INTRODUCTION
Cancer remains a major challenge in modern medicine [1–6]. Despite improved diagnostics, an arsenal of FDA-approved anti-cancer drugs and targeted therapies, cancer remains a leading cause of death, a major economic burden and a severe limitation on patient quality-of-life. Moreover, in developed countries, cancer mortality has remained relatively constant for the past 35 years. Almost all of the ~500,000 annual cancer deaths in the U.S. are due, at least in part, to drug resistance [4–7]. Drug resistance, either de novo or acquired, still accounts for the majority of tumor relapses contributing to poor outcomes [6–10]. Drug combinations may prevent the emergence of resistance since a clone resistant to a single agent is more readily eradicated by multiple agents from different drug classes. Such a strategy has improved survival in certain cancers, e.g., acute lymphocytic leukemia, diffuse large B cell lymphoma, Hodgkin’s lymphoma and germ cell tumors. However, drug resistance inevitably emerges in most cancer types to hinder patient response [11–21].
Numerous mechanisms may explain the molecular basis of drug resistance and treatment failure. Resistance results from genetic variations within the patient’s tumor cell. Recently, the dormant state of cancer stem cells and the epithelial-mesenchymal transition have been linked to chemotherapeutic resistance [22–27]. While the design of cancer chemotherapy has become increasingly sophisticated, few agents universally prevent disease relapse. A common mechanism for the acquisition of resistance is expression of energy-dependent transporters that eject anticancer drugs from cells [28–30]. Induction of drug detoxification, reduced sensitivity to apoptotic signals, uncoupling of growth control and inactivation of cell death pathways all promote drug resistance in cancer cells. Recent evidence points to a relationship between the drug-resistant phenotype and epigenetic alterations within cancer cells [31–33]. Non-coding (ncRNAs) are major regulators of epigenetic, transcriptional and post-transcriptional gene expression. NcRNAs have also been reported to play a role in chemoresistance by impairing the response through cell cycle arrest, inhibition of apoptosis and enhanced DNA damage repair [34–37]. Here, we describe the rapidly emerging role of long ncRNAs (lncRNAs) in cancer drug resistance and highlight a prominent example, the lncRNA HOTAIR, to correlate lncRNA expression with metastasis, drug resistance and clinical outcome [38–40].
NcRNAs AND CANCER
Studies have revealed that protein-coding and non-protein-coding genes are extensively transcribed within the human genome [41–44]. Protein-coding and non-protein-coding RNAs, such as long and short ncRNAs, are found on all human chromosomes (Figure 1A). This has led to the identification of novel classes of ncRNAs that regulate gene expression through diverse mechanisms. NcRNAs are deregulated or mutated in diseased cells from many human cancers [45–48]. NcRNAs represent a broad class of structurally and functionally distinct RNAs. These include long and short ncRNAs that represent a significant proportion (~60%) of the RNAs distributed throughout the human genome (Figure 1B) [49–52]. The distribution of lncRNAs and different short ncRNA types within the human genome is shown (Figure 1C).
For nearly four decades, mutation, amplification or deletions within protein-coding oncogenes and/or tumor suppressors was considered the primary cause of tumorigenesis [53–57]. This thinking has recently been revisited with the discovery of ncRNAs [58–60]. Thousands of genes that produce long or short ncRNA transcripts without any significant open reading frame have demonstrated that the complexity of cancer cell genetics is far greater than expected. These results led to a paradigm shift in addressing cancer biology and approaches in cancer treatment.
NcRNAs do not encode proteins but studies have documented their role during the steps of tumorigenesis and the development of therapeutic resistance [61–64]. NcRNAs are differentially expressed in a number of cancer types relative to the surrounding healthy (normal) tissue [65–67]. Importantly, many of the same ncRNAs appear to be commonly deregulated in multiple tumor types. While ncRNAs are newcomers in genome biology, these findings indicate that ncRNAs directly modulate key pathways that promote cancer growth. MiRNAs range from 18–25 nucleotides (nt) while lncRNAs extend up to >10,000nt. MiRNAs typically are excised from a 60 to 110nt hairpin precursor (pre-miRNA) that is transcribed from a larger primary transcript (pri-miRNA). MiRNAs bind target messenger RNA (mRNA) transcripts through sequence complementarity to either promote mRNA degradation or to prevent mRNA translation. While much is known about miRNAs, little exists regarding the biology and function of lncRNAs. While the precise functions of individual lncRNAs in cancer is only beginning to emerge, it is evident that ncRNAs are essential for tumorigenesis [68–71].
LncRNA HOTAIR AND HUMAN CANCERS
LncRNAs are a heterogeneous group of RNAs that regulate expression at the epigenetic, transcriptional or post-transcriptional level [72–76]. LncRNAs bind specific gene clusters to not only prevent the binding of transcriptional activators but also recruit chromatin remodeling proteins as a molecular scaffold to silence gene expression. LncRNA can also interact with chromatin modifying proteins in trans to epigenetically silence genes at distant loci. LncRNA transcription may open the chromatin structure and permit access to the transcriptional machinery and enhance expression of neighboring protein-coding genes. Similarly, transcription of lncRNAs near protein-coding loci can repress transcription since the presence of the transcriptional machinery on the lncRNA gene locus physically prevents binding to protein-coding genes. The post-transcriptional roles of lncRNAs are more diverse. Since many lncRNAs are complementary to protein-coding genes, lncRNAs may function in mRNA splicing, editing, transport, translation and degradation. Although lncRNAs are best known for modulating transcription, their post-transcriptional influence on mRNA translation is emerging. Recently, HOTAIR was shown to be a post-translational inducer of ubiquitin-mediated proteolysis. HOTAIR associates with certain E3 ubiquitin ligases bearing RNA-binding domains, such as Dzip3 and Mex3b, as well as with their respective ubiquitination substrates, Ataxin-1 and Snurportin-1 [88]. HOTAIR facilitates the ubiquitination of Ataxin-1 by Dzip3 and Snurportin-1 by Mex3b and accelerates their degradation. These studies have uncovered a fascinating and unforeseen role for HOTAIR in protein degradation.
The hallmarks of cancer comprise biological capabilities acquired during the multistep development of tumors [78, 79]. These features constitute an organizing principle for rationalizing the complexities of cancer and include sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis, and activating invasion and metastasis. Underlying these hallmarks is the common thread of genomic instability to generate the diversity that expedites tumor survival. Importantly, the results collectively indicate that lncRNAs regulate many genes that contribute to tumorigenesis and promote the hallmark features of cancer [Figure 2] [79–82].
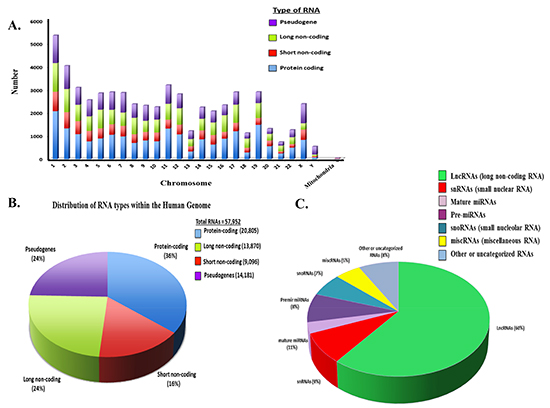
Figure 1: Distribution of Coding and Non-coding RNAs in the Human Genome. (A) Distribution of different RNA types on the individual human chromosomes. Number of coding and ncRNAs is based upon http://useast.ensembl.org and Gencode19. It is noted that the most recent ensembl version indicates a total putative ~140,000 ncRNAs based upon whole genome sequencing. Numbers used here reflect on those RNAs that have been validated. (B) Distribution of RNA types within the human genome. (C) Distribution of non-coding RNA types within the human genome.
LncRNA expression and metastasis, drug resistance and overall survival
The Hox transcript antisense intergenic RNA HOTAIR is expressed from the developmental HOXC locus located on chromosome 12q13.13 and cooperates with the polycomb repressive complex PRC2 [83]. This 2.2kb spliced RNA transcript interacts with the PRC2 to modify chromatin and repress transcription of the HOX genes [84, 85]. Although less than 2% of a mammalian genome codes for protein, at least 50% is transcribed [85]. Recently it was shown that HOTAIR serves as a modular scaffold, assembling a molecular cargo of specific combinations of enzymes that are equipped to regulate target genes [86]. Further studies then demonstrated that HOTAIR reprograms chromatin organization and in breast cancer cells promotes metastatic potential [87].
LncRNAs that correlate with metastasis
HOTAIR expression has been investigated in tumor samples from many different cancer types. Healthy and cancer tissue was compared to demonstrate that HOTAIR is deregulated in many tumor types (Figure 3A). HOTAIR expression from patients diagnosed with stage IV colorectal cancer (CRC) and liver metastases was shown to correlate with a poor prognosis [89]. In addition, high expression of HOTAIR correlated with the presence of liver metastases. MALAT1 (metastasis-associated lung adenocarcinoma transcript 1) is another lncRNA that has been associated with metastasis and poor prognosis for patients with NSCLC [90]. MALAT1 resides on chromosome 11q13.1 which has been found to harbor chromosomal translocation breakpoints linked to cancer [91, 92]. Other in vitro studies, have implicated MALAT1 in the regulation of the invasive potential of cancer cells, in cervical [93] and lung cancers [94].
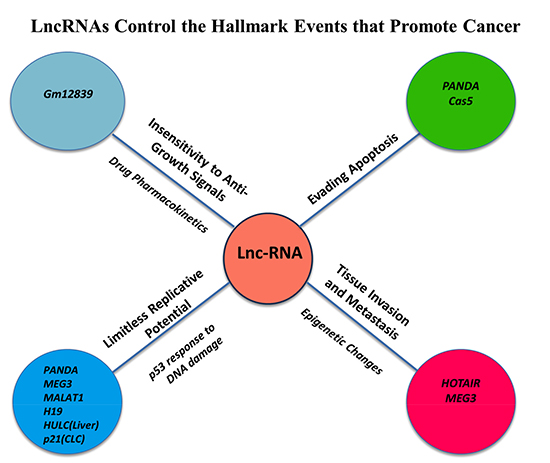
Figure 2: LncRNAs Control the Hallmark Events that Promote Cancer Initiation and Progression. LncRNAs have been linked to the promotion of hallmark features of cancer cells including the insensitivity to anti-growth signals, evading apoptosis, limitless replicative potential and tissue invasion and metastasis.
LncRNAs that correlate with therapeutic resistance
The lncRNA cancer upregulated drug resistance gene (CUDR) is a 2.2kb RNA transcript that antagonizes the apoptotic effect of cisplatin in bladder cancer cells [95]. CUDR expression correlates with tumorigenesis as well as cellular growth [96]. CUDR has been shown to render resistance to doxorubicin and etoposide, commonly used for squamous cell cancer. The growth arrest-specific 5 lncRNA Gas5 contributes to glucocorticoid resistance [97]. Gas5 exerts its effects by assuming a secondary structure that is similar to a sequence within the steroid-responsive gene promoter. The result is that Gas5 competitively inhibits steroid receptor binding to the steroid-responsive region of the promoter to prevent transcriptional activation. A role as a tumor suppressor in renal cell carcinoma has also recently been reported [98]. PANDA is a lncRNA increased in a subset of breast cancer cells that contributes to anthracycline resistance, a crucial component of breast cancer chemotherapy [99]. PANDA acts by interacting with the nuclear transcription factor NF-YA to decrease the apoptotic effect. PANDA-depleted fibroblasts have been shown to display increased sensitivity to chemotherapy [100]. A large intergenic ncRNA induced by p53 mediates global gene repression and many p53-regulated lncRNAs are induced in response to DNA damage and promote chemoresistance. H19 knockdown by transfection with antisense H19 oligonucleotides suppresses the multi-drug resistance gene and promotes doxorubicin sensitivity [101, 102]. Finally, HULC is a 1.6 kb lncRNA that may act as an endogenous sponge to reduce miRNA levels and inhibit their functional activity [103, 104].
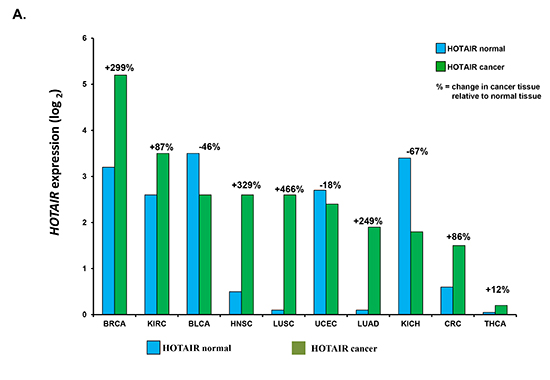
Figure 3: HOTAIR expression in cancer tissues and correlation with patient outcome. (A) Correlation of HOTAIR expression in healthy vs. cancer tissue samples. HOTAIR expression was determined from indicated tissue samples and correlated to that in healthy (normal) tissue. Number of samples (n) = 5599. Data was obtained from the starBase Pan-Cancer Analysis Platform (http://starbase.sysu.edu.cn/panCancer.php) to explore Pan-Cancer Networks of lncRNAs, miRNAs, ceRNAs and RNA-binding proteins (RBPs) by mining clinical and expression profiles of 14 cancer types including tumor and normal samples from the Cancer Genome Atlas (TCGA) data portal. Cancer types used were from urothelial bladder (BLCA), breast (BRCA), colon and rectal adenocarcinoma (CRC), glioblastoma multiforme (GBM), head and neck squamous cell carcinoma (HNSC), chromophobe renal cell carcinoma (KIRH), clear cell kidney carcinoma (KIRC), acute myeloid leukemia (LAML), lung adenocarcinoma (LUAD), lung squamous cell carcinoma (LUSC), ovarian serous cystadenocarcinoma (OV), cutaneous melanoma (SKCM), papillary thyroid carcinoma (THCA) and uterine corpus endometrial carcinoma (UCEC). B. Correlation of HOTAIR expression with patient 5 year survival. Shown is the correlation of HOTAIR expression in tissue from non-small cell lung, colorectal, breast and cervical cancer patients.
LncRNAs in metabolic inactivation of anti-cancer drugs
The Drosophila sechellia lncRNA Gm12839 is located ~40kb downstream of a cytochrome P450 system gene [105]. Gm12839 exhibits a reverse expression pattern with the P450 genes to suggest an inhibitory effect on the cytochrome P450 drug metabolism and may be crucial in the inactivation of chemotherapeutics.
Hotair expression correlates with overall survival
HOTAIR expression in tissue from patients diagnosed with either non-small cell lung, colorectal, breast or cervical cancers was the correlated with five year overall survival (Figure 3b.) We found that patients with increased HOTAIR expression had a worse survival in all four cancer types studied. These and related studies may soon lead to the development of lncRNA signatures that classify patients into high and low-risk groups with significantly different survival rates. The prognostic value of lncRNA signatures needs to be confirmed in independent data sets but the identification of prognostic lncRNAs indicates their potential.
HOTAIR REGULATION OF CHROMATIN REMODELING
In a subset of 32 CRC specimens, gene set enrichment analysis revealed a correlation between HOTAIR expression and the PRC2 complex members, namely SUZ12, EZH2 and H3K27me3 [106]. HOTAIR may be associated with the genome-wide reprogramming of PRC2 in a broad number of tumor types. Approximately 20% of the identified lncRNAs regulate the transcriptional activity of protein-coding genes by guiding the histone methyltransferase PRC2 to specific genomic loci. Chemotherapy has been shown to induce epigenetic changes which contribute to the emergence of drug resistance [107]. We correlated the cross-cancer alteration of HOTAIR through either mutation, deletion, amplification with a broad number of cancer types (Figure 4A) [108]. HOTAIR was significantly deregulated in a significant percentage of patients from the different cancer types examined.
HOTAIR contains 6,232bp and encodes a 2.2kb lncRNA located within a HOXC gene cluster. It is shuttled from chromosome 12 to chromosome 2 by Suz-Twelve. The 5' end of HOTAIR interacts with PRC2 and as a result regulates chromatin state. The 3' end of HOTAIR interacts with the histone demethylase LSD1. PRC2 consists of four subunits: Suz12, Eed, Ezh1 or Ezh2 (SET domain with histone methyltransferase activity and RbAp48 (histone-binding domain). HOTAIR may contribute to the regulation and functioning of the PRC2 subunits. In addition to HOTAIR, a correlation between alterations in EED1 was also seen across the different cancer types [Figure 4B]. Future studies will further define the role of HOTAIR on PRC2, chromatin remodeling and individual components, e.g., EED1, of PRC2.
LncRNA THERAGNOSTICS
Theragnostics, a portmanteau of therapeutics and diagnostics, describes a system to customize healthcare using molecular and genetic tools for treatment decisions tailored to the individual patient. LncRNAs are therefore perfectly tailored as cancer theragnostics to design personalized medicines based upon molecular features of each tumor [109, 110]. Novel cancer diagnostics, prognostics and anti-cancer strategies based upon ncRNA biology are rapidly emerging. For example, lncRNAs isolated from tumor cells or circulating within the bloodstream may provide for readily-available, inexpensive and stable blood-borne diagnostics to more readily detect cancers and cancer subtypes. Interference of RNA expression to treat cancer has recently gained momentum as a treatment modality since growth-promoting lncRNAs may be inactivated through antisense technologies. LncRNAs may also function as mRNA or miRNA sponges to inactivate growth-promoting, pro-tumorigenic signaling pathway. Therefore, synthetically-engineered lncRNAs may be employed through replacement therapy to inhibit tumor cells [111]. Cancer cells have increased the protein synthesizing machinery as well as enhanced ncRNA transcription rates making RNA interference an attractive strategy. NcRNAs offer potential as therapeutic targets is exemplified by Onconase, a non-specific ribonuclease which neutralizes tRNA, rRNA, mRNAs and ncRNAs. Onconase, currently in phase II trials, induces a G1 block in the cell cycle and increased apoptosis [112]. Delivery vehicles, e.g., liposomal membranes, carry growth-inhibitory ncRNAs into tumor cells as another lncRNA-based treatment strategy. MRX34 is a ncRNA-based therapeutic that targets miR-34 is being evaluated in clinical trials for hepatocellular cancers [113, 114]. The absence of nucleases within the cerebral spinal fluid (CSF) has made intrathecal injection of RNA-based agents an attractive approach to treat neurodegenerative diseases. Primary brain tumors and brain metastases may similarly be targeted. Nanoparticles that deliver RNA inhibitors are also emerging [115–117]. Dextran nanoparticles can deliver chemotherapy to the nucleus and may be used to attach cytotoxic agents to lncRNAs. The plasmid DTA-H19 was designed by express a diphtheria toxin subunit controlled under the H19 promoter [118, 119]. Intratumoral injection of this compound can lead to a H19-dependent activation of diphtheria toxin within the tumor. Another emerging modality is to increase tumor suppressor activity by neutralizing the inhibitory of effect of lncRNAs through antisense oligonucleotides. These compounds do not need any delivery vehicle and can be injected subcutaneously and are directly taken up by cells. They can cross endosomal membranes, enter the nucleus and inhibit PRC2 interaction with lncRNAs. These compounds may also cross the blood brain barrier without the need of a lipid carrier.
CONCLUSIONS
LncRNAs are rapidly being recognized as important regulators of gene expression in cancer. The evolutionary conservation, diversity and complexity of lncRNAs indicates that they exert significant regulatory control on cell growth. While understanding the mechanistic role of lncRNA in cancer is still early, general principles are emerging. LncRNAs, e.g., HOTAIR, are deregulated in many cancer types, affect signaling pathways that promote cancer and modulate gene expression. Future studies will identify similar lncRNAs leading to signatures to better diagnose cancers. LncRNAs should allow for better stratification of cancer subtypes to correlate lncRNA expression with therapeutic response, drug resistance and overall survival. Similar to miRNA expression profiling in multiple myeloma and related B cell malignancies, lncRNAs profiling should lead to the identification of genetic subtypes and their association with treatment response and survival [120–122]. Finally, the structural and functional novelty of lncRNAs offers promise as anticancer therapeutics that may avoid the emergence of drug resistance commonly seen with the currently used agents. Studies to better understand the molecular mechanisms of lncRNAs in cancer offers promise for the development of more effective cancer therapies.
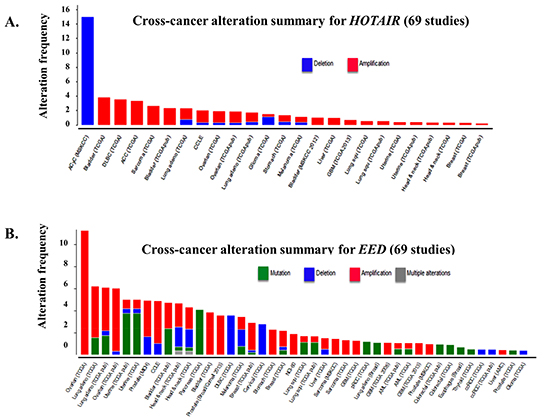
Figure 4: Frequency of HOTAIR alteration and putative HOTAIR-controlled genes across cancer types. (A) Cross-cancer alteration summary for HOTAIR from 69 independent studies to correlate HOTAIR expression with alteration frequency. Potential alterations included mutation, deletion, amplification or multiple alterations. Data was obtained from the cBioportal for cancer genomics (http://www.cbioportal.org/public-portal/, Memorial Sloan-Kettering Cancer Center, NY, NY). (B) Shown is the correlation between the alteration of EED1, a putative target of HOTAIR, across different cancer types. Potential alterations included mutation, deletion, amplification or multiple alterations. Data was obtained from the cBioportal for cancer genomics (http://www.cbioportal.org/public-portal/, Memorial Sloan-Kettering Cancer Center, NY, NY).
Conflicts of interest
The authors declare no potential conflicts of interest.
REFERENCES
1. Jemal A, Simard EP, Dorell C, Noone AM, Markowitz LE, Kohler B, Eheman C, Saraiya M, Bandi P, Saslow D, Croni KA, Watson M, Schiffman M and et al. Annual report to the nation on the status of cancer, 1975–2009, featuring the burden and trends in HPV-associated cancers and HIV vaccination levels. J Natl Cancer Inst. 2014; 105:175–201.
2. Trent L, The American Society of Clinical Oncology. The state of cancer care in America, 2014: A report by the Amercian Society of Clinical Oncology. 2014; 10:119–142.
3. Jemal A, Bray F, Center MM, Ferlay J, Ward E and Forman D. Global cancer statistics. CA: a cancer journal for clinicians. 2013; 61(2):69–90.
4. Guy G, Ekwueme DU, Yabroff R, Dowling EC, Li C, Rodriguez JL, de Moor JS and Virgo KS. Economic burden of cancer survivorship among adults in the United States. J Clin Oncol. 2013; 31:3749–3757.
5. Smith B, Smith GL, Hortobagyi GH and Buchholz TA. Future of cancer incidence in the United States: Burdens upon an aging, changing nation. J Clin Oncol. 2009; 27:2758–2765.
6. McKoy JM, Samaras AT and Bennet CL. Providing Cancer Care to a Graying and Diverse Cancer Population in the 21st Century: Are We Prepared? J Clin Oncol. 2009; 27:2745–2746.
7. SEER Stat Fact Sheets: All Cancer Sites. National Cancer Institute website.
8. Siegel R, Ward E, Brawley O and Jemal A. Cancer statistics, 2011: The impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J Clin. 2011; 61(4):212–236.
9. GLOBOCAN v1.2. Cancer Incidence and Mortality Worldwide: IARC CancerBase No. 10: 2008.
10. Gottesman MM, Fojo T and Bates SE. Multidrug resistance in cancer: role of ATP-dependent transporters. Nat Rev Cancer. 2002; 2(1):48–58.
11. Jeha S and Pui CH. Clinical manifestations and treatment of acute lymphoblastic leukemia in children. In: Hoffman R, Benz EJ Jr, Silberstein LE eds. Hematology: Basic Principles and Practice. 6th ed. Philadelphia, Pa: Elsevier Saunders; 2012: chap 64.
12. National Cancer Institute: PDQ Adult Acute Lymphoblastic Leukemia Treatment. National Cancer Institute. 2014; http://www.cancer.gov/cancertopics/pdq/adultALL.
13. Dunleavy K, Pittaluga S, Czuczman MS, Dave SS, Wright S, Shovlin M, Jaffe ES, Janik JE, Staudt LM and Wilson WH. Differential efficacy of bortezomib plus chemotherapy within molecular subtypes of diffuse large B-cell lymphoma. Blood. 2009; 113(24):6069–6076.
14. Lenz G, Wright G, Dave SS, Xiao W, Powell J, Zhao H, Xu W, Tan B, Goldschmidt N, Iqbal J, Vose J, Bast M, Fu K and et al. Stromal Gene Signatures in Large-B-Cell Lymphomas. New England Journal of Medicine. 2008; 359(22):2313–2323.
15. Gobbi PG, Levis A, Chisesi T, Broglia C, Vitolo U, Stelitano C, Pavone V, Cavanna L, Santini G, Merli F, Liberati M, Baldini L and Deliliers GL. ABVD vs. modified Stanford V vs. MOPPEBVCAD with optional and limited radiotherapy in intermediate- and advanced-stage Hodgkin's lymphoma: final results of a multicenter randomised trial by the Intergruppo Italiano Linfomi. J. Clin. Oncol. 2005; 23(36):9198–9207.
16. Herbst C, Rehan FA, Skoetz N, Bohlius J, Brillant C and Schulz H. Chemotherapy alone versus chemotherapy plus radiotherapy for early stage Hodgkin lymphoma. Cochrane Database Syst Rev. 2011; (2):CD007110.
17. Szakács G, Paterson JK, Ludwig JA, Booth-Genthe C and Gottesman MM. Targeting multidrug resistance in cancer. Nature Reviews Drug Discovery. 2006; 5(3):219–234.
18. Williams S, Blessing JA, Liao SY, Ball H and Hanjani P. Adjuvant therapy of ovarian germ cell tumors with cisplatin, etoposide, and bleomycin: a trial of the Gynecologic Oncology Group. J Clin Oncol. 1994; 12(4):701–6.
19. Williams SD, Kauderer J, Burnett AF, Lentz SS, Aghajanian C and Armstrong DK. Adjuvant therapy of completely resected dysgerminoma with carboplatin and etoposide: a trial of the Gynecologic Oncology Group. Gynecol Oncol. 2004; 95(3):496–499.
20. Pavet V, Portal MM, Moulin JC, Herbrecht R and Gronemeyer H. Towards novel paradigms for cancer therapy. Oncogene. 2011; 30:1–20.
21. Clozel T, Yang SN, Elstrom RL, Tam W, Martin P, Kormaksson M, Banerjje S, Vasanthakumar A, Culjkovic B, Scott DW, Wyman S, Leser Mm, and et al. Shaknovich R. Mechanism-based epigenetic chemosensitization therapy of diffuse large B-cell lymphoma. Cancer Discovery. 2013; 3(9):1002–1019.
22. Al-Hajj M, Becker MW, Wicha M, Weissman I and Clarke MF. Therapeutic implications of cancer stem cells. Curr. Opin. Genet. Dev. 2004; 14:43–47.
23. Miyake K, Mickley L, Litman T, Zhan Z, Robey R, Cristensen B, Brangi B, Greenberger L, Dean M, Fojo T and Bates SE. Molecular cloning of cDNAs which are highly overexpressed in mitoxantroneresistant cells: demonstration of homology to ABC transport genes. Cancer Res. 1999; 59:8–13.
24. Doyle LA, Yang W, Abruzzo LV, Krogmann T, Gao Y, Rishi AK and Ross DR. A multidrug resistance transporter from human MCF-7 breast cancer cells. Proc. Natl Acad. Sci. USA. 1998; 95:15665–15670.
25. Singh A and Settleman J. EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene. 2010; 29:4741–4751.
26. Bao B, Ahmad A, Li Y, Azmi AS, Ali S, Banerjee S, Kong D and Sarkar FH. Targeting CSC-related miRNAs for cancer therapy by natural agents. Curr Drug Targets. 2012; 13(14):1858–68.
27. Chen K and Rajewsky N. The evolution of gene regulation by transcription factors and microRNAs. Nature Reviews Genetics. 2007; 8(2):93–103.
28. Xu C, Li CY and Kong AN. Induction of phase I, II and III drug metabolism/transport by xenobiotics. Arch. Pharm. Res. 2005; 28:249–268.
29. Calcagno AM, Kim IW, Wu CP, Shukla S and Ambudkar SV. ABC drug transporters as molecular targets for the prevention of multidrug resistance and drug–drug interactions. Curr. Drug Deliv. 2007; 4:324–333.
30. Tukey RH and Strassburg CP. Human UDP-glucuronosyltransferases: metabolism, expression, and disease. Annu. Rev. Pharmacol. Toxicol. 2000; 40:581–616.
31. Ivanov M, Kacevska M and Ingelman-Sundberg M. 2012; Epigenomics and interindividual differences in drug response. Clin Pharmacol Ther. 2012; 92:727–736.
32. Shen DW, Pouliot LM, Hall MD and Gottesman MM. Cisplatin resistance: a cellular self-defense mechanism resulting from multiple epigenetic and genetic changes. Pharmacol Rev. 2012; 64:706–721.
33. Haenisch S and Cascorbi I. miRNAs as mediators of drug resistance. Epigenomics. 2012; 4:369–381.
34. Lipovich L, Johnson R and Lin CY. MacroRNA underdogs in a microRNA world: evolutionary, regulatory, and biomedical significance of mammalian long non-protein-coding RNA. Biochim Biophys Acta. 2010; 1799:597–615.
35. Harries LW. Long non-coding RNAs and human disease. Biochem Soc Trans. 2012; 40:902–906.
36. Goldman B. Multidrug resistance: can new drugs help chemotherapy score against cancer? Journal of the National Cancer Institute. 2003; 95(4):255–257.
37. d’Adda di Fagagna F. Living on a break: cellular senescence as a DNA-damage response. Nat. Rev. Cancer. 2008; 8:512–522.
38. Jeffares DC, Poole AM and Penny D. Relics from the RNA world. J Mol Evol. 2008; 46:18–36.
39. Morris KV. Non-coding RNAs and Epigenetic Regulation of Gene Expression: Drivers of Natural Selection. 2012; Caister Academic Press. ISBN 978-1-904455-94-3.
40. Hüttenhofer A, Schattner P and Polacek N. Non-coding RNAs: hope or hype? Trends Genet. 2012; 21(5):289–97.
41. Ulitsky I and Bartel DP. lncRNAs: genomics, evolution and mechanisms. Cell. 2013; 136:215–33.
42. Bentwich I., Avniel A., Karov Y., Aharonov R., Gilad S., Barad O., Barzilai A., Einat P., Einav U., Meiri E and et al. Identification of hundreds of conserved and nonconserved human microRNAs. Nat. Genet. 2005; 37:766–770.
43. Lewis BP, Burge CB and Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2004; 120(1):15–20.
44. Lee CT, Risom T and Strauss WM. Evolutionary conservation of microRNA regulatory circuits: an examination of microRNA gene complexity and conserved microRNA-target interactions through metazoan phylogeny. DNA Cell Biol. 2005; 26(4):209–18.
45. Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E and et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2002; 99(24):15524–15529.
46. Alsiwadi S, Malek E and Driscoll JJ. MicroRNAs in Brain Metastases: Potential Role as Diagnostics and Therapeutics. Int. J. Mol. Sci. 2014; 15:10508–10526.
47. Esteller M. Non-coding RNAs in human disease. Nature Rev. Genetics. 2011; 12:861–874.
48. Nagano T and Fraser P. No-nonsense functions for long noncoding RNAs. Cell. 2011; 145(2):178–181.
49. Wang KC and Chang HY. Molecular mechanisms of long noncoding RNAs. Molecular Cell. 2011; 43(6):904–914.
50. Wright MW and Bruford EA. Naming 'junk': human non-protein coding RNA (ncRNA) gene nomenclature. Human Genomics. 2011; 5(2):90–98.
51. Bachellerie JP, Cavaillé J and Hüttenhofer A. The expanding snoRNA world. Biochimie. 2002; 84(8):775–790.
52. Kiss-László Z, Henry Y and Kiss T. Sequence and structural elements of methylation guide snoRNAs essential for site-specific ribose methylation of pre-rRNA. EMBO J. 1998; 17(3):797–807.
53. Bonasio R, Tu S and Reinberg D. Molecular signals of epigenetic states. Science. 2010; 330:612–616.
54. Croce CM. Oncogenes and cancer. N Engl J Med. 2008; 358(5):502–511.
55. Todd R and Wong DT. Oncogenes. Anticancer Res. 1998; 19(6A):4729–46.
56. Weinberg RA. The Biology of Cancer. 2014; Garland Science.
57. Sherr CJ. Principles of tumor suppression. Cell. 2004; 116(2):235–46.
58. ENCODE Project Consortium: Birney E, Stamatoyannopoulos JA, Dutta A, Guigó R, Gingeras TR, Margulies EH, Weng Z, Snyder M, Dermitzakis ET, Thurman RE, Kuehn MS and Taylor CM. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007; 447(7146):799–816.
59. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermüller J, Hofacker IL, Bell I, Cheung E, Drenkow J and et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007; 316(5830):1484–1488.
60. Hutchinson J.N., Ensminger A.W., Clemson C.M., Lynch C.R., Lawrence J.B. and Chess A. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics. 2007; 8:39.
61. Yang Y, Li H, Hu B, Liu J and Wang J. The noncoding RNA expression profile and the effect of lncRNA AK126698 on cisplatin resistance in non-small-cell lung cancer cell. Plos One. 2013; 31:8(5):e65309.
62. Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD, Horlings HM, Shah N, Umbricht C, Wang P, Wang Y, Kong B, Langerød A and et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nature Genetics. 2011; 43(7):621–629.
63. Gibb EA, Vucic EA, Enfield KSS, Stewart GL, Lonergan KM, Kennett JY, Becker-Santos DD, MacAulay CE, Lam S, Brown CJ and Lam WL. Human cancer long non-coding RNA transcriptomes. PLoS One. 2011; 6(10):e25915
64. Esquela-Kerscher A and Slack FJ. OncomiRs-microRNAs with a role in cancer. Nature Rev. Cancer. 2006; 6:259–269.
65. Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nature Rev. Genet. 2009; 10:704–714.
66. Nicoloso MS, Spizzo R, Shimizu M, Rossi S and Calin GA. MicroRNAs-the micro steering wheel of tumour metastases. Nature Rev. Cancer. 2009; 9:293–302.
67. Calin G. A and et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl Acad. Sci. USA. 2004; 101:2999–3004.
68. Ponting CP, Oliver PL and Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009; 136(4):629–641.
69. Guttman M, Donaghey J, Carey B, Garber M, Grenier JK, Munson G and et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature. 2011; 477295–300.
70. Spizzo R, Almeida MI, Colombatti A and Calin GA. Long non-coding RNAs and cancer: a new frontier of translational research. Oncogene. 2012; 31:4577–4587.
71. Li X, Wu Z, Fu X and Han W. Long noncoding RNAs: insights from biological features and functions to diseases. Med Res Rev. 2013; 33(3):517–553.
72. Chu C, Qu K, Zhong FL, Artandi and Chang H. Genomic maps of long noncoding RNA occupancy reveal principles of RNA-chromatin interactions. Molecular Cell. 2011; 44:667–678.
73. Simon MD, Wang CI, Kharchenko PV, West JA, Chapman BA, Alekseyenko AA, Borowsky ML, Kuroda MI and Kingston RE. The genomic binding sites of a noncoding RNA. Proceedings of the National Academy of Sciences. 2011; 108(51):20497–20502.
74. Gutschner T, Baas M and Diederichs S. Noncoding RNA gene silencing through genomic integration of RNA destabilizing elements using zinc finger nucleases. Genome Research. 2011; 21(11):1944–1954.
75. Tsai M.C and et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010; 329(5992):689–693.
76. Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A, Regev A, Lander ES and Rinn JL. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proceedings of the National Academy of Sciences. 2009; 106(28):11667–11672.
77. Yoon JH, Abdelmohsen K, Kim J, Yang X, Martindale JL, Tominaga-Yamanaka K, White EJ, Orjalo AV, Rinn JL, Kreft SG, Wilson GM and Gorospe M. Scaffold function of long non-coding RNA HOTAIR in protein ubiquitination. Nature Communications. 11(4):2939
78. Hanahan D and Weinberg RA. The Hallmarks of Cancer. Cell;. 2000; 100(1):57–70.
79. Hanahan D and Weinberg RA. Hallmarks of Cancer: The Next Generation. Cell. 2011; 144(5):646–674.
80. Lee JT. Epigenetic regulation by long noncoding RNAs. Science. 2012; 338(6113):1435–9.
81. Fu X, Ravindranath L, Tran N, Petrovics G and Srivastava S. Regulation of apoptosis by a prostate-specific and prostate cancer-associated noncoding gene, PCGEM1. DNA and Cell Biology. 2006; 25(3):135–41.
82. Reis EM, Nakaya HI, Louro R, Canavez FC, Flatschart AV, Almeida GT, Egidio CM, Paquola AC, Machado AA, Festa F, Yamamoto D, Alvarenga R, da Silva CC, Brito GC and et al. Antisense intronic non-coding RNA levels correlate to et al. Antisense intronic non-coding RNA levels correlate to et al. Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer. Oncogene. 2004; 23(39):6684–6692.
83. Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C, Fabbri M, Cimmino A, Lee EJ, Wojcik SE, Shimizu M, Tili E, Rossi S and et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell. 2007; 12(3):215–29.
84. Petherick A. Genetics: The production line. Nature. 2008; 454(7208):1042–5.
85. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E and Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007; 129:1311–1323.
86. Baker M. Long noncoding RNAs: the search for function. Nat. Methods. 2011; 8:379–383.
87. Tsai MC, Wan Y, Manor O, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E and Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010; 329(5992):689–693.
88. Rinn JL and Chang HY. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012; 81:145–166.
89. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B and et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464(7291):1071–1076.
90. Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H and Müller-Tidow C. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003; 22(39):8031–41.
91. Ren S, Wang F, Shen J, Sun Y, Xu W, Lu J, Wei M, Xu C, Wu C, Zhang Z, Gao X, Liu Z, Hou J, Huang J and Sun Y. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer. 2013; 49:2949–2959.
92. Ren S, Liu Y, Xu W, Sun Y, Lu J, Wang F, Wei M, Shen J, Hou J, Gao X, Xu C, Huang J, Zhao Y and Sun Y. Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. Journal of Urology. 2013; 190(6):2278–2287.
93. Guo F, Li Y, Liu Y, Wang J and Li G. Inhibition of metastasis-associated lung adeno-carcinoma transcript 1 in CaSki human cervical cancer cells suppresses proliferation and invasion. Acta Biochimica et Biophysica Sinica (Shanghai). 2010; 42:224–229.
94. Tano K, Mizuno R, Okada T, Rakwal R, Shibato J, Masuo Y, Ijiri K and Akimitsu N. MALAT-1 enhances cell motility of lung adenocarcinoma cells by influencing the expression of motility-related genes. FEBS Lett. 2010; 584:4575–4580.
95. Wang Y, Chen W, Yang C, Wu W, Wu S, Qin X and Li X. Long non-coding RNA UCA1a (CUDR) promotes proliferation and tumorigenesis of bladder cancer. International Journal of Oncology. 2012; 41(1):276.
96. Tsang WP, Wong TWL, Cheung AHH, Co CNN and Kwok TT. Induction of drug resistance and transformation in human cancer cells by the noncoding RNA CUDR. Rna. 2007; 13(6):890–898.
97. Kino T, Hurt DE, Ichijo T, Nader N and Chrousos GP. Noncoding RNA gas5 is a growth arrest-and starvation-associated repressor of the glucocorticoid receptor. Science Signaling. 2010; 3(107):ra8.
98. Qiao H-P, Gao WS, Huo JX and Yang ZS. Long non-coding RNA GAS5 functions as a tumor suppressor in renal cell carcinoma. Asian Pac J Cancer Prev. 2013; 14(2):1077–1082.
99. Sotillo E and Thomas-Tikhonenko A. The long reach of noncoding RNAs. Nature Genetics. 2011; 43:616–617.
100. Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD, Horlings HR, Shah N, Umbricht C, Wang P, Wang U, Kong B, Langered A and et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nature Genetics. 2011; 43:621–629.
101. Tsang W and Kwok T. Riboregulator H19 induction of MDR1-associated drug resistance in human hepatocellular carcinoma cells. Oncogene. 2007; 26(33):4877–4881.
102. Amit D and Hochberg A. Development of targeted therapy for bladder cancer mediated by a double promoter plasmid expressing diphtheria toxin under the control of H19 and IGF2-P4 regulatory sequences. J Transl Med. 2010; 8(1):134.
103. Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM, Buck CR, Denk H, Schroeder R, Trauner M and Zatloukal K. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007; 132(1):330–342.
104. Wang J, Liu X, Wu H, Ni P, Gu Z, Qiao Y, Chen N, Sun F and Fan Q. CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic acids research. 2010; 38(16):5366–5383.
105. Drosophila 12 Genomes Consortium1: Clark AG, Eisen MB, Smith DR, Bergman CM, Oliver B, Markow TA, Kaufman TC, Kellis M, Gelbart W, Iyer VN, Pollard DA, Sackton TB and et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature. 2007; 450(7167):203–18.
106. Zhao J, Ohsumi TK, Kung JT, Ogawa Y, Grau DJ, Sarma K, Song JJ, Kingston RE, Borowsky M and Lee JT. Genome-wide identification of polycomb-associated RNAs by RIP-seq. Molecular Cell. 2010; 40(6):939–953.
107. Wu X, Shao X-Q, Meng XX, Zhang XN, Zhu L, Liu SX, Lin J and X HS. Genome-wide analysis of microRNA and mRNA expression signatures in hydroxycamptothecin-resistant gastric cancer cells. Acta Pharmacologica Sinica. 2011; 32(2):259–269.
108. http://www.cbioportal.org/public-portal/Memorial Sloan-Kettering Cancer Center
109. Ahmad N, Haider S, Anaissie E and Driscoll JJ. MicroRNA theragnostics for the clinical management of multiple myeloma. Leukemia. 2013; 10:1038/leu.2013.262.
110. Redis RS, Berindan-Neagoe I, Pop VI and Calin GA. Non-coding RNAs as theranostics in human cancers. J. Cell Biochem. 2012; 113:1451–1459.
111. Roberts TC, Morris KV and Weinberg MS. Perspectives on the mechanism of transcriptional regulation by long non-coding RNAs. Epigenetics. 2014; 1; 9(1):13–20.
112. Beck AK, Pass HI, Carbone M and Yang H. Ranpirnase as a potential antitumor ribonuclease treatment for mesothelioma and other malignancies. Future Oncol. 2008; 4:341–349.
113. Bouchie A. First microRNA mimic enters clinic. Nature Biotechnology. 2013; 31:577.
114. Craig VJ, Tzankov A, Flori M, Schmid CA, Bader AG and Muller A. Systemic microRNA-34a delivery induces apoptosis and abrogates growth of diffuse large B-cell lymphoma in vivo. Leukemia. 2012; 26:2421–2424.
115. Huang L and Liu Y. In vivo delivery of RNAi with lipid-based nanoparticles. Annual Review of Biomedical Engineering. 2011; 13:507–530.
116. Bartlett DW, Su H, Hildebrant IJ, Weber WA and Davis ME. Impact of tumor-specific targeting on the biodistribution and efficacy of siRNA nanoparticles measured by multimodality in vivo imaging. Proceedings of the National Academy of Sciences. 2007; 104(39):15549–15554.
117. Li YL, Zhu L, Liu Z, Cheng R, Meng F, Cui JH, Ji SJ and Zhong Z. Reversibly stabilized multifunctional dextran nanoparticles efficiently deliver doxorubicin into the nuclei of cancer cells. Angewandte Chemie International Edition. 2009; 48(52):9914–9918.
118. Sidi AA, Sidi AA, Ohana P, Benjamin S, Shalev M, Ransom JH, Lamm D, Hochberg A and Leibovitch I. Phase I/II marker lesion study of intravesical BC-819 DNA plasmid in H19 over expressing superficial bladder cancer refractory to bacillus Calmette-Guerin. Journal of Urology. 2008; 180(6):2379–2383.
119. Scaiewicz V, Sorin V, Fellig Y, Birman T, Mizrahi A, Galula J, Abu-Lail R, Shneider T, Ohana P, Buscail L, Hochberg A and Czerniak A. Use of H19 gene regulatory sequences in DNA-based therapy for pancreatic cancer. Journal of Oncology. 2010; 178174.
120. Chi J, Ballabio E, Chen XH, Kusec R, Taylor S, Hay D, Tramonti D, Saunders NJ, Littlewood T, Pezzella F, Boultwood J, Wainscoat J.S, Hatton C.S and et al. McroRNA expression in multiple myeloma is associated with genetic subtype, isotype and survival. Biology Direct. 2011; 6:23.
121. Roccaro AM, Sacco A, Chen C, Runnels J, Leleu X, Azab F, Azab AK, Jia X, Ngo HT, Melhem MR, Burwick N, Varticovski L, Novina CD, Rollins BJ, Anderson KC and Ghobrial IM. MicroRNA expression in the biology, prognosis, and therapy of Waldenstrom macroglobulinemia. Blood. 2009; 113:4391–402.
122. Roccaro AM, Sacco A, Thompson B, Azab F, Runnels J, Jia X, Ngo HT, Melhem MR, Lin CP, Ribatti D, Rollins BJ, Witzig TE, Anderson KC and Ghobrial IM. MicroRNAs 15a and 16 regulate tumor proliferation in multiple myeloma. Blood. 2009; 113:6669–80.

