 cells receiving TGF-β1, when compared with those cells under galunisertib treatment (a selective inhibitor of TGF-β signaling [75]), suggesting an association between intracellular miR-21 and TGF-β signaling (Figure 6A).
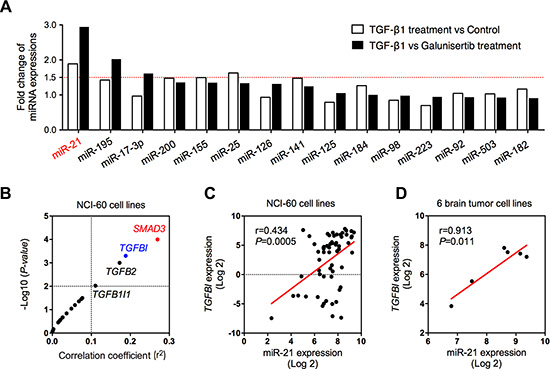
Figure 6: Association between miR-21 and TGF-β signaling pathway. (A) The expression changes of 15 cancer-related miRNAs after TGF-β1 treatment, comparing with control or TβR inhibitor (galunisertib) treatment group; (B) Expression correlation between miR-21 and genes in TGF-β signaling pathway in 60 cancer cell lines; (C) Expression correlation between miR-21 and TGFBI in 60 cancer cell lines; (D) Expression correlation between miR-21 and TGFBI in 6 brain tumor cell lines.
To further explore the factors mediating TGF-β-miR-21 pathway, we evaluated the expression correlation between 19 genes in TGF-β signaling pathway and miR-21 expression using NCI-60 expression profiling data (GSE5846 and GSE26375). TGFBI, a secreted protein induced by TGF-β, has been considered to be tightly associated with extracellular levels of TGF-β [76]. We therefore selected TGFBI as a mediator to see the relationship between extracellular TGF-β levels and intracellular miR-21 expression. As demonstrated in Figure 6B, TGFBI was one of the four top genes in close correlation with miR-29b expression. The expression association between TGFBI and miR-21 was shown to be significant with a P value of lower than 0.05 either in the NCI-60 cells (Figure 6C) or in 6 brain cancer cell lines (including SF-268, SF-295, SF-539, SNB-19, SNB-75 and U251 cell lines, Figure 6D). Above data once again confirmed the inductive activity of TGF-β in intracellular miR-21 expression.
Secretion of miR-21 in glioma cells depends on Smad3 activity
As demonstrated in Figure 6B, we found that SMAD3 was significantly associated with intracellular miR-21 expression. Previous studies suggested that Smad3 was identified as key mediator for transducing TGF-β signals from the cell membrane to the nucleus, and was highly expressed in the brain. Therefore, to Figure out whether activated Smad3 was involved in production and secretion of miR-21, we detected the intracellular and extracellular levels of miR-21 in different activation status of Smad3. Our data showed that TGF-β1 induced phosphorylation of Smad3 and increased intracellular and extracellular levels of miR-21. In contrast, galunisertib significantly inhibited the phosphorylation of Smad3 induced by TGF-β1 and depressed miR-21 levels (Figure 7A, 7D and 7G). When blocking Smad3 activity using small interference RNA (siRNA) of Smad3, the intracellular and extracellular levels of miR-21 were significantly decreased, compared with TGF-β1 treated cells (Figure 7B, 7E and 7H). Furthermore, we also investigated the effect of Smad3 over-expression on miR-21 levels. We found that transfection of Smad3 expressing vector increased the intracellular and extracellular levels of miR-21, which was not inhibited by galunisertib treatment (Figure 7C, 7F and 7I). Above results suggested that Smad3 activity was essential for production and release of miR-21 induced by TGF-β1 in glioma cells.
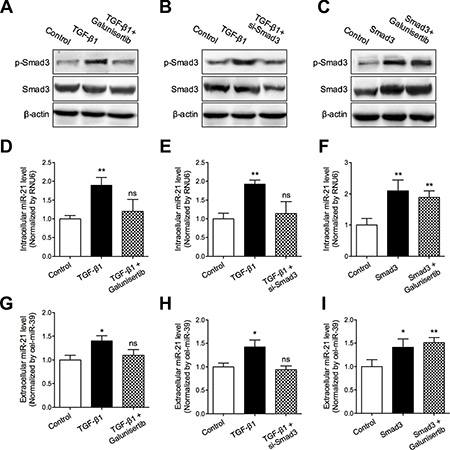
Figure 7: TGF-β/Smad3 signaling pathway mediated miR-21 secretion. (A–C) Smad3 phosphorylation status in the corresponding subgroups were detected using Western blotting assay; (D–F) Intracellular miR-21 levels in the corresponding subgroups were detected using qRT-PCR assay; (G–I) Extracellular miR-21 level in the corresponding subgroups were detected using qRT-PCR assay. *P < 0.05 and **P < 0.01.
DISCUSSION
In this two-phase study, we systematically investigated the expression of extracellular miR-21. In the first phase, by conducting a comprehensive meta-analysis, we discussed the diagnostic efficiency of extracellular miR-21 in various cancer types and different sample types. Additionally, by using a validation study comprised of matched tissues and CSF samples, we evaluated the clinical significance of miR-21 as a potential diagnostic and prognostic biomarker for glioma patients. In the second phase, we demonstrated that cultured glioma cells secreted miR-21 into extracellular medium, and TGF-β/Smad3 signaling pathway was involved in the secretory regulation of miR-21 in glioma cells.
Recent studies in growing numbers have shown that extracellular miRNAs might be a new potential biomarker for early diagnose of cancers [5, 6]. MiR-21, one of the most studied miRNAs, has been widely reported for its over-expression in various cancer types, and has also been found in extracellular environment [10, 12–14, 77]. In the present study, we performed a meta-analysis of 7494 participants derived from 81 published studies worldwide. Compared with other published meta-analyses involving diagnostic accuracy of extracellular miR-21 [78–82], our analysis included totally 81 studies that covered multiple cancer types and sample types, and therefore provided more comprehensive and innovative results. To the best of our knowledge, this is the largest meta-analysis exploring evaluate the diagnostic efficiency of extracellular miR-21 in cancer diagnosis. Our meta-analysis demonstrated that the pooled sensitivity, specificity and AUC of extracellular miR-21 in cancer diagnosis was 0.79, 0.83 and 0.88, respectively, indicating a high accuracy in diagnosing cancers. Interestingly, we further found that miR-21 showed an outstanding efficiency in diagnosis of brain cancer, with a sensitivity of 0.89, specificity of 0.89, and AUC of 0.94. In glioma, the pooled AUC of extracellular miR-21 was increased to 0.95. In addition, we found that sample sources also exerted an impact on the diagnostic accuracy of extracellular miRNA-21. Our results showed that CSF-based miR-21 had a much higher efficiency than miR-21 derived from other sample types, with sensitivity of 0.88, specificity of 0.89, and AUC of 0.94. Malignant glioma has a high mortality with no curative therapies available. Thus, it is urgent to find novel biomarkers for the early diagnosis of glioma. In contrast to blood samples, the miRNA profiles of CSF are mainly affected by CNS diseases and are less affected by blood miRNA concentrations due to blood-brain barrier. Considering above clear strengths, CSF sample is more predictable for real-time monitoring cancer burden and therapeutic response in patients with glioma. Therefore, to evaluate the diagnostic accuracy of CSF-based miR-21 in glioma, we further conducted a validation cohort recruiting 35 glioma patients and 10 non-cancer patients. Our validation results confirmed the findings that CSF-based miR-21 had a high diagnostic accuracy in glioma diagnosis, with an AUC of 0.81 [52]. Furthermore, in our validation cohort, we also found that CSF-based miR-21 exhibited a higher accuracy than tissue-based miR-21 for prognostic prediction of glioma, consistent with a previous report [56]. Taken together, our meta-analysis and clinical validation strongly suggested the potential application of CSF-based miR-21 in glioma, and large-scale multicenter studies are warranted to confirm or refuse our findings.
Despite the strong association between CSF-based miR-21 and glioma, it has not been investigated whether or not the miR-21 existed in CSF was secreted from glioma cells. Previous studies have clearly demonstrated high intercellular miR-21 expression in glioma cells in vitro and in vivo [83–87]. Recently, Baraniskin A et al. firstly observed an increased miR-21 level in CSF samples from patients with glioma [52]. Shi et al. further identified that miR-21 existed in CSF derived exosomes, suggesting that CSF-based miR-21 might be secreted by glioma cells. In this study, by conducting a high miR-21 expression glioma cell, we investigated the secretion of miR-21 in glioma cells. Our data showed that the relative extracellular miR-21 level elevated in a time-dependent manner, which provided evidence for the secretory process of miR-21 from glioma cells into extracellular environment.
TGF-β signaling pathway has gained a special interest in recent years because of its essential roles in regulating cell growth, proliferation, differentiation, and apoptosis. Previous studies have suggested an association between TGF-β signaling pathway and miR-21 [73]. Recently, it is reported that TGF-β1 is capable of inducing numerous miRNAs, including miR-21, in renal fibrosis and lung cancer [88, 89]. Wang et al. [90] also observed the expression correlation between TGF-β and intracellular miR-21 in ursolic acid-treated U251 cells. However, it was still uncertain whether or not TGF-β signaling pathway was involved in miR-21 secretion in glioma. In this study, we clearly demonstrated that both intracellular and extracellular expressions of miR-21 were induced by TGF-β. Meanwhile, inhibition of TGF-β signaling pathway using specific TGF-β receptor inhibitor can significantly suppress both intracellular and extracellular miR-21 expression. To explore the detailed secretory mechanisms of miR-21 in glioma, we further screened the expression correlation between miR-21 and 19 members of TGF-β signaling pathway. Our data revealed that Smad3, one of the most important mediators of TGF-β signaling pathway, was strongly associated with miR-21 expression. Furthermore, we also found that over-expressed Smad3 induced secretion of miR-21 from glioma cells into extracellular environment. Meanwhile, inhibition of Smad3 attenuated the TGF-β induced miR-21 secretion. Above findings provided evidence for the essential roles of TGF-β/Smad3 signaling pathway in mediating miR-21 secretion in glioma (Figure 8).
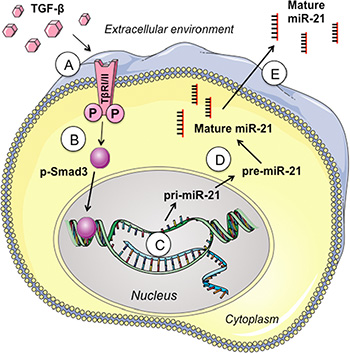
Figure 8: Overview of TGF-β/Smad3 signaling pathways in regulating miR-21 secretion. (A) Exogenous TGF-β binds to TGF-β receptors (TβRI/II); (B) Activated TGF-β receptors phosphorylate and thus activate the intracellular signaling mediator Smad3; (C and D) Phosphorylated Smad3 promotes the processing and maturation of pri-miR-21; (E) Mature miR-21 is secreted into extracellular environment.
Despite the clear strengths including a meta-analysis based on a large number of studies and independent validation using clinical and experimental data, several possible limitations in our study should be noted. First, we only conducted a meta-analysis to investigate extracellular miR-21 in glioma diagnosis, but did not meta-analyze its prognostic potentials in survivorship of glioma patients, mainly due to the lack of literature on this topic. Second, because of the small sample size recruited in our study, tissue and CSF-based miR-21 expression was compared between glioma patients and healthy volunteers, and we failed to take different tumor origin, tumor stages and histological classification into consideration. Third, in the experimental validation, we only provided evidence that TGF-β/Smad3 signaling pathway is essential in regulating miR-21 secretion, but the detailed mechanisms involving TGF-β/Smad3 mediated miR-21 production and secretion is still unknown and warrants further investigation.
In conclusion, by conducting meta-analysis, clinical and experimental investigations, we demonstrated that extracellular miR-21 level, especially CSF-based miR-21 level, could serve as a potential biomarker for glioma patients. Our results further suggest that CSF-based miR-21 might be secreted by glioma cells and TGF-β/Smad3 signaling pathway was responsible for regulating miR-21 secretion.
MATERIALS AND METHODS
Search strategy
A literature search for studies that explored the diagnostic value of circulating miR-21 for cancers was conducted among several computerized databases, including PubMed, Embase, Google Scholar, Cochrane Library, Scopus, Chinese Biomedical Literature Database (CBM) and Chinese National Knowledge Infrastructure (CNKI). The search terms we used were as follows: [“microRNA-21” or “miRNA-21” or “miR-21”] AND [“cancer” or “carcinoma” or “neoplasms” or “tumor”] AND [“sensitivity” or “specificity” or “ROC curve” or “diagnosis”]. Additionally, citations in retrieved articles as well as systemic reviews or meta-analysis on the same topic were also searched where relevant.
Inclusion and exclusion criteria
Three investigators (Kai Qu, Tian Liu and Wenquan Niu) independently scanned the titles and abstracts of all retrieved articles to evaluate their eligibility. The articles fulfilling the following criteria were included for further analyses: (1) investigation of the diagnostic potential of circulating (blood, serum, plasma, digestive juice and CSF) miR-21 for human cancers; (2) the diagnosis of cancer patients was confirmed by pathological detection; (3) studies provided sufficient data, including case and control number, sensitivity and specificity. The exclusion criteria were as follows: (1) studies were obviously unrelated to diagnostic value of circulating miR-21; (2) studies were in forms of letters, editorials, case reports, meta-analyses or reviews.
Data extraction and meta-analysis
Data from all eligible studies were extracted as followings: basic characteristics of articles (including author name, publication year, country of the study, ethnicity, number of cases and controls, cancer and sample types), and diagnostic results [including sensitivity, specificity, AUC, true positive (TP), true negative (TN), false positive (FP), and false negative (FN)], respectively. All the meta-analysis were carried out using above data by the STATA12.0 software. To evaluate diagnostic effects, bivariate meta-analysis models were employed to calculate the pooled sensitivity, specificity, PLR, NLR, DOR and AUC. The summary receiver operator characteristic (ROC) curve was also conducted by pooling the sensitivity and specificity of each study. We performed Deek’s funnel plot to explore whether there was publication bias in our included studies; the P value was set at 0.10, which meant if P < 0.10, publication bias was significant.
Patients and sample collection
Overall, 35 patients pathologically diagnosed with glioma were included in the study. All included individuals were Chinese Han People and were recruited from the First Affiliated Hospital of Xi’an Jiaotong University. All tumor specimens obtained intra-operatively were collected, and were immediately fresh-frozen on dry ice and stored at −80°C until further detection. At least 2 mL of CSF sample was collected from each patient. All CSF samples were immediately cleared of cells and debris after collection by brief centrifugation and then stored at −80°C. The clinical-pathological characteristics of all participants recruited in the study are summarized in Table S2. This study has been approved by the Institutional Review Board for Human Research of the First Affiliated Hospital of Xi’an Jiaotong University.
Cell culture and reagents
Human glioma cell line U251 was obtained from Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China). U251 cells (5.0 × 104 cells/mL) were cultured in DMEM medium supplemented with 10% fetal calf serum (FCS) and penicillin/streptomicyn in a humidified atmosphere of 5% CO2/95% air at 37°C. Cell pellets and conditioned medium were harvested every 12 h during cell culture and stored at −80°C for further detection.
TGF-β1 was purchased from R&D Systems (Minneapolis, MN) and was dissolved at working concentration of 5.0 ng/ml. Galunisertib (TGFβ receptor I kinase inhibitor, LY2157299) was purchased from Selleck Chemicals (Houston, TX, USA) and was dissolved in PBS to make a 50 mmol/L stock solution. All above reagents were stored at −20°C.
Generation of miR-21 stably overexpressing cells
The expression plasmid for miRNA-21 (pCMV -miR-21) and the corresponding empty vector were purchased from OriGene (Rockville, MD). U251 cells were transfected with pCMV-miR-21 vector or empty vector in 24-well dishes using Lipofectamine reagent (Invitrogen, Carlsbad, CA) according to the manufacturer’s protocol. After 24 h transfection, the cells were replaced in a 10-cm dish followed by a 14-day selection using G418 (1 mg/ml). Surviving colonies were picked up from each transfectant and were then cultured for another 2 weeks in the presence of 0.3 mg/ml G418. The cells expressing the largest amount of pCMV-miR-21 were used as miR-21 stably expressing cells.
Smad3 siRNA and expression vector transfection
The siRNA sequences corresponding to the cDNA sequences of Smad3 (NM_005902) was as following: Smad3 siRNA, 5′-TCCGCATGAGCTTCGTCAA-3′; scramble negative control (SNC) siRNA, 5′-TTCTCCGA ACGTGTCACGTTT-3′. Smad3 expression vector (pMD-SMAD3) was purchased from Sino Biological Inc (Beijing, China). Equal amounts of an empty vector pcDNA3.1 serve as control. Transfection of siRNA (100 nM) or plasmid (4 μg) was carried out using a Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA) according to the procedure recommended by the manufacturer.
Total RNA extraction and quantitative real-time PCR
Total RNA derived from tumor tissues and cultured cells was extracted using Trizol Reagent (Invitrogen, Carlsbad, CA, USA), and total RNA derived from CSF and medium were isolated using Trizol LS Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. As previously reported [91, 92], miRNA expression levels were determined by quantitative real time-PCR applying the SYBR® PrimeScriptTM miRNA RT-PCR Kit and SYBR® Premix Ex TaqTM (TaKaRa Biotechnology, Dalian, China). The miRNA expression was assayed in triplicate and normalized to corresponding housekeeping miRNAs, RNU6 (for intracellular level of miRNAs) and cel-miR-39 (for extracellular level of miRNAs). The relative miRNA levels were calculated using the Comparative-Ct method (Ct method). All primers were synthesized by TaKaRa Biotechnology.
Western blotting
Cultured cells were lysed in RIPA buffer (Beyotime Inc., NanTong, China). Protein concentration was identified using the Bradford reagent (Beyotime Inc.). We then performed electrophoresis of protein extracts and subsequent blotting as previously described [91, 92]. In brief, Equivalent amounts of protein (30 μg) were separated by SDS-PAGE gels and transferred to polyvinylidene difluoride (PVDF) membranes (Bio-Rad, Hercules, CA, USA). The membranes were then immunoblotted with the appropriate primary antibodies against Smad3 or p-Smad3 (Santa Cruz Biotechnology, Santa Cruz, CA), at 4°C for overnight, and subsequently were incubated with HRP conjugated anti-mouse or anti-rabbit secondary antibodies at room temperature. Signals were detected using an enhanced chemiluminescence (ECL) system (Beyotime, China) on Kodak X-ray film. Equal protein loading was assessed by the expression of β-actin. The protein bands were quantified using the BioRad Quantity One software package.
Statistical analysis
All experiments in this study were repeated three times. The data were analyzed by SPSS 11.0 software (SPSS Inc, Chicago, IL, USA) and expressed as mean ± standard error of measurement (SEM). Analysis of continuous variables were performed using a Student’s t-test and P < 0.05 was considered statistically significant.
CONFLICTS OF INTEREST
There is no competing financial interest among the authors.
GRANT SUPPORT
This study was supported by National Science Foundation of China (Nos. 81071876, 81472247 and 81201549), the Project of Innovative Research Team for Key Science and Technology in Xi’an Jiaotong University, the Program for Innovative Research Team of Shaanxi Province (No. 2013KCJ-23), the Clinical Research Award of the First Affiliated Hospital of Xi’an Jiaotong University, China (No. XJTU1AF-CRF-2015–011) and the Fundamental Research Funds for the Central Universities.
Authors’ contributions
Kai Qu, Chang Liu and Shunbin Dong: Designed the research; Kai Qu and Tian Liu: Drafted the manuscript; Kai Qu, Chang Liu and Shunbin Dong: Revised the paper; Qing Pang and Ting Lin: Performed meta-analysis; Zhixin Wang, Minghui Tai and Jingyao Zhang: Participated in research work; Fandi Meng, Yong Wan and Ping Mao: Collected samples; Kai Qu, Qing Pang and Wenquan Niu: Analyzed data and constructed figures.
REFERENCES
1. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004; 116:281–297.
2. Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009; 136:642–655.
3. Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006; 6:857–866.
4. Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011; 39:7223–7233.
5. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, Peterson A, Noteboom J, O’Briant KC, Allen A, Lin DW, Urban N, Drescher CW, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008; 105:10513–10518.
6. Hsu CM, Lin PM, Wang YM, Chen ZJ, Lin SF, Yang MY. Circulating miRNA is a novel marker for head and neck squamous cell carcinoma. Tumour Biol. 2012; 33:1933–1942.
7. Yang Y, Gu X, Zhou M, Xiang J, Chen Z. Serum microRNAs: A new diagnostic method for colorectal cancer. Biomed Rep. 2013; 1:495–498.
8. Li S, Liang Z, Xu L, Zou F. MicroRNA-21: a ubiquitously expressed pro-survival factor in cancer and other diseases. Mol Cell Biochem. 2012; 360:147–158.
9. Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY. miR-21-mediated tumor growth. Oncogene. 2007; 26:2799–2803.
10. Kanaan Z, Rai SN, Eichenberger MR, Roberts H, Keskey B, Pan J, Galandiuk S. Plasma miR-21: a potential diagnostic marker of colorectal cancer. Ann Surg. 2012; 256:544–551.
11. Kumar S, Keerthana R, Pazhanimuthu A, Perumal P. Overexpression of circulating miRNA-21 and miRNA-146a in plasma samples of breast cancer patients. Indian J Biochem Biophys. 2013; 50:210–214.
12. Liao Q, Han P, Huang Y, Wu Z, Chen Q, Li S, Ye J, Wu X. Potential Role of Circulating microRNA-21 for Hepatocellular Carcinoma Diagnosis: A Meta-Analysis. PloS one. 2015; 10:e0130677.
13. Tomimaru Y, Eguchi H, Nagano H, Wada H, Kobayashi S, Marubashi S, Tanemura M, Tomokuni A, Takemasa I, Umeshita K, Kanto T, Doki Y, Mori M. Circulating microRNA-21 as a novel biomarker for hepatocellular carcinoma. J Hepatol. 2012; 56:167–175.
14. Shen L, Wan Z, Ma Y, Wu L, Liu F, Zang H, Xin S. The clinical utility of microRNA-21 as novel biomarker for diagnosing human cancers. Tumour Biol. 2015; 36:1993–2005.
15. Wang J, Chen J, Chang P, LeBlanc A, Li D, Abbruzzesse JL, Frazier ML, Killary AM, Sen S. MicroRNAs in plasma of pancreatic ductal adenocarcinoma patients as novel blood-based biomarkers of disease. Cancer Prev Res (Phila). 2009; 2:807–813.
16. Tsujiura M, Ichikawa D, Komatsu S, Shiozaki A, Takeshita H, Kosuga T, Konishi H, Morimura R, Deguchi K, Fujiwara H, Okamoto K, Otsuji E. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010; 102:1174–1179.
17. Komatsu S, Ichikawa D, Takeshita H, Tsujiura M, Morimura R, Nagata H, Kosuga T, Iitaka D, Konishi H, Shiozaki A, Fujiwara H, Okamoto K, Otsuji E. Circulating microRNAs in plasma of patients with oesophageal squamous cell carcinoma. Br J Cancer. 2011; 105:104–111.
18. Liu J, Gao J, Ren Y, Wang X, Wang W, Lu H. [Diagnostic value of plasma miR-21 in pancreatic cancer]. World Chin J Digestol. 2011; 19:860–863.
19. Shen J, Todd NW, Zhang H, Yu L, Lingxiao X, Mei Y, Guarnera M, Liao J, Chou A, Lu CL, Jiang Z, Fang H, Katz RL, et al. Plasma microRNAs as potential biomarkers for non-small-cell lung cancer. Lab Invest. 2011; 91:579–587.
20. Wei J, Liu LK, Gao W, Zhu CJ, Liu YQ, Cheng T, Shu YQ. Reduction of Plasma MicroRNA-21 is Associated with Chemotherapeutic Response in Patients with Non-small Cell Lung Cancer. Chin J Cancer Res. 2011; 23:123–128.
21. Li BS, Zhao YL, Guo G, Li W, Zhu ED, Luo X, Mao XH, Zou QM, Yu PW, Zuo QF, Li N, Tang B, Liu KY, et al. Plasma microRNAs, miR-223, miR-21 and miR-218, as novel potential biomarkers for gastric cancer detection. PLoS One. 2012; 7:e41629.
22. Kishimoto T, Eguchi H, Nagano H, Kobayashi S, Akita H, Hama N, Wada H, Kawamoto K, Tomokuni A, Tomimaru Y, Umeshita K, Doki Y, Mori M. Plasma miR-21 is a novel diagnostic biomarker for biliary tract cancer. Cancer Sci. 2013; 104:1626–1631.
23. Liu X, Luo HN, Tian WD, Lu J, Li G, Wang L, Zhang B, Liang BJ, Peng XH, Lin SX, Peng Y, Li XP. Diagnostic and prognostic value of plasma microRNA deregulation in nasopharyngeal carcinoma. Cancer Biol Ther. 2013; 14:1133–1142.
24. Luo X, Stock C, Burwinkel B, Brenner H. Identification and evaluation of plasma microRNAs for early detection of colorectal cancer. PLoS One. 2013; 8:e62880.
25. Ma J, Li N, Guarnera M, Jiang F. Quantification of Plasma miRNAs by Digital PCR for Cancer Diagnosis. Biomark Insights. 2013; 8:127–136.
26. Mozzoni P, Banda I, Goldoni M, Corradi M, Tiseo M, Acampa O, Balestra V, Ampollini L, Casalini A, Carbognani P, Mutti A. Plasma and EBC microRNAs as early biomarkers of non-small-cell lung cancer. Biomarkers. 2013; 18:679–686.
27. Ouyang L, Liu P, Yang S, Ye S, Xu W, Liu X. A three-plasma miRNA signature serves as novel biomarkers for osteosarcoma. Med Oncol. 2013; 30:340.
28. Tang D, Shen Y, Wang M, Yang R, Wang Z, Sui A, Jiao W, Wang Y. Identification of plasma microRNAs as novel noninvasive biomarkers for early detection of lung cancer. Eur J Cancer Prev. 2013; 22:540–548.
29. Abue M, Yokoyama M, Shibuya R, Tamai K, Yamaguchi K, Sato I, Tanaka N, Hamada S, Shimosegawa T, Sugamura K, Satoh K. Circulating miR-483–3p and miR-21 is highly expressed in plasma of pancreatic cancer. Int J Oncol. 2015; 46:539–47. doi: 10.3892/ijo.2014.2743.
30. Jones K, Nourse JP, Keane C, Bhatnagar A, Gandhi MK. Plasma microRNA are disease response biomarkers in classical Hodgkin lymphoma. Clin Cancer Res. 2014; 20:253–264.
31. Liu SS, Wang YS, Sun YF, Miao LX, Wang J, Li YS, Liu HY, Liu QL. Plasma microRNA-320, microRNA-let-7e and microRNA-21 as novel potential biomarkers for the detection of retinoblastoma. Biomed Rep. 2014; 2:424–428.
32. Ye M, Ye P, Zhang W, Rao J, Xie Z. [Diagnostic values of salivary versus and plasma microRNA-21 for early esophageal cancer]. [Article in Chinese]. Nan Fang Yi Ke Da Xue Xue Bao. 2014; 34:885–889.
33. Zanutto S, Pizzamiglio S, Ghilotti M, Bertan C, Ravagnani F, Perrone F, Leo E, Pilotti S, Verderio P, Gariboldi M, Pierotti MA. Circulating miR-378 in plasma: a reliable, haemolysis-independent biomarker for colorectal cancer. Br J Cancer. 2014; 110:1001–1007.
34. Asaga S, Kuo C, Nguyen T, Terpenning M, Giuliano AE, Hoon DS. Direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin Chem. 2011; 57:84–91.
35. Xu J, Wu C, Che X, Wang L, Yu D, Zhang T, Huang L, Li H, Tan W, Wang C, Lin D. Circulating microRNAs, miR-21, miR-122, and miR-223, in patients with hepatocellular carcinoma or chronic hepatitis. Mol Carcinog. 2011; 50:136–142.
36. Le HB, Zhu WY, Chen DD, He JY, Huang YY, Liu XG, Zhang YK. Evaluation of dynamic change of serum miR-21 and miR-24 in pre- and post-operative lung carcinoma patients. Med Oncol. 2012; 29:3190–3197.
37. Liu AM, Yao TJ, Wang W, Wong KF, Lee NP, Fan ST, Poon RT, Gao C, Luk JM. Circulating miR-15b and miR-130b in serum as potential markers for detecting hepatocellular carcinoma: a retrospective cohort study. BMJ Open. 2012; 2:e000825.
38. Wang B, Zhang Q. The expression and clinical significance of circulating microRNA-21 in serum of five solid tumors. J Cancer Res Clin Oncol. 2012; 138:1659–1666.
39. Abd-El-Fattah AA, Sadik NA, Shaker OG, Aboulftouh ML. Differential microRNAs expression in serum of patients with lung cancer, pulmonary tuberculosis, and pneumonia. Cell Biochem Biophys. 2013; 67:875–884.
40. Egidi MG, Cochetti G, Serva MR, Guelfi G, Zampini D, Mechelli L, Mearini E. Circulating microRNAs and kallikreins before and after radical prostatectomy: are they really prostate cancer markers? Biomed Res Int. 2013; 2013:241780.
41. Gao J, Zhang Q, Xu J, Guo L, Li X. Clinical significance of serum miR-21 in breast cancer compared with CA153 and CEA. Chin J Cancer Res. 2013; 25:743–748.
42. Liu GH, Zhou ZG, Chen R, Wang MJ, Zhou B, Li Y, Sun XF. Serum miR-21 and miR-92a as biomarkers in the diagnosis and prognosis of colorectal cancer. Tumour Biol. 2013; 34:2175–2181.
43. Mar-Aguilar F, Mendoza-Ramirez JA, Malagon-Santiago I, Espino-Silva PK, Santuario-Facio SK, Ruiz-Flores P, Rodriguez-Padilla C, Resendez-Perez D. Serum circulating microRNA profiling for identification of potential breast cancer biomarkers. Dis Markers. 2013; 34:163–169.
44. Que R, Ding G, Chen J, Cao L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J Surg Oncol. 2013; 11:219.
45. Si H, Sun X, Chen Y, Cao Y, Chen S, Wang H, Hu C. Circulating microRNA-92a and microRNA-21 as novel minimally invasive biomarkers for primary breast cancer. J Cancer Res Clin Oncol. 2013; 139:223–229.
46. Toiyama Y, Takahashi M, Hur K, Nagasaka T, Tanaka K, Inoue Y, Kusunoki M, Boland CR, Goel A. Serum miR-21 as a diagnostic and prognostic biomarker in colorectal cancer. J Natl Cancer Inst. 2013; 105:849–859.
47. Kotb S, Mosharafa A, Essawi M, Hassan H, Meshref A, Morsy A. Circulating miRNAs 21 and 221 as biomarkers for early diagnosis of prostate cancer. Tumour Biol. 2014; 35:12613–7. doi: 10.1007/s13277-014-2584-7.
48. Mao X, Sun Y, Tang J. Serum miR-21 is a diagnostic and prognostic marker of primary central nervous system lymphoma. Neurol Sci. 2014; 35:233–238.
49. Wang J, Zhou Y, Lu J, Sun Y, Xiao H, Liu M, Tian L. Combined detection of serum exosomal miR-21 and HOTAIR as diagnostic and prognostic biomarkers for laryngeal squamous cell carcinoma. Med Oncol. 2014; 31:148.
50. Wu J, Li G, Wang Z, Yao Y, Chen R, Pu X, Wang J. Circulating MicroRNA-21 Is a Potential Diagnostic Biomarker in Gastric Cancer. Dis Markers. 2015; 2015:435656.
51. Baraniskin A, Kuhnhenn J, Schlegel U, Chan A, Deckert M, Gold R, Maghnouj A, Zollner H, Reinacher-Schick A, Schmiegel W, Hahn SA, Schroers R. Identification of microRNAs in the cerebrospinal fluid as marker for primary diffuse large B-cell lymphoma of the central nervous system. Blood. 2011; 117:3140–3146.
52. Baraniskin A, Kuhnhenn J, Schlegel U, Maghnouj A, Zollner H, Schmiegel W, Hahn S, Schroers R. Identification of microRNAs in the cerebrospinal fluid as biomarker for the diagnosis of glioma. Neuro Oncol. 2012; 14:29–33.
53. Teplyuk NM, Mollenhauer B, Gabriely G, Giese A, Kim E, Smolsky M, Kim RY, Saria MG, Pastorino S, Kesari S, Krichevsky AM. MicroRNAs in cerebrospinal fluid identify glioblastoma and metastatic brain cancers and reflect disease activity. Neuro Oncol. 2012; 14:689–700.
54. Akers JC, Ramakrishnan V, Kim R, Skog J, Nakano I, Pingle S, Kalinina J, Hua W, Kesari S, Mao Y, Breakefield XO, Hochberg FH, Van Meir EG, et al. MiR-21 in the extracellular vesicles (EVs) of cerebrospinal fluid (CSF): a platform for glioblastoma biomarker development. PLoS One. 2013; 8:e78115.
55. Yuan P, He XY, Li XG. [Identification of miR-128 and miR-21 in the cerebrospinal fluid as biomarker for the diagnosis of glioma]. Int J Lab Med. 2014; 35:2464–2468.
56. Shi R, Wang PY, Li XY, Chen JX, Li Y, Zhang XZ, Zhang CG, Jiang T, Li WB, Ding W, Cheng SJ. Exosomal levels of miRNA-21 from cerebrospinal fluids associated with poor prognosis and tumor recurrence of glioma patients. Oncotarget. 2015; 6:26971–26981. doi: 10.18632/oncotarget.4699.
57. Xie Z, Chen G, Zhang X, Li D, Huang J, Yang C, Zhang P, Qin Y, Duan Y, Gong B, Li Z. Salivary microRNAs as promising biomarkers for detection of esophageal cancer. PLoS One. 2013; 8:e57502.
58. Humeau M, Vignolle-Vidoni A, Sicard F, Martins F, Bournet B, Buscail L, Torrisani J, Cordelier P. Salivary MicroRNA in Pancreatic Cancer Patients. PloS one. 2015; 10:e0130996.
59. Cui L, Zhang X, Ye G, Zheng T, Song H, Deng H, Xiao B, Xia T, Yu X, Le Y, Guo J. Gastric juice MicroRNAs as potential biomarkers for the screening of gastric cancer. Cancer. 2013; 119:1618–1626.
60. Sadakari Y, Ohtsuka T, Ohuchida K, Tsutsumi K, Takahata S, Nakamura M, Mizumoto K, Tanaka M. MicroRNA expression analyses in preoperative pancreatic juice samples of pancreatic ductal adenocarcinoma. JOP. 2010; 11:587–592.
61. Shen J, Liao J, Guarnera MA, Fang H, Cai L, Stass SA, Jiang F. Analysis of MicroRNAs in sputum to improve computed tomography for lung cancer diagnosis. J Thorac Oncol. 2014; 9:33–40.
62. Ryu JK, Matthaei H, Dal Molin M, Hong SM, Canto MI, Schulick RD, Wolfgang C, Goggins MG, Hruban RH, Cope L, Maitra A. Elevated microRNA miR-21 levels in pancreatic cyst fluid are predictive of mucinous precursor lesions of ductal adenocarcinoma. Pancreatology. 2011; 11:343–350.
63. Li Y, Li W, Ouyang Q, Hu S, Tang J. Detection of lung cancer with blood microRNA-21 expression levels in Chinese population. Oncol Lett. 2011; 2:991–994.
64. Zheng Y, Cui L, Sun W, Zhou H, Yuan X, Huo M, Chen J, Lou Y, Guo J. MicroRNA-21 is a new marker of circulating tumor cells in gastric cancer patients. Cancer Biomark. 2011; 10:71–77.
65. Ng EKO, Li R, Shin VY, Jin HC, Leung CPH, Ma ESK, Pang R, Chua D, Chu K-M, Law WL, Law SYK, Poon RTP, Kwong A. Circulating microRNAs as Specific Biomarkers for Breast Cancer Detection. Plos One. 2013; 8.
66. Basati G, Razavi AE, Abdi S, Mirzaei A. Elevated level of microRNA-21 in the serum of patients with colorectal cancer. Med Oncol. 2014; 31.
67. Du M, Liu S, Gu D, Wang Q, Zhu L, Kang M, Shi D, Chu H, Tong N, Chen J, Adams TS, Zhang Z, Wang M. Clinical potential role of circulating microRNAs in early diagnosis of colorectal cancer patients. Carcinogenesis. 2014; 35:2723–30. doi: 10.1093/carcin/bgu189.
68. Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, Gunji T, Ohta H, Okamoto H, Sonoda H, Watanabe M, Nakagama H, Yokota J, et al. Circulating Exosomal microRNAs as Biomarkers of Colon Cancer. Plos One. 2014; 9.
69. Zhang H, Li P, Ju H, Pesta M, Kulda V, Jin W, Cai M, Liu C, Wu H, Xu J, Ye Y, Zhang G, Xu E, et al. Diagnostic and prognostic value of micro RNA-21 in colorectal cancer: An original study and individual participant data meta-analysis. Cancer Epidemiol Biomarkers Prev. 2014; 23:2783–92. doi: 10.1158/1055-9965.EPI-14-0598.
70. Erbes T, Hirschfeld M, Rucker G, Jaeger M, Boas J, Iborra S, Mayer S, Gitsch G, Stickeler E. Feasibility of urinary microRNA detection in breast cancer patients and its potential as an innovative non-invasive biomarker. BMC cancer. 2015; 15:193.
71. Kosaka N, Iguchi H, Ochiya T. Circulating microRNA in body fluid: a new potential biomarker for cancer diagnosis and prognosis. Cancer sci. 2010; 101:2087–2092.
72. Aguirre-Gamboa R, Trevino V. SurvMicro: assessment of miRNA-based prognostic signatures for cancer clinical outcomes by multivariate survival analysis. Bioinformatics. 2014; 30:1630–1632.
73. Qian B, Katsaros D, Lu L, Preti M, Durando A, Arisio R, Mu L, Yu H. High miR-21 expression in breast cancer associated with poor disease-free survival in early stage disease and high TGF-beta1. Breast Cancer Res Treat. 2009; 117:131–140.
74. Wang T, Zhang L, Shi C, Sun H, Wang J, Li R, Zou Z, Ran X, Su Y. TGF-beta-induced miR-21 negatively regulates the antiproliferative activity but has no effect on EMT of TGF-beta in HaCaT cells. Int J Biochem Cell Biol. 2012; 44:366–376.
75. Serova M, Tijeras-Raballand A, Dos Santos C, Albuquerque M, Paradis V, Neuzillet C, Benhadji KA, Raymond E, Faivre S, de Gramont A. Effects of TGF-beta signalling inhibition with galunisertib (LY2157299) in hepatocellular carcinoma models and in ex vivo whole tumor tissue samples from patients. Oncotarget. 2015; 6:21614–21627. doi: 10.18632/oncotarget.4308.
76. Thapa N, Lee BH, Kim IS. TGFBIp/betaig-h3 protein: a versatile matrix molecule induced by TGF-beta. Int J Biochem Cell Biol. 2007; 39:2183–2194.
77. Chan SH, Wu CW, Li AF, Chi CW, Lin WC. miR-21 microRNA expression in human gastric carcinomas and its clinical association. Anticancer Res. 2008; 28:907–911.
78. Wang Y, Gao X, Wei F, Zhang X, Yu J, Zhao H, Sun Q, Yan F, Yan C, Li H, Ren X. Diagnostic and prognostic value of circulating miR-21 for cancer: a systematic review and meta-analysis. Gene. 2014; 533:389–397.
79. Yang X, Guo Y, Du Y, Yang J, Li S, Liu S, Li K, Zhang D. Serum microRNA-21 as a diagnostic marker for lung carcinoma: a systematic review and meta-analysis. PloS one. 2014; 9:e97460.
80. Yang Y, Qian J, Chen Y, Pan Y. Prognostic role of circulating microRNA-21 in cancers: evidence from a meta-analysis. Tumour biol. 2014; 35:6365–6371.
81. Ye TT, Yang YL, Liu XY, Ji QQ, Pan YF, Xiang YQ. Prognostic value of circulating microRNA-21 in digestive system cancers: a meta-analysis. Int J Clin Exp Med. 2014; 7:873–878.
82. Zhu X, Lv M, Wang H, Guan W. Identification of circulating microRNAs as novel potential biomarkers for gastric cancer detection: a systematic review and meta-analysis. Dig Dis Sci. 2014; 59:911–919.
83. Conti A, Aguennouz M, La Torre D, Tomasello C, Cardali S, Angileri FF, Maio F, Cama A, Germano A, Vita G, Tomasello F. miR-21 and 221 upregulation and miR-181b downregulation in human grade II-IV astrocytic tumors. J Neurooncol. 2009; 93:325–332.
84. Hermansen SK, Dahlrot RH, Nielsen BS, Hansen S, Kristensen BW. MiR-21 expression in the tumor cell compartment holds unfavorable prognostic value in gliomas. J Neurooncol. 2013; 111:71–81.
85. Tao T, Wang Y, Luo H, Yao L, Wang L, Wang J, Yan W, Zhang J, Wang H, Shi Y, Yin Y, Jiang T, Kang C, et al. Involvement of FOS-mediated miR-181b/miR-21 signalling in the progression of malignant gliomas. Eur J Cancer. 2013; 49:3055–3063.
86. Wu L, Li G, Feng D, Qin H, Gong L, Zhang J, Zhang Z. MicroRNA-21 expression is associated with overall survival in patients with glioma. Diagn Pathol. 2013; 8:200.
87. Yang CH, Yue J, Pfeffer SR, Fan M, Paulus E, Hosni-Ahmed A, Sims M, Qayyum S, Davidoff AM, Handorf CR, Pfeffer LM. MicroRNA-21 Promotes Glioblastoma Tumorigenesis by Down-regulating Insulin-like Growth Factor-binding Protein-3 (IGFBP3). J Biol Chem. 2014; 289:25079–25087.
88. Chung AC, Dong Y, Yang W, Zhong X, Li R, Lan HY. Smad7 suppresses renal fibrosis via altering expression of TGF-beta/Smad3-regulated microRNAs. Mol Ther. 2013; 21:388–398.
89. Jiang Y, Chen X, Tian W, Yin X, Wang J, Yang H. The role of TGF-beta1-miR-21-ROS pathway in bystander responses induced by irradiated non-small-cell lung cancer cells. Br J Cancer. 2014; 111:772–780.
90. Wang J, Li Y, Wang X, Jiang C. Ursolic acid inhibits proliferation and induces apoptosis in human glioblastoma cell lines U251 by suppressing TGF-beta1/miR-21/PDCD4 pathway. Clin Pharmacol Toxicol. 2012; 111:106–112.
91. Xu X, Chen W, Miao R, Zhou Y, Wang Z, Zhang L, Wan Y, Dong Y, Qu K, Liu C. miR-34a induces cellular senescence via modulation of telomerase activity in human hepatocellular carcinoma by targeting FoxM1/c-Myc pathway. Oncotarget. 2015; 6:3988–4004. doi: 10.18632/oncotarget.2905.
92. Qu K, Xu X, Liu C, Wu Q, Wei J, Meng F, Zhou L, Wang Z, Lei L, Liu P. Negative regulation of transcription factor FoxM1 by p53 enhances oxaliplatin-induced senescence in hepatocellular carcinoma. Cancer Lett. 2013; 331:105–114.

