INTRODUCTION
The gut microbiota plays a crucial role in promoting cell proliferation. For examples, germ-free mice have reduced intestinal epithelial cell turnover due to decreased proliferation, apoptosis, and crypt-to-tip cellular migration [1]. Gut microbiota-generated toll-like receptor signals are required for injured tissues to regenerate and regain homeostasis [2]. Moreover, conventional mice have reduced cancer incidence compared with germ-free mice [3], and an increased bacterial load is found in colonic biopsies from patients with colorectal adenoma or cancer [4]. One mechanism by which bacteria promote cell proliferation is through the stimulatory effect of gram negative bacteria-generated lipopolysaccharide (LPS) on DNA synthesis [5]. It has been shown in mice that hepatic DNA synthesis is impaired when gut-derived LPS is blocked from reaching the liver [6]. Furthermore, both germ-free and LPS-resistant mice exhibit delayed liver regeneration after liver resection, which can be rescued by LPS administration [7]. In addition to proliferation, the gut microbiota modulates host metabolic phenotype and participates in microbial-host co-metabolism [8]. Alterations in gut bacterial communities are associated with metabolic disorders [9], metabolic syndrome [10], obesity [11-13], and nonalcoholic steatohepatitis [14]. Taken together, there is a close relationship between microbial-host metabolism and tissue injury as well as regeneration [15, 16].
Besides LPS, another gut-derived signaling that contributes to liver regeneration is bile acid (BA). There is substantial metabolic demand during liver regeneration, and proper function of intestinal nutrient absorption-mediated by BAs is essential for normal liver repair. The hydrophobic nature of BAs facilitates lipid absorption while conjugated hydrophilic BAs are more effective in lipid emulsification [17]. Fine-tuned lipid and BA homeostasis is important in supporting liver regeneration. Steatosis dampens liver repair capability [18, 19], and leptin-inhibited hepatic fat accumulation also impairs partial hepatectomy (PHx)-induced liver regeneration [20]. Additionally, BA overload induces liver injury in regenerating liver and increases mortality [21-23]. However, BA depletion by cholestyramine reduces liver regeneration capability, and the restored liver mass is reduced by 50% [24]. Together, maintaining BA homeostasis is crucial for normal progression of liver regeneration [15, 16].
Hepatic enzymes together with bacteria enzymes are responsible for generating various kinds of BAs. Microbiota plays a pivotal role in BA synthesis by increasing BA composition diversity via de-conjugation, dehydrogenation, dihydroxylation, and sulfation of primary BAs in the gastrointestinal (GI) tract [25]. Bile salt hydrolase (BSH) detected from Bacteroides, Bifidobacterium, and Lactobacillus catalyzes the de-conjugation of BAs to liberate free primary BAs. Furthermore, several bacteria belonging to the Firmicutes phylum have hydroxysteroid dehydrogenase, which mediates BA oxidation and epimerization. The compositions of BAs between germ-free and conventional rats are markedly different [26]. A cross-sectional study of patients with cirrhosis showed elevated primary BAs and Enterobacteriaceae and diminished 7α-dehydroxylating bacteria including Lachonospiraceae, Ruminococcaceae, and Blautia, which convert free BAs to secondary BAs [27]. Gut microbiota-generated deoxycholic acid (DCA) and lithocholic acid (LCA) induce cell proliferation and GI cancers [28]. These findings clearly indicate that gut microbiota modulate host BA synthesis and diversity, which in turn influence liver function.
Similar to BAs, all-trans retinoic acid (RA), naturally presented in the GI tract, has a profound effect in regulating lipid homeostasis [29, 30]. We have previously shown that RA can facilitate PHx-induced liver regeneration via inducing cell cycle gene expression, which was comparable to the liver regeneration effect of cholic acid (CA) [24, 31]. Because of the intimate relationship between gut-derived signaling and liver regeneration, we hypothesize that RA may regulate gut microbiota and BA composition thereby promoting hepatocyte proliferation in regenerating livers. To test this hypothesis, we analyzed the gut microbiota and BA composition to understand their potential role in PHx-induced liver regeneration in response to RA treatment. Our data showed that priming mice with RA resulted in a lean microbiota composition and hydrophilic BA profile, which were associated with facilitated metabolism and accelerated hepatocyte proliferation.
RESULTS
Priming mice with RA accelerated liver regeneration
Wild type mice received an oral gavage of RA or vehicle, and PHx was performed 48 hours later. RA-primed mouse livers had higher induction of cell cycle gene expressions, including Cyclin D (0-1, 2 day), Cyclin E (0-2 day), Cyclin A (0-1.5 day), and Cyclin B (0-2 day) (Figure 1A). The hepatic protein levels of CYCLIN D (0-0.5 day) and CYCLIN A (0-1 day) also showed increased induction (Figure 1B). RA-induced cell cycle gene expression correlated with enhanced liver regrowth and proliferation, as shown by greater liver-to-body weight ratio and numbers of Ki67-positive cells at all studied times (Figure 1C). Together, RA administration accelerated liver regeneration. Interesting, lipid droplets, usually occurring 1-1.5 day after PHx [18], were absent in RA-primed mouse livers (Figure 1D), indicating a regulatory effect on lipid metabolism by RA during liver regeneration.
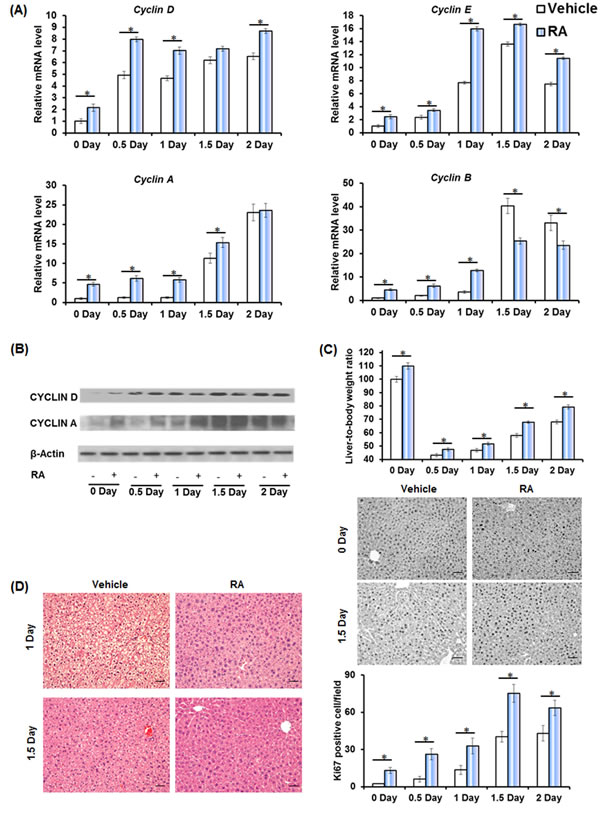
Figure 1: Accelerated liver regeneration in RA-primed mice. Wild type (WT) mice were treated with all-trans retinoic acid (RA) or vehicle by oral gavage 48 hours prior to partial hepatectomy (n = 4). A. Quantitative RT-PCR analyses of hepatic mRNA level of Cyclin D, Cyclin E, Cyclin A, Cyclin B. B. Immunoblotting of liver extracts using anti-CYCLIN D and CYCLIN A antibodies. C. Liver-to-body weight ratios and immunohistochemistry of liver sections for Ki67 staining (20×) are shown. The number of Ki67-labeled nuclei was counted in at least 10 low-magnification (20×) microscope fields for each liver section. D. H&E staining of liver sections. Representative images are shown (20×). Means ± SD are graphed with * indicating p < 0.05.
Priming mice with RA altered gut microbiota during liver regeneration
Firmicutes and Bacteroidetes phyla dominated the microbial community ( > 85%) in both control and RA-primed mice during liver regeneration (Figure 2A). In control mice, there was a transient increase in the abundance of Verrucomicrobia which rose to 7% on day 1, continued increasing to 15% on day 1.5, and contracted to 0.6% on day 2. However, in RA-primed mice, the Verrucomicrobia increase was only detected on 1.5 day after PHx (Figure 2A). RA-treated mice had a reduced ratio of Firmicutes to Bacteroidetes 1 day after surgery (2-fold increase in Bacteroidetes) (Figure 2B). The Ruminococcaceae and Lachnospiraceae are two of the most abundant families from the Firmicutes phyla found in the mammalian intestine, and are associated with intestinal health [32]. Higher levels of Ruminococcaceae (day 2) and Lachnospiraceae (0, 0.5, 2 days) were observed in RA-primed mice (Figure 2C). At the genus level, Akkermansia from the Verrucomicrobia phyla appeared on day 1 (7%), increased to 14.6% on day 1.5, and became undetectable on day 2 in the control mice. The appearance of Akkermansia was delayed in RA-treated mice (Figure 2D). RA increased the abundance of genus Lactobacillus by four folds on day zero (Figure 2D). The abundance of Bifidobacterium was also dramatically higher in RA-primed mice on day zero and 1 day after PHx ( > 9 folds) (Figure 2D).
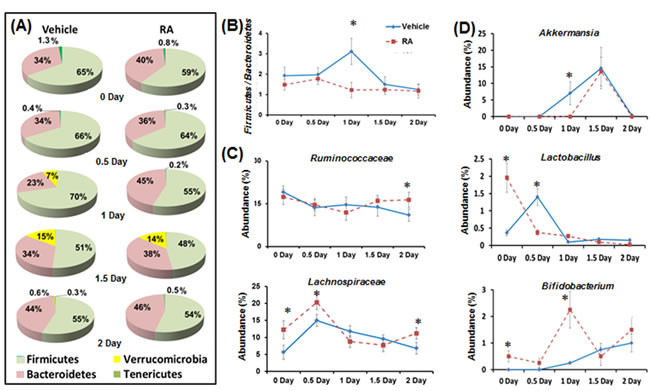
Figure 2: The effect of RA on gut microbiota during liver regeneration. RA-primed mice showed altered gut microbiota composition A., the ratio of Firmicutes to Bacteroidetes B., family levels of Ruminococcaceae and Lachnospiraceae C., and genus levels of Akkemansia, Lactobacillus, and Bifidobacterium D.. Means ± SD are graphed with * indicating p < 0.05.
RA regulates the expression of BA homeostasis genes during liver generation
Studies of the hepatic BA synthesis pathway revealed that priming mice with RA reduced the expression of hepatic cholesterol 7α-hydroxylase (Cyp7a1) and sterol 12α-hydroxylase (Cyp8b1) on day zero. At the protein level, RA reduced CYP7A1 (0-1 day) and CYP8B1 (0, 1, 2 day), but increased CYP8B1 on day 1.5 (Figure 3A).
In the ileum, PHx itself induced small heterodimer partner (Shp) and fibroblast growth factor (Fgf15) expression immediately after surgery (day 0.5) and this induction was sustained throughout the active phase of hepatocyte proliferation. Priming mice with RA further increased the expression of Shp (0.5 day) and Fgf15 (0-0.5 day) indicating an effect of RA in modulating BA-mediated FXR signaling in the ileum during liver regeneration (Figure 3B). SHP and FGF15 protein levels showed a consistent increase with the biggest difference on 0.5 day (Figure 3B). Similar patterns were also observed for the expression of ileal organic anion transporting polypeptide-2 (Oatp2), apical sodium-dependent bile acid transporter (Asbt), and ileal bile acid-binding protein (Ibabp). The expression of these transporters was induced after PHx, indicating increased intestinal BA uptake during liver regeneration [33]. Furthermore, RA enhanced the expression of Oatp2 (0-0.5 day), Asbt (0.5, 2 day), and Ibabp (0-1 day) (Figure 3C). Organic solute transporter beta (Ostβ), a basolateral BA efflux transporter, was also induced in the ileum during liver regeneration, and its induction was further enhanced by RA-priming (day 0.5) (Figure 3C). RA had no effect in regulating ATP-binding cassette, sub-family g, member 5 (Abcg5), Abcg8, Ostα, and Abca1 expression. However, these genes were induced in regenerating livers (Figure 3D).
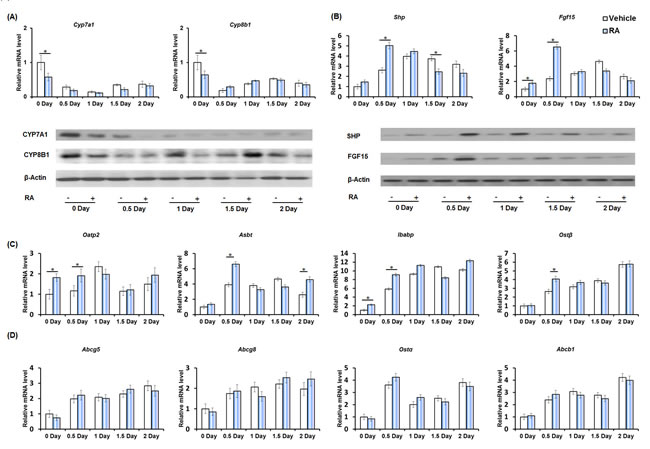
Figure 3: The effect of RA on the expression of genes regulating bile acid homeostasis during liver regeneration. A. Quantitative RT-PCR analyses of hepatic mRNA abundance of BA synthesis genes Cyp7a1 and Cyp8b1, and immunoblotting of liver extracts using CYP7A1 and CYP8B1 antibodies. B. Quantitative RT-PCR analyses of ileal mRNA level of BA regulator Shp and Fgf15, and immunoblotting of ileal extracts using anti-SHP and FGF15 antibodies. C. Quantitative RT-PCR analyses of ileal mRNA level of BA transporter Oatp2, Abst, Ibabp, and Ostβ. D. RA had no effects on ileal mRNA level of BA transporter Abcg5, Abcg8, Ostα, and Abcb1. Means ± SD are graphed with * indicating p < 0.05.
RA alters hepatic BA profile during liver regeneration
Hepatic BAs quantification showed that priming mice with RA had no effect on the amount of total bile acids (TBA) (Figure 4A). However, RA treatment shifted the composition of free BAs (Figure 4B). Tauro-conjugation of BAs increases their hydrophilic/hydrophobic ratio, solubility and lipid emulsification [17], and the taurocholic acid (TCA)/CA ratio was significantly higher in RA-treated mice compared to controls (Figure 4C). The tauro-β-muricholic acid (T-β-MCA) to β-muricholic acid (β-MCA) ratio, associated with increasing lipid emulsion efficiency [8, 17], was also higher in RA-primed mice compared to controls on day zero and 1 day after PHx (Figure 4D).
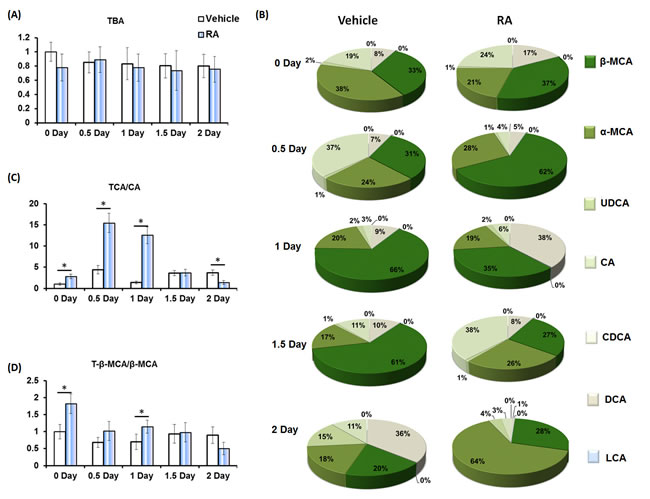
Figure 4: The effect of RA on hepatic bile acid composition during liver regeneration. A. RA had no effect on hepatic total bile acids (TBA); B. RA shifted the percentages of indicated free bile acids in vehicle and RA-treated mice after PHx; C. the ratio of taurocholic acid (TCA) to cholic acid (CA), and D. the ratio of tauro-β-muricholic acid (T-β-MCA) to β-muricholic acid (β-MCA). Means ± SD are graphed with * indicating p < 0.05.
RA regulates energy homeostasis by inducing FGF21 signaling
Fibroblast growth factor 21 (FGF21) is a master metabolic regulator that has a striking ability to reverse obesity, diabetes, and injury as well as prolong life span [34]. FGF21 is transiently induced during the early phase of liver regeneration [23]. RA treatment resulted in early induction of FGF21 mRNA and protein levels after PHx (Figure 5A). Serine/threonine kinase 11 (LKB1), a downstream target of FGF21, also showed earlier induction in RA-primed mice after PHx. FGF21 increases the phosphorylation of AMP-activated protein kinase (AMPK) through LKB1 [35]. Higher phospho-AMPK level was found in RA-treated mice on day 0, 0.5, and 1.5, but RA did not alter total AMPK protein level (Figure 5A). AMPK has been shown to induce NAD+-dependent deacetylase sirtuin 1 (SIRT1) activity (day 1 and 2), resulting in modulation of downstream targets including peroxisome proliferator-activated receptor-γ coactivator 1α (Pgc1α), carnitine acyltransferase 1 (Cpt1), and sterol regulatory element-binding transcription factor 1c (Srebp1c) [35]. Increased FGF21 was accompanied by elevated SIRT1, Pgc1α, and Cpt1 0-1 day after PHx (Figure 5B). In addition, PHx did not affect the total extracellular-signal-regulated kinase 1 and 2 (ERK1/2) level, but rather induced ERK1/2 phosphorylation, which was further increased by RA on day 0 and 0.5 day (Figure 5B).
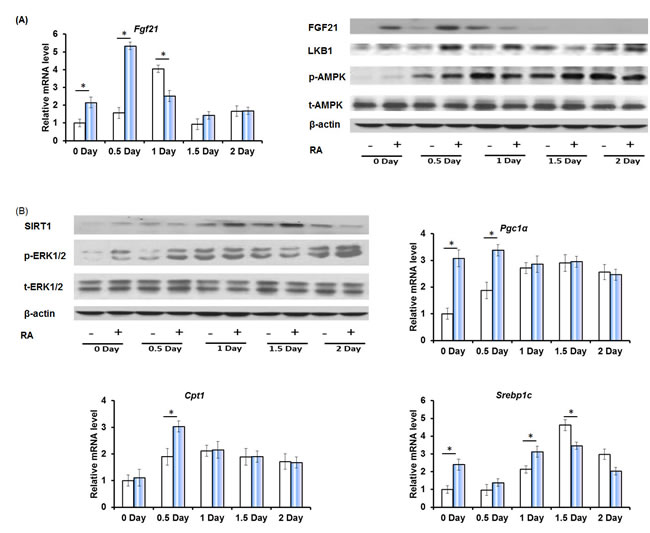
Figure 5: The effect of RA on the expression of factors that regulate hepatic metabolism and cell proliferation during liver regeneration. A. Quantitative RT-PCR analyses of hepatic mRNA abundance of Fgf21, and immunoblotting of liver extracts using FGF21, LKB1, p-AMPK, and t-AMPK antibodies. B. Immunoblotting of liver extracts using SIRT1, p-ERK1/2, and t-ERK1/2 antibodies, and Quantitative RT-PCR analyses of hepatic mRNA abundance of Pgc1α, Cpt1, and Srebp1c. Means ± SD are graphed with * indicating p < 0.05.
DISCUSSION
Emerging evidence indicates that bidirectional communication exists between the gut microbiota and BAs through a “gut-liver axis” [1, 25]. The gut microbiome has increasingly been acknowledged as a novel contributor to host metabolism [25, 27]. Mounting evidence in mice and humans supports that the gut microbiota is linked with metabolic disorders such as diabetes and obesity [1, 27]. Liver resection changes lipid and BA metabolism during a period when increased energy production is needed to meet the metabolic demands of cell proliferation [36]. We demonstrated, for the first time, that RA altered the gut microbiota composition and BA profile, which was accompanied by an increase in energy metabolism, beneficial for liver regeneration.
Alterations in gut microbiota composition have been implicated in metabolic disorders, and specific bacterial phylotypes or metabolic activities may be beneficial or detrimental to patients with obesity [1, 8]. Firmicutes and Bacteroidetes are the two most dominant bacterial phyla that affect the efficiency of host energy extraction and are linked with excess adiposity in both mice and humans [13, 37]. Obese mice have more Firmicutes and fewer Bacteroidetes compared to lean mice. The role of RA in regulating lipid homoestasis had been defined by genomic binding and transcriptome profiling [29]. Decreased lipid accumulation and the ratio of Firmicutes to Bacteroidetes presented in RA-primed mice are associated with accelerated liver regeneration, indicating beneficial lipid metabolism induced by RA during liver regeneration. Phyla Verrucomicrobia is occasionally observed, and is capable of pathogenic activity with growth inhibition toward the host [38, 39]. Lachnospiraceae and Ruminococcaceae are the most abundant Firmicute families in the intestines, accounting for roughly 50% and 30% of phylotypes, respectively in healthy adults [40, 41]. Lachnospiraceae and Ruminococcaceae, associated with the production of butyrate necessary for colonic epithelial health, have been shown to be depleted in inflammatory bowel disease patients [42, 43]. The gut microbiota exerts a strong influence on energy homeostasis due to bacteria-produced butyrate which serves as an energy source for colonocytes [44]. The increased Lachnospiraceae and Ruminococcaceae due to RA treatment may increase butyrate production, which may benefit liver regeneration.
BAs are efficiently conserved under normal conditions by a process termed enterohepatic recirculation [33]. Conjugated and unconjugated BAs are reabsorbed by passive diffusion along the entire gut and by active transport in the terminal ileum. BA overload has been observed after liver resection, and Cyp7a1 and Cyp8b1 inhibition indicate the necessity to down-regulate BA synthesis during liver regeneration [23, 24]. Cyp7a1 expression can be regulated by a negative feedback loop involving intestinal BA signaling [22, 33]. RA-mediated induction of SHP and Fgf15 has been shown to mediate this suppression [30]. During liver regeneration, RA can boost this Fxr-mediated pathway to further inhibit hepatic BA synthesis. Asbt absorbs the majority of BAs available in the adjacent lumen of the ileum and is critical for intestinal BA reabsorption during enterohepatic recirculation [45]. Mutation or inhibition of Asbt causes BA malabsorption and interruption of the enterohepatic circulation of BAs resulting in chronic diarrhea, steatorrhea, and fat-soluble vitamin malabsorption [45, 46]. In the intestines, BAs are transported by Ibabp, after which they are secreted back into circulation by several transporters including Ostα/β. The up regulation of these genes found in regenerating livers can accelerate BA circulation, thereby protecting hepatocytes against high concentration of BAs. Together, the presented data illustrated that in response to liver resection, the enterohepatic recirculation adapted to the immediate BA overload by inhibiting hepatic BA synthesis and inducing intestinal BAs transporters, both of which were enhanced by RA.
It is possible that through gut microbiota-modulated BA composition, RA facilitated metabolism, which contributed to accelerated liver regeneration. Metabolism of BAs by gut microbes increases BA diversity, thus not only affecting BA and lipid homeostasis, but also other aspects of host health [8]. For example, BSH activity, produced in significant amounts by Lactobacillus and Bifidobacterium, could be a way for bacterial species to detoxify BAs [8]. In addition, the dehydroxylation of chenodeoxycholic acid (CDCA) forms LCA, which is toxic to hepatocytes and has been linked to colon carcinogenesis [28]. Conversely, epimerization of CDCA generates ursodeoxycholic acid (UDCA), which is chemoprotective and is used to treat cholesterol gallstone [47]. Furthermore, conjugated BAs reduce the hydrophobic/hydrophilic ratio and are more effective in lipid emulsification and micelle formation. The presented data revealed a shift towards hydrophilic and conjugated BAs in RA-treated mice. Together, RA treatment may exert beneficial BA metabolism by intestinal microbes for liver regeneration.
FGF21 is required for activation of hepatic lipid oxidation, triglyceride clearance, and ketogenesis during fasting [48]. Administration of FGF21 to rodents with diet-induced or genetic obesity and diabetes resulted in potent antihyperglycemic and triglyceride lowering effects [35]. Forced FGF21 expression in partial hepatectomized hPPARαPAC mice reduced hepatic steatosis, prevented focal necrosis, and restored liver mass [23]. LKB1 and AMPK are required for FGF21-mediated energy regulation of mitochondrial oxidative function [35] via Pgc1α, Cpt1, and Srebp1c [49, 50]. Inhibition of AMPK activity attenuated FGF21-stimulated increase in Cpt1 and Pgc1α gene expression [35]. Both AMPK and SIRT1 act together with Pgc1α, the master regulator of mitochondrial biogenesis, to modulate energy homeostasis in response to environmental and nutritional stimuli [35]. The sustained activation of ERK1/2, can also be regulated by FGF21 to induce stimulatory cell cycle regulators [51]. Transient accumulation of hepatocellular fat during the early stage of liver regeneration has been proposed to serve as an β-oxidation energy source for hepatocellular proliferation [52]. Consistently, infusion of octanoylcarnitine, an inhibitor of β-oxidation, decreased DNA synthesis and energy charge levels in regenerating livers [53]. Mice treated with RA seem to have increased efficiency in lipid transport and metabolism, which is suggested by a lack of lipid accumulation and accelerated hepatocyte proliferation.
In summary, our data showed that RA-accelerated liver regeneration was accompanied by a decreased ratio of Firmicutes to Bacteroidetes, which is known to be associated with a lean phenotype. Consistently, RA-primed mice lacked the transient lipid accumulation normally found in regenerating mouse livers. In addition, RA altered BA signaling, enhanced enterohepatic recirculation of BAs, and increased the hydrophilic/hydrophobic ratio of BAs in the regenerating livers, potentially yielding higher solubility and lipid emulsification capability in RA-primed mice. RA-priming afforded more efficient lipid circulation and induction of the FGF21-LKB1-AMPK pathway, which boosted the energy metabolism necessary for liver regeneration. A better mechanistic understanding of the proliferative effects exerted by RA through modulating gut microbiota composition may allow for the development of therapeutics to promote liver regeneration in the clinical setting.
MATERIALS AND METHODS
Animal
Male C57BL/6 mice (3-5 months old) were housed in steel microisolator cages at 22°C with a 12-hour light/dark cycle. Food and water were provided ad libitum throughout study. RA (25 μg/gram of body weight, one time) (Sigma-Aldrich Corp., St. Louis, MO) or vehicle control (carboxymethyl cellulose) (Sigma-Aldrich Corp., St. Louis, MO) was administered to the mice by gavage. Forty-eight hours after treatment, 2/3 PHx was performed [31, 54, 55]. Mice were killed 0.5, 1, 1.5, and 2 days after surgery (n = 4) covering the time when hepatocytes are actively proliferating. Zero time point mice, which were killed immediately after PHx, served as controls (0 day). All animal experiments were conducted in accordance with the National Institutes of Health Guide for the Care and Use of Laboratory Animals under protocols approved by the Institutional Animal Care and Use Committee of the University of California, Davis.
Sequencing and analysis of gut microbial communities in the cecal contents
Cecal DNA was extracted using the MoBio power soil kit (MoBio) according to the manufacturer’s protocol. Multiplexed DNA libraries were prepared according to a previously described protocol [56]. In brief, the V4 domain was amplified using primers 515F and 806R, a set of prokaryotic universal primers that was well-validated and used as the standard protocol for the Earth Microbiome Project (http://www.earthmicrobiome.org/), with a unique identifier sequence for each sample. Polymerase chain reaction (PCR) mixtures contained 1 Unit Kapa2G Robust Hot Start Polymerase (Kapa Biosystems), 1.5 mM MgCl2, 10 pmol of each primer, and 2 ng of DNA. PCR included 95°C for 2 minutes, followed by 30 cycles of 95°C for 15 seconds, 55°C for 30 seconds, and 72°C for 45 seconds and a final extension of 72°C for 3 minutes. The amplicon libraries were sequenced using 2 × 250-bp paired-end protocol on an Illumina Miseq system. Sequencing reads were trimmed of their barcodes, demultiplexed and combined using custom Perl scripts and aligned to the Greengenes (release 13_5) database using Qiime with default parameters.
Quantification of bile acids
BAs were analyzed using a ProminenceTM UFLC system (Shimadzu, Kyoto) coupled to an API 4000 QTRAPTM mass spectrometer (AB Sciex, CA) operated in negative ionization mode based on published methods [23, 57, 58]. Chromatography was performed on a Kinetex C18 column (50 mm X 2.1 mm, 2.6 μm) maintained at 40°C preceded by high pressure column prefilter. The mobile phase consisted of methanol gradient delivered at a flow rate of 0.4 ml/min.
Real-time quantitative PCR (qPCR)
Hepatic RNA isolated by TRIzol (Invitrogen, Carlsbad, CA) was reverse transcribed to generate cDNA followed by amplification using the ABI Prism 7900HT sequence detection system (Applied Biosystems, Foster City, CA). The hepatic mRNA levels were normalized to Gpadh mRNA levels.
Western blot
Hepatic and ileal proteins (40 µg) were electrophoresed on SDS-polyacrylamide gels and transferred to polyvinylidene fluoride membranes. Anti-SHP, FGF15, FGF21, CYP7A1, CYP8B1, LKB1, AMPK, and β-actin (Santa Cruz, CA) antibodies were used for detection the specific proteins.
Statistical analysis
Data are given as mean ± SD. Statistical analysis was performed using Student’s t test or one-way analysis of variance. Significance was defined by p < 0.05.
ACKNOWLEDGMENTs
The authors thank the Host-Microbe Systems Biology Core facility at the University of California, Davis for microbiota sequencing and data analysis. The authors thank Thinh Chau and Lisa Teixeria for editing the manuscript. This study is supported by grants funded by National Institutes of Health CA53596, DK092100, and U01CA179582.
Financial Support
This study is supported by grants funded by National Institutes of Health CA53596, DK092100, and U01CA179582.
Conflicts of Interest
The authors declare no conflict of interest.
REFERENCES
1. Sommer F and Backhed F. The gut microbiota - masters of host development and physiology. Nat Rev Microbiol. 2013; 11:227-238.
2. Rakoff-Nahoum S, Paglino J, Eslami-Varzaneh F, Edberg S and Medzhitov R. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell. 2004; 118:229-241.
3. Dove WF, Clipson L, Gould KA, Luongo C, Marshall DJ, Moser AR, Newton MA and Jacoby RF. Intestinal neoplasia in the Apc(Min) mouse: Independence from the microbial and natural killer (beige locus) status. Cancer Res. 1997; 57:812-814.
4. Swidsinski A, Khilkin M, Kerjaschki D, Schreiber S, Ortner M, Weber J and Lochs H. Association between intraepithelial Escherichia coli and colorectal cancer. Gastroenterology. 1998; 115:281-286.
5. Cornell RP. Gut-Derived Endotoxin Elicits Hepatotrophic Factor Secretion for Liver-Regeneration. Am J Physiol. 1985; 249:R551-R562.
6. Cornell RP. Restriction of Gut-Derived Endotoxin Impairs DNA-Synthesis for Liver-Regeneration. Am J Physiol. 1985; 249:R563-R569.
7. Cornell RP, Liljequist BL and Bartizal KF. Depressed Liver-Regeneration after Partial-Hepatectomy of Germ-Free, Athymic and Lipopolysaccharide-Resistant Mice. Hepatology. 1990; 11:916-922.
8. Martin FPJ, Dumas ME, Wang YL, Legido-Quigley C, Yap IKS, Tang HR, Zirah S, Murphy GM, Cloarec O, Lindon JC, Sprenger N, Fay LB, Kochhar S, et al. A top-down systems biology view of microbiome-mammalian metabolic interactions in a mouse model. Mol Syst Biol. 2007; 3. 112.
9. Larsen N, Vogensen FK, van den Berg FWJ, Nielsen DS, Andreasen AS, Pedersen BK, Abu Al-Soud W, Sorensen SJ, Hansen LH and Jakobsen M. Gut Microbiota in Human Adults with Type 2 Diabetes Differs from Non-Diabetic Adults. Plos One. 2010; 5:e9085.
10. Zupancic ML, Cantarel BL, Liu ZQ, Drabek EF, Ryan KA, Cirimotich S, Jones C, Knight R, Walters WA, Knights D, Mongodin EF, Horenstein RB, Mitchell BD, Steinle N, Snitker S, Shuldiner AR, et al. Analysis of the Gut Microbiota in the Old Order Amish and Its Relation to the Metabolic Syndrome. Plos One. 2012; 7:e43052.
11. Ley RE, Backhed F, Turnbaugh P, Lozupone CA, Knight RD and Gordon JI. Obesity alters gut microbial ecology. P Natl Acad Sci USA. 2005; 102:11070-11075.
12. Backhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A, Semenkovich CF and Gordon JI. The gut microbiota as an environmental factor that regulates fat storage. P Natl Acad Sci USA. 2004; 101:15718-15723.
13. Ley RE, Turnbaugh PJ, Klein S and Gordon JI. Microbial ecology - Human gut microbes associated with obesity. Nature. 2006; 444:1022-1023.
14. Moschen AR, Kaser S and Tilg H. Non-alcoholic steatohepatitis: a microbiota-driven disease. Trends Endocrin Met. 2013; 24:537-545.
15. Liu HX, Keane R, Sheng L and Wan YY. Implications of microbiota and bile acid in liver injury and regeneration. J Hepatol. 2015; 63:1502-10.
16. Tsuei J, Chau T, Mills D and Wan YJY. Bile acid dysregulation, gut dysbiosis, and gastrointestinal cancer. Exp Biol Med. 2014; 239:1489-1504.
17. Heuman DM. Quantitative Estimation of the Hydrophilic Hydrophobic Balance of Mixed Bile-Salt Solutions. J Lipid Res. 1989; 30:719-730.
18. Liu HX, Fang YP, Hu Y, Gonzalez FJ, Fang JW and Wan YJY. PPAR beta Regulates Liver Regeneration by Modulating Akt and E2f Signaling. Plos One. 2013; 8:e65644.
19. DeAngelis RA, Markiewski MM, Taub R and Lambris JD. A high-fat diet impairs liver regeneration in C57BL/6 mice through overexpression of the NF-kappa B inhibitor, I kappa B alpha. Hepatology. 2005; 42:1148-1157.
20. Shteyer E, Liao YJ, Muglia LJ, Hruz PW and Rudnick DA. Disruption of hepatic adipogenesis is associated with impaired liver regeneration in mice. Hepatology. 2004; 40:1322-1332.
21. Uriarte I, Fernandez-Barrena MG, Monte MJ, Latasa MU, Chang HCY, Carotti S, Vespasiani-Gentilucci U, Morini S, Vicente E, Concepcion AR, Medina JF, Marin JJG, Berasain C, et al. Identification of fibroblast growth factor 15 as a novel mediator of liver regeneration and its application in the prevention of post-resection liver failure in mice. Gut. 2013; 62:899-910.
22. Pean N, Doignon I, Garcin I, Besnard A, Julien B, Liu BK, Branchereau S, Spraul A, Guettier C, Humbert L, Schoonjans K, Rainteau D and Tordjmann T. The Receptor TGR5 Protects the Liver From Bile Acid Overload During Liver Regeneration in Mice. Hepatology. 2013; 58:1451-1460.
23. Liu HX, Hu Y, French SW, Gonzalez FJ and Wan YJ. Forced expression of fibroblast growth factor 21 reverses the sustained impairment of liver regeneration in hPPARalphaPAC mice due to dysregulated bile acid synthesis. Oncotarget. 2015; 6:9686-9700. Doi: 10.18632/oncotarget.3531.
24. Huang WD, Ma K, Zhang J, Qatanani M, Cuvillier J, Liu J, Dong BN, Huang XF and Moore DD. Nuclear receptor-dependent bile acid signaling is required for normal liver regeneration. Science. 2006; 312:233-236.
25. Sayin SI, Wahlstrom A, Felin J, Jantti S, Marschall HU, Bamberg K, Angelin B, Hyotylainen T, Oresic M and Backhed F. Gut Microbiota Regulates Bile Acid Metabolism by Reducing the Levels of Tauro-beta-muricholic Acid, a Naturally Occurring FXR Antagonist. Cell Metab. 2013; 17:225-235.
26. Swann JR, Want EJ, Geier FM, Spagou K, Wilson ID, Sidaway JE, Nicholson JK and Holmes E. Systemic gut microbial modulation of bile acid metabolism in host tissue compartments. P Natl Acad Sci USA. 2011; 108:4523-4530.
27. Kakiyama G, Pandak WM, Gillevet PM, Hylemon PB, Heuman DM, Daita K, Takei H, Muto A, Nittono H, Ridlon JM, White MB, Noble NA, Monteith P, Fuchs M, Thacker LR, Sikaroodi M, et al. Modulation of the fecal bile acid profile by gut microbiota in cirrhosis. J Hepatol. 2013; 58:949-955.
28. Hu Y, Chau T, Liu HX, Liao DG, Keane R, Nie YQ, Yang H and Wan YJY. Bile Acids Regulate Nuclear Receptor (Nur77) Expression and Intracellular Location to Control Proliferation and Apoptosis. Mol Cancer Res. 2015; 13:281-292.
29. He YQ, Gong L, Fang YP, Zhan Q, Liu HX, Lu YL, Guo GL, Lehman-McKeeman L, Fang JW and Wan YJY. The role of retinoic acid in hepatic lipid homeostasis defined by genomic binding and transcriptome profiling. Bmc Genomics. 2013; 14:575.
30. Yang F, He YQ, Liu HX, Tsuei J, Jiang XY, Yang L, Wang ZT and Wan YJY. All-trans retinoic acid regulates hepatic bile acid homeostasis. Biochem Pharmacol. 2014; 91:483-489.
31. Liu HX, Ly I, Hu Y and Wan YJY. Retinoic acid regulates cell cycle genes and accelerates normal mouse liver regeneration. Biochem Pharmacol. 2014; 91:256-265.
32. Tap J, Mondot S, Levenez F, Pelletier E, Caron C, Furet JP, Ugarte E, Munoz-Tamayo R, Le Paslier D, Nalin R, Dore J and Leclerc M. Towards The Healthy Human Intestinal Microbiota Phylogenetic Core? Microb Ecol. 2009; 57:580-581.
33. Halilbasic E, Claudel T and Trauner M. Bile acid transporters and regulatory nuclear receptors in the liver and beyond. J Hepatol. 2013; 58:155-168.
34. Li Y, Wong K, Walsh K, Gao B and Zang MW. Retinoic Acid Receptor beta Stimulates Hepatic Induction of Fibroblast Growth Factor 21 to Promote Fatty Acid Oxidation and Control Whole-body Energy Homeostasis in Mice. J Biol Chem. 2013; 288:10490-10504.
35. Chau MDL, Gao JP, Yang Q, Wu ZD and Gromada J. Fibroblast growth factor 21 regulates energy metabolism by activating the AMPK-SIRT1-PGC-1 alpha pathway. P Natl Acad Sci USA. 2010; 107:12553-12558.
36. Fausto N, Campbell JS and Riehle KJ. Liver regeneration. Hepatology. 2006; 43:S45-S53.
37. Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER and Gordon JI. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature. 2006; 444:1027-1031.
38. Dubourg G, Lagier JC, Armougom F, Robert C, Audoly G, Papazian L and Raoult D. High-level colonisation of the human gut by Verrucomicrobia following broad-spectrum antibiotic treatment. Int J Antimicrob Ag. 2013; 41:149-155.
39. Sait M, Kamneva OK, Fay DS, Kirienko NV, Polek J, Shirasu-Hiza MM and Ward NL. Genomic and experimental evidence suggests that Verrucomicrobium spinosum interacts with eukaryotes. Front Microbiol. 2011; 2:211.
40. Jalanka-Tuovinen J, Salonen A, Nikkila J, Immonen O, Kekkonen R, Lahti L, Palva A and de Vos WM. Intestinal Microbiota in Healthy Adults: Temporal Analysis Reveals Individual and Common Core and Relation to Intestinal Symptoms. Plos One. 2011; 6:e23035.
41. Tap J, Mondot S, Levenez F, Pelletier E, Caron C, Furet JP, Ugarte E, Munoz-Tamayo R, Paslier DLE, Nalin R, Dore J and Leclerc M. Towards the human intestinal microbiota phylogenetic core. Environ Microbiol. 2009; 11:2574-2584.
42. Vital M, Howe AC and Tiedje JM. Revealing the Bacterial Butyrate Synthesis Pathways by Analyzing (Meta) genomic Data. Mbio. 2014; 5:e00889.
43. Frank DN, Amand ALS, Feldman RA, Boedeker EC, Harpaz N and Pace NR. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. P Natl Acad Sci USA. 2007; 104:13780-13785.
44. Donohoe DR, Garge N, Zhang XX, Sun W, O’Connell TM, Bunger MK and Bultman SJ. The Microbiome and Butyrate Regulate Energy Metabolism and Autophagy in the Mammalian Colon. Cell Metab. 2011; 13:517-526.
45. Oelkers P, Kirby LC, Heubi JE and Dawson PA. Primary bile acid malabsorption caused by mutations in the ileal sodium-dependent bile acid transporter gene (SLC10A2). J Clin Invest. 1997; 99:1880-1887.
46. Heubi JE, Balistreri WF, Fondacaro JD, Partin JC and Schubert WK. Primary bile acid malabsorption: defective in vitro ileal active bile acid transport. Gastroenterology. 1982; 83:804-811.
47. Azcarate-Peril MA, Sikes M and Bruno-Barcena JM. The intestinal microbiota, gastrointestinal environment and colorectal cancer: a putative role for probiotics in prevention of colorectal cancer? Am J Physiol-Gastr L. 2011; 301:G401-G424.
48. Badman MK, Pissios P, Kennedy AR, Koukos G, Flier JS and Maratos-Flier E. Hepatic fibroblast growth factor 21 is regulated by PPAR alpha and is a key mediator of hepatic lipid metabolism in ketotic states. Cell Metab. 2007; 5:426-437.
49. Inagaki T, Dutchak P, Zhao GX, Ding XS, Gautron L, Parameswara V, Li Y, Goetz R, Mohammadi M, Esser V, Elmquist JK, Gerard RD, Burgess SC, et al. Endocrine regulation of the fasting response by PPAR alpha-mediated induction of fibroblast growth factor 21. Cell Metab. 2007; 5:415-425.
50. Xu J, Lloyd DJ, Hale C, Stanislaus S, Chen M, Sivits G, Vonderfecht S, Hecht R, Li YS, Lindberg RA, Chen JL, Jung DY, Zhang ZY, et al. Fibroblast Growth Factor 21 Reverses Hepatic Steatosis, Increases Energy Expenditure, and Improves Insulin Sensitivity in Diet-Induced Obese Mice. Diabetes. 2009; 58:250-259.
51. Fisher FM, Chui PC, Antonellis PJ, Bina HA, Kharitonenkov A, Flier JS and Maratos-Flier E. Obesity Is a Fibroblast Growth Factor 21 (FGF21)-Resistant State. Diabetes. 2010; 59:2781-2789.
52. Holecek M. Nutritional modulation of liver regeneration by carbohydrates, lipids, and amino acids: A review. Nutrition. 1999; 15:784-788.
53. Nakatani T, Ozawa K, Asano M, Ukikusa M, Kamiyama Y and Tobe T. Differences in Predominant Energy Substrate in Relation to the Resected Hepatic Mass in the Phase Immediately after Hepatectomy. J Lab Clin Med. 1981; 97:887-898.
54. Hu Y, Zhan Q, Liu HX, Chau T, Li YY and Wan YJY. Accelerated Partial Hepatectomy-Induced Liver Cell Proliferation Is Associated with Liver Injury in Nur77 Knockout Mice. Am J Pathol. 2014; 184:3272-3283.
55. Liu HX, Fang Y, Hu Y, Gonzalez FJ, Fang J and Wan YJ. PPARbeta Regulates Liver Regeneration by Modulating Akt and E2f Signaling. Plos One. 2013; 8:e65644.
56. Meyer M and Kircher M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harbor protocols. 2010; 2010:pdb prot5448.
57. Yang F, Xu Y, Xiong AZ, He YG, Yang L, Wan YJY and Wang ZT. Evaluation of the protective effect of Rhei Radix et Rhizoma against alpha-naphthylisothiocyanate induced liver injury based on metabolic profile of bile acids. J Ethnopharmacol. 2012; 144:599-604.
58. Garcia-Canaveras JC, Donato MT, Castell JV and Lahoz A. Targeted profiling of circulating and hepatic bile acids in human, mouse, and rat using a UPLC-MRM-MS-validated method. J Lipid Res. 2012; 53:2231-2241.

