Introduction
Aldehydes, which accumulate during the metabolism and biotransformation of chemicals and drugs, are reactive electrophilic compounds which are harmful to the organism [1]. Aldehyde dehydrogenases (ALDH) are a family of enzymes that catalyze aldehyde conversion into carboxylic acids via NAD(P)+-dependent oxidation [2]. In addition to detoxifying aldehydes, ALDH enzymes have multiple other functions, such as nitrate reductase activity [1]. These enzymes are present in various human tissues, with the highest concentration in the liver, and are also found in stem cells [3]. High ALDH activity has been detected in hematopoietic stem/progenitor cells [4, 5], and inhibition of ALDH activity has been shown to impair the differentiation of hematopoietic stem cells [6]. Within cells, ALDH is found in cytosols, nuclei, mitochondria, and endoplasmic reticulum. Nineteen ALDH family members have been identified in humans, including ALDH1A1, ALDH1A3, ALDH2, ALDH3A1, and ALDH4A1 [1].
ALDH1A1 has been suggested as a breast cancer stem cell marker [7, 8]. However, contradictory findings on the role of ALDH1A1 in predicting the prognosis of breast cancer patients have been reported. Some studies have shown that ALDH1A1 protein expression is associated with late-stage cancer, large tumor size, chemoresistance, and poor prognosis [7, 9, 10], while other studies have found that ALDH1A1 protein levels do not predict breast cancer survival [11, 12]. In addition, one study reported that high levels of ALDH1A1 in tumor stromal tissues are associated with better clinical outcomes [13]. Considered together, these study findings are inconclusive in determining whether the expression of ALDH1A1, either as mRNA or as a protein, can predict clinical outcomes in breast cancer patients. In addition, no studies have specifically evaluated the role of ALDH1A1 in predicting prognosis of triple-negative breast cancer (TNBC: estrogen receptor negative (ER-), progesterone receptor negative (PR-), human epidermal growth factor receptor 2 negative (HER2-)) .
In this study, we analyzed the association between mRNA expression of the ALDH1A1 gene in tumor tissues, and the clinical outcomes in patients with TNBC in three cohorts of breast cancer patients. We also evaluated the association between ALDH1A3 gene expression and TNBC prognosis because one study had suggested that ALDH1A3 expression can predict metastasis in breast cancer patients [11]. In addition, we validated our findings using 4 publicly-available gene expression data sets.
Results
In the SBCSS, TNBC patients with expression levels of the ALDH1A1 gene above the median had better disease-free survival (DFS) (P = 0.01) and overall survival (OS) (P = 0.048) than those with expression levels of ALDH1A1 below the median (Figure 1). In the multivariate analysis, adjusted for age at diagnosis and TNM stage, one log unit increment of the ALDH1A1 gene expression was associated with DFS (HR = 0.87, 95% CI: 0.80-0.95) and OS (HR = 0.85, 95% CI: 0.78-0.93) (Table 2). The association remained largely unchanged after further adjustment for radiotherapy treatment, chemotherapy treatment, and basal-like breast cancer subtype (Table 2). Analyses by quartile or median cut points of expression levels of the ALDH1A1 gene revealed a similar pattern: higher ALDH1A1 expression was associated with better DFS and OS, although not all associations were statistically significant, particularly when subtypes of TNBC were adjusted for. No association of ALDH1A3 gene expression levels with DFS and OS was observed in the SBCSS (Table 2). In addition, we found that patients with higher grade tumors had a lower level of ALDH1A1 mRNA expression (Figure 2A); no association was found for TNM stages (Figure 2B). SBCSS participants with basal-like TNBC had lower ALDH1A1 expression levels than participants with non-basal-like TNBC (Figure 2C).
Table 1: Characteristics of participants in the SBCSS, NBHS, and SCCS cohorts.
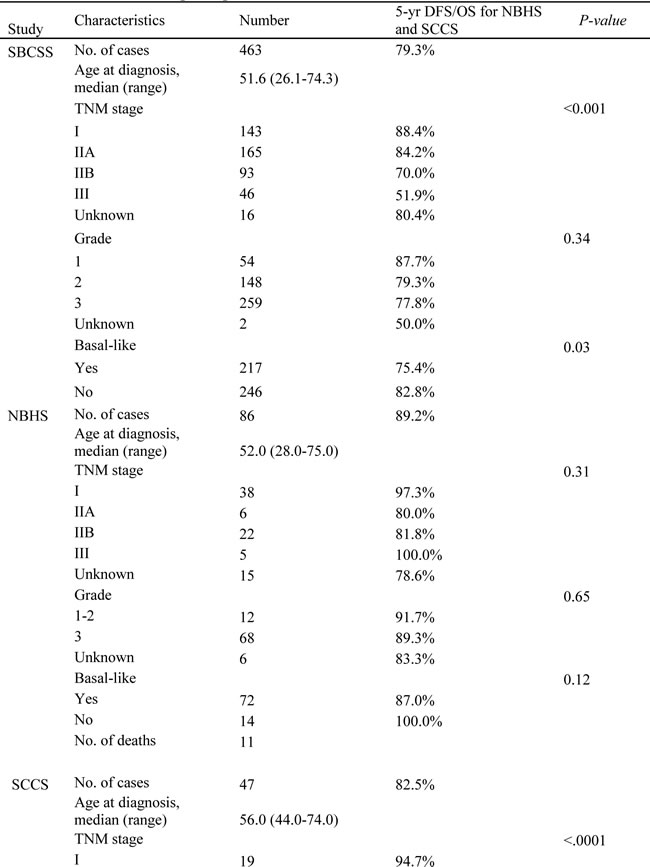
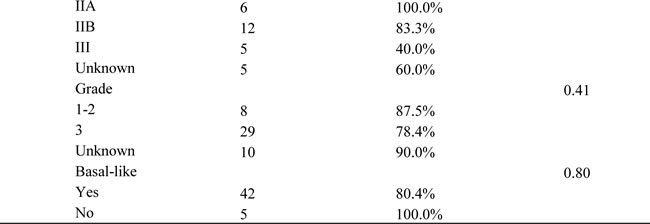
Table 2: Association of ALDH1A1/ALDH1A3 with disease-free survival and overall survival in the SBCSS.
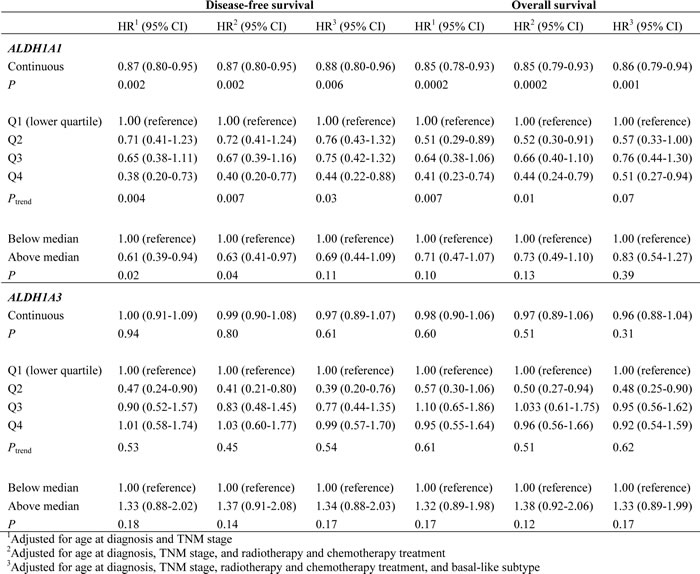
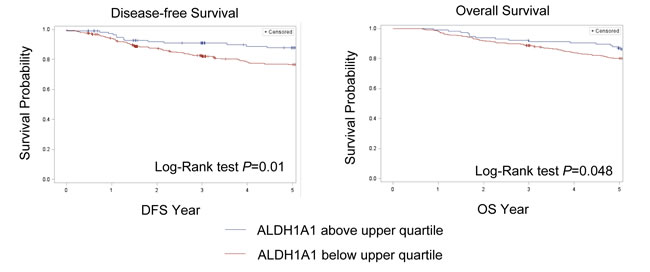
Figure 1: Kaplan-Meier curves. Participants with ALDH1A1 mRNA expression levels below the upper quartile have a significantly worse prognosis compared to participants with ALDH1A1 mRNA expression levels in the upper quartile.
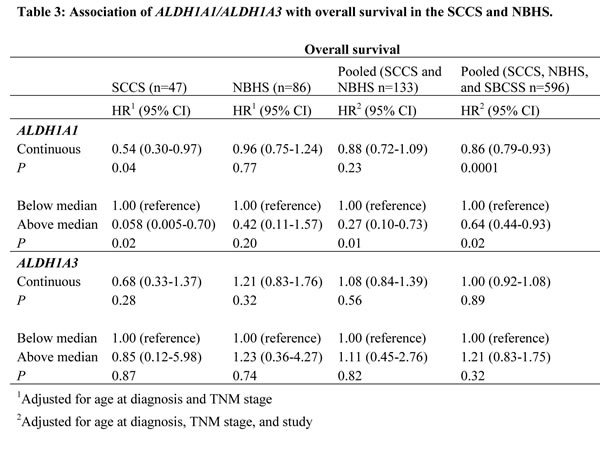
A similar association pattern was observed in the SCCS and NBHS, although not all point estimates were statistically significant, likely due to the small sample sizes of the individual studies (Table 3). When the two studies were combined, ALDH1A1 gene expression level was positively, but not statistically significantly, associated with OS (HR = 0.88, 95% CI: 0.72-1.09). When categorized into two groups, participants with ALDH1A1 expression levels above the median had a reduced risk of OS (HR = 0.27, 95% CI: 0.10-0.73). Similarly, no association of ALDH1A3 gene expression level with OS was observed in the SCCS and NBHS (Table 3). In an analysis of overall survival based on the combined data from all three cohorts, we found that ALDH1A1 gene expression level was statistically significantly associated with OS regardless of whether it was treated as a continuous variable (HR = 0.86, 95% CI: 0.79-0.93) or categorized by median cuts (HR = 0.64, 95% CI: 0.44-0.93). Similarly, we found no association between ALDH1A3 gene expression level and OS in the combined analysis (Table 3). Information on DFS was not available for SCCS and NBHS participants.
Meta-analyses on the association between ALDH1A1/ALDH1A3 gene expression and DFS in 4 publicly-available TNBC datasets with 347 samples were conducted based on continuous variable and median cut points. Again, we found that expression level of the ALDH1A1 gene was positively associated with DFS (HR = 0.86, 95% CI: 0.76-0.97, based on analysis of continuous scale; and HR = 0.58, 95% CI: 0.39-0.85, based on median cut point). ALDH1A3 gene expression level was not associated with DFS (Table 4).
Finally, in the combined data from the SCCS and NBHS studies, we found that TNBC patients had lower expression levels of the ALDH1A1 gene than did non-TNBC patients (Figure 2D).
Table 4: Meta-analysis of the association of ALDH1A1/ALDH1A3 with disease-free survival in four publicly available TNBC datasets.
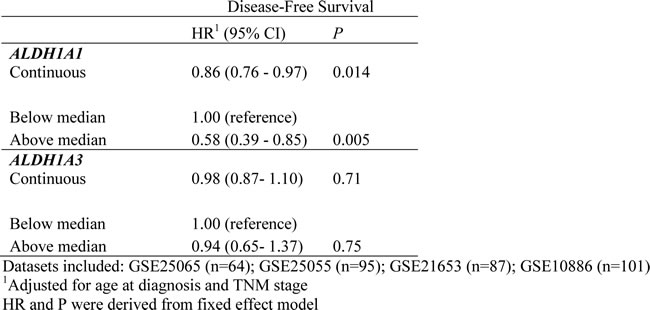
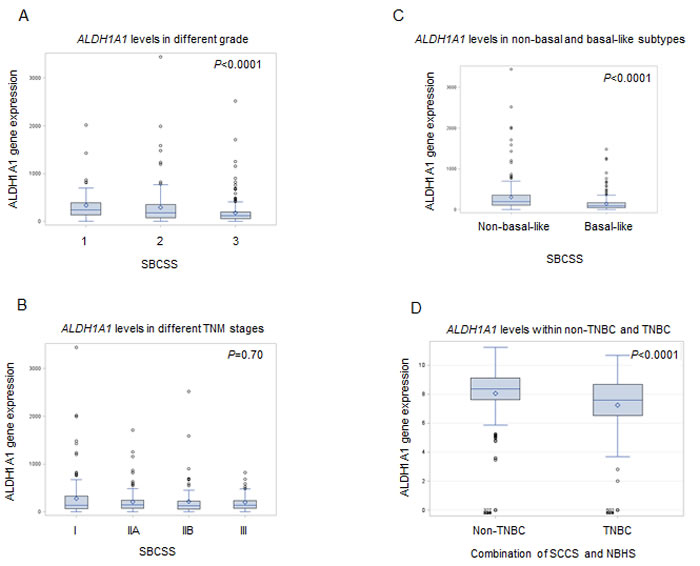
Figure 2: Correlation of ALDH1A1 gene expression with tumor grade A., stage B., basal-like breast cancer subtype C. and TNBC D.
Discussion
TNBC is an aggressive breast cancer subtype with limited treatment options. Identification of new biomarkers for prognosis is urgently needed. It has been suggested that ALDH1 is a biomarker for normal and malignant mammary stem cells [7]. Human mammary epithelial cells with high ALDH activity have stem cell characteristics and have the potential to form tumors in vivo. In a study of 577 cancer tissues of all types of breast cancer combined, ALDH1, detected by immunohistochemical (IHC) staining, was correlated with poorer survival [7]. In our study, we found that ALDH1A1 expression was higher for receptor-positive, low-grade, and non-basal like TNBC tumor tissue. Not taking those clinical predictors into consideration in analysis could result in a false inverse association between ALDH1A1 and cancer prognosis.
ALDH1A1 can inactivate integral agents of chemotherapy; therefore, it has been postulated that breast cancer patients with high ALDH1A1 expression may have an increased risk of recurrence [14]. Two other studies on all types of breast cancer combined also reported that the ALDH1A1 protein was a potential predictive marker of early local tumor recurrence and distant metastasis [9, 10]. However, different results were reported by other studies. These include one study reporting that ALDH1A3, rather than ALDH1A1, contributes to the ALDH activity of cancer stem cells in tissues and cell lines [11], and another study observing no correlation between ALDH and breast cancer stem cells [15]. High expression of ALDH1 in stromal tissues was found to be associated with better DFS and OS in another study [13]. Key limitations of previous studies include not adjusting for confounding factors, such as ER/PR status or TNM stage, and not taking into consideration the positive staining in tumor cells and stromal tissues.
In our study, we sought to evaluate the association between ALDH1A1/ALDH1A3 mRNA expression levels and TNBC outcomes using three population-based cohorts, following an identical lab protocol for each. Total RNA was extracted from breast cancer tissues with at least 80% of the tissues from tumor cells, to reduce the effects of stromal cells on the analyses. We adjusted for age at diagnosis, TNM stage, chemotherapy and radiotherapy treatments, and basal-like breast cancer subtype. Results from these three independent studies demonstrated that high ALDH1A1 gene expression level is associated with reduced breast cancer recurrence and total mortality in patients with TNBC, independent of age at diagnosis and TNM stage. In the SBCSS, additional adjustment for common treatment types and breast cancer basal-like subtypes did not materially alter the observed associations. However, the association for the dichotomized ALDH1 level (by median cut) lost its significance when adjustment for TNBC subtype was made, probably due to reduced statistical power from collapsing the top two quartiles that are associated with different effect sizes. Furthermore, meta-analysis using 4 publicly-available TNBC datasets validated the association between high ALDH1A1 gene expression level and improved DFS. Our results suggest that ALDH1A1 mRNA expression in tumor tissue may be an independent predictor of TNBC recurrence and mortality.
Although we have attempted to include tissue with at least 80% of tumor cells in our study, we cannot completely remove the stromal cells. If cancer stem cells, which presumably have a high mRNA expression level of ALDH1A1 and account for 3-4% of breast cancer cells, are the main source of elevated ALDH1A1 in tumor tissue [7], then the vast majority of ALDH1A1 mRNA measured in our study could still come from stromal cells. Simultaneous measurement of mRNA and protein levels in the same tissue sample would help answer this question and should be considered in the future studies. Other limitations of our study include lack of information on recurrence and the relatively small sample sizes from the NBHS and SCCS. The strengths of our study are its inclusion of multiple independent cohorts and its collection of a large number of TNBC cases. In addition, we adjusted for a wide array of potential confounding factors, including age at diagnosis, TNM stage, chemotherapy and radiotherapy treatments, and basal-like breast cancer subtype.
In summary, our data indicate that tumor tissue ALDH1A1 mRNA expression level may be an independent biomarker of prognosis in TNBC patients.
Materials and methods
Study population
Participants in this study were drawn from three studies: the Shanghai Breast Cancer Survival Study (SBCSS, n = 463), the Nashville Breast Health Study (NBHS, n = 86), and the Southern Community Cohort Study (SCCS, n = 47) (Table 1). Only patients with TNM stage I-III TNBC were included in the present study. A description of the participants has been published elsewhere [16-19].
Briefly, the SBCSS is a population-based cohort study of 5,042 incident breast cancer survivors, aged 20 to 75 years, recruited to the study approximately 6 months following cancer diagnosis [16]. In-person interviews and record linkages were conducted to collect information on demographics, lifestyle factors, clinical characteristics, and disease outcome (recurrence and morality). The demographic and clinical predictors for breast cancer among these participants with TNBC were previously reported [17]. Medical charts from each patient’s initial diagnostic hospital were reviewed to gather information on tumor characteristics (including stage and grade), first-line treatments, and ER/PR status. HER2 status was assessed in the Vanderbilt Molecular Epidemiology Laboratory [17, 20]. Tumor sections were collected from the diagnostic hospitals, resulting in tumor tissue samples from 463 participants being included in the current study.
The NBHS is a population-based, case-control study of incident breast cancer among 2,726 women, aged 25 to 75 years, who were newly diagnosed with primary breast cancer between 2001 and 2011 [18, 21]. Information on demographic, anthropometric, medical, reproductive, and other characteristics was ascertained through telephone interview by trained interviewers using a structured questionnaire. Breast cancer diagnosis information was derived from medical and pathology records, including types and results of diagnostic tests, histopathology, tumor stage, tumor grade, and hormone receptor status. Mortality information was obtained by linkage to the National Death Index through December 31st, 2011. Eighty-six TNBC cases with tissue samples were included in the current study.
The SCCS is a population-based, prospective cohort study of 85,806 participants, aged 40 to 79 years, who were recruited between 2002 and 2009 from 12 southeastern states in the US [19, 22]. Ascertainment of incident breast cancer cases among SCCS participants was obtained through annual linkage of the cohort with the 12 state cancer registries that cover the SCCS catchment area. Mortality information was obtained through linkage with the National Death Index. Information on ER, PR, and HER2 status, as well as first-line treatment, was obtained from these tumor registries. Forty-seven TNBC cases with tissue samples were included in the current study.
Gene expression analysis
Participants’ hematoxylin and eosin (H&E) slides were reviewed by a study pathologist. Tumor tissue was dissected to ensure that samples contained more than 80% tumor cells for RNA extraction [17]. Total RNA was isolated and purified using miRNeasy FFPE Kit (Qiagen, Valencia, CA), and quality and quantity were checked with Nanodrop and an Agilent BioAnalyzer. Expression levels of ALDH1A1 and ALDH1A3 genes were measured as part of a large gene expression effort. A custom-designed nCounter Gene Expression CodeSet profiling of 311 selected gene targets using NanoString nCounter technology was performed following the NanoString standard protocol. Quality control and normalization of gene expression data protocol has been described in detail elsewhere [17]. Briefly, the R package NanoStringNorm (version 1.1.16) was used for quality control and expression normalization with five housekeeping genes (ACTB, RPLP0, MRPL19, SF3A1 and PSMC4). The expression data was log2 transformed. We classified tumors into subgroups most resembling Basal-like, Luminal A, Luminal B, HER2-enriched or Normal-like breast cancer based on PAM50 genes by applying the calling algorithm developed by Parker et al. [17].
Statistical analysis
Outcomes of the study were defined as recurrence/breast cancer-specific mortality (disease-free survival: DFS) and/or all-cause mortality (overall survival: OS). (Note: recurrence is not collected in NBHS or SCCS, and therefore DFS can only be investigated in the SBCSS.) Event-free participants were censored at the date of last follow-up. The associations between ALDH1A1 expression levels and DFS and OS were evaluated using a Cox regression model with adjustment for age at diagnosis, TNM stage, chemotherapy treatment, radiotherapy treatment, and basal-like breast cancer subtype in the SBCSS and in other studies whenever available. The Kruskal-Wallis test was used to compare ALDH1A1 mRNA expression among different tumor grades and stages. The Wilcoxon-Mann-Whitney test was used to analyze ALDH1A1 mRNA expression in patients with TNBC/non-TNBC and patients with basal-like TNBC and non-basal-like TNBC.
In addition, 4 publicly-available TNBC microarray datasets: GSE25065 (n = 64) [23], GSE25055 (n = 95) [23], GSE21653 (n = 87) [24], and a combined data set (n = 101) of GSE10886, GSE6128, GSE3165 and GSE3521 [25-28], were included in the meta-analysis. Original gene expression data were log2 transformed. Clinical data and gene expression data were obtained from publicly-available data sets from previous publications and from the Gene Expression Omnibus (GEO) deposited at the National Center for Biotechnology Information (NCBI). To ensure that we derived only high-quality survival data sets from the published breast cancer studies, we applied the “rule of fifty” [29-31] as an inclusion criterion. Specifically, to be included in our study, each dataset was required to have at least 50 TNBC samples with survival data and a minimum of 10 events, as well as 60% or more of its samples with survival information. In total, 10 datasets with molecular subtype information and survival information were identified from the NCBI database. Among them were 3 independent data sets, GSE21653 [24], GSE25055 [23], GSE25065 [23], and 1 combined data set of GSE10886 [25], GSE6128 [26], GSE3165 [27], and GSE3521 [28] that met our study criteria and were thus included in our first-stage screening analysis. Cox regression models were used to derive hazard ratios (HRs) for breast cancer recurrence/breast cancer-specific mortality in association with each mRNA, with adjustment for age at diagnosis and TNM stage. The mRNAs, including ALDH1A1 and ALDH1A3, from the 3 independent data sets and the combined data set, were used as exposure factors in the Cox model, respectively. Each gene was categorized into two categories: < median (reference) and ≥median. Four HRs from each mRNA were used in the meta-analyses. The weighted average HR was calculated using an inverse variance of each HR as the weight. All tests were two-tailed with a significance level of P < 0.05. All analyses were performed using SAS statistical software (version 9.3; SAS Institute Inc. NC).
Acknowledgments
We thank Regina Courtney and Bo Huang for their help with RNA sample preparation. Sample preparation was conducted at the Survey and Biospecimen Shared Resources, which is supported in part by the Vanderbilt-Ingram Cancer Center (P30CA068485). NanoString nCounter assays were conducted at the University of Miami Sylvester Comprehensive Cancer Center. Michelle Baglia is funded by a CTSA TL1 fellowship (award No. TL1TR000447).
Funding
This study was supported by grants from the US Department of Defense (DOD) Breast Cancer Research Program (DAMD 17-02-1-0607 to X.-O. Shu), and the National Institutes of Health (NIH; R01 CA118229 to X.-O. Shu, P50CA098131 to C. Arteaga). Dr. Yan Liu was supported by the Vanderbilt Molecular and Genetic Epidemiology of Cancer training program (R25CA160056). Ms. Michelle Baglia is supported by CTSA award TL1TR000447 from the National Center for Advancing Translational Sciences.
Conflicts of Interests
The authors declare no conflict of interests.
References
1. Vasiliou V, Nebert DW. Analysis and update of the human aldehyde dehydrogenase (ALDH) gene family. Hum Genomics. 2005; 2:138–43.
2. Marchitti SA, Brocker C, Stagos D, Vasiliou V. Non-P450 aldehyde oxidizing enzymes: the aldehyde dehydrogenase superfamily. Expert Opin Drug Metab Toxicol. 2008; 4:697–720.
3. Crabb DW, Matsumoto M, Chang D, You M. Overview of the role of alcohol dehydrogenase and aldehyde dehydrogenase and their variants in the genesis of alcohol-related pathology. Proc Nutr Soc. 2004; 63:49–63.
4. Avigdor A, Goichberg P, Shivtiel S, Dar A, Peled A, Samira S, Kollet O, Hershkoviz R, Alon R, Hardan I, Ben-Hur H, Naor D, Nagler A, Lapidot T. CD44 and hyaluronic acid cooperate with SDF-1 in the trafficking of human CD34+ stem/progenitor cells to bone marrow. Blood. 2004; 103:2981–9.
5. Oswald J, Boxberger S, Jørgensen B, Feldmann S, Ehninger G, Bornhäuser M, Werner C. Mesenchymal stem cells can be differentiated into endothelial cells in vitro. Stem Cells Dayt Ohio. 2004; 22:377–84.
6. Patrawala L, Calhoun T, Schneider-Broussard R, Li H, Bhatia B, Tang S, Reilly JG, Chandra D, Zhou J, Claypool K, Coghlan L, Tang DG. Highly purified CD44+ prostate cancer cells from xenograft human tumors are enriched in tumorigenic and metastatic progenitor cells. Oncogene. 2006; 25:1696–708.
7. Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG, Liu S, Schott A, Hayes D, Birnbaum D, Wicha MS, Dontu G. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007; 1:555–67.
8. Charafe-Jauffret E, Ginestier C, Bertucci F, Cabaud O, Wicinski J, Finetti P, Josselin E, Adelaide J, Nguyen T-T, Monville F, Jacquemier J, Thomassin-Piana J, Pinna G, Jalaguier A, Lambaudie E, Houvenaeghel G, Xerri L, Harel-Bellan A, Chaffanet M, Viens P, Birnbaum D. ALDH1-positive cancer stem cells predict engraftment of primary breast tumors and are governed by a common stem cell program. Cancer Res. 2013; 73:7290–300.
9. Zhong Y, Lin Y, Shen S, Zhou Y, Mao F, Guan J, Sun Q. Expression of ALDH1 in breast invasive ductal carcinoma: an independent predictor of early tumor relapse. Cancer Cell Int. 2013; 13:60.
10. Zhong Y, Shen S, Zhou Y, Mao F, Guan J, Lin Y, Xu Y, Sun Q. ALDH1 is a better clinical indicator for relapse of invasive ductal breast cancer than the CD44+/CD24- phenotype. Med Oncol Northwood Lond Engl. 2014; 31:864.
11. Marcato P, Dean CA, Pan D, Araslanova R, Gillis M, Joshi M, Helyer L, Pan L, Leidal A, Gujar S, Giacomantonio CA, Lee PWK. Aldehyde dehydrogenase activity of breast cancer stem cells is primarily due to isoform ALDH1A3 and its expression is predictive of metastasis. Stem Cells Dayt Ohio. 2011; 29:32–45.
12. Tan EY, Thike AA, Breast Surgical Team at Outram, Tan PH. ALDH1 expression is enriched in breast cancers arising in young women but does not predict outcome. Br J Cancer. 2013; 109:109–13.
13. Resetkova E, Reis-Filho JS, Jain RK, Mehta R, Thorat MA, Nakshatri H, Badve S. Prognostic impact of ALDH1 in breast cancer: a story of stem cells and tumor microenvironment. Breast Cancer Res Treat. 2010; 123:97–108.
14. Khoury T, Ademuyiwa FO, Chandrasekhar R, Chandraseekhar R, Jabbour M, Deleo A, Ferrone S, Wang Y, Wang X. Aldehyde dehydrogenase 1A1 expression in breast cancer is associated with stage, triple negativity, and outcome to neoadjuvant chemotherapy. Mod Pathol Off J U S Can Acad Pathol Inc. 2012; 25:388–97.
15. Liu Y, Nenutil R, Appleyard MV, Murray K, Boylan M, Thompson AM, Coates PJ. Lack of correlation of stem cell markers in breast cancer stem cells. Br J Cancer. 2014; 110:2063–71.
16. Shu XO, Zheng Y, Cai H, Gu K, Chen Z, Zheng W, Lu W. Soy food intake and breast cancer survival. JAMA J Am Med Assoc. 2009; 302:2437–43.
17. Baglia ML, Cai Q, Zheng Y, Wu J, Su Y, Ye F, Bao P-P, Cai H, Zhao Z, Balko J, Zheng W, Lu W, Shu X-O. Dual specificity phosphatase 4 gene expression in association with triple-negative breast cancer outcome. Breast Cancer Res Treat. 2014; 148:211–20.
18. Han M-R, Deming-Halverson S, Cai Q, Wen W, Shrubsole MJ, Shu X-O, Zheng W, Long J. Evaluating 17 breast cancer susceptibility loci in the Nashville breast health study. Breast Cancer. 2015; 22:544-51.
19. Signorello LB, Hargreaves MK, Blot WJ. The Southern Community Cohort Study: investigating health disparities. J Health Care Poor Underserved. 2010; 21:26–37.
20. Su Y, Zheng Y, Zheng W, Gu K, Chen Z, Li G, Cai Q, Lu W, Shu XO. Distinct distribution and prognostic significance of molecular subtypes of breast cancer in Chinese women: a population-based cohort study. BMCCancer. 2011; 11:292.
21. Zheng W, Cai Q, Signorello LB, Long J, Hargreaves MK, Deming SL, Li G, Li C, Cui Y, Blot WJ. Evaluation of 11 breast cancer susceptibility loci in African-American women. Cancer EpidemiolBiomarkers Prev. 2009; 18:2761–4.
22. Signorello LB, Hargreaves MK, Steinwandel MD, Zheng W, Cai Q, Schlundt DG, Buchowski MS, Arnold CW, McLaughlin JK, Blot WJ. Southern community cohort study: establishing a cohort to investigate health disparities. J Natl Med Assoc. 2005; 97:972–9.
23. Hatzis C, Pusztai L, Valero V, Booser DJ, Esserman L, Lluch A, Vidaurre T, Holmes F, Souchon E, Wang H, Martin M, Cotrina J, Gomez H, Hubbard R, Chacón JI, Ferrer-Lozano J, Dyer R, Buxton M, Gong Y, Wu Y, Ibrahim N, Andreopoulou E, Ueno NT, Hunt K, Yang W, Nazario A, DeMichele A, O’Shaughnessy J, Hortobagyi GN, Symmans WF. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA. 2011; 305:1873–81.
24. Sabatier R, Finetti P, Cervera N, Lambaudie E, Esterni B, Mamessier E, Tallet A, Chabannon C, Extra J-M, Jacquemier J, Viens P, Birnbaum D, Bertucci F. A gene expression signature identifies two prognostic subgroups of basal breast cancer. Breast Cancer Res Treat. 2011; 126:407–20.
25. Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, Davies S, Fauron C, He X, Hu Z, Quackenbush JF, Stijleman IJ, Palazzo J, Marron JS, Nobel AB, Mardis E, Nielsen TO, Ellis MJ, Perou CM, Bernard PS. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009; 27:1160–7.
26. Hoadley KA, Weigman VJ, Fan C, Sawyer LR, He X, Troester MA, Sartor CI, Rieger-House T, Bernard PS, Carey LA, Perou CM. EGFR associated expression profiles vary with breast tumor subtype. BMC Genomics. 2007; 8:258.
27. Herschkowitz JI, Simin K, Weigman VJ, Mikaelian I, Usary J, Hu Z, Rasmussen KE, Jones LP, Assefnia S, Chandrasekharan S, Backlund MG, Yin Y, Khramtsov AI, et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol. 2007; 8:R76.
28. Hu Z, Fan C, Livasy C, He X, Oh DS, Ewend MG, Carey LA, Subramanian S, West R, Ikpatt F, Olopade OI, van de Rijn M, Perou CM. A compact VEGF signature associated with distant metastases and poor outcomes. BMC Med. 2009; 7:9.
29. Hsieh FY, Lavori PW. Sample-size calculations for the Cox proportional hazards regression model with nonbinary covariates. Control Clin Trials. 2000; 21:552–60.
30. Shih JH. Sample size calculation for complex clinical trials with survival endpoints. Control Clin Trials. 1995; 16:395–407.
31. Cantor AB. Sample size calculations for the log rank test: a Gompertz model approach. J Clin Epidemiol. 1992; 45:1131–6.

