INTRODUCTION
Non-small cell lung cancer (NSCLC) has a dismal prognosis with one of the highest mortality rates among cancer types [1, 2]. Patients with NSCLC harboring epidermal growth factor receptor (EGFR) mutation respond to treatment of EGFR tyrosine kinase inhibitors (TKIs) [3, 4]. Large randomized phase III trials comparing EGFR TKIs such as gefitinib, erlotinib or afatinib with cytotoxic chemotherapy consistently demonstrated higher response rates and prolonged progression-free survival (PFS) in EGFR mutant NSCLC patients, leading to the standard treatment of EGFR TKI as first line therapy [5-8]. However, most patients with NSCLC treated with EGFR TKIs eventually develop acquired resistance [9, 10]. Although multiple mechanisms are involved, EGFR T790M mutation accounts for more than 50% of the acquired TKI resistance [11, 12]. Detection of resistance requires acquisition and analysis of tumor tissue at the time of resistance to understand the underlying mechanism. However, the invasive nature of repeated biopsies makes it difficult to obtain from patients particularly those with poor performance. Other limitations include tumor heterogeneity [13], inaccessibility of re-biopsy due to tumor location or tumor containing blood vessel or air bronchogram, high incidence of tumor necrosis, and a single snapshot in time unable to track dynamic changes in the mutation.
The circulating tumor DNA in plasma with its non-invasive technology has been used as a surrogate for tumor tissues in detecting genetic alterations, thus providing molecular evolution of the tumors [14-17]. Bai et al. showed the difference in response rates (59.5% and 23.1%, P = 0.002) for gefitinib between patients with and without activating EGFR mutation in plasma [18]. Recently, droplet digital polymerase chain reaction (ddPCR), a highly sensitive technology has been developed for detecting low frequency cancer-associated mutations [19]. We hypothesized that plasma-based EGFR mutation analysis using ddPCR may be feasible in monitoring response and resistance to EGFR TKIs. Therefore, we conducted a multi-center prospective study to assess dynamic changes in EGFR mutation profile using a ddPCR method in longitudinally collected plasma samples from NSCLC patients harboring activating EGFR mutations treated with EGFR TKIs.
RESULTS
Patient characteristics
From January 2012 to October 2014, 81 EGFR mutant NSCLC patients who were treated with gefitinib or erlotinib and eventually developed acquired resistance were enrolled. The median age was 58.2 years (range, 32.1-81.1 years), 61.7% were female, and 63.0% were never smokers. Over four-fifths of patients (84.0%) had stage IV disease, 59.3% had ex19del and 59.3% were treated with EGFR TKIs as first-line therapy (Table 1).
Table 1: Baseline characteristics (n = 81) |
||
No. of patients |
% |
|
Age, years |
||
Median |
58.2 |
|
Range |
32.1-81.1 |
|
Sex |
||
Male |
31 |
38.3 |
Female |
50 |
61.7 |
Smoking history |
||
Never smoker |
51 |
63.0 |
Former smoker |
18 |
22.2 |
Current smoker |
12 |
14.8 |
Tumor type |
||
Adenocarcinoma |
80 |
98.8 |
Adenosquamous |
1 |
1.2 |
Tumor stage |
||
IV |
68 |
84.0 |
Postoperative relapse |
13 |
16.0 |
Type of EGFR mutation |
||
Exon 19 deletion |
48 |
59.3 |
L858R |
33 |
40.7 |
Line of EGFR TKIs |
||
First line |
48 |
59.3 |
Second line |
33 |
40.7 |
EGFR TKIs |
||
Gefitinib |
58 |
71.6 |
Erlotinib |
23 |
28.4 |
Abbreviations: EGFR, epidermal growth factor receptor; TKIs, kinase inhibitors
Clinical outcomes
Of total 81 patients, 59 (72.8%) patients achieved an objective response. With a median follow-up of 18.8 months, the median PFS and overall survival (OS) were 8.1 months (95% CI, 6.2-10.0) and 23.5 months (95% CI, 19.4-27.6), respectively.
The technical sensitivity and specificity of L858R and ex19del assays in the cfDNA
The sensitivity and specificity of the ddPCR assays in measuring EGFR mutations in cfDNA were determined from 367 plasma samples collected from baseline to every 8 weeks after EGFR TKI treatment from 81 patients. We applied two QC criteria for the results of these plasma samples: 1) the droplet number must be greater than 9000; 2) the wild type (WT) levels to be greater than 100 copies/mL. After QC, we excluded two patients and 6 time points, resulting in 361 samples from 79 patients (Figure 1). Among the qualified samples, the lowest level of WT cfDNA was 1040 copies/mL, and the median was 4568 copies/mL (data not shown), suggesting that good quality of cfDNA is obtained with the current sample preparation procedures.
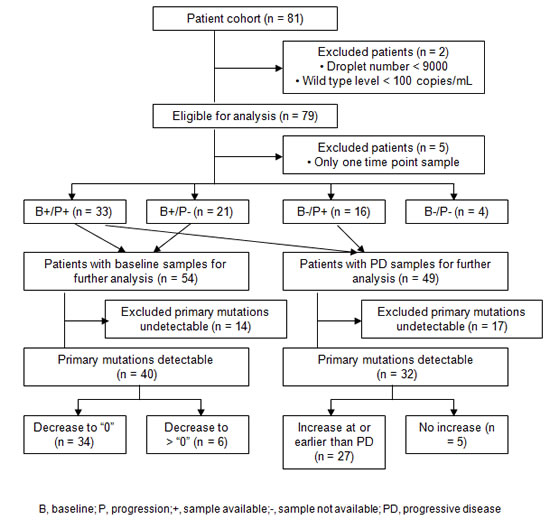
Figure 1: Patients exclusion in the analysis based on the availability of the baseline and PD samples, as well as primary mutations (L858R, ex19del) detection status (“detectable” vs. “undetectable”).
Since ddPCR assays are very sensitive in quantifying rare mutation events, we were able to detect very low MT allele frequency (AF) in the cfDNA for both L858R and ex19del mutations. The lowest MT AF detected among 57 L858R+ samples was 0.003%, while among 113 ex19del+ samples was 0.005% (Figure 2A). To test the specificity, we analyzed the results based on the mutual exclusivity of the two mutations in the same sample. For the L858R assay, we observed no L858L mutant copies in the 110 ex19del+ plasma samples (Figure 2B); for ex19del assay, no ex19del mutant copies were detected in the 57 L858R+ samples (Figure 2C), which demonstrated 100% of specificity.
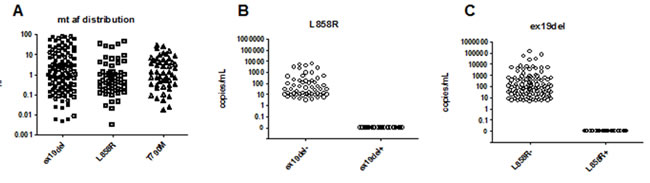
Figure 2: Technical sensitivity and specificity of cell-free plasma DNA using ddPCR assay (n = 79). A. distribution of frequency of alleles according to the EGFR mtuations (ex19del, L858R, and T790), B. detection of L858R mutant alleles in ex19del- negative and ex19del- positive population, and C. detection of ex19del mutant alleles in L858R-negative and L858R-positive population.
Clinical sensitivity of plasma-based EGFR mutation assays and their concordances with tissue mutation status
The clinical sensitivity of the EGFR mutation detection in plasma was assessed among 58 patients whose baseline samples were available. L858R or ex19del mutations were detected in 43 patients (Table 2) yielding sensitivity of 74.1% (43/58); 70.8% (17/24) for L858R and 76.5% (26/34) for ex19del. We assessed the specificity based on the mutual exclusivity of L858R mutation and ex19del mutation in EGFR mutant tumors. No L858R mutation was detected in the ex19del patient plasmas, and vice versa. Therefore, the specificity of cfDNA L858R assay and ex19del assay was 100% (34/34) and 100% (24/24) respectively. The concordance rate between cfDNA and tissue was 87.9% for L858R and 86.2% for ex19del before treatment.
We then examined the EGFR mutation detection rate in plasma from patients who experienced progressive disease (PD). Of 79 patients, 49 patients had plasma samples available at the time of progression. Among 49 patients, L858R and ex19del mutations were detected in 32 patients, resulting in 65.4% (32/49) clinical sensitivity in plasma at disease progression (Figure 1, Table 2).
We subsequently assessed the detection rate of resistance mutation T790M among patients who developed PD. As described previously [20], the technical background of T790M assay in fresh and FFPE gDNA is 0.03% and 0.05%, respectively. The quadruplicate repeats of the test were implemented in determining the true positives of plasma T790M measurements. Among the four data points for each sample, only those with at least three non-zero readings were considered as positives. With these criteria, 50 out of 361 samples were positive in the T790M detection. The lowest allele frequency detected was 0.017% and the median was 1.24% (Figure 2A). Among 49 patients with available samples at the time of progression, 14 patients (28.6%) had detectable T790M in their plasma.
Table 2: Clinical sensitivity and concordance between tissue and cfDNA measurement results |
||||||||||||
Prior Treatment (n = 58) |
Disease Progression (n = 49) |
|||||||||||
cfDNA |
cfDNA |
|||||||||||
Tissue |
L858R+ |
L858R- |
Total |
Ex19del+ |
Ex19del- |
Total |
L858R+ |
L858R- |
Total |
Ex19del+ |
Ex19del- |
Total |
Positive |
17 |
7 |
24 |
26 |
8 |
34 |
11 |
10 |
21 |
21 |
7 |
28 |
Negative |
0 |
34 |
34 |
0 |
24 |
24 |
0 |
28 |
28 |
0 |
21 |
21 |
Total |
17 |
41 |
58 |
26 |
32 |
58 |
11 |
38 |
49 |
21 |
28 |
49 |
Sensitivity |
70.8 |
76.5 |
73.7 |
75.0 |
||||||||
Specificity |
100 |
100 |
100 |
100 |
||||||||
Concordance |
87.9 |
86.2 |
79.6 |
85.7 |
||||||||
Abbreviations: cfDNA, cell-free DNA; Ex19del, exon 19 deletion
Longitudinal monitoring for primary and resistance EGFR mutations in patients treated with EGFR TKIs
To investigate whether cfDNA mutation level can be used to monitor the treatment response and resistance longitudinally, we analyzed plasma samples at specified times over the course of treatment cycle. Among 79 patients, five patients who had only one time point and four who had neither baseline nor PD data points were excluded for analysis (Figure 1). In addition, 24 samples collected beyond disease progression were excluded. Among 70 patients with more than 1 available time point, 33 patients had both baseline and PD points, 21 patients had only baseline, and 16 had only PD (Figure 1, Table 3).
The level of L858R copies at baseline were shown to be significantly (P = 0.0068) associated with shorter PFS. But we did not see the correlation in ex19del (Supplementary Figure 1). Among the 54 patients whose baseline samples were available for analysis, L858R or ex19del mutations were detected in 40 patients’ plasmas at the baseline. All 40 patients showed decreased mutant DNAs in circulation after EGFR TKI treatment. Of note, 34 patients (85.0%) had plasma EGFR mutations decreased to the undetectable level (0 copy) during the treatment cycle compared to the baseline (Table 3, Supplementary Figure 2). Intriguingly, there was a significant difference in PFS between patients with undetectable EGFR mutation and detectable EGFR mutation (10.1 months vs 6.3 months; HR 3.88, 95% CI 1.48-10.19, P = 0.006; Figure 3A). Similarly, the median OS of patients with undetectable EGFR and detectable EGFR were 23.7 and 11.2 months, respectively (HR 5.97, 95% CI 2.04-17.47, P = 0.001; Figure 3B). Of 49 patients with PD plasma samples, 15 patients (30.6%) showed increased mutation level at the PD time point, and 12 patients (24.5%) showed increased mutation level about 2 to 12 months earlier than the PD time point, as judged by the primary mutation L858R or ex19del (Figure 1, Table 3).
Table 3: Dynamic monitoring of patient response to therapy by cfDNA |
||||||||
Patient |
Total (n=70*) |
cfDNA mutation at baseline (n=40) L858R/ex19del |
Dynamic monitoring of cfDNA mutation |
|||||
L858R/ex19del |
T790M |
|||||||
Decrease not zero |
Decrease to zero |
Increase at progression |
Increase earlier than progression |
Increase at progression |
Increase earlier than progression |
|||
B+P+ |
33 |
24 |
3 |
21 |
10 |
10 |
4 |
5 |
B+P- |
21 |
16 |
3 |
13 |
||||
B-P+ |
16 |
5 |
2 |
2 |
3 |
|||
Abbreviations: cfDNA, cell-free DNA; Ex19del, exon 19 deletion
Available samples: B+, available at baseline; P+, available at progression
* Among the 79 patients, 5 patients who had only one time point and 4 patients who had neither baseline nor PD data point data excluded for time course analysis.
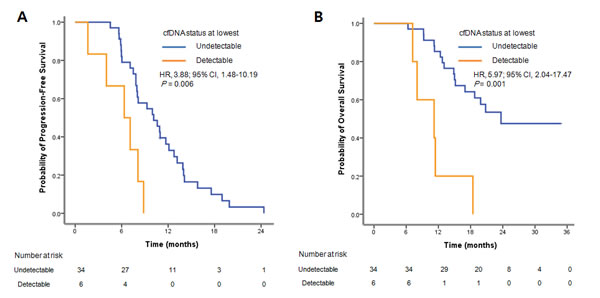
Figure 3: Survival curves for the 40 patients treated with EGFR TKIs. A. Progression-free survival (PFS) and B. overall survival (OS) by cfDNA status at lowest level during the treatment.
Among 49 patients with available samples at the time of progression, the T790M mutation was detected in 8 patients (16.3%) as early as 2 to 12 months prior to radiological progression: the remaining 6 patients (12.2%) developed T790M mutation at the time of PD (Supplementary Figure 3). It is noteworthy that one patient had detectable prior treatment T790M mutation. Although plasma ex19del and T790M levels were decreased with clinical response after gefitinib treatment in patients with de novo T790M mutation, brain and bone metastases occurred after 6 months of gefitinib treatment.
The tumor volume was assessed using Chest CT scans by radiologists (HY Lee, and SH Bak) for each patient every eight weeks. Figure 4 shows the levels of circulating two EGFR primary mutations and the corresponding tumor volumes over the course of treatment to disease progression. In patients whose plasma T790M mutation was detected at the time of progression, T790M was generally at slightly lower levels than the primary mutations. In one patient (#2) with a response to gefitinib, an increase in plasma ex19del and T790M levels was seen during the development of bone metastasis (Figure 4). Two patients (#10 and #88) showed an increase in copy number of both plasma L858R and T790M before RECIST-defined progression, 11 months and 2 months, respectively (Figure 4).
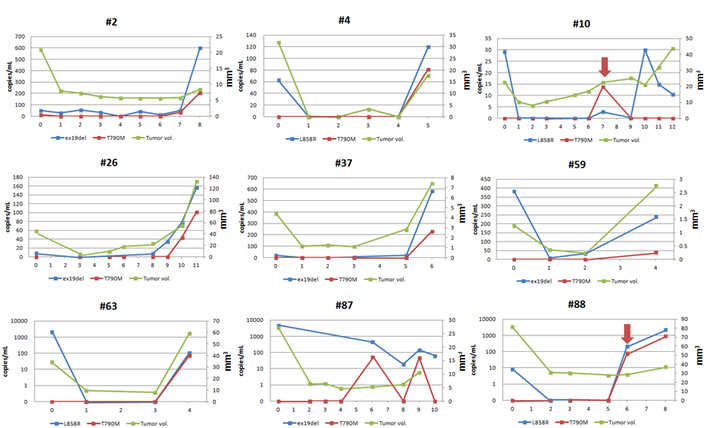
Figure 4: Longitudinal L858R, ex19del and T790M levels in plasma along with tumor volume measured by CT scan. Arrow (red) shows increased copy number of both plasma L858R and T790M before RECIST-defined progression in two patients (#10 and #88). (Unit of x-axis: time point of acquisition of plasma (every 8 weeks). 0 = baseline).
DISCUSSION
The development of acquired resistance to anticancer therapy derives from clonal evolution and selection [21, 22]. Thus, serial biopsies for the tumor are required to identify the underlying drug resistance mechanism and potential therapeutics to overcome or prevent it. Moreover, monitoring treatment response accompanied by mutations in the tumor genome may be needed to evaluate the potential benefits of targeted cancer therapies and subsequently determine and tailor optimal treatment strategies for patients. However, detection of EGFR mutations in NSCLC patients using sampling tumor tissues has significant limitations including tumor heterogeneity and difficulty in obtaining samples. As a non-invasive liquid biopsy capable of identifying molecular changes, genomic analysis of circulating tumor DNA released from cancer cells into blood has been proposed to determine real-time genomic landscapes of the tumor without additional burden and risk to patients [23].
Although several studies have suggested that cfDNA from patients with cancer can be used to detect mutations with high sensitivity [24-26], the detection of samples with low mutation fractions has uncertain clinical significance [27]. The study by Zhang et al. [20] demonstrated the feasibility of applying the ddPCR system to detect EGFR mutation and the advantage of ddPCR in the detection of samples with low EGFR mutation fractions.
In the present study, we demonstrated 74.1% sensitivity and 100% specificity of a plasma assay for EGFR mutations using ddPCR. The concordance rates of L858R and ex19del were 87.9% and 86.2%, respectively. Our results are consistent with those from previous studies [28, 29] and support the use of this technology in clinical practice for making treatment decisions especially for patients not amenable to tissue biopsy. The relatively lower clinical sensitivity of EGFR mutations in patients at the time of disease progression (65%) is likely attributed to the tumor shrinkage after drug treatment and the collection time of plasma samples near or at the PD stage.
In the current study with a total of 40 patients whose plasma collections from baseline were available for a longitudinal analysis, we observed a marked decrease in plasma EGFR mutations during the first 2 months in all patients after EGFR TKI treatment. More intriguingly, 34 patients (85%) had EGFR level decreased to undetectable level with longer PFS and OS than those with detectable EGFR in blood. In addition, the levels of both EGFR primary and resistance mutations in plasma increased progressively during disease progression. These findings suggest that longitudinal monitoring of plasma EGFR mutation levels can be a good predictive biomarker. Sorensen et al. [30] and Mok et al. [31] also showed the predictive value of serial assessment of cfDNA during treatment. To the best of our knowledge, this is the first study that evaluates longitudinal measurements of tumor volume with serial assessments of tumor DNA in blood for NSCLC patients treated with EGFR TKIs. The trend of mutation titer in plasma was well correlated with the tumor volume as measured by CT scan.
EGFR T790M is one of the most common mutations associated with acquired resistance to EGFR TKIs in NSCLC patients, which accounts for 50-60% [11, 32]. At present, the presence of T790M mutation is determined upon taking a new biopsy of the tumor from the patients at the time of progression. Given the limitations of repeated invasive biopsies, several studies were investigated to analyze acquired resistance to cancer therapies by non-invasive methods using plasma DNA [29, 33, 34]. We found that 14 patients (28.6%) among a total of 49 patients harbored resistance mutation T790M in the blood during EGFR TKIs treatment, which is similar to the study by Sakai et al. [35] in which T790M mutation was detected in 21 (30%) of 75 plasma samples from patients with clinical PD using the single allele base extension reaction (SABER) assay. It is noteworthy that the T790M mutant alleles were detectable in the blood of EGFR TKIs-treated patients as early as 12 months before radiographic documentation of disease progression in our study. This suggests that cfDNA may have important clinical implications as a useful surrogate biomarker of therapeutic response and be used to identify the development of resistant mutation, especially in early detection before it becomes clinically detectable. Furthermore, the detection of T790M mutation in cfDNA would be clinically useful for patients who are not amendable to repeated biopsies. Given the promising clinical results of several 3rd generation EGFR TKIs in patients with T790M mutation [36-38], these treatment therapies will soon be available and accessible warranting effective clinical diagnostics to evaluate drug resistance. However, it still remains undetermined whether T790M mutation detected in cfDNA prior to clinically overt progression will change treatment decisions because patients harboring this mutation are usually indolent and EGFR TKI can be continued beyond progression [39]. Therefore, the clinical impact of early detection of the resistance mutation in plasma DNA needs to be elucidated in the near future.
In conclusion, this study demonstrated that plasma EGFR assay using ddPCR can be feasible and effective in monitoring treatment response to EGFR TKIs, and detecting the underlying mechanism of resistance in a noninvasive method. Determining the genetic landscapes of tumors through noninvasive real-time mutational profiling may have significant clinical implications in identifying and guiding optimal treatment strategies, particularly when T790M mutation is detected in the blood.
MATERIALS AND METHODS
Study design
This was an exploratory study to evaluate the potential utilization of EGFR mutations in plasma using ddPCR in monitoring treatment response and resistance among NSCLC patients harboring EGFR activating mutations treated with EGFR TKIs. Eligible patients had a diagnosis of advanced or recurrent NSCLC harboring activating EGFR mutation (exon 19 deletion or L858R) who had progressed on gefitinib or erlotinib. The analysis of EGFR (exon 18-21) mutations for tumor tissues obtained from either diagnostic or surgical procedures was performed prior to TKI treatment by direct sequencing or PNA-clamp method (Panagene Inc., Daejeon, Korea). Patients were treated with gefitinib (250 mg po daily) or erlotinib (150 mg po daily) at the discretion of treating physicians. Appropriate imaging measures including CT scans of the chest were performed every two cycles (8 weeks) and evaluated treatment response according to Response Evaluation Criteria in Solid Tumors (RECIST version 1.1). All patients provided written informed consent. This study was performed in accordance with the Declaration of Helsinki and approved by Institutional Review Board for Human Research at each institute.
The primary objective of this exploratory analysis was to assess the diagnostic utility of the blood test for sensitivity, specificity, concordance rate, and comparison with tumor tissue EGFR mutation status. The secondary objectives included identifying the time difference between development of genetic aberrancy associated with resistance in plasma and overt clinical progression.
Sample collection and DNA extraction
Plasma samples were prospectively obtained before treatment and every 8 weeks post-treatment until development of disease progression for cell-free DNA (cfDNA) EGFR ddPCR testing. Blood was collected into one 10-mL EDTA-containing vacuum tube and was centrifuged at 1500 rpm for 15 minutes within 4 hours of collection. The supernatants were collected and centrifuged at 1500 rpm for 15 minutes. The plasmas were transferred to 1.5-mL Eppendorf tubes and stored at -70°C until DNA extraction. The cfDNA was extracted from 1-2 mL of plasma using QIAamp Circulating Nucleic Acid Kit (Qiagen) according to the manufacturer’s instructions.
Droplet digital PCR detection of EGFR mutations in plasma cfDNA
Primers and probes were custom synthesized by Thermo Fisher and the sequences were reported previously [28]. Briefly, the ddPCR assay was conducted by droplets generation using QX200 generator (Bio-Rad Laboratories, Inc., Hercules, CA, USA), followed by endpoint PCR reactions using C1000 (Bio-Rad) and droplet flow cytometry readings using QX200 reader (Bio-Rad). Analyzed data were processed using QuantaSoft (Bio-Rad) software. Ex19del assay covers all deletion mutations in the exon 19. Detailed procedures can be found elsewhere [20].
Measurement of tumor volume
Considering the recent studies that the conventional RECIST-based assessment alone does not fully characterize response and progression in genomically characterized patients [40, 41], we further explored the volumetric tumor response during TKI therapy. All CT image data were reconstructed with a section thickness of 1.25 mm using soft-tissue algorithms. For each measurable lesion, volume measurements were performed with a semi-automated segmentation method using MRIcro (version 1.40, Chris Rorden, University of Nottingham, Great Britain). Using in-house software, the computer automatically calculated the volume (cm3) by multiplying the number of voxels segmented by the unit volume of a voxel. A sum of the volume for all target lesions at each time-point was used for the analysis.
Statistical analysis
Sensitivity was calculated as the number of samples positive in both tissue and plasma out of the positive tissue samples, whereas specificity was calculated from the number of plasma- and tissue-negative samples out of the total negative tissue samples. Concordance rate was calculated as the number of samples positive in both tissue and plasma, plus the number of samples negative in both tissue and plasma, out of the total number of matched samples. Survival estimates were calculated according to Kaplan-Meier method. Cox regression methods were used for biomarker at baseline associated with PFS. Statistical analysis was performed using SAS 9.4 (SAS Institute Inc, Cary, NC) and R version 3.0.3 (Vienna, Austria; http://www.R-project.org/). A 2-sided P-value of < 0.05 was considered statistically significant.
Acknowledgments
This study was supported in part by AstraZeneca and by the Samsung Biomedical Research Institute Grant (GE1-B3-081-1).
Conflicts of interests
The authors declare no conflicts of interest.
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E and Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61: 69-90.
2. Jung KW, Park S, Kong HJ, Won YJ, Lee JY, Park EC and Lee JS. Cancer statistics in Korea: incidence, mortality, survival, and prevalence in 2008. Cancer Res Treat. 2011; 43: 1-11.
3. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004; 350: 2129-2139.
4. Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004; 304: 1497-1500.
5. Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N, Sunpaweravong P, Han B, Margono B, Ichinose Y, Nishiwaki Y, Ohe Y, Yang JJ, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009; 361: 947-957.
6. Han JY, Park K, Kim SW, Lee DH, Kim HY, Kim HT, Ahn MJ, Yun T, Ahn JS, Suh C, Lee JS, Yoon SJ, Han JH, et al. First-SIGNAL: first-line single-agent iressa versus gemcitabine and cisplatin trial in never-smokers with adenocarcinoma of the lung. J Clin Oncol. 2012; 30: 1122-1128.
7. Zhou C, Wu YL, Chen G, Feng J, Liu XQ, Wang C, Zhang S, Wang J, Zhou S, Ren S, Lu S, Zhang L, Hu C, et al. Erlotinib versus chemotherapy as first-line treatment for patients with advanced EGFR mutation-positive non-small-cell lung cancer (OPTIMAL, CTONG-0802): a multicentre, open-label, randomised, phase 3 study. Lancet Oncol. 2011; 12: 735-742.
8. Wu YL, Zhou C, Hu CP, Feng J, Lu S, Huang Y, Li W, Hou M, Shi JH, Lee KY, Xu CR, Massey D, Kim M, et al. Afatinib versus cisplatin plus gemcitabine for first-line treatment of Asian patients with advanced non-small-cell lung cancer harbouring EGFR mutations (LUX-Lung 6): an open-label, randomised phase 3 trial. Lancet Oncol. 2014; 15: 213-222.
9. Kosaka T, Yatabe Y, Endoh H, Yoshida K, Hida T, Tsuboi M, Tada H, Kuwano H and Mitsudomi T. Analysis of epidermal growth factor receptor gene mutation in patients with non-small cell lung cancer and acquired resistance to gefitinib. Clin Cancer Res. 2006; 12: 5764-5769.
10. Ohashi K, Maruvka YE, Michor F and Pao W. Epidermal growth factor receptor tyrosine kinase inhibitor-resistant disease. J Clin Oncol. 2013; 31: 1070-1080.
11. Pao W, Miller VA, Politi KA, Riely GJ, Somwar R, Zakowski MF, Kris MG and Varmus H. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med. 2005; 2: e73.
12. Balak MN, Gong Y, Riely GJ, Somwar R, Li AR, Zakowski MF, Chiang A, Yang G, Ouerfelli O, Kris MG, Ladanyi M, Miller VA and Pao W. Novel D761Y and common secondary T790M mutations in epidermal growth factor receptor-mutant lung adenocarcinomas with acquired resistance to kinase inhibitors. Clin Cancer Res. 2006; 12: 6494-6501.
13. Weber B, Meldgaard P, Hager H, Wu L, Wei W, Tsai J, Khalil A, Nexo E and Sorensen BS. Detection of EGFR mutations in plasma and biopsies from non-small cell lung cancer patients by allele-specific PCR assays. BMC Cancer. 2014; 14: 294.
14. Sozzi G, Conte D, Leon M, Ciricione R, Roz L, Ratcliffe C, Roz E, Cirenei N, Bellomi M, Pelosi G, Pierotti MA and Pastorino U. Quantification of free circulating DNA as a diagnostic marker in lung cancer. J Clin Oncol. 2003; 21: 3902-3908.
15. Bremnes RM, Sirera R and Camps C. Circulating tumour-derived DNA and RNA markers in blood: a tool for early detection, diagnostics, and follow-up? Lung Cancer. 2005; 49: 1-12.
16. Kimura H, Suminoe M, Kasahara K, Sone T, Araya T, Tamori S, Koizumi F, Nishio K, Miyamoto K, Fujimura M and Nakao S. Evaluation of epidermal growth factor receptor mutation status in serum DNA as a predictor of response to gefitinib (IRESSA). Br J Cancer. 2007; 97: 778-784.
17. Yeo CD, Kim JW, Kim KH, Ha JH, Rhee CK, Kim SJ, Kim YK, Park CK, Lee SH, Park MS and Yim HW. Detection and comparison of EGFR mutations in matched tumor tissues, cell blocks, pleural effusions, and sera from patients with NSCLC with malignant pleural effusion, by PNA clamping and direct sequencing. Lung Cancer. 2013; 81: 207-212.
18. Bai H, Mao L, Wang HS, Zhao J, Yang L, An TT, Wang X, Duan CJ, Wu NM, Guo ZQ, Liu YX, Liu HN, Wang YY, et al. Epidermal growth factor receptor mutations in plasma DNA samples predict tumor response in Chinese patients with stages IIIB to IV non-small-cell lung cancer. J Clin Oncol. 2009; 27: 2653-2659.
19. Hindson BJ, Ness KD, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, Bright IJ, Lucero MY, Hiddessen AL, Legler TC, Kitano TK, Hodel MR, Petersen JF, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem. 2011; 83: 8604-8610.
20. Zhang BO, Xu CW, Shao Y, Wang HT, Wu YF, Song YY, Li XB, Zhang Z, Wang WJ, Li LQ and Cai CL. Comparison of droplet digital PCR and conventional quantitative PCR for measuring gene mutation. Exp Ther Med. 2015; 9: 1383-1388.
21. Aparicio S and Caldas C. The implications of clonal genome evolution for cancer medicine. N Engl J Med. 2013; 368: 842-851.
22. Gerlinger M, Rowan AJ, Horswell S, Larkin J, Endesfelder D, Gronroos E, Martinez P, Matthews N, Stewart A, Tarpey P, Varela I, Phillimore B, Begum S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012; 366: 883-892.
23. Diaz LA, Jr. and Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol. 2014; 32: 579-586.
24. Misale S, Yaeger R, Hobor S, Scala E, Janakiraman M, Liska D, Valtorta E, Schiavo R, Buscarino M, Siravegna G, Bencardino K, Cercek A, Chen CT, et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature. 2012; 486: 532-536.
25. Higgins MJ, Jelovac D, Barnathan E, Blair B, Slater S, Powers P, Zorzi J, Jeter SC, Oliver GR, Fetting J, Emens L, Riley C, Stearns V, et al. Detection of tumor PIK3CA status in metastatic breast cancer using peripheral blood. Clin Cancer Res. 2012; 18: 3462-3469.
26. Diaz LA, Jr., Williams RT, Wu J, Kinde I, Hecht JR, Berlin J, Allen B, Bozic I, Reiter JG, Nowak MA, Kinzler KW, Oliner KS and Vogelstein B. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature. 2012; 486: 537-540.
27. Zhou Q, Zhang XC, Chen ZH, Yin XL, Yang JJ, Xu CR, Yan HH, Chen HJ, Su J, Zhong WZ, Yang XN, An SJ, Wang BC, et al. Relative abundance of EGFR mutations predicts benefit from gefitinib treatment for advanced non-small-cell lung cancer. J Clin Oncol. 2011; 29: 3316-3321.
28. Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O’Connell A, Messineo MM, Luke JJ, Butaney M, Kirschmeier P, Jackman DM and Janne PA. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res. 2014; 20: 1698-1705.
29. Yung TK, Chan KC, Mok TS, Tong J, To KF and Lo YM. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non-small cell lung cancer patients. Clin Cancer Res. 2009; 15: 2076-2084.
30. Sorensen BS, Wu L, Wei W, Tsai J, Weber B, Nexo E and Meldgaard P. Monitoring of epidermal growth factor receptor tyrosine kinase inhibitor-sensitizing and resistance mutations in the plasma DNA of patients with advanced non-small cell lung cancer during treatment with erlotinib. Cancer. 2014; 120: 3896-3901.
31. Mok T, Wu YL, Lee JS, Yu CJ, Sriuranpong V, Sandoval-Tan J, Ladrera G, Thongprasert S, Srimuninnimit V, Liao M, Zhu Y, Zhou C, Fuerte F, et al. Detection and Dynamic Changes of EGFR Mutations from Circulating Tumor DNA as a Predictor of Survival Outcomes in NSCLC Patients Treated with First-line Intercalated Erlotinib and Chemotherapy. Clin Cancer Res. 2015; 21: 3196-3203.
32. Kobayashi S, Boggon TJ, Dayaram T, Janne PA, Kocher O, Meyerson M, Johnson BE, Eck MJ, Tenen DG and Halmos B. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med. 2005; 352: 786-792.
33. Murtaza M, Dawson SJ, Tsui DW, Gale D, Forshew T, Piskorz AM, Parkinson C, Chin SF, Kingsbury Z, Wong AS, Marass F, Humphray S, Hadfield J, et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature. 2013; 497: 108-112.
34. Nakamura T, Sueoka-Aragane N, Iwanaga K, Sato A, Komiya K, Kobayashi N, Hayashi S, Hosomi T, Hirai M, Sueoka E and Kimura S. Application of a highly sensitive detection system for epidermal growth factor receptor mutations in plasma DNA. J Thorac Oncol. 2012; 7: 1369-1381.
35. Sakai K, Horiike A, Irwin DL, Kudo K, Fujita Y, Tanimoto A, Sakatani T, Saito R, Kaburaki K, Yanagitani N, Ohyanagi F, Nishio M and Nishio K. Detection of epidermal growth factor receptor T790M mutation in plasma DNA from patients refractory to epidermal growth factor receptor tyrosine kinase inhibitor. Cancer Sci. 2013; 104: 1198-1204.
36. Janne PA, Ramalingam SS, Yang JC-H, Ahn MJ, Kim DW, Kim SW, Planchard D, Ohe Y, Felip E, Watkins C, Cantarini M, Ghiorghiu S and Ranson M. Clinical activity of the mutant-selective EGFR inhibitor AZD9291 in patients (pts) with EGFR inhibitor–resistant non-small cell lung cancer (NSCLC). J Clin Oncol. 2014; 32: abstr 8009.
37. Sequist LV, Soria J-C, Gadgeel SM, Wakelee HA, Camidge DR, Varga A, Solomon BJ, Papadimitrakopoulou V, Jaw-Tsai SS, Caunt L, Kaur P, Rolfe L, Allen AR, et al. First-in-human evaluation of CO-1686, an irreversible, highly selective tyrosine kinase inhibitor of mutations of EGFR (activating and T790M). J Clin Oncol. 2014; 32: abstr 8010.
38. Kim DW, Lee DH, Kang JH, Park K, Han J-Y, Lee J-S, Jang I-J, Kim H-Y, Son J and Kim J-H. Clinical activity and safety of HM61713, an EGFR-mutant selective inhibitor, in advanced non-small cell lung cancer (NSCLC) patients (pts) with EGFR mutations who had received EGFR tyrosine kinase inhibitors (TKIs). J Clin Oncol. 2014; 32: Suppl; abstr 8011.
39. Oxnard GR, Arcila ME, Chmielecki J, Ladanyi M, Miller VA and Pao W. New strategies in overcoming acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors in lung cancer. Clin Cancer Res. 2011; 17: 5530-5537.
40. Nishino M, Jagannathan JP, Krajewski KM, O’Regan K, Hatabu H, Shapiro G and Ramaiya NH. Personalized tumor response assessment in the era of molecular medicine: cancer-specific and therapy-specific response criteria to complement pitfalls of RECIST. AJR Am J Roentgenol. 2012; 198: 737-745.
41. Nishino M, Jackman DM, Hatabu H, Janne PA, Johnson BE and Van den Abbeele AD. Imaging of lung cancer in the era of molecular medicine. Acad Radiol. 2011; 18: 424-436.

