INTRODUCTION
Chronic inflammation is an important mediator of nasopharyngeal carcinoma (NPC) [1-3] and gastric cancer (GC) [4]. A large number of inflammatory cells, including tumor-associated macrophages (TAMs), are found in NPC and GC biopsies. TAM infiltration is tightly associated with poor prognosis in NPC [5, 6]. TAMs were shown, together with tumor-derived FasL, to serve as a barrier against the infiltration of CD8+ T cells into GC [7, 8]. Furthermore, our previous studies demonstrated that TAM-derived inflammatory factors, such as IL-6, stimulate NPC cell proliferation [9, 10].
Epstein-Barr virus (EBV) infection is closely associated with the development and progression of NPC and GC [11-13]. EBV encodes 44 mature microRNAs (miRNAs), divided into two clusters, BHRFs and BARTs [14-16]. BART miRNAs were shown to affect the malignant phenotype of some lymphomas [17], GC [18], and NPC [19-22], including viral latency [23-25], immune escape [26], cell proliferation [27], cell apoptosis [28, 29], cell cycle regulation, and cancer metabolism [27-30]. These findings suggest that EBV miRNAs may exert a variety of important regulatory functions in EBV-mediated tumorigenesis and cancer progression. Nevertheless, the functions of most EBV-encoded miRNAs remain to be elucidated.
We previously profiled all 44 EBV-encoded mature miRNAs in NPC biopsies and non-cancerous nasopharyngeal tissues, and found that EBV miRNAs located in the BART region were highly expressed in NPC biopsies [31-33]. Additionally, we constructed an EBV miRNA-dependent gene regulatory network and determined that both the 3′- and 5′-arms of the mature EBV-miR-BART11 hairpin precursor may specifically target forkhead box P1 (FOXP1) mRNA. FOXP1 has been reported to be involved in monocyte differentiation to macrophage, where its downregulation represents a key molecular event [34, 35]. However, the biological functions of both EBV-miR-BART11 and FOXP1 in EBV-associated carcinogenesis have not been defined yet. We hypothesized that EBV may participate in TAM differentiation and promote inflammation-induced EBV-associated carcinogenesis via EBV-miRNA-BART11 expression and subsequent FOXP1 downregulation.
RESULTS
FOXP1 is a target of EBV-miR-BART11
Bioinformatic analysis identified three putative EBV-miR-BART11 binding sites (two for EBV-miR-BART11-3p and one for EBV-miR-BART11-5p) in the FOXP1 3′-untranslated region (UTR) (Figure 1A). To determine the effect of EBV-miR-BART11 on FOXP1 expression, EBV-miR-BART11 precursor vector expressing both mature EBV-miR-BART11-3p and EBV-miR-BART11-5p was constructed and transfected into three different EBV-negative cancer cell lines (5-8F, HK-1, and AGS). The expression of mature EBV-miR-BART11-3p and EBV-miR-BART11-5p was measured using qRT-PCR (Figure 1B). These results demonstrated that the EBV-miR-BART11 precursor vector can successfully express mature EBV-miR-BART11-3p and EBV-miR-BART11-5p in EBV-negative cancer cell lines. Further analysis revealed that EBV-miR-BART11 significantly inhibited FOXP1 expression at the mRNA and protein levels when compared with empty vector controls in 5-8F, HK-1, and AGS cells (Figure 1C–1D).
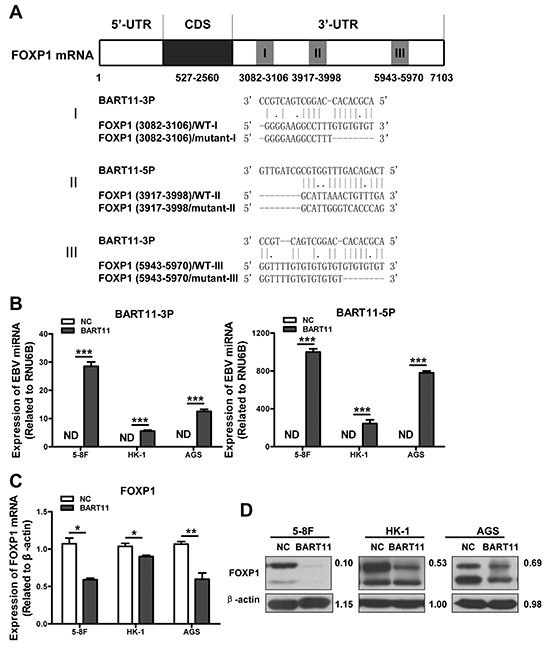
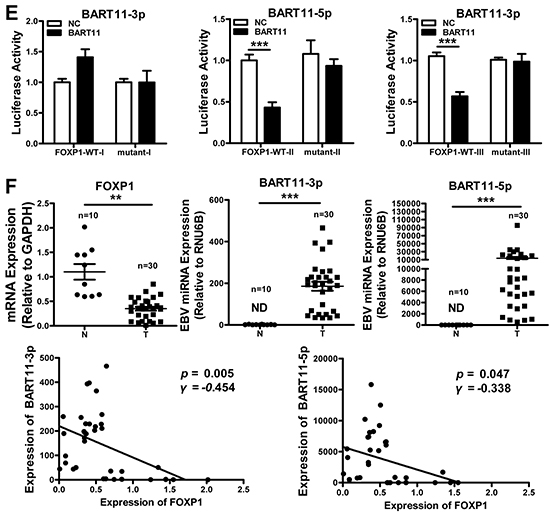
Figure 1: FOXP1 is a direct target of EBV-miR-BART11. A. Three binding sites of EBV-miR-BART11-3p and EBV-miR-BART11-5p were predicted in the FOXP1 3’-UTR, including 3082 bp-3106 bp (I), 3917 bp-3998 bp (II), and 5943 bp-5970 bp (III). Wild-type (FOXP1-WT) and mutant (FOXP1-mutant) sequences were used to validate these predictions. B. The expression of exogenous EBV-miR-BART11-3p (left) and EBV-miR-BART11-5p (right) was detected by qRT-PCR. FOXP1 C. mRNA and D. protein expression levels in 5-8F, HK-1, and AGS cells after EBV-miR-BART11 treatment. β-actin served as loading control. E. Luciferase reporter assay, using reporter vectors containing either wild-type (FOXP1-WT) or mutant (FOXP1-mutant) FOXP1 3′-UTR, and EBV-miR-BART11 or non-targeting control, was performed in order to identify the direct binding of EBV-miR-BART11 to the FOXP1 3′-UTR in 5-8F cells. F. EBV-miR-BART11 and FOXP1 expression levels in NPC and control specimens were detected by qRT-PCR. N, non-tumor nasopharyngeal epithelium (n = 10); T, NPC (n = 30). Representative images or data expressed as mean ± SD of the measurements obtained in three separate experiments are presented (ND: not detected; *p < 0.05; **p < 0.01;***p < 0.001).
To elucidate if FOXP1 is a direct target of EBV-miR-BART11-3p and EBV-miR-BART11-5p, three pairs of luciferase reporter vectors containing either wild-type (WT-I, WT-II, and WT-III) EBV-miR-BART11 binding or mutant sequences of the FOXP1 3′-UTR were co-transfected with the EBV-miR-BART11 precursor expression vector in 5-8F cells. EBV-miR-BART11 significantly attenuated the luciferase activity of FOXP1-WT vectors II and III, but exhibited no effects on the FOXP1-WT-I vector or the FOXP1-mutant vectors (Figure 1E). These results suggested that EBV-miR-BART11 is able to inhibit FOXP1 expression by targeting the binding sites II and III in the FOXP1 3′-UTR.
To explore the relationship between EBV-miR-BART11 and FOXP1, EBV-miR-BART11-(3p and 5p) and FOXP1 mRNA expression was assessed in 30 NPC biopsies and 10 non-tumor nasopharyngeal epithelial tissues. As expected, EBV-miR-BART11-3p and EBV-miR-BART11-5p expression levels were significantly higher in NPC samples than in normal nasopharyngeal epithelial samples, and these levels were negatively correlated with FOXP1 expression (p < 0.05, Figure 1F).
EBV-miR-BART11 promotes monocyte differentiation by attenuating FOXP1 expression
In order to define the relationship between EBV-miR-BART11 and FOXP1 in monocyte to macrophage differentiation further, we monitored temporal FOXP1 expression in THP-1 monocytes subjected to PMA-induced macrophage differentiation. This revealed that FOXP1 is dramatically downregulated during monocyte to macrophage transformation (Figure 2A), which is consistent with the previous reports [34, 35]. Therefore, we hypothesized that EBV-miR-BART11 may stimulate monocyte differentiation. To test this theory, THP-1 cells were infected with lentivirus encoding FOXP1 or EBV-miR-BART11, and treated with PMA to induce differentiation. The results revealed that EBV-miR-BART11 downregulated both FOXP1 mRNA and protein expression (Figure 2B). In addition, we found that FOXP1 expression hindered PMA-induced THP-1 differentiation, whereas EBV-miR-BART11 overexpression was shown to induce this process, compared with the negative control (Figure 2C).
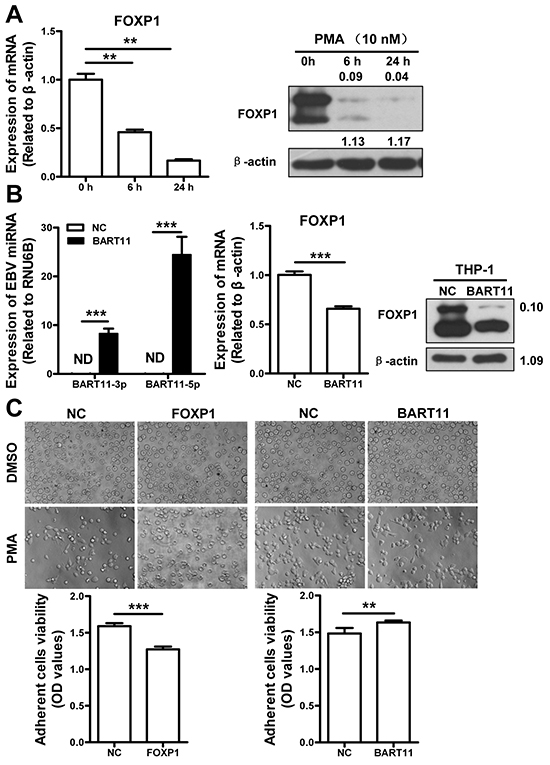
Figure 2: EBV-miR-BART11 promotes monocyte differentiation of THP-1 cells by attenuating FOXP1 expression. A. FOXP1 expression at mRNA (left) and protein (right) levels, during the PMA-induced differentiation of monocytic THP-1 cells. B. The expression of EBV-miR-BART11 (left) and FOXP1 (middle, mRNA; right, protein) was examined by qRT-PCR and western blotting, respectively, in THP-1 cells infected with lentivirus encoding EBV-miR-BART11. C. The effects of EBV-miR-BART11 and FOXP1 on monocyte differentiation. Morphological changes were monitored in PMA-induced THP-1 cells following FOXP1 overexpression vector or EBV-miR-BART11 precursor vector transfection. THP-1 cell differentiation was determined by the viability of adherent cells, using MTT assay. Data represent mean ± SD of OD values obtained in three separate experiments (ND: not detected; **p < 0.01; ***p < 0.001).
EBV-miR-BART11-expressing macrophages are hyperresponsive to LPS
Macrophages play a key role in chronic inflammation and can trigger a pro-inflammatory response by secreting inflammatory factors [36, 37]. The expression of several prototypical pro-inflammatory cytokines (IL-1β, IL-6, and IL-8) markedly increased in PMA-induced THP-1 monocytes (D-THP-1), compared with the untreated controls (Figure 3A). In order to investigate the effects of FOXP1 and EBV-miR-BART11 on LPS-induced pro-inflammatory cytokine production in D-THP-1 cells, the investigated cells were infected with lentivirus encoding FOXP1 or EBV-miR-BART11. Notably, IL-1β, IL-6, and IL-8 mRNA expression in FOXP1-overexpressing cells was significantly lower than in controls. Elevated cytokine levels were detected in both the control and FOXP1-overexpressing cells stimulated with LPS (1 μg/mL) for 24 h; however, the levels in FOXP1-overexpressing cells were shown to be significantly lower than those in controls. Together, these data demonstrated that FOXP1 attenuates spontaneous and LPS-stimulated IL-1β, IL-6, and IL-8 expression in D-THP-1 cells, whereas EBV-miR-BART11 has the opposite effect (Figure 3B).
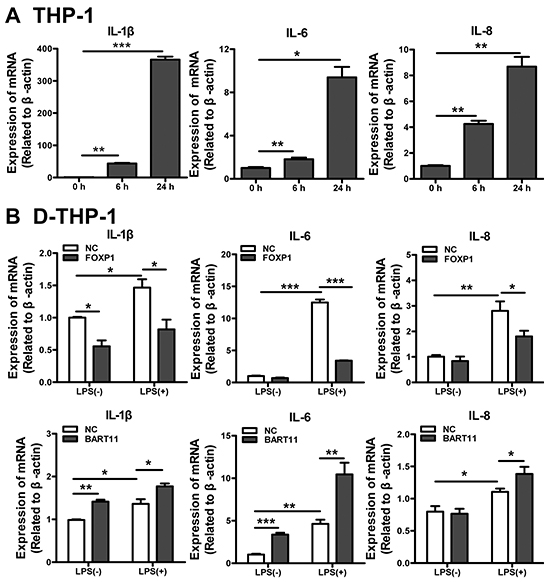
Figure 3: EBV-miR-BART11-transfected macrophages are hyperresponsive to LPS. A. Pro-inflammatory cytokine (IL-1β, IL-6, and IL-8) expression in PMA-treated THP-1 cells. B. Differentiated THP-1 (D-THP-1) cells, stimulated with LPS in the presence of exogenous FOXP1 overexpression (upper panel) or EBV-miR-BART11 (lower panel). Data are expressed as mean ± SD of the results obtained from each group of cells in three separate experiments (*p < 0.05; **p < 0.01; ***p < 0.001).
EBV-miR-BART11 enhances the local inflammatory response in carcinoma
TAMs are a major component of the immune infiltrates present in the NPC microenvironment, where they produce cytokines, growth factors, and angiogenic inducers that amplify the inflammatory response and promote tumor survival and growth [38]. The respective contributions of FOXP1 and EBV-miR-BART11 to this process in nasopharyngeal epithelial cells remain unclear. Therefore, we examined IL-1β, IL-6, and IL-8 production in EBV-negative cell lines (5-8F, HK-1, and AGS) following LPS treatment. Naïve EBV-negative cells were treated with conditioned media collected from LPS-treated D-THP-1 cells, in order to mimic the tumor microenvironment. IL-1β, IL-6, and IL-8 mRNA expression increased in EBV-negative epithelial cells following LPS treatment, which was potentiated after culturing in conditioned media. Consistent with our earlier findings [6], exogenous expression of FOXP1 or EBV-miR-BART11 was shown to significantly reduce or potentiate the local inflammatory response, respectively (Figure 4A–4B).
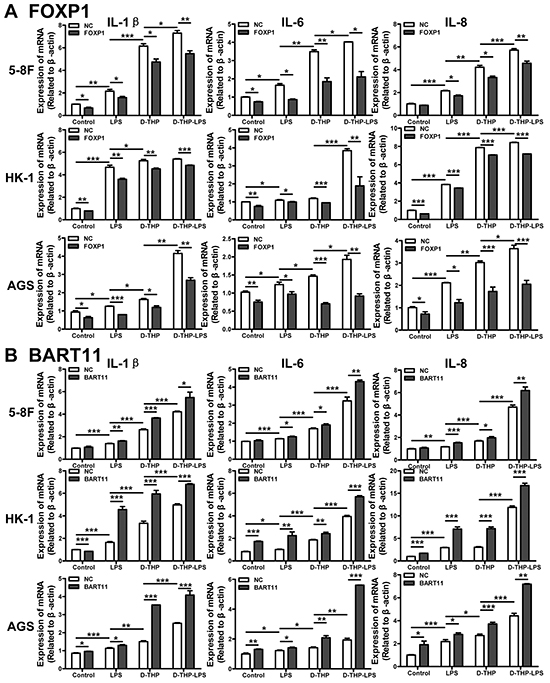
Figure 4. Effect of FOXP1 and EBV-miR-BART11 expression on the local inflammatory response in epithelial cells. The expression of inflammatory factors (IL-1β, IL-6, and IL-8) , measured in 5-8F and HK-1 NPC and AGS GC epithelial cells transfected with FOXP1 overexpression vector A. or EBV-miR-BART11 precursor vector B. and treated with LPS or conditioned media collected from the untreated differentiated-THP-1 cells (D-THP) or LPS-treated D-THP-1 cells (D-THP-LPS). Data are represented as mean ± SD of three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001.
EBV-miR-BART11 accelerates cell proliferation by attenuating FOXP1 expression
Cytokines stimulated by the local inflammatory response promote epithelial cell carcinogenesis through the induction of cell proliferation [5, 9]. In order to understand the roles of EBV-miR-BART11 and FOXP1 in epithelial cell proliferation, EBV-negative cell lines were transfected with EBV-miR-BART11, FOXP1, or FOXP1 siRNAs (siFOXP1), and cell proliferation was examined by the MTT assay. siFOXP1 was demonstrated to induce FOXP1 knockdown specifically, at both mRNA and protein levels (Supplementary Figure S1). Both EBV-miR-BART11 overexpression and FOXP1 silencing enhanced cell proliferation, while FOXP1 overexpression had the opposite result (Figure 5A). These findings were validated using the colony formation assay as well, which showed that FOXP1 expression significantly reduced colony forming potential, in terms of both colony number and size. Similarly, EBV-miR-BART11 and siFOXP1 exhibited the opposite effects on colony formation potential (Figure 5B). Furthermore, flow cytometric analysis revealed that FOXP1 overexpression results in a higher percentage of cell accumulation in G0/G1 phase, compared with the non-transfected controls (Figure 5C).
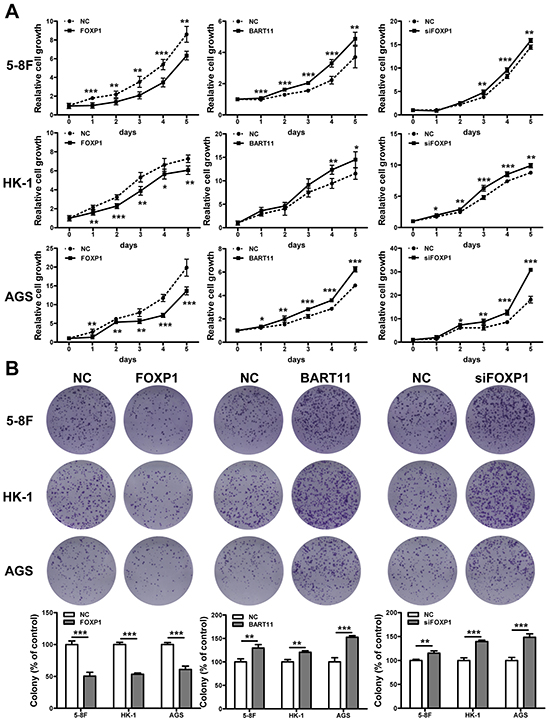
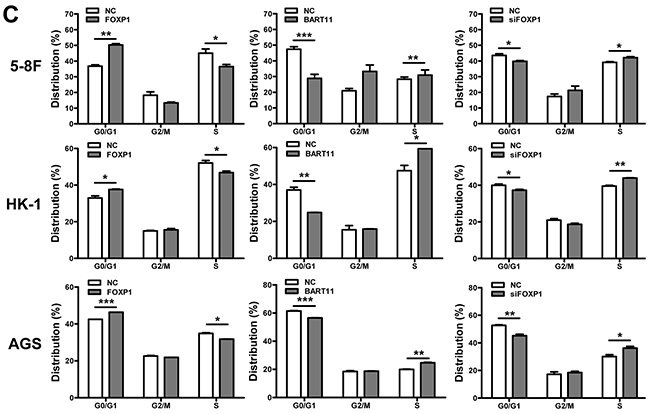
Figure 5: Effect of EBV-miR-BART11 and FOXP1 on cancer cell proliferation. The effect of FOXP1 or EBV-miR-BART11 overexpression, or FOXP1 siRNA knockdown (siFOXP1) on cell proliferation was investigated by the A. MTT assay and B. colony formation assay in 5-8F, HK-1, and AGS cells. C. Cell cycle analysis of cell cycle progression by flow cytometric analysis. All data were normalized to the vector-only control cell proliferation rate, and the results are expressed as mean ± SD of the results obtained in three independent experiments (*p < 0.05; **p < 0.01; ***p < 0.001).
EBV-miR-BART11 promotes inflammation-induced cell proliferation by suppressing FOXP1 expression
Our previous study indicated that LPS-stimulated TAMs can promote epithelial cell proliferation [5]. In order to understand the functions of EBV-miR-BART11 and FOXP1 in inflammation-induced cell proliferation, 5-8F, HK-1, and AGS cells with exogenous EBV-miR-BART11 or FOXP1 expression, or FOXP1 knockdown cells were cultured in conditioned media obtained from LPS-stimulated D-THP-1 cells. These results showed that EBV-miR-BART11 overexpression (Figure 6A) or FOXP1 knockdown (Figure 6B) can promote inflammation-induced cell proliferation, whereas FOXP1 overexpression inhibited this effect (Figure 6C).
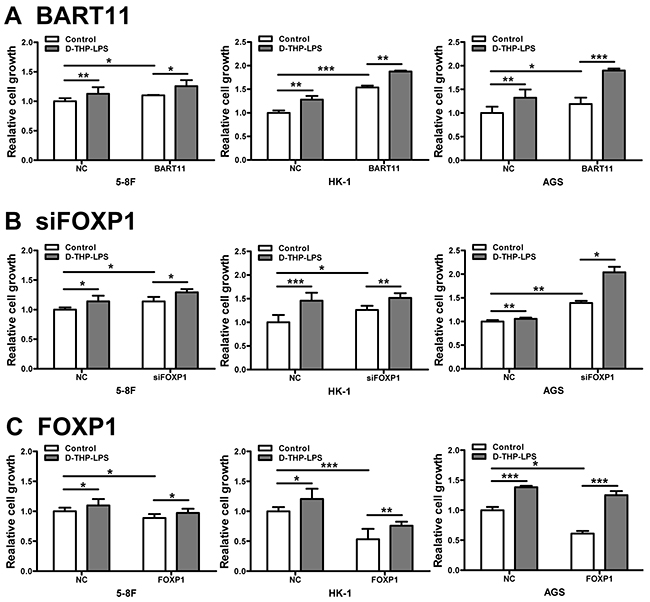
Figure 6: Effect of EBV-miR-BART11 and FOXP1 on inflammation-induced cancer cell proliferation. The proliferation of 5-8F, HK-1, and AGS cells transfected with EBV-miR-BART11 precursor vector A. FOXP1 siRNA B. or FOXP1 overexpression vector C. cultured in the conditioned media collected from LPS-treated differentiated THP-1 cells (D-THP-LPS). Data are expressed as mean ± SD of cell growth following different treatments, of three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001.
EBV-miR-BART11 activates NF-κB signaling through FOXP1
Nuclear factor κB (NF-κB) orchestrates the inflammatory response by regulating the expression of numerous genes [5, 36]. Therefore, we investigated the effects of FOXP1 and EBV-miR-BART11 on NF-κB p65 expression in THP-1, 5-8F, HK-1, and AGS cells. FOXP1 overexpression markedly decreased NF-κB p65 protein expression, while EBV-miR-BART11 overexpression or FOXP1 knockdown led to an increase in NF-κB p65 expression (Figure 7A). The analysis of NF-κB transcriptional activity using the reporter assay in 5-8F cells revealed that exogenous FOXP1 expression led to the inhibition of NF-κB transcriptional activity, while EBV-miR-BART11 or FOXP1 siRNA had the opposite effect (Figure 7B).
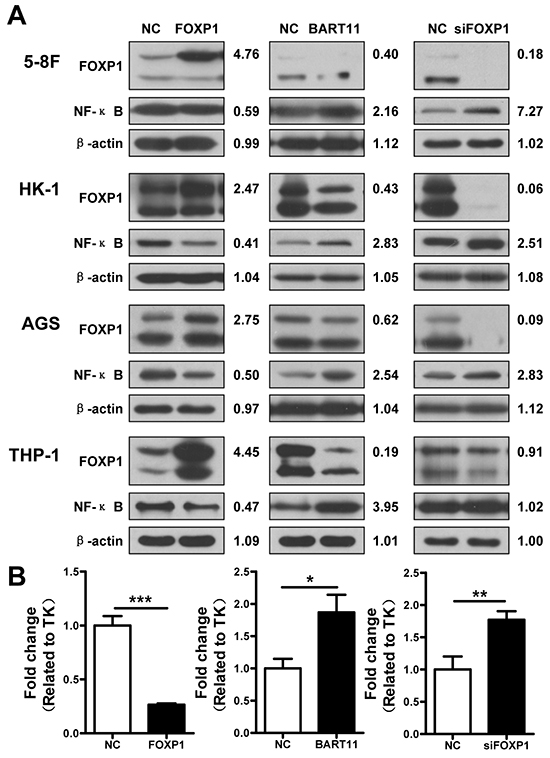
Figure 7: EBV-miR-BART11-mediated NF-κB activation via FOXP1 inhibition. A. The expression of FOXP1 and NF-κB in 5-8F, HK-1, and AGS epithelial cancer cells and THP-1 monocytes, transfected with FOXP1, EBV-miR-BART11, or siFOXP1, and determined by western blotting and densitometry. B. NF-κB transcriptional activity, assessed by luciferase reporter assay in 5-8F cells transfected with the FOXP1 overexpression vector, EBV-miR-BART11 precursor vector, or siFOXP1. Data shown are representative images or expressed as mean ± SD of different groups of cells from three independent experiments. *p < 0.05; **p < 0.01; ***p < 0.001.
DISCUSSION
Chronic, non-resolving inflammation plays an important role in the initiation of a variety of tumors, including NPC [38, 39] and GC [40-42]. While various immune cells are involved in chronic inflammation, TAMs represent the key mediators of chronic inflammation within the tumor microenvironment [37]. We previously found that TAM infiltration promotes NPC cell proliferation by the secretion of inflammatory factors, such as IL-6 [5, 9], and induce NF-κB and STAT3 transcription factor activation [39, 43], ultimately resulting in poor disease prognosis.
EBV-infected macrophages induce an inflammatory response in EBV-related human tumors [44-46], but the relationships between EBV-encoded miRNAs and TAMs in EBV-associated solid tumors have not been reported. In our previous study [2], we constructed an EBV miRNA-dependent gene regulatory network and predicted that the EBV-encoded miRNAs, BART11-3p and BART11-5p, may regulate FOXP1 expression [2]. FOXP1 belongs to the FOX transcription factor family, which includes a variety of ‘winged helix’ transcription factors that play crucial roles in immune homeostasis [47]. Several members of the FOX family, for example FOXF1, FOXP3, FOXN1, FOXO1, and FOXO3, were shown to execute diverse functions during the regulation of inflammation and adaptive immune response [48-51]. The FOXP1 locus is located on 3p14.1 [47], a region known for its high-frequency allele imbalance [52-54], and it is a defined genetic susceptibility region [55, 56] in NPC. Generally, FOXP1 acts as a transcriptional repressor that regulates the differentiation of B [57] and T lymphocytes [58], monocytes [34, 59], and other immune cells. Therefore, we speculated that EBV may alter immune cell differentiation and the inflammatory process, in order to promote inflammation-induced carcinogenesis in EBV-associated cancers by inhibiting FOXP1 expression.
We confirmed FOXP1 as a target of EBV-miR-BART11 in both monocytes and epithelial cells, and showed that FOXP1 downregulation was necessary for monocyte differentiation. We further investigated the effects of EBV-miR-BART11 and FOXP1 on macrophage cytokine secretion. EBV-miR-BART11 overexpression in TAMs resulted in LPS hyperresponsiveness and the increased secretion of cytokines, chemokines, and growth factors that support tumor survival, proliferation, and invasion. Taken together, our results indicate that EBV-miR-BART11 is sufficient to maintain and potentiate the inflammatory environment by enhancing macrophage differentiation and responsiveness through the downregulation of FOXP1.
Under certain conditions, tumor cells can induce their own inflammatory factor secretion, in order to intensify the local inflammatory microenvironment, resulting in further TAM and lymphocyte infiltration [60]. Therefore, we investigated the effects of EBV-miR-BART11 and FOXP1 on the local inflammatory response in epithelial cells. Our results showed that LPS stimulation significantly enhances cytokine secretion in both macrophages and epithelial cells, and that the conditioned media collected from LPS-treated macrophages is able to stimulate epithelial cells to produce an even higher level of cytokines.
FOXP1 acts as a transcriptional repressor in lymphocytes where its downregulation plays important roles in immune cell differentiation. It is also frequently downregulated in solid tumors, including breast cancer [61-63], non-small cell lung cancer [64], oral squamous cell carcinoma [65], ovarian cancer [66, 67], renal cell carcinoma [68, 69], hepatocellular carcinoma [70] and prostate cancer [71-73], which suggests that FOXP1 may act as a tumor suppressor. However, its role in EBV-associated solid tumors, such as NPC and GC, is yet to be completely understood. Our results demonstrate that tumor-suppressive effects of FOXP1 inhibit epithelial cell proliferation, which can be counteracted with EBV-miR-BART11 expression.
NF-κB transcriptional regulation plays a critical role in tumorigenic inflammation. FOXP1 and NF-κB interact through a complex regulatory mechanism involved in several biological processes that vary in a cell type-dependent manner. In lymphocytes, FOXP1 is able to inhibit directly apoptotic gene transcription and cooperate with NF-κB, promoting human B cell survival, while FOXP1 expression is dependent on NF-κB transcriptional activity and plays a synergistic role in NF-κB self-activation [74]. NF-κB activation in malignant B cells can increase FOXP1 expression [75], but some results indicate that full-length FOXP1 may function as a NF-κB transcriptional repressor in follicular lymphoma [76, 77]. Conversely, NF-κB has also been shown to downregulate FOXP1 in monocyte differentiation and macrophage function [34], and therefore, downregulation of the NF-κB signaling pathway is an effective mechanism of FOXP1-mediated inhibition of monocyte differentiation and macrophage function. The regulatory relationships between EBV-miR-BART11, FOXP1, NF-κB, and their downstream signaling pathways, and their detailed biological functions, require further investigations.
In conclusion, we revealed EBV-miR-BART11 plays an important role in the inflammatory microenvironment and inflammation-induced carcinogenesis in EBV-associated cancers through the direct inhibition of FOXP1 and NF-κB activation. Collectively, these data demonstrate that EBV-miR-BART11 or FOXP1 may serve as potential diagnostic or prognostic markers in NPC or GC, and may represent important targets for EBV-related cancer immunotherapy.
MATERIALS AND METHODS
Cell lines and reagents
Two EBV-negative human NPC cell lines (5-8F and HK-1), AGS GC cells, and THP-1 monocytes were grown in RPMI-1640 medium supplemented with 10% fetal calf serum (FCS) at 37°C in the atmosphere with 5% CO2. THP-1 cells were treated with 10 nM phorbol 12-myristate 13-acetate (PMA, Sigma, St Louis, MO, USA) to induce monocyte differentiation. Lipopolysaccharide (LPS) from Escherichia coli 0111:B4 was obtained from Sigma (10 ng/mL). 293T cells used to produce the lentiviral stock were grown in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% FCS, at 37°C in the atmosphere containing 5% CO2.
siRNAs, EBV-miR-BART11 and FOXP1 overexpression vectors, and cell transfection
FOXP1-specific siRNAs and their corresponding controls were synthesized by Ruibo (Guangzhou, China) (Supplementary Table S1). EBV-miR-BART11 precursor sequence was synthesized (Invitrogen, Shanghai, China) and cloned into the pSUPER.neo/GFP vector (Invitrogen, Shanghai, China). The FOXP1 full-length coding sequence (CDS, NM_032682) was amplified and cloned into the pIRESneo3 vector (Invitrogen). Cell transfection was performed in 70-80% confluent cells using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s protocol.
Lentiviral vectors and THP-1 cell infection
Lentiviral vector construction was constructed using the ViraPower™ Lentiviral Expression System (Invitrogen). First, EBV-miR-BART11 precursor or full-length FOXP1 CDS was subcloned into the pLenti6/V5-D-TOPO vector. pLenti6/V5-D-TOPO/BART11 or pLenti6/V5-D-TOPO/FOXP1 vectors and ViraPower Packaging Mix were co-transfected using a gene carrier kit (Epoch-Biolabs, Missouri City, TX, USA) into 293T cells to produce a lentiviral stock, and 48 h after transfection, virus-containing supernatant was collected and used to infect THP-1 at a ratio of 1:1 with fresh medium, as described previously [78].
Quantitative real time polymerase chain reaction (qRT-PCR)
Total RNA was harvested using the TRIzol Extraction Kit (Invitrogen) and cDNA samples were prepared using a QuantiTect Reverse Transcription Kit (Qiagen, Hilden, Germany). Stem-loop real-time qRT-PCR for mature miRNAs was done with the Qiagen QuantiTect SYBR Green PCR Kits (Qiagen), using RNU6B (U6) as an international control. qRT-PCR for mRNA expression was performed using the SYBR Premix Ex Taq II kit (Takara, Dalian, China) according to the manufacturer instructions with β-actin or GAPDH as the internal controls. The primers used are shown in Supplementary Table S1.
Western blotting
Whole cell lysates were extracted using RIPA lysis buffer (50 mM Tris-HCl pH 7.4, 250 mM NaCl, 0.1% SDS, 0.5% NP-40, 2 mM DTT, 1 mg/mL protease inhibitors). Protein concentration was quantified using a BCA protein assay kit (Pierce, Grand Island, NY). Afterward, 50 μg of protein was resolved by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and electroblotted onto a PVDF membrane (Millipore, Billerica, MA, USA). Immunodetection using antibodies against FOXP1 (Cell Signaling, Danvers, MA), NF-κB p65 (Abcam, MA, USA), or β-actin (Millipore) was done for 1 h at room temperature or overnight at 4°C. Antibody-antigen complexes were detected using the enhanced chemiluminescence (ECL) system (Amersham Bioscience, Piscataway, NJ, USA). β-actin served as a loading control.
Methylthiazol tetrazolium assay (MTT) and colony formation assay
Cell proliferation was measured using an MTT (Millipore) assay. After transfection or treatment, 800 cells per well were seeded in 96-well plates, and viability was assessed in five replicates at designed times. The formation of colored formazan dye was measured at 490 nm on a spectrophotometric plate reader.
For the clonogenic assays, 1000 cells were plated in six-well plates at 24 h post-transfection and incubated for 14 days. The number of colonies was counted with Image-Pro Plus (Syngene, Frederick, MD, USA) and normalized to the control group. The experiments were performed at least three times.
Flow cytometry
Cell cycle analysis was performed using flow cytometry with propidium iodide (PI) staining (5 μg/mL) 48 h post-transfection. Events were recorded from samples using a FACSCalibur Flow Cytometer (BD Biosciences, New Jersey, USA) equipped with Cell Quest 3.3 software and analyzed using ModFit software (Verity Software House, Topsham, ME).
Luciferase reporter assay
5-8F cells were co-transfected with synthetic EBV-miR-BART11 and luciferase reporter vectors (FOXP1-WT or FOXP1-mutant) along with pRL-TK Renilla luciferase vector (Promega, Madison, WI). For NF-κB reporter assay, 5-8F cells were transiently co-transfected with the pNF-κB-Luc construct (Promega), pRL-TK vector, and EBV-miR-BART11 or the FOXP1 expression vector or siFOXP1. Luciferase activity was determined by the Dual Luciferase Assay Kit (Promega) 48 h after transfection [79-81]. Transfections were performed in duplicates and repeated in three independent experiments.
Statistical analysis
Statistical analyses were performed using SPSS 13.0 (SPSS, Chicago, IL) and Graph Pad Prism 5 (GraphPad, La Jolla, CA). Data are presented as mean ± standard deviation (SD). Comparisons between two groups were performed using Student’s t-test or one-way ANOVA, unless otherwise indicated. The association between EBV-miR-BART11-(3p and 5p) and FOXP1 gene expression was analyzed using Spearman’s correlation coefficient. p < 0.05 was considered statistically significant.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest in this work.
GRANT SUPPORT
This study was supported in part by grants from the National Natural Science Foundation of China (81272298, 81372907, 81472531 and 81572787), the Natural Science Foundation of Hunan Province (14JJ1010 and 2015JJ1022), the Free Exploration Program of Central South University of China (2013zzts282), and the Hunan Provincial Innovation Foundation for Postgraduate (CX2014B125).
REFERENCES
1. Xiong W, Zeng ZY, Xia JH, Xia K, Shen SR, Li XL, Hu DX, Tan C, Xiang JJ, Zhou J, Deng H, Fan SQ, Li WF, et al. A susceptibility locus at chromosome 3p21 linked to familial nasopharyngeal carcinoma. Cancer Res. 2004; 64:1972-1974.
2. Zeng Z, Huang H, Zhang W, Xiang B, Zhou M, Zhou Y, Ma J, Yi M, Li X, Xiong W and Li G. Nasopharyngeal carcinoma: advances in genomics and molecular genetics. Sci China Life Sci. 2011; 54:966-975.
3. Zhou HD, Li GY, Yang YX, Li XL, Sheng SR, Zhang WL and Zhao J. Intracellular co-localization of SPLUNC1 protein with nanobacteria in nasopharyngeal carcinoma epithelia HNE1 cells depended on the bactericidal permeability increasing protein domain. Mol Immunol. 2006; 43:1864-1871.
4. Song X, Chen HX, Wang XY, Deng XY, Xi YX, He Q, Peng TL, Chen J, Chen W, Wong BC and Chen MH. H. pylori-encoded CagA disrupts tight junctions and induces invasiveness of AGS gastric carcinoma cells via Cdx2-dependent targeting of Claudin-2. Cell Immunol. 2013; 286:22-30.
5. Liao Q, Zeng Z, Guo X, Li X, Wei F, Zhang W, Chen P, Liang F, Xiang B, Ma J, Wu M, Tang H, Deng M, et al. LPLUNC1 suppresses IL-6-induced nasopharyngeal carcinoma cell proliferation via inhibiting the Stat3 activation. Oncogene. 2014; 33:2098-2109.
6. Zhou Y, Liao Q, Li X, Wang H, Wei F, Chen J, Yang J, Zeng Z, Guo X, Chen P, Zhang W, Tang K, Li X, Xiong W and Li G. HYOU1, Regulated by LPLUNC1, Is Up-Regulated in Nasopharyngeal Carcinoma and Associated with Poor Prognosis. J Cancer. 2016; 7:367-376.
7. Zheng H, Li M, Ren W, Zeng L, Liu HD, Hu D, Deng X, Tang M, Shi Y, Gong J and Cao Y. Expression and secretion of immunoglobulin alpha heavy chain with diverse VDJ recombinations by human epithelial cancer cells. Mol Immunol. 2007; 44:2221-2227.
8. Duan Z, Zheng H, Xu S, Jiang Y, Liu H, Li M, Hu D, Li W, Bode AM, Dong Z and Cao Y. Activation of the Ig Ialpha1 promoter by the transcription factor Ets-1 triggers Ig Ialpha1-Calpha1 germline transcription in epithelial cancer cells. Cell Mol Immunol. 2014; 11:197-205.
9. Liao Q, Guo X, Li X, Chen P, Liang F, Tang H, Deng M, Wu M, Ma J, Xiong W and Li G. Analysis of the contribution of nasopharyngeal epithelial cancer cells to the induction of a local inflammatory response. J Cancer Res Clin Oncol. 2012; 138:57-64.
10. Yang Y, Zhou H, Li W, Zhou M, Zeng Z, Xiong W, Wu M, Huang H, Zhou Y, Peng C, Huang C, Li X and Li G. Lipopolysaccharide (LPS) regulates TLR4 signal transduction in nasopharynx epithelial cell line 5-8F via NFkappaB and MAPKs signaling pathways. Mol Immunol. 2007; 44:984-992.
11. Zheng H, Li LL, Hu DS, Deng XY and Cao Y. Role of Epstein-Barr virus encoded latent membrane protein 1 in the carcinogenesis of nasopharyngeal carcinoma. Cell Mol Immunol. 2007; 4:185-196.
12. Zeng Z, Fan S, Zhang X, Li S, Zhou M, Xiong W, Tan M, Zhang W and Li G. Epstein-Barr virus-encoded small RNA 1 (EBER-1) could predict good prognosis in nasopharyngeal carcinoma. Clin Transl Oncol. 2016; 18:206-211.
13. Xiao K, Yu Z, Li X, Li X, Tang K, Tu C, Qi P, Liao Q, Chen P, Zeng Z, Li G and Xiong W. Genome-wide Analysis of Epstein-Barr Virus (EBV) Integration and Strain in C666-1 and Raji Cells. J Cancer. 2016; 7:214-224.
14. Barth S, Meister G and Grasser FA. EBV-encoded miRNAs. Biochim Biophys Acta. 2011; 1809:631-640.
15. Chen SJ, Chen GH, Chen YH, Liu CY, Chang KP, Chang YS and Chen HC. Characterization of Epstein-Barr virus miRNAome in nasopharyngeal carcinoma by deep sequencing. PLoS One. 2010; 5.
16. Pfeffer S, Zavolan M, Grasser FA, Chien M, Russo JJ, Ju J, John B, Enright AJ, Marks D, Sander C and Tuschl T. Identification of virus-encoded microRNAs. Science. 2004; 304:734-736.
17. Ramakrishnan R, Donahue H, Garcia D, Tan J, Shimizu N, Rice AP and Ling PD. Epstein-Barr virus BART9 miRNA modulates LMP1 levels and affects growth rate of nasal NK T cell lymphomas. PLoS One. 2011; 6:e27271.
18. Kim do N, Chae HS, Oh ST, Kang JH, Park CH, Park WS, Takada K, Lee JM, Lee WK and Lee SK. Expression of viral microRNAs in Epstein-Barr virus-associated gastric carcinoma. J Virol. 2007; 81:1033-1036.
19. Xie YJ, Long ZF and He XS. Involvement of EBV-encoded BART-miRNAs and dysregulated cellular miRNAs in nasopharyngeal carcinoma genesis. Asian Pac J Cancer Prev. 2013; 14:5637-5644.
20. Yang HJ, Huang TJ, Yang CF, Peng LX, Liu RY, Yang GD, Chu QQ, Huang JL, Liu N, Huang HB, Zhu ZY, Qian CN and Huang BJ. Comprehensive profiling of Epstein-Barr virus-encoded miRNA species associated with specific latency types in tumor cells. Virol J. 2013; 10:314.
21. Yan Q, Zeng Z, Gong Z, Zhang W, Li X, He B, Song Y, Li Q, Zeng Y, Liao Q, Chen P, Shi L, Fan S, Xiang B, Ma J, Zhou M, et al. EBV-miR-BART10-3p facilitates epithelial-mesenchymal transition and promotes metastasis of nasopharyngeal carcinoma by targeting BTRC. Oncotarget. 2015; 6:41766-41782. doi: 10.18632/oncotarget.6155.
22. Zeng Z, Huang H, Huang L, Sun M, Yan Q, Song Y, Wei F, Bo H, Gong Z, Zeng Y, Li Q, Zhang W, Li X, et al. Regulation network and expression profiles of Epstein-Barr virus-encoded microRNAs and their potential target host genes in nasopharyngeal carcinomas. Sci China Life Sci. 2014; 57:315-326.
23. Barth S, Pfuhl T, Mamiani A, Ehses C, Roemer K, Kremmer E, Jaker C, Hock J, Meister G and Grasser FA. Epstein-Barr virus-encoded microRNA miR-BART2 down-regulates the viral DNA polymerase BALF5. Nucleic Acids Res. 2008; 36:666-675.
24. Lo AK, To KF, Lo KW, Lung RW, Hui JW, Liao G and Hayward SD. Modulation of LMP1 protein expression by EBV-encoded microRNAs. Proc Natl Acad Sci U S A. 2007; 104:16164-16169.
25. Lung RW, Tong JH, Sung YM, Leung PS, Ng DC, Chau SL, Chan AW, Ng EK, Lo KW and To KF. Modulation of LMP2A expression by a newly identified Epstein-Barr virus-encoded microRNA miR-BART22. Neoplasia. 2009; 11:1174-1184.
26. Nachmani D, Stern-Ginossar N, Sarid R and Mandelboim O. Diverse herpesvirus microRNAs target the stress-induced immune ligand MICB to escape recognition by natural killer cells. Cell Host Microbe. 2009; 5:376-385.
27. Cai LM, Lyu XM, Luo WR, Cui XF, Ye YF, Yuan CC, Peng QX, Wu DH, Liu TF, Wang E, Marincola FM, Yao KT, Fang WY, Cai HB and Li X. EBV-miR-BART7-3p promotes the EMT and metastasis of nasopharyngeal carcinoma cells by suppressing the tumor suppressor PTEN. Oncogene. 2014; 34:2156-2166.
28. Choi H, Lee H, Kim SR, Gho YS and Lee SK. Epstein-Barr virus-encoded microRNA BART15-3p promotes cell apoptosis partially by targeting BRUCE. J Virol. 2013; 87:8135-8144.
29. Choy EY, Siu KL, Kok KH, Lung RW, Tsang CM, To KF, Kwong DL, Tsao SW and Jin DY. An Epstein-Barr virus-encoded microRNA targets PUMA to promote host cell survival. J Exp Med. 2008; 205:2551-2560.
30. Ye Y, Zhou Y, Zhang L, Chen Y, Lyu X, Cai L, Lu Y, Deng Y, Wang J, Yao K, Fang W, Cai H and Li X. EBV-miR-BART1 is involved in regulating metabolism-associated genes in nasopharyngeal carcinoma. Biochem Biophys Res Commun. 2013; 436:19-24.
31. Zeng ZY, Zhou YH, Zhang WL, Xiong W, Fan SQ, Li XL, Luo XM, Wu MH, Yang YX, Huang C, Cao L, Tang K, Qian J, Shen SR and Li GY. Gene expression profiling of nasopharyngeal carcinoma reveals the abnormally regulated Wnt signaling pathway. Hum Pathol. 2007; 38:120-133.
32. Zeng Z, Zhou Y, Xiong W, Luo X, Zhang W, Li X, Fan S, Cao L, Tang K, Wu M and Li G. Analysis of gene expression identifies candidate molecular markers in nasopharyngeal carcinoma using microdissection and cDNA microarray. J Cancer Res Clin Oncol. 2007; 133:71-81.
33. Zhang W, Zeng Z, Zhou Y, Xiong W, Fan S, Xiao L, Huang D, Li Z, Li D, Wu M, Li X, Shen S, Wang R, Cao L, Tang K and Li G. Identification of aberrant cell cycle regulation in Epstein-Barr virus-associated nasopharyngeal carcinoma by cDNA microarray and gene set enrichment analysis. Acta Biochim Biophys Sin (Shanghai). 2009; 41:414-428.
34. Shi C, Sakuma M, Mooroka T, Liscoe A, Gao H, Croce KJ, Sharma A, Kaplan D, Greaves DR, Wang Y and Simon DI. Down-regulation of the forkhead transcription factor Foxp1 is required for monocyte differentiation and macrophage function. Blood. 2008; 112:4699-4711.
35. Shi C, Zhang X, Chen Z, Sulaiman K, Feinberg MW, Ballantyne CM, Jain MK and Simon DI. Integrin engagement regulates monocyte differentiation through the forkhead transcription factor Foxp1. J Clin Invest. 2004; 114:408-418.
36. Hanahan D and Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646-674.
37. Sica A, Larghi P, Mancino A, Rubino L, Porta C, Totaro MG, Rimoldi M, Biswas SK, Allavena P and Mantovani A. Macrophage polarization in tumour progression. Semin Cancer Biol. 2008; 18:349-355.
38. Shimakage M and Sakamoto H. Macrophage involvement in Epstein-Barr virus-related tumors. Exp Ther Med. 2010; 1:285-291.
39. Coussens LM and Werb Z. Inflammation and cancer. Nature. 2002; 420:860-867.
40. Ding SZ and Zheng PY. Helicobacter pylori infection induced gastric cancer; advance in gastric stem cell research and the remaining challenges. Gut pathogens. 2012; 4:18.
41. Howlader N, Ries LA, Mariotto AB, Reichman ME, Ruhl J and Cronin KA. Improved estimates of cancer-specific survival rates from population-based data. Journal of the National Cancer Institute. 2010; 102:1584-1598.
42. Peek RM, Jr. and Crabtree JE. Helicobacter infection and gastric neoplasia. The Journal of pathology. 2006; 208:233-248.
43. Oshima H and Oshima M. The inflammatory network in the gastrointestinal tumor microenvironment: lessons from mouse models. J Gastroenterol. 2012; 47:97-106.
44. Lamouille S, Xu J and Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014; 15:178-196.
45. Mantovani A, Allavena P, Sica A and Balkwill F. Cancer-related inflammation. Nature. 2008; 454:436-444.
46. Talbot LJ, Bhattacharya SD and Kuo PC. Epithelial-mesenchymal transition, the tumor microenvironment, and metastatic behavior of epithelial malignancies. Int J Biochem Mol Biol. 2012; 3:117-136.
47. Banham AH, Beasley N, Campo E, Fernandez PL, Fidler C, Gatter K, Jones M, Mason DY, Prime JE, Trougouboff P, Wood K and Cordell JL. The FOXP1 winged helix transcription factor is a novel candidate tumor suppressor gene on chromosome 3p. Cancer Res. 2001; 61:8820-8829.
48. Birkenkamp KU and Coffer PJ. FOXO transcription factors as regulators of immune homeostasis: molecules to die for? J Immunol. 2003; 171:1623-1629.
49. Jonsson H and Peng SL. Forkhead transcription factors in immunology. Cellular and molecular life sciences. 2005; 62:397-409.
50. Peng SL. Foxo in the immune system. Oncogene. 2008; 27:2337-2344.
51. Yang XF. Factors regulating apoptosis and homeostasis of CD4+ CD25(high) FOXP3+ regulatory T cells are new therapeutic targets. Frontiers in bioscience. 2008; 13:1472-1499.
52. Deng L, Jing N, Tan G, Zhou M, Zhan F, Xie Y, Cao L and Li G. A common region of allelic loss on chromosome region 3p25.3-26.3 in nasopharyngeal carcinoma. Genes Chromosomes Cancer. 1998; 23:21-25.
53. Lo KW, Teo PM, Hui AB, To KF, Tsang YS, Chan SY, Mak KF, Lee JC and Huang DP. High resolution allelotype of microdissected primary nasopharyngeal carcinoma. Cancer Res. 2000; 60:3348-3353.
54. Shao JY, Wang HY, Huang XM, Feng QS, Huang P, Feng BJ, Huang LX, Yu XJ, Li JT, Hu LF, Ernberg I and Zeng YX. Genome-wide allelotype analysis of sporadic primary nasopharyngeal carcinoma from southern China. Int J Oncol. 2000; 17:1267-1275.
55. Ng CC, Yew PY, Puah SM, Krishnan G, Yap LF, Teo SH, Lim PV, Govindaraju S, Ratnavelu K, Sam CK, Takahashi A, Kubo M, Kamatani N, Nakamura Y and Mushiroda T. A genome-wide association study identifies ITGA9 conferring risk of nasopharyngeal carcinoma. J Hum Genet. 2009; 54:392-397.
56. Zeng Z, Zhou Y, Zhang W, Li X, Xiong W, Liu H, Fan S, Qian J, Wang L, Li Z, Shen S and Li G. Family-based association analysis validates chromosome 3p21 as a putative nasopharyngeal carcinoma susceptibility locus. Genet Med. 2006; 8:156-160.
57. Hu H, Wang B, Borde M, Nardone J, Maika S, Allred L, Tucker PW and Rao A. Foxp1 is an essential transcriptional regulator of B cell development. Nat Immunol. 2006; 7:819-826.
58. Feng X, Wang H, Takata H, Day TJ, Willen J and Hu H. Transcription factor Foxp1 exerts essential cell-intrinsic regulation of the quiescence of naive T cells. Nat Immunol. 2011; 12:544-550.
59. Xue ZH, Zhao CQ, Chua GL, Tan SW, Tang XY, Wong SC and Tan SM. Integrin alphaMbeta2 clustering triggers phosphorylation and activation of protein kinase C delta that regulates transcription factor Foxp1 expression in monocytes. J Immunol. 2010; 184:3697-3709.
60. Yang Y, Zhou H, Yang Y, Li W, Zhou M, Zeng Z, Xiong W, Wu M, Huang H, Zhou Y, Peng C, Huang C, Li X and Li G. Lipopolysaccharide (LPS) regulates TLR4 signal transduction in nasopharynx epithelial cell line 5-8F via NFkappaB and MAPKs signaling pathways. Mol Immunol. 2007; 44:984-992.
61. Ijichi N, Ikeda K, Horie-Inoue K and Inoue S. FOXP1 and estrogen signaling in breast cancer. Vitam Horm. 2013; 93:203-212.
62. Ijichi N, Shigekawa T, Ikeda K, Horie-Inoue K, Shimizu C, Saji S, Aogi K, Tsuda H, Osaki A, Saeki T and Inoue S. Association of double-positive FOXA1 and FOXP1 immunoreactivities with favorable prognosis of tamoxifen-treated breast cancer patients. Horm Cancer. 2012; 3:147-159.
63. Shigekawa T, Ijichi N, Ikeda K, Horie-Inoue K, Shimizu C, Saji S, Aogi K, Tsuda H, Osaki A, Saeki T and Inoue S. FOXP1, an estrogen-inducible transcription factor, modulates cell proliferation in breast cancer cells and 5-year recurrence-free survival of patients with tamoxifen-treated breast cancer. Horm Cancer. 2011; 2:286-297.
64. Feng J, Zhang X, Zhu H, Wang X, Ni S and Huang J. High expression of FoxP1 is associated with improved survival in patients with non-small cell lung cancer. Am J Clin Pathol. 2012; 138:230-235.
65. Yang MH, Lin BR, Chang CH, Chen ST, Lin SK, Kuo MY, Jeng YM, Kuo ML and Chang CC. Connective tissue growth factor modulates oral squamous cell carcinoma invasion by activating a miR-504/FOXP1 signalling. Oncogene. 2011; 31:2401-2411.
66. Hu Z, Zhu L, Gao J, Cai M, Tan M, Liu J and Lin B. Expression of FOXP1 in epithelial ovarian cancer (EOC) and its correlation with chemotherapy resistance and prognosis. Tumour Biol. 2015; 36:7269-7275.
67. Hu Z, Zhu L, Tan M, Cai M, Deng L, Yu G, Liu D, Liu J and Lin B. The expression and correlation between the transcription factor FOXP1 and estrogen receptors in epithelial ovarian cancer. Biochimie. 2015; 109:42-48.
68. Toma MI, Weber T, Meinhardt M, Zastrow S, Grimm MO, Fussel S, Wirth MP and Baretton GB. Expression of the Forkhead transcription factor FOXP1 is associated with tumor grade and Ki67 expression in clear cell renal cell carcinoma. Cancer Invest. 2011; 29:123-129.
69. Wan L, Huang J, Chen J, Wang R, Dong C, Lu S and Wu X. Expression and significance of FOXP1, HIF-1a and VEGF in renal clear cell carcinoma. J BUON. 2015; 20:188-195.
70. Zhang Y, Zhang S, Wang X, Liu J, Yang L, He S, Chen L and Huang J. Prognostic significance of FOXP1 as an oncogene in hepatocellular carcinoma. J Clin Pathol. 2012; 65:528-533.
71. Banham AH, Boddy J, Launchbury R, Han C, Turley H, Malone PR, Harris AL and Fox SB. Expression of the forkhead transcription factor FOXP1 is associated both with hypoxia inducible factors (HIFs) and the androgen receptor in prostate cancer but is not directly regulated by androgens or hypoxia. Prostate. 2007; 67:1091-1098.
72. Takayama K, Horie-Inoue K, Ikeda K, Urano T, Murakami K, Hayashizaki Y, Ouchi Y and Inoue S. FOXP1 is an androgen-responsive transcription factor that negatively regulates androgen receptor signaling in prostate cancer cells. Biochem Biophys Res Commun. 2008; 374:388-393.
73. Takayama K, Suzuki T, Tsutsumi S, Fujimura T, Takahashi S, Homma Y, Urano T, Aburatani H and Inoue S. Integrative analysis of FOXP1 function reveals a tumor-suppressive effect in prostate cancer. Mol Endocrinol. 2014; 28:2012-2024.
74. van Keimpema M, Gruneberg LJ, Mokry M, van Boxtel R, Koster J, Coffer PJ, Pals ST and Spaargaren M. FOXP1 directly represses transcription of proapoptotic genes and cooperates with NF-kappaB to promote survival of human B cells. Blood. 2014; 124:3431-3440.
75. Mraz M, Chen L, Rassenti LZ, Ghia EM, Li H, Jepsen K, Smith EN, Messer K, Frazer KA and Kipps TJ. miR-150 influences B-cell receptor signaling in chronic lymphocytic leukemia by regulating expression of GAB1 and FOXP1. Blood. 2014; 124:84-95.
76. Brown PJ, Ashe SL, Leich E, Burek C, Barrans S, Fenton JA, Jack AS, Pulford K, Rosenwald A and Banham AH. Potentially oncogenic B-cell activation-induced smaller isoforms of FOXP1 are highly expressed in the activated B cell-like subtype of DLBCL. Blood. 2008; 111:2816-2824.
77. Wang B, Lin D, Li C and Tucker P. Multiple domains define the expression and regulatory properties of Foxp1 forkhead transcriptional repressors. J Biol Chem. 2003; 278:24259-24268.
78. Zhang W, Zeng Z, Fan S, Wang J, Yang J, Zhou Y, Li X, Huang D, Liang F, Wu M, Tang K, Cao L, Li X, Xiong W and Li G. Evaluation of the prognostic value of TGF-beta superfamily type I receptor and TGF-beta type II receptor expression in nasopharyngeal carcinoma using high-throughput tissue microarrays. J Mol Histol. 2012; 43:297-306.
79. Xu K, Xiong W, Zhou M, Wang H, Yang J, Li X, Chen P, Liao Q, Deng H, Li X, Li G and Zeng Z. Integrating ChIP-sequencing and digital gene expression profiling to identify BRD7 downstream genes and construct their regulating network. Mol Cell Biochem. 2016; 411:57-71.
80. Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F, Shi L, Yang J, Zhang W, Zhou Y, Zeng Y, Li X, Xiang B, et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. PLoS One. 2014; 9:e110674.
81. Bo H, Gong Z, Zhang W, Li X, Zeng Y, Liao Q, Chen P, Shi L, Lian Y, Jing Y, Tang K, Li Z, Zhou Y, et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 2015; 6:20404-20418. doi: 10.18632/oncotarget.4057.

