INTRODUCTION
The budding Saccharomyces cerevisiae is the most scientifically and industrially exploited species among the Saccharomyces sensu stricto complex as it is widely used as a model organism and in the fermentation processes such as the production of food and alcoholic beverages [1, 2]. There are at least seven natural Saccharomyces sensu stricto species (S. cerevisiae, S. paradoxus, S. mikatae, S. kudriavzevii, S. arboricola, S. eubayanus and S. uvarum) and numerous related industrial hybrids of a biotechnological interest (e.g., S. cerevisiae x S. kudriavzevii, S. pastorianus, S. bayanus, S. cerevisiae x S. mikatae) [1, 3-11]. More recently, S. paradoxus has been also established as a main yeast component in Croatian wines that may suggest a potentially important enological characteristics for this species [12].
The domestication within the Saccharomyces sensu stricto complex has led to the evolution of special phenotypic features via hybridization, polyploidization, gene duplication and gene transfer [2]. The best example of how fermentative conditions can shape the yeast genome is the acquiring SSU1-R allele-based resistance to sulfite by wine yeasts [13]. This adaptation is a result of a reciprocal translocation between chromosomes VIII and XVI due to unequal crossing-over mediated by microhomology between very short sequences on the 5’ upstream regions of the SSU1 and ECM34 genes that provokes the induction of the SSU1 transporter and increases the ability of yeast cells to expulse sulfite from the cytoplasm [13]. This genetic change can be found in 50% of the wine strains, whereas it has not been observed among wild strains suggesting that the use for millennia of sulfite as a preservative in wine production could have favored its selection [14].
In contrast to the best studied genomes of wine and brewing yeast strains, the information on genetic and genomic diversity of yeast isolates involved in the production of distilled spirits is limited. In the present study, array-CGH-based genome-wide analysis of twenty two commercially available distillery yeasts was conducted. We have revealed four groups with different pattern of the gene copy number variants that in the case of the YRF1 gene dosage diversity may provoke changes in genetic stability.
RESULTS
Electrophoretic karyotyping of distillery yeasts reveals four species categories
As there are limited number of published data on genomic and genetic characteristics of distillery yeasts [15, 16], the karyotype and the genome of, commercially available and widely used in food industry, twenty two distillery yeast strains were comprehensively investigated (Table 1).
On the basis of PFGE separation (electrophoretic karyotyping), one can conclude that all yeasts examined belonging to the Saccharomyces sensu stricto complex [17]. In general, the chromosome number of analyzed yeasts is 16 (Figure 1). However, an additional band was observed between chromosomes IV and VII in strains from 1 to 6 and strain 16 and between chromosomes I and VI in strain 19 (Figure 1). In almost all strains examined, chromosomes IV and XII migrated together (Figure 1).
The strains from 1 to 6 and strain 16 had the S. bayanus-like chromosome pattern, whereas strains 7, 8, 10, 11, 12, 13, 14, 15, 17, 18, 20, 21 and 22 were classified as S. paradoxus, strain 9 as S. cerevisiae and strain 19 as S. kudriavzevii (Figure 1). A chromosomal band of about 1300 kb (between chromosomes IV and VII) observed in strains from 1 to 6 and strain 16 is a characteristic feature of S. bayanus karyotype [18]. Chromosome similarity between analyzed strains was also further evaluated using UPGMA clustering (Figure 1). Strains from 2 to 6 were the most similar within assigned S. bayanus group, whereas strains 1 and 16 differed from other S. bayanus strains (Figure 1). Similarly, strains 21 and 22 were more distant from other S. paradoxus strains (Figure 1).
Table 1: Distillery yeast strains used in this study.
No. |
Trade name |
Company |
1 |
Samogon turbo |
CBF Drinkit |
2 |
Superyeast T48 Dual Use |
CBF Drinkit |
3 |
Spiritferm Extreme 8 kg Turbo |
Spiritferm |
4 |
Spiritferm T3 |
Spiritferm |
5 |
Spiritferm turbo fruit |
Spiritferm |
6 |
Spiritferm Moskva style |
Spiritferm |
7 |
Coobra 24 Snabbsats |
CBF Drinkit |
8 |
Coobra 6 Magnum Snabbsats |
Vendor |
9 |
Coobra 8 Snabbsats |
Vendor |
10 |
Coobra 48 Turbo Yeast |
CBF Drinkit |
11 |
Coobra RUM YEAST |
CBF Drinkit |
12 |
Double Snake Turbo Yeast C3 Extra |
Hambleton Bard Ltd. |
13 |
Alcotec Pure Turbo Super Yeast 48 |
Hambleton Bard Ltd. |
14 |
Drożdże gorzelnicze Turbo 72h |
BIOWIN |
15 |
Black Bull Turbo Yeast |
Avedore Trading |
16 |
Gozdawa 1410 Turbo |
Gozdawa |
17 |
Superyeast T Vodka Star |
CBF Drinkit |
18 |
Alcotec Vodka Star Turbo Yeast |
Hambleton Bard Ltd. |
19 |
Alcotec Single Strain Whisky with Amyloglucosidase |
Hambleton Bard Ltd. |
20 |
Fermiol drożdże gorzelnicze |
BIOWIN/FERMIOL |
21 |
BIOWIN Turbo Super Yeast 48h |
BIOWIN |
22 |
Alcotec Pure Turbo Super Yeast 24h |
Hambleton Bard Ltd. |
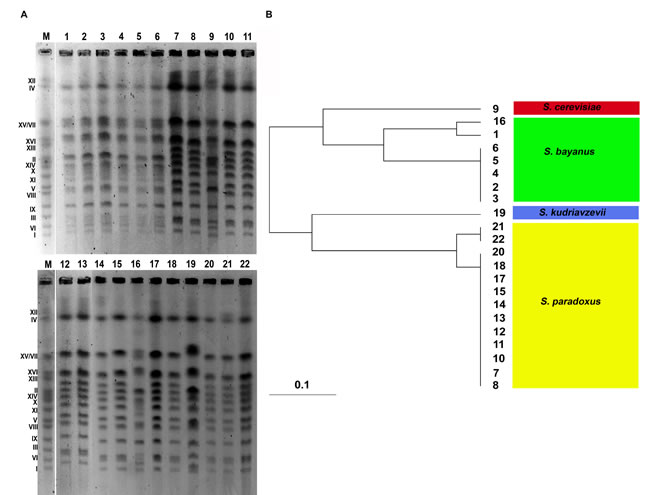
Figure 1: Electrophoretic karyotyping of twenty two distillery yeast strains (A, lanes from 1 to 22). The yeast S. cerevisiae chromosome marker YNN295 (BIORAD) is shown (A., lane M). The dendrogram of chromosome band-based similarity is also presented B. The species classification within the Saccharomyces sensu stricto complex is provided.
Distillery yeasts are diploid
The ploidy of distillery strains was then analyzed using fluorescence-activated cell sorting (FACS) (Figure 2).
We found that all strains used were diploid when compared to reference laboratory yeast cells with known ploidy (haploid, diploid, triploid and tetraploid cells) (Figure 2).
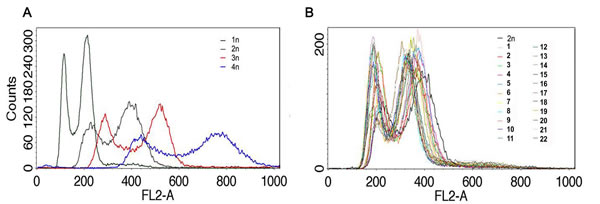
Figure 2: The ploidy analysis. Fluorescence-activated cell sorting (FACS)-based analysis of DNA content of distillery strains B. Haploid, diploid, triploid and tetraploid reference strains are also shown A.
The diversity of gene copy number and loci-specific gains and losses involve mainly the subtelomeric regions
After electrophoretic karyotyping, the genome of distillery strains was characterized using array-based comparative genomic hybridization (array-CGH) (Figure 3).
The analysis of array-CGH profiles revealed variabilities in the gene copy number exclusively within the subtelomeric regions of all analyzed chromosomes and two short intrachromosomal regions of chromosomes IV and XII (Figures 3A and 4).
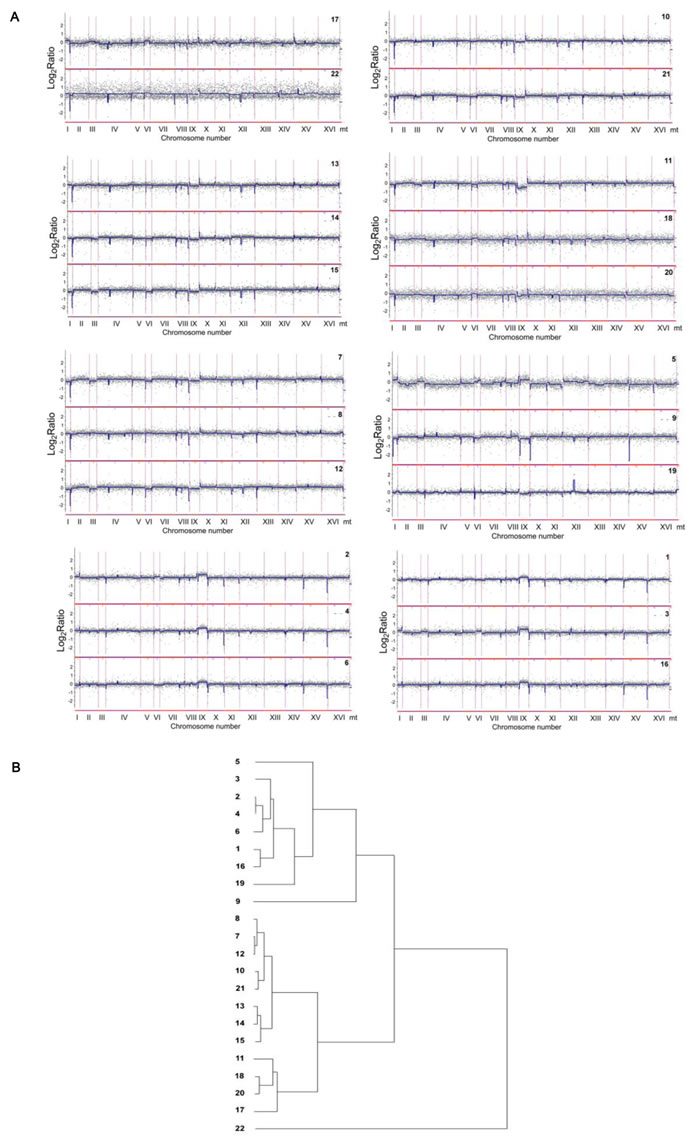
Figure 3: Comparison of the gene copy number between analyzed distillery yeasts using array-CGH. A. The strains with similar array-CGH profiles were grouped together. Each grey dot represents the value of the log2 ratio for an individual gene. Blue lines were provided to emphasize the most accented differences (DNA losses and gains). B. The relatedness of distillery strains as determined by cluster analysis. Similarity tree is shown (see Materials and Methods section for the details).
The differences between strains were more accented including the losses and/or gains of fragments of chromosomes I, III, VI and IX, and in the case of strain 5 also the changes within chromosome XII (Figure 3A). The gain of chromosomes I and VI in strains 3, 5 and 17, and the loss of chromosomes I and VI in strains 7, 9, 11, 12 and 15 were revealed (Figure 3A). The gain of chromosome III in strains 3, 5 and 17, and the loss of chromosome III in strains 7, 9, 11, 12, 14, 15 and 21 were observed (Figure 3A). The most variable chromosome was chromosome IX. The gains of chromosome IX were shown in strains 1, 2, 3, 4, 5, 6 and 16, whereas the losses of chromosome IX were documented in strains 7, 9, 11, 12, 15 and 19 (Figure 3A). The gains of chromosome XII was exclusively reported in strain 5. Interestingly, small chromosomes were frequently affected and changes in one small chromosome were accompanied by changes in other small chromosomes. However, these gains and losses were too small to be interpreted as duplications or deletions of chromosomal regions or whole chromosome aneuploidy events within the whole population of particular strain. Perhaps, the chromosome variations may suggest the cellular heterogeneity within a population. Additionally, array-CGH profiles were used to estimate the level of similarity (relatedness) between distillery strains on the basis of observed diversity in subtelomeric regions and chromosome IX (Figure 3B). Array-CGH-based relationships between analyzed strains were comparable with electrophoretic karyotyping-based relationships (Figures 1 and 3B). The strains from 1 to 6 and strain 16 already classified as S. bayanus (Figure 1) were clustered together (Figure 3B). According to both similarity analyses used, strains 2, 4 and 6, and strains 1 and 16 were closely located (Figures 1 and 3B). The strains belonging to S. paradoxus species (Figure 1), were grouped into several categories using array-CGH-based analysis, namely the group of the strains 7, 8 and 12; 10 and 21; 13, 14 and 15; 11, 17, 18 and 20 (Figure 3B). The most variable was strain 22 (S. paradoxus species, Figure 1) with its own category (Figure 3B).
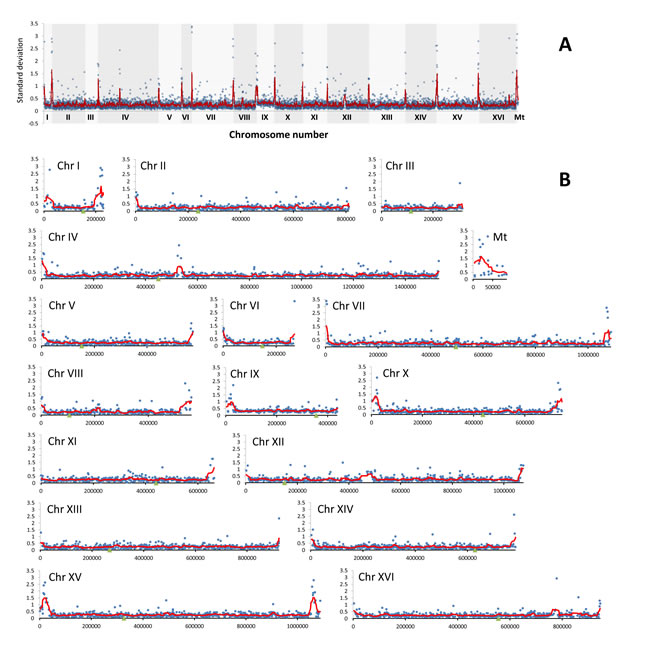
Figure 4: The divergence of relative abundance of genes as determined by array-CGH analysis represented by standard deviation (SD) of log2 ratio values for each gene in all analyzed strains. A. The summary plot for the whole genome. B. Individual plots for each chromosome. Blue dots indicate the SD values for individual genes, the red line denotes the smoother trend calculated by moving average of SD values to expose the genome regions of higher log2 ratio divergence and green triangles indicate centromere position.
Gene ontology overrepresentation profiles are species-specific
As the observed differences in the gene copy number and loci-specific gains and losses may affect the functional properties of distillery strains, the genes that were most divergent according to array-CGH-based analysis were then subjected to gene ontology overrepresentation analysis (Figure 5).
The selected gene-set consisted of 257 genes, for which, in at least one strain, the log2 ratio value was greater than four standard deviations of log2 ratio calculated for all genes in all strains. Six functional categories overrepresented in the group of selected genes were revealed, namely 1) telomere maintenance via recombination, DNA helicase activity or DNA binding; 2) maltose metabolism process, glucose transmembrane transporter activity; 3) asparagine catabolism, cellular response to nitrogen starvation, localized in cell wall-bounded periplasmic space; 4) siderophore transport; 5) response to copper ion, cadmium ion binding and 6) L-iditol 2- dehydrogenase activity (p < 0.05) and are presented as a heat map in Figure 5. Species-dependent variability in the gene copy number within functional categories of selected genes were revealed, e.g., similar genetic features were observed among strains belonging to S. bayanus species that differed from genetic features in the strains of S. paradoxus species (Figure 5). Moreover, strains 9 (S. cerevisiae) and 19 (S. kudriavzevii) had their own overrepresentation profiles (Figure 5). Interestingly, within functional category of genes involved in the telomere maintenance via recombination, DNA helicase activity or DNA binding, the gains of YRF1 genes (helicases encoded by the Y’ element of subtelomeric regions) were exclusively shown in the S. bayanus strain group and strain 19 (S. kudriavzevii), whereas the losses of YRF1 genes were observed in the S. paradoxus strain group (Figure 5). A heat map generated from array-CGH data reflecting the variability in the gene copy number of the whole genome of all analyzed distillery strains is also presented in Supplementary File 1.
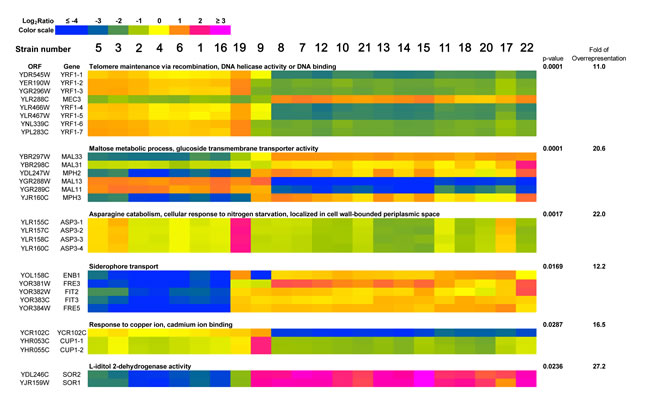
Figure 5: A heat map generated from array-CGH data. Functional categories overrepresented in the group of genes that were the most divergent among analyzed strains are shown. The strains were ordered according to the result of clustering analysis (Figure 3B) and the selected genes were grouped according to their functional assignment. Positive and negative log2 ratio values represent higher and lower than average abundance of the gene, as determined by array-CGH analysis (see Materials and Methods section for the details).
The YRF1 gene copy number corresponds to the presence of Y’ telomeric sequences
Since array-CGH-based analysis revealed that the majority of genomic differences can be found in subtelomeric regions of the genome of distillery strains and Y’ element ATP-dependent helicase activity may be affected in the opposite direction in the S. bayanus and S. paradoxus strain groups (Figures 3, 4 and 5), we then evaluated the presence of Y’ telomeric sequences in all examined strains (Figure 6).
Y’ telomeric sequences were the most accented in the S. bayanus strain group, whereas they were marginally noticeable in the S. paradoxus strain group (Figure 6). Southern blot data using Y’ telomeric probes are in agreement with array-CGH results (Figure 5). The same relationship was observed for strain 19 (S. kudriavzevii) with the highest log2 ratios of YRF1 genes (Figure 5) and rich in Y’ telomeric sequences (Figure 6).
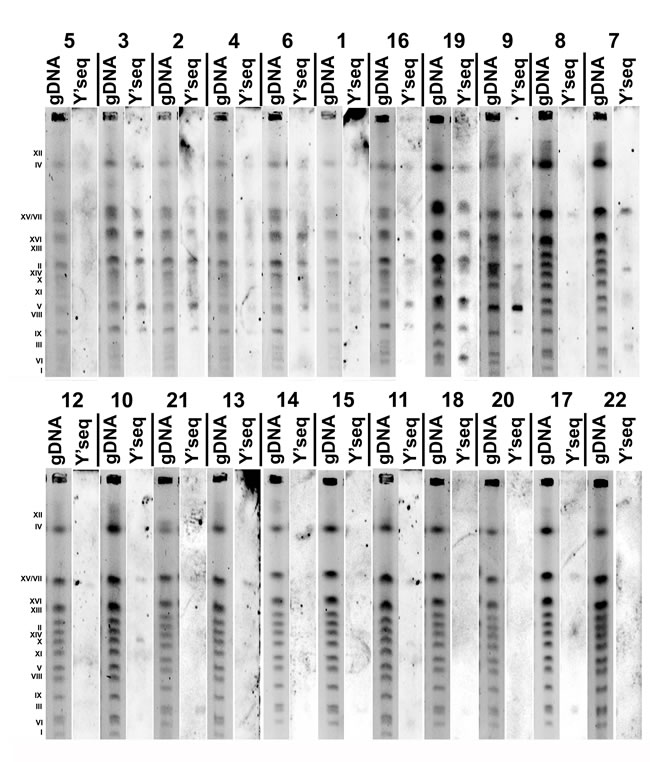
Figure 6: The presence of Y’ telomeric sequences in twenty two distillery yeasts (categories from 1 to 22, lane gDNA: chromosome pattern of an individual strain, lane Y’ seq: Y’ telomeric sequences) detected using Southern blot using Y’ telomeric sequence probes.
The YRF1 gene copy number modulates genetic stability
We hypothesized that altered Y’ telomeric sequence-dependent helicase activity may modulate genetic stability in distillery strains. Thus, we also evaluated the strain-dependent susceptibility to DNA damage (Figures 7 and 8).
Indeed, the S. bayanus strain group (p < 0.001) and strains 9 (S. cerevisiae) (p < 0.001) and 19 (S. kudriavzevii) with the abundance of Y’ telomeric sequences and higher number of YRF1 gene copies were less affected by DNA double strand breaks (DSBs) and DNA single strand breaks (SSBs) than the S. paradoxus strain group (Figure 7). Moreover, the level of oxidative DNA damage (8-hydroxy-2’-deoxyguanosine, 8-oxo-dG, content) was increased in the S. paradoxus group compared to the S. bayanus group (Figure 8). However, the effect was statistically insignificant. The intracellular production of reactive oxygen species (ROS) was also elevated in the S. paradoxus group (p < 0.001) but no clear-cut relationship between ROS production and the 8-oxo-dG level was observed in this group, e.g., strains 21 and 22 with the most imbalanced redox equilibrium were characterized by relatively low level of 8-oxo-dG (Figure 8). Thus, it might not be concluded that the elevation in 8-oxo-dG level was a result of increased ROS production in the S. paradoxus group.
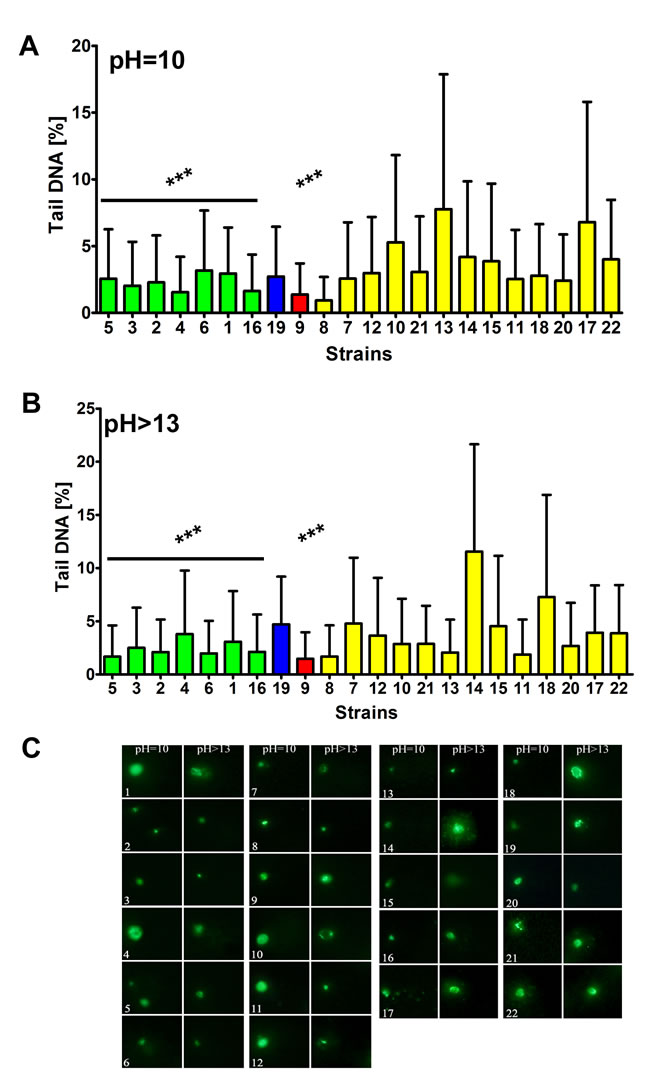
Figure 7: The susceptibility to DNA double strand breaks (DSBs) A. and DNA single strand breaks (SSBs) B. DSBs and SSBs were assessed using neutral and alkaline comet assay, respectively. The strains belonging to the same species were grouped together and the data were marked in different colors (S. bayanus: green, S. kudriavzevii: blue, S. cerevisiae: red and S. paradoxus: yellow). As a DNA damage marker, the % tail DNA was used. The bars indicate SD, n = 150, ***p < 0.001 compared to the S. paradoxus group (ANOVA and Tukey’s a posteriori test). C. The typical micrographs are shown. DNA was visualized using YOYO-1 staining (green).
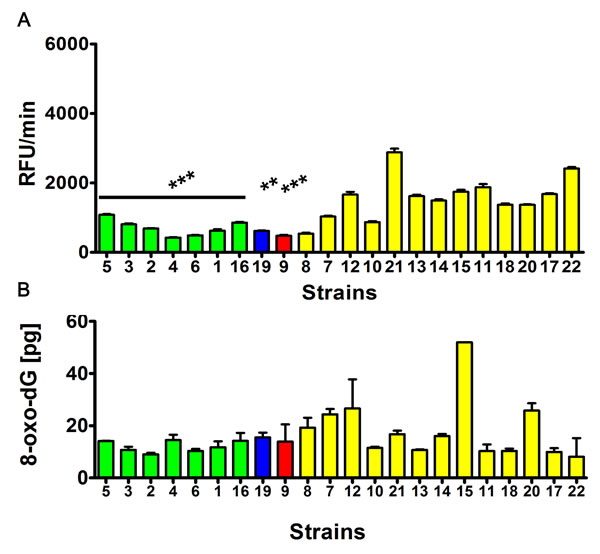
Figure 8: The intracellular reactive oxygen species (ROS) production A. and the level of oxidative DNA damage (8-hydroxy-2’-deoxyguanosine, 8-oxo-dG, level) B. ROS production was assessed using H2DCF-DA fluorogenic probe and the results are presented as relative fluorescence units per minute (RFU/min). The level of 8-oxo-dG was analyzed using ELISA-based assay. The strains belonging to the same species were grouped together and the data were marked in different colors (S. bayanus: green, S. kudriavzevii: blue, S. cerevisiae: red and S. paradoxus: yellow). The bars indicate SD, n = 5, ***p < 0.001, **p < 0.01 compared to the S. paradoxus group (ANOVA and Tukey’s a posteriori test).
DISCUSSION
This is the first report on detailed evaluation of genomic features of twenty two distillery yeast strains used in food industry to produce distilled spirits such as vodka and whisky. To date, one paper has been published on molecular genetic characteristics of thirty six distillery yeast strains belonging to the S. cerevisiae species [15]. The authors performed PCR-RFLP analysis of rDNA 5.8S-ITS fragment, molecular karyotyping (PFGE separation), and Southern blot-based detection of MAL, SUC and MEL genes [15]. Analyzed strains were aneuploid and rich in polymeric genes SUC and MAL important for sucrose and maltose fermentation, respectively [15]. As we have purchased the strains from multiple suppliers, we are aware that our analyzed “distillery group” may be more heterogeneous. Indeed, the strains examined in the present study were more diverse and belonged to four species of the Saccharomyces sensu stricto complex, namely S. bayanus (n = 7), S. paradoxus (n = 13), S. cerevisiae (n = 1) and S. kudriavzevii (n = 1) according to electrophoretic karyotyping. The obtained species-specific chromosome patterns were in agreement with previously reported data on karyotypic characteristics of reference yeast strains [9, 17-19]. Similar chromosome profiles were observed within the S. bayanus group (strains 1 to 6 and strain 16). However, one should remember that some karyotypic variants may also occur within the yeast species. This is particularly true for the S. bayanus group [6, 19]. S. bayanus var. uvarum isolates are typically characterized by only two small chromosomal bands in the range of 245-370 kb (between chromosomes I and III) instead of three or more in S. bayanus var. bayanus [6, 19]. The strains assigned to the S. bayanus group (this study) exhibited karyotypic features of the S. bayanus var. bayanus. In general, the analyzed strains were diploid but aneuploid events (the presence of some additional chromosome bands) were also observed. It is widely accepted the industrially relevant yeast strains, e.g., brewer’s and wine yeasts, are aneuploid with disomies, trisomies and tetrasomies [20, 21]. Alloploidy is also a common phenomenon [1]. The best and most well-known example of industrial hybrid is the lager yeast S. pastorianus (syn. S. carlsbergensis), which is the cold-adapted S. cerevisiae x S. eubayanus allotetraploid [22]. Under certain conditions, e.g., during fermentation-associated biotic and abiotic stresses, aneuploidy events and changes in the ploidy may be adaptive and advantageous by increasing the number of copies of beneficial genes or by protecting the yeasts against recessive lethal or deleterious mutations that may confer resistance to low temperature or high ethanol levels [20, 23].
Genome-wide array-CGH analysis reveals variations in the gene copy number almost exclusively in the subtelomeric regions of the genome of distillery yeasts, and the most affected chromosomes were the chromosome I, III, VI and IX. It is worthwhile to note that the strain relatedness based on array-CGH data was comparable with electrophoretic karyotyping-based similarities among strains. Statistically significant differences in the gene dosage were observed in six functional gene categories, namely 1) telomere maintenance via recombination, DNA helicase activity or DNA binding, 2) maltose metabolism process, glucose transmembrane transporter activity; 3) asparagine catabolism, cellular response to nitrogen starvation, localized in cell wall-bounded periplasmic space, 4) siderophore transport, 5) response to copper ion, cadmium ion binding and 6) L-iditol 2- dehydrogenase activity. The effects were species-dependent that may suggest that strains within distillery group analyzed may differently respond to changing environments and may have diverse adaptation strategies. Surprisingly, in almost all gene categories, the effects observed in the S. bayanus and S. paradoxus groups were opposite, e.g., increased and decreased copy number of YRF1 genes (YRF1-1 to YRF1-7) in the S. bayanus and S. paradoxus group was shown, respectively. The YRF1 genes (YRF1-1 to YRF1-7) are localized on different yeast chromosomes within the Y’ element of subtelomeric regions and encoded Y’ element ATP-dependent helicase (Y’-Help1, Y’-HELicase Protein 1) implicated in telomerase-independent telomere maintenance [24]. In laboratory yeasts, Y’-Help1 is highly induced in the survivors of telomerase deficient cells [24]. It has been speculated that Y’-Help1 may enhance homologous DNA recombination among Y’ elements and, as a consequence, may induce Y’ amplification to prevent chromosomal loss and cell death [24]. We hypothesized that altered YRF1 gene copy number and the presence of Y’ elements may affect genetic stability in distillery strains. Indeed, the strains from the S. paradoxus group with decreased YRF1 gene dosage and the lack of Y’ sequences were more prone to DNA double and single strand breaks and oxidative DNA damage than the S. bayanus group that may influence the biotechnological processes using distillery strains. The opposite effect, namely increased copy number of MEC3 gene encoded a DNA damage and meiotic pachytene checkpoint protein [25, 26] was observed in the S. paradoxus group that may have implications for DNA damage response and adaptations to DNA-damaging conditions.
The other genes with affected copy number were mainly involved in carbohydrate and amino acid metabolism, and ion transport that may also modulate a biotechnological process. The dosage of numerous genes implicated in maltose metabolism was affected (e.g., MAL11, MAL13, MAL31, MAL33, MPH2 and MPH3). The MAL gene family of Saccharomyces is comprised of five multigene complexes, MAL1, MAL2, MAL3, MAL4 and MAL6, located at or near the telomere of a different chromosome, any one of which is sufficient for yeast to metabolize the disaccharide maltose and encodes maltose permease (GENE l), maltase (GENE 2) and the trans-acting MAL-activator (GENE 3) [27]. MAL11 and MAL13 are part of the MAL1 complex locus located on chromosome VII and encode high-affinity maltose transporter (α-glucoside transporter) and MAL-activator protein, respectively, whereas MAL31 and MAL33 are part of the MAL3 complex located on chromosome II and encode maltose permease and MAL-activator protein, respectively [28, 29]. It has been suggested that the MAL loci have been translocated to different chromosomes via a mechanism that involved the rearrangement(s) of chromosome termini [30]. MPH2 and MPH3 genes (maltose permease homologs) encode α-glucoside permeases that transport maltose, maltotriose, α-methylglucoside, and turanose [31].
The distillery strains also differed in the copy number of ASP3 genes, especially highly elevated ASP3 gene copy number was revealed in strain 19 (S. kudriavzevii). ASP3 contains a gene cluster located on chromosome XII comprised of four identical genes, ASP3-1, ASP3-2, ASP3-3, and ASP3-4, which encode for cell wall-associated L-asparaginase II that catalyzes the conversion of L-asparagine to aspartate and ammonia [32]. Asp3p is induced in response to nitrogen starvation and regulated by Gln3p/Ure2p [33]. More recently, the ASP3 locus has been shown to be originated by horizontal gene transfer from Wickerhamomyces [34]. It has been speculated that ASP3 acquisition may have aided yeast adaptation to artificial environments and may further highlight the importance of gene sharing between yeasts in the evolution of their remarkable metabolic diversity [34].
The most accented differences were observed in the copy number of SOR1 and SOR2 genes. The SOR1 gene encode a NAD-dependent sorbitol dehydrogenase that is a member of the polyol dehydrogenase branch of the medium-chain dehydrogenase/reductase (MDR) superfamily of enzymes [35]. It has been reported that the expression of SOR1 gene is elevated in the presence of sorbitol or xylose, though S. cerevisiae is a non-xylose-utilizing microorganism [35, 36]. Similarly, high variability in the gene copy number of genes involved in the siderophore transport, namely ENB1, FRE3, FRE5, FIT2 and FIT3, was observed. They represent two genetically separable systems for the uptake of siderophore-bound iron in S. cerevisiae. One system is based on family of transporters that is expressed as part of the AFT1 regulon and are termed ARN1, ARN2 (TAF1), ARN3 (SIT1) and ARN4 (ENB1) [37, 38]. These transporters are expressed in intracellular vesicles [39]. The second system relies on the high affinity ferrous iron transport complex, which is encoded by FET3 and FTR1 and is located on the plasma membrane [40, 41]. Ferric reductases encoded by FRE genes take part in iron uptake by the reduction of siderophore-bound iron prior to uptake by transporters [42, 43]. There are also three cell wall mannoproteins (Fit1, Fit2, Fit3) that facilitate the uptake of iron [44]. Low iron levels stimulate the expression of components of both systems [45]. Perhaps, increased copy number of genes involved in the transport of siderophore-bound iron in the S. paradoxus group may be advantageous in the certain growth conditions, e.g., during iron deprivation. Additionally, in all groups analyzed, the metallothionein gene dosage CUP1-1 and CUP1-2 was increased that was the most accented in strain 9 (S. cerevisiae). This may be also beneficial as may confer resistance to copper and cadmium [46].
In conclusion, we have provided for the first time array-CGH-based comprehensive genomic characterization of commercially available twenty two distillery yeast strains. We have documented the naturally occurring diversity in the gene copy number within six functional gene categories and revealed that the variations in the YRF1 gene copies may be accompanied by altered genetic stability in the analyzed yeast groups. Our genomic data may be helpful for better understanding of the fermentative environment-mediated changes in the yeast genome and accompanying phenotypic features. Thus, the knowledge on genetic diversity of distillery strains may be further exploited in economically important biotechnological processes.
MATERIALS AND METHODS
Reagents
All reagents, if not otherwise mentioned, were purchased from Sigma (Poland) and were of analytical grade.
Yeast strains and growth conditions
All distillery yeast strains used in this study are listed in Table 1. Yeast from one single colony was grown either on liquid YPD medium (1% w/v Difco Yeast Extract, 2% w/v Difco Yeast Bacto-Peptone, 2% w/v dextrose) or on solid YPD medium containing 2% w/v Difco Bacto-agar, at 28 °C.
Pulsed-field gel electrophoresis (PFGE)
Preparation of agarose-embedded yeast DNA and PFGE separation of yeast DNA were conducted as described elsewhere [47]. The dendrogram of chromosomal DNA-based similarity was created using Free-Tree software [48] using UPGMA (Unweighted Pair Group Method with Arithmetic Mean) algorithm, Jaccard similarity coefficient and Java TreeView 1.1.6.r2 (http://jtreeview.sourceforge.net/).
FACS-based ploidy analysis
The DNA content was measured via flow cytometry as previously described [49] except that a total of 3x104 cells were counted in a single assay.
Array-based comparative genomic hybridization (array-CGH)
Genomic DNA (0.5 μg) was labeled with SureTag DNA Labeling Kit and either Cy3- or Cy5-dUTP. Equal amounts of labeled DNA of tested and of the reference strain (BY4741) were combined and hybridized to Yeast (V2) Gene Expression Microarray, 8x15K using Oligo aCGH Hybridization Kit. All components were supplied by Agilent Technologies Inc. (Santa Clara, CA, USA) and all steps of the experiment were performed according to manufacturer’s protocols. Following hybridization and washing, the slides were scanned using Axon GenePix 4000B. Feature extraction was conducted using GenePix Pro 6.1 and normalization using Acuity 4.0 (Molecular Devices, Sunnyvale, CA, USA). CGH profiles with superimposed piecewise regression plots to highlight aneuploidies, were generated using CGH-Explorer v3.2 [50]. The original CGH profiles obtained after the comparison of analyzed strains to BY4741 gave consistently high noise due most probably to genomic DNA sequence differences between BY4741 and the industrial strains, which influenced the hybridization strength of individual probes. Therefore to obtain final CGH profiles, the data for each strain were compared to the average of all industrial strains used in the experiment.
Gene analysis after array-CGH
The analysis of over-representation of functional categories was performed using Cytoscape v. 2.8.2 with BiNGO v. 2.44 plug-in and hypergeometric test using Benjamini and Hochberg False Discovery Rate (FDR) correction and significance level of 0.05.
Cluster analysis
The array-CGH data for all strains were subjected to complete linkage clustering with Cluster 3.0 software using Euclidean distance similarity metrics [51]. To obtain the tree graph of similarity, the clustering output was visualized using Java TreeView 1.1.6.r2 (http://jtreeview.sourceforge.net/).
Detection of telomeric Y’ sequences
Y’ element telomeric probe was obtained according to [52] with minor modifications. After standard PFGE separation, Y’ sequences within particular yeast chromosomes were detected using digoxigenin labeling, anti-digoxigenin antibody and phosphate alkaline-based chemiluminescence [53].
Comet assay
Yeast spheroplasts were obtained [47] and DNA double-strand breaks (DSBs) and DNA single-strand breaks (SSBs) were assessed by neutral and alkaline single-cell microgel electrophoresis (comet assay), respectively, as described elsewhere [54]. The percentage of tail DNA was used as a parameter of DNA damage.
Oxidative stress parameters
Intracellular reactive oxygen species (ROS) production was measured using 2’,7’-dichlorodihydrofluorescein diacetate (H2DCF-DA) as described elsewhere [53]. Oxidative DNA damage as a level of 8-hydroxy-2’-deoxyguanosine (8-OHdG, 8-oxo-dG) was measured using Epigentek EpiQuik 8-OHdG DNA Damage Quantification Direct Kit (Gentaur, Poland) using the standard protocol according to the manufacturer’s instructions.
Statistical analysis
The results represent the mean ± SD from at least three independent experiments. Statistical significance was assessed by 1-way ANOVA using GraphPad Prism 5, and with the Tukey’s multiple comparison test.
ACKNOWLEDGMENTS
We are indebted to Prof. Martin Kupiec and Dr Yaniv Harari (Tel Aviv University, Israel) for sharing with us the protocol on the construction of Y’ telomeric probes.
CONFLICTS OF INTEREST
No potential conflicts of interest were disclosed.
GRANT SUPPORT
This work was supported by European Union, within Regional Operational Programme of Subcarpathia Voivodeship (2007-2013), Priority 1: Competitive and Innovative Economy, Action 1.3, Regional Innovation System, grant WND-RPPK-01.03.00-18-038/13.
Authors' contributions
Conceived and designed the experiments: MW. Performed the experiments: AD MS JA AK ER AS AL MW. Analyzed the data: MS AL MW. Contributed reagents/materials/analysis tools: AL MW. Wrote the paper: AL MW.
REFERENCES
1. Hittinger CT. Saccharomyces diversity and evolution: a budding model genus. Trends Genet. 2013; 29:309-317.
2. Sicard D, Legras JL. Bread, beer and wine: yeast domestication in the Saccharomyces sensu stricto complex. C R Biol. 2011; 334:229-236.
3. Naumov GI, James SA, Naumova ES, Louis EJ, Roberts IN. Three new species in the Saccharomyces sensu stricto complex: Saccharomyces cariocanus, Saccharomyces kudriavzevii and Saccharomyces mikatae. Int J Syst Evol Microbiol. 2000; 50 Pt 5:1931-1942.
4. Nguyen HV, Gaillardin C. Evolutionary relationships between the former species Saccharomyces uvarum and the hybrids Saccharomyces bayanus and Saccharomyces pastorianus; reinstatement of Saccharomyces uvarum (Beijerinck) as a distinct species. FEMS Yeast Res. 2005; 5:471-483.
5. Naumov GI, Masneuf I, Naumova ES, Aigle M, Dubourdieu D. Association of Saccharomyces bayanus var. uvarum with some French wines: genetic analysis of yeast populations. Res Microbiol. 2000; 151:683-691.
6. Naumov GI, Nguyen HV, Naumova ES, Michel A, Aigle M, Gaillardin C. Genetic identification of Saccharomyces bayanus var. uvarum, a cider-fermenting yeast. Int J Food Microbiol. 2001; 65:163-171.
7. Naumov GI, Naumova ES, Antunovics Z, Sipiczki M. Saccharomyces bayanus var. uvarum in Tokaj wine-making of Slovakia and Hungary. Appl Microbiol Biotechnol. 2002; 59:727-730.
8. Erny C, Raoult P, Alais A, Butterlin G, Delobel P, Matei-Radoi F, Casaregola S, Legras JL. Ecological success of a group of Saccharomyces cerevisiae/Saccharomyces kudriavzevii hybrids in the northern european wine-making environment. Appl Environ Microbiol. 2012; 78:3256-3265.
9. Masneuf I, Hansen J, Groth C, Piskur J, Dubourdieu D. New hybrids between Saccharomyces sensu stricto yeast species found among wine and cider production strains. Appl Environ Microbiol. 1998; 64:3887-3892.
10. Gonzalez SS, Barrio E, Gafner J, Querol A. Natural hybrids from Saccharomyces cerevisiae, Saccharomyces bayanus and Saccharomyces kudriavzevii in wine fermentations. FEMS Yeast Res. 2006; 6:1221-1234.
11. Bellon JR, Schmid F, Capone DL, Dunn BL, Chambers PJ. Introducing a new breed of wine yeast: interspecific hybridisation between a commercial Saccharomyces cerevisiae wine yeast and Saccharomyces mikatae. PLoS One. 2013; 8:e62053.
12. Redzepovic S, Orlic S, Sikora S, Majdak A, Pretorius IS. Identification and characterization of Saccharomyces cerevisiae and Saccharomyces paradoxus strains isolated from Croatian vineyards. Lett Appl Microbiol. 2002; 35:305-310.
13. Perez-Ortin JE, Querol A, Puig S, Barrio E. Molecular characterization of a chromosomal rearrangement involved in the adaptive evolution of yeast strains. Genome Res. 2002; 12:1533-1539.
14. Yuasa N, Nakagawa Y, Hayakawa M, Iimura Y. Distribution of the sulfite resistance gene SSU1-R and the variation in its promoter region in wine yeasts. J Biosci Bioeng. 2004; 98:394-397.
15. Naumova ES, Sadykova AZ, Martynenko NN, Naumov GI. Molecular genetic characteristics of Saccharomyces cerevisiae distillers’ yeasts. Microbiology. 2013; 82(2):175-185.
16. Brown SD, Klingeman DM, Johnson CM, Clum A, Aerts A, Salamov A, Sharma A, Zane M, Barry K, Grigoriev IV, Davison BH, Lynd LR, et al. Genome Sequences of Industrially Relevant Saccharomyces cerevisiae Strain M3707, Isolated from a Sample of Distillers Yeast and Four Haploid Derivatives. Genome Announc. 2013; 1.
17. Fischer G, James SA, Roberts IN, Oliver SG, Louis EJ. Chromosomal evolution in Saccharomyces. Nature. 2000; 405:451-454.
18. Naumov GI, Naumova ES, Lantto RA, Louis EJ, Korhola M. Genetic homology between Saccharomyces cerevisiae and its sibling species S. paradoxus and S. bayanus: electrophoretic karyotypes. Yeast. 1992; 8:599-612.
19. Naumova ES, Naumov GI, Masneuf-Pomarede I, Aigle M, Dubourdieu D. Molecular genetic study of introgression between Saccharomyces bayanus and S. cerevisiae. Yeast. 2005; 22:1099-1115.
20. Bakalinsky AT, Snow R. The chromosomal constitution of wine strains of Saccharomyces cerevisiae. Yeast. 1990; 6:367-382.
21. Mortimer RK. Evolution and variation of the yeast (Saccharomyces) genome. Genome Res. 2000; 10:403-409.
22. Dunn B, Sherlock G. Reconstruction of the genome origins and evolution of the hybrid lager yeast Saccharomyces pastorianus. Genome Res. 2008; 18:1610-1623.
23. Salmon JM. Enological fermentation kinetics of an isogenic ploidy series derived form an industrial Saccharomyces cerevisiae strain. J Ferment Bioeng. 1997; 83:253-260.
24. Yamada M, Hayatsu N, Matsuura A, Ishikawa F. Y’-Help1, a DNA helicase encoded by the yeast subtelomeric Y’ element, is induced in survivors defective for telomerase. J Biol Chem. 1998; 273:33360-33366.
25. Majka J, Burgers PM. Yeast Rad17/Mec3/Ddc1: a sliding clamp for the DNA damage checkpoint. Proc Natl Acad Sci U S A. 2003; 100:2249-2254.
26. Kondo T, Matsumoto K, Sugimoto K. Role of a complex containing Rad17, Mec3, and Ddc1 in the yeast DNA damage checkpoint pathway. Mol Cell Biol. 1999; 19:1136-1143.
27. Chow TH, Sollitti P, Marmur J. Structure of the multigene family of MAL loci in Saccharomyces. Mol Gen Genet. 1989; 217:60-69.
28. Charron MJ, Dubin RA, Michels CA. Structural and functional analysis of the MAL1 locus of Saccharomyces cerevisiae. Mol Cell Biol. 1986; 6:3891-3899.
29. Cheng Q, Michels CA. MAL11 and MAL61 encode the inducible high-affinity maltose transporter of Saccharomyces cerevisiae. J Bacteriol. 1991; 173:1817-1820.
30. Charron MJ, Read E, Haut SR, Michels CA. Molecular evolution of the telomere-associated MAL loci of Saccharomyces. Genetics. 1989; 122:307-316.
31. Day RE, Higgins VJ, Rogers PJ, Dawes IW. Characterization of the putative maltose transporters encoded by YDL247w and YJR160c. Yeast. 2002; 19:1015-1027.
32. Kim KW, Kamerud JQ, Livingston DM, Roon RJ. Asparaginase II of Saccharomyces cerevisiae. Characterization of the ASP3 gene. J Biol Chem. 1988; 263:11948-11953.
33. Bon EP, Carvajal E, Stanbrough M, Rowen D, Magasanik B. Asparaginase II of Saccharomyces cerevisiae. GLN3/URE2 regulation of a periplasmic enzyme. Appl Biochem Biotechnol. 1997; 63-65:203-212.
34. League GP, Slot JC, Rokas A. The ASP3 locus in Saccharomyces cerevisiae originated by horizontal gene transfer from Wickerhamomyces. FEMS Yeast Res. 2012; 12:859-863.
35. Sarthy AV, Schopp C, Idler KB. Cloning and sequence determination of the gene encoding sorbitol dehydrogenase from Saccharomyces cerevisiae. Gene. 1994; 140:121-126.
36. Toivari MH, Salusjarvi L, Ruohonen L, Penttila M. Endogenous xylose pathway in Saccharomyces cerevisiae. Appl Environ Microbiol. 2004; 70:3681-3686.
37. Heymann P, Ernst JF, Winkelmann G. Identification and substrate specificity of a ferrichrome-type siderophore transporter (Arn1p) in Saccharomyces cerevisiae. FEMS Microbiol Lett. 2000; 186:221-227.
38. Heymann P, Ernst JF, Winkelmann G. A gene of the major facilitator superfamily encodes a transporter for enterobactin (Enb1p) in Saccharomyces cerevisiae. Biometals. 2000; 13:65-72.
39. Yun CW, Ferea T, Rashford J, Ardon O, Brown PO, Botstein D, Kaplan J, Philpott CC. Desferrioxamine-mediated iron uptake in Saccharomyces cerevisiae. Evidence for two pathways of iron uptake. J Biol Chem. 2000; 275:10709-10715.
40. Askwith C, Eide D, Van Ho A, Bernard PS, Li L, Davis-Kaplan S, Sipe DM, Kaplan J. The FET3 gene of S. cerevisiae encodes a multicopper oxidase required for ferrous iron uptake. Cell. 1994; 76:403-410.
41. Stearman R, Yuan DS, Yamaguchi-Iwai Y, Klausner RD, Dancis A. A permease-oxidase complex involved in high-affinity iron uptake in yeast. Science. 1996; 271:1552-1557.
42. Yun CW, Bauler M, Moore RE, Klebba PE, Philpott CC. The role of the FRE family of plasma membrane reductases in the uptake of siderophore-iron in Saccharomyces cerevisiae. J Biol Chem. 2001; 276:10218-10223.
43. Martins LJ, Jensen LT, Simon JR, Keller GL, Winge DR. Metalloregulation of FRE1 and FRE2 homologs in Saccharomyces cerevisiae. J Biol Chem. 1998; 273:23716-23721.
44. Protchenko O, Ferea T, Rashford J, Tiedeman J, Brown PO, Botstein D, Philpott CC. Three cell wall mannoproteins facilitate the uptake of iron in Saccharomyces cerevisiae. J Biol Chem. 2001; 276:49244-49250.
45. Philpott CC, Protchenko O, Kim YW, Boretsky Y, Shakoury-Elizeh M. The response to iron deprivation in Saccharomyces cerevisiae: expression of siderophore-based systems of iron uptake. Biochem Soc Trans. 2002; 30:698-702.
46. Jeyaprakash A, Welch JW, Fogel S. Multicopy CUP1 plasmids enhance cadmium and copper resistance levels in yeast. Mol Gen Genet. 1991; 225:363-368.
47. Lewinska A, Miedziak B, Wnuk M. Assessment of yeast chromosome XII instability: single chromosome comet assay. Fungal Genet Biol. 2014; 63:9-16.
48. Pavlicek A, Hrda S, Flegr J. Free-Tree-freeware program for construction of phylogenetic trees on the basis of distance data and bootstrap/jackknife analysis of the tree robustness. Application in the RAPD analysis of genus Frenkelia. Folia Biol (Praha). 1999; 45:97-99.
49. Krol K, Brozda I, Skoneczny M, Bretne M, Skoneczna A. A genomic screen revealing the importance of vesicular trafficking pathways in genome maintenance and protection against genotoxic stress in diploid Saccharomyces cerevisiae cells. PLoS One. 2015; 10:e0120702.
50. Lingjaerde OC, Baumbusch LO, Liestol K, Glad IK, Borresen-Dale AL. CGH-Explorer: a program for analysis of array-CGH data. Bioinformatics. 2005; 21:821-822.
51. de Hoon MJ, Imoto S, Nolan J, Miyano S. Open source clustering software. Bioinformatics. 2004; 20:1453-1454.
52. Romano GH, Harari Y, Yehuda T, Podhorzer A, Rubinstein L, Shamir R, Gottlieb A, Silberberg Y, Pe’er D, Ruppin E, Sharan R, Kupiec M. Environmental stresses disrupt telomere length homeostasis. PLoS Genet. 2013; 9:e1003721.
53. Lewinska A, Miedziak B, Kulak K, Molon M, Wnuk M. Links between nucleolar activity, rDNA stability, aneuploidy and chronological aging in the yeast Saccharomyces cerevisiae. Biogerontology. 2014; 15:289-316.
54. Dworak N, Wnuk M, Zebrowski J, Bartosz G, Lewinska A. Genotoxic and mutagenic activity of diamond nanoparticles in human peripheral lymphocytes in vitro. Carbon. 2014; 68:763-776.

