INTRODUCTION
Spermiogenesis is a highly complex morphological process that lasts from the end of the meiosis to the release of mature sperm within the seminiferous tubule of the testis. During the process of spermiogenesis, the spermatids undergo a structural reorganization involving development of the acrosome, condensation of the nuclear material, rearrangement of the mitochondria and removal of unnecessary cytoplasm [1, 2]. All of the above events are interconnected by different cellular processes, and failure of any of these processes during the differentiation of spermatids can cause various abnormalities within the morphology of spermatozoa [3, 4]. Although these morphological changes during spermiogenesis have been well documented, the cellular mechanisms of unnecessary cytoplasm shedding and acrosomal changes are still unknown [5, 6]. Thus, we hypothesized that autophagy may be involved in spermiogenesis within male haploid germ cells.
Autophagy is an intercellular pathway that delivers cytoplasmic components or organelles to lytic compartments such as lysosomes (in mammals) or vacuoles (in yeast) for breakdown and recycling [7, 8]. First, a double-membraned cupped structure known as phagophore develops; it engulfs some portion of cytoplasm and then closes to form a double membrane-bound vesicle called the autophagosome. This process of autophagosome development consists of three major steps: initiation of membrane biogenesis, elongation of the isolation membrane, and closure of the membrane [9]. Autophagy is generally induced by starvation, but recent findings have linked it to numerous physiological and pathological conditions, such as normal development, placental detachment during birth in mammals, cancer, programmed cell death, neurodegenerative disorders, diabetes and infections [10–13].
The current biology of autophagy was revolutionized following the identification of so-called Atg (autophagy) genes within yeast, most of which are conserved in mammals [14, 15]. ATG7 is one particular member of the ATG protein family that acts as an E1-like activating enzyme that facilitates both LC3 and Atg12 [16]. LC3 is a microtubule-associated light protein chain 3, upon induction of autophagy LC3-I is conjugated to phosphatidylethanolamine to form LC3-II, which is tightly bound to the membrane of the autophagosome. Microtubule-associated protein 1 light chain 3-II is a marker of autophagosomes [17].
The origins of autophagosome membrane biogenesis may involve various sources [18, 19], including ER exit sites (ERES) [20, 21], the ER–Golgi intermediate compartment (ERGIC) [22], the Golgi [23, 24], the recycling endosome [19, 25, 26] and the plasma membrane [9]. Despite significant progress in identifying molecules (Atg genes) responsible for autophagosome formation, the origin and exact source of the autophagosomal membrane are still unclear [9]. Thus, the objectives of the current study were to determine the in vivo role of autophagy and origin of autophagosome membrane biogenesis within male haploid germ cells during spermiogenesis in the rat testis.
RESULTS
LC3 and ATG7 are mainly localized in the male haploid cells
Light microscopy revealed that the seminiferous tubules (STs) of rat testes contained different stages of developing germ cells and Sertoli cells. Spermatogonia were found near the basal membrane and primary spermatocytes, including some early round spermatids that were located in the basal compartments of the STs. Elongated spermatids were observed near the lumens of STs. The interstitial space between STs contained numerous Leydig cells and blood vessels (Figure 1). IHC for LC3 within the seminiferous tubules showed no positive localization around the basal membranes within the spermatogonia or primary spermatocytes or on the Sertoli cells. While weak immunoreactivity was noted in the round spermatids and high expression was found on elongated spermatids, localization increased from the basal to the luminal compartments, corresponding with the progression of spermiogenesis (Figure 2A and 2B). Similarly, immunofluorescence for GPF-LC3 revealed a strong focal localization around spermatids and progressively increased in elongated spermatids towards the lumen, this focal localization (light green spots) indicates the autophagosomes (Figure 2C and 2D). Furthermore, ATG7 showed no localization on spermatogonia or Sertoli cells; weak positive expression was noted in the primary spermatocytes and in the round spermatids around their basal compartments. Immunoreactivity was highly increased within the elongated spermatids near their lumens, as observed for LC3 (Figure 3A and 3B).
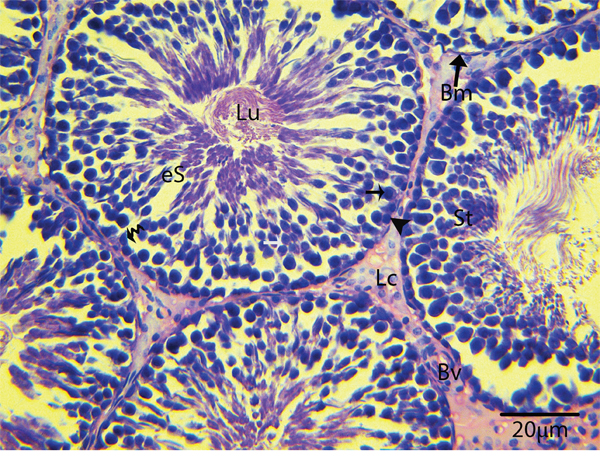
Figure 1: Light micrograph showing the histological structure of rat testis. St: seminiferous tubule; Bm: basal membrane; Lu: lumen; eS: elongated spermatid; Lc: Leydig cell; Bv: blood vessels; (▲): spermatogonia; (black arrow): primary spermatocytes. H & E stain. Scale bar = 20μm.
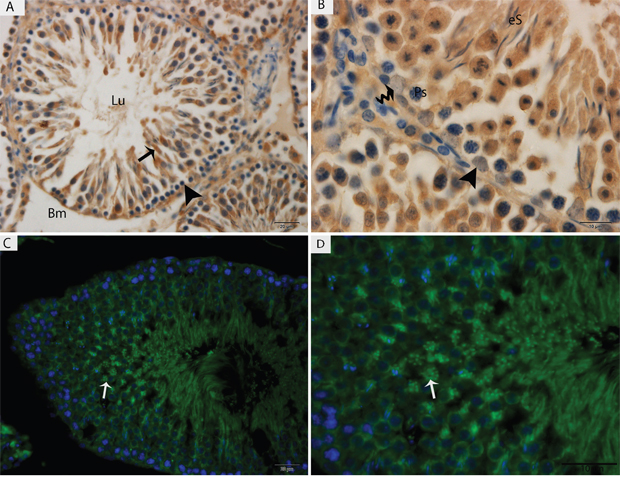
Figure 2: Light micrograph of LC3 localization in the rat testis. (A, B) Immunohistochemistry, (C, D) immunofluorescence. Lu: lumen; Bm: basal membrane; Ps: primary spermatocytes; eS: elongated spermatid; (▲): spermatogonia. Scale bars = 20μm (A, C) and 10μm (B, D).
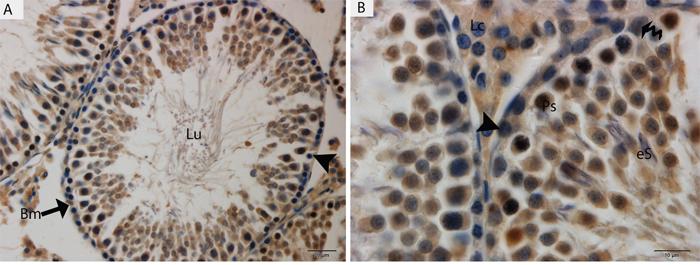
Figure 3: Light micrograph of ATG7 localization in the rat testis. Lu: lumen; Bm: basal membrane; Ps: primary spermatocytes; eS: elongated spermatid; Lc: Leydig cell; (▲): spermatogonia. Scale bars = 20μm (A) and 10μm (B).
Lysosomes and autophagosomes appear within the male haploid germ cells
TEM findings revealed that the STs of rat testes consist of Sertoli cells, spermatogonia, primary spermatocytes and round spermatids within the basal compartment, while elongated spermatids and several lysosomes are located near the luminal compartment (Figure 4A). Spermatogonia and Sertoli cells were found to rest directly on the basal membranes, and their cytoplasm contained extremely long mitochondria. Mitochondria within the round spermatids were vacuolated and either round or oval; moreover, these mitochondria lined up within the cytoplasm near the periphery of cell membrane. Acrosomal vesicles were attached to the nuclei of round spermatids, indicating the beginning of acrosome formation; meanwhile, Golgi apparatuses and endoplasmic reticula were also observed (Figure 4B, 4C and 4D). During further formation of the acrosome and progression of nuclear elongation, mitochondria were found to be scattered within the cytoplasm and moving toward the mid-piece of the elongated spermatids. At this stage, autophagosomes and some lysosomes appeared within the cytoplasm of elongated spermatids (Figure 5A, 5B, 5C and 5D). The elongated spermatids, which were located near the lumens of the STs, contained large size lysosomes. Meanwhile, the number of autophagosomes also increased within the cytoplasm of these elongated spermatids as spermiogenesis progresses, corresponding to the latter stages of spermiogenesis. At this stage, mitochondria lined up around the mid-piece and appeared dense (Figures 6–8).
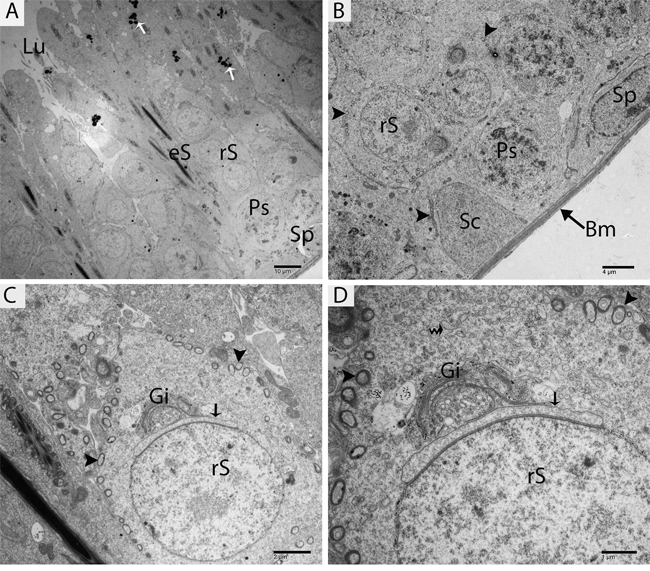
Figure 4: TEM micrograph of a germ cell at an early stage of development and Sertoli cells. Sp: spermatogonia; Ps: primary spermatocytes; rS: round spermatid; eS: elongated spermatid; SC: Sertoli cell; Bm: basal membrane; Lu: lumen; Gi: Golgi apparatus; (▲): mitochondrion; (arrow): acrosomal vesicle; (curved arrow): endoplasmic reticulum; (white arrow): lysosome. Scale bars = 10μm (A), 4μm (B), 2μm (C) and 1μm (D).
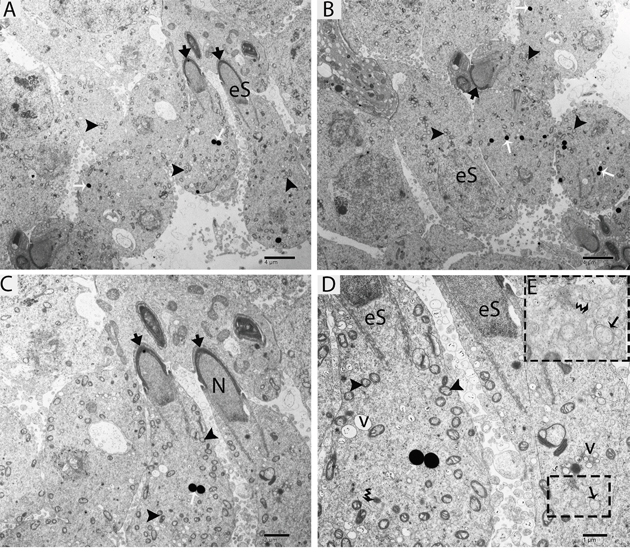
Figure 5: TEM micrograph of an elongated spermatid at an early stage showing lysosomes and autophagosomes. eS: elongated spermatid; (arrowhead): acrosome; N: nucleus; v: vesicle; (▲): mitochondrion; (black arrow): autophagosome; (white arrow): lysosome; (curved arrow): endoplasmic reticulum. Scale bars =4μm (A, B), 2μm (C) and 1μm (D).
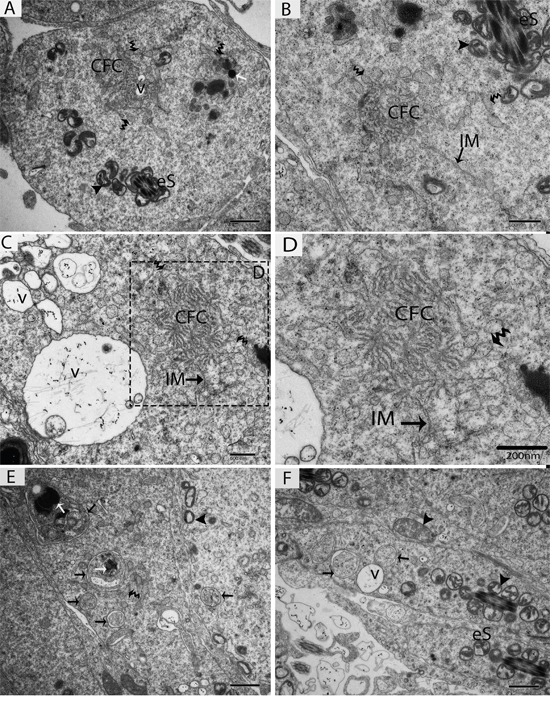
Figure 6: TEM micrograph of a late stage of spermiogenesis showing autophagosome biogenesis. eS: elongated spermatid; CFC: chrysanthemum flower center; IM: isolation membranes; v: vesicle; (▲): mitochondrion; (black arrow): autophagosome; (white arrow): lysosome; (curved arrow): endoplasmic reticulum. Scale bars = 1μm (A, E, F) and 600nm (B, C,) and 200nm (D).
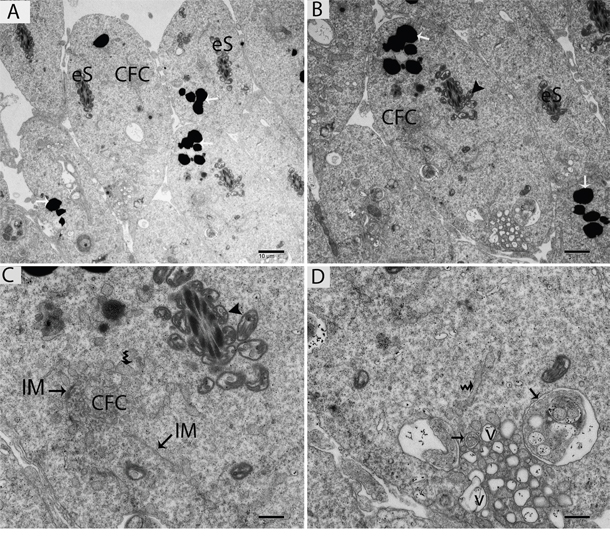
Figure 7: TEM micrograph of an elongated spermatid showing large numbers of lysosomes and autophagosomes near the lumen of a seminiferous tubule. eS: elongated spermatid; CFC: chrysanthemum flower center; v: vesicle; (▲): mitochondrion; (black arrow): autophagosome; (white arrow): lysosome; (curved arrow): endoplasmic reticulum; IM: isolation membranes. Scale bars = 10μm (A), 5μm (B), 2μm (C) and 1μm (D).
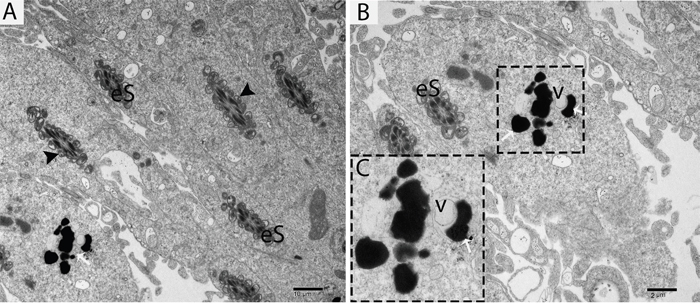
Figure 8: TEM micrograph of an elongated spermatid showing a vesicle attached to a lysosome. eS: elongated spermatid; v: vesicle; (▲): mitochondrion; (white arrow): lysosome. Scale bars = 10μm (A), 2μm (B) and 500nm (C).
Origin of autophagosome membrane biogenesis
Further TEM analysis revealed that autophagosome membrane formation originated in the endoplasmic reticulum (ER) of differentiating spermatids. From the ER, a structure developed whose name we coined as the “chrysanthemum flower center” (CFC) because it resembled a chrysanthemum flower. Numerous branches of the ER were distributed around the CFC, and the ER ends attached to the CFC through short narrow tubules that formed many daisy petal-like structures. Meanwhile, several vesicles were found between and around the CFC (Figures 6B, 6C, 6D, 6E, 7B and 7C). Denser double-layer isolation membranes (IM) appeared in the CFC, and these membranes elongate and enwrap cytoplasm along with different structures to develop into autophagosomes as previously reported by our research group in the turtle [5]. Autophagosomes were attached to vesicles and lysosome; lysosomes and vesicles were also found inside the autophagosome along with other structures (Figures 6E, 6F and 7D). The lysosome was found closely attached to the vesicle membranes located near the lumen of ST in elongated spermatids (Figure 8A and 8B).
DISCUSSION
During spermiogenesis, efficient removal of cytoplasm is thought to be important for the generation of motile and functional spermatozoa, but this mechanism remains poorly understood [27]. Previously proposed mechanisms suggest that the elimination of residual bodies and cytoplasm are mediated by the Sertoli cells via engulfment [28, 29]. However, the role of the germ cells in this process is largely understudied [30]. In the current work, we found that in vivo autophagy developed within haploid germ cells as elongation progressed. Our immunohistochemistry for LC3 and ATG7 showed expression increased dramatically in elongated spermatids as spermiogenesis progresses from basal toward the luminal compartment of ST. This expression pattern suggests that autophagy might be involved in some process in elongated spermatids. Shang and his colleagues [30] reported similar patterns of expression of LC3A/B and ATG7 in germ cell-specific atg7 knockout mice, which exhibit multiple abnormalities in their spermatozoa. In addition, we also found some positive signals of ATG7 within the primary spermatocytes. Recently, reported that the depletion of ATG7 in germ cells did not affect the early stages of development of germ cells in the Atg7-knockout mice [6]. The cytoskeleton-based structures, such as F-actin-containing cytoplasmic manchettes, are also disrupted after the ablation of ATG7, consequently impairing acrosome formation and preventing complete cytoplasm removal. ATG7 is an E1-like activating enzyme that plays an essential role in the 2 ubiquitin-like conjugation system of autophagy. It is essential for ATG12 conjugation and for LC3 association with the membrane [12].
Further, our transmission electron microscopy showed that lysosomes and autophagosomes appeared within elongated spermatids during acrosome formation and that their number increased as elongation progressed toward the lumens of seminiferous tubules. Similar to our immunohistochemistry (LC3 and ATG7) findings, TEM showed no consistent evidence of autophagy within the Sertoli cells or early germ cell stages (spermatogonia, primary spermatocytes and early round spermatids). These results suggest that these autophagosomes and lysosomes might have role in removal of extra cytoplasm from the spermatids during spermiogenesis. Moreover, autolysosomes are essential for acrosome development, while mere lysosomes are not. Acrosome biogenesis is highly disrupted during early stages of spermiogenesis in spermatids, as the proacrosomal vesicles are unable to fuse into a single acrosomal vesicle during the Golgi phase secondary to ATG7 depletion [6]. Together, these findings collectively suggest that the autophagy is an essential for the process of spermiogenesis.
Autophagy depends on the formation of double-membraned autophagosomes. Unlike other organelles that appear to stably exist in the cell, autophagosomes only appear when required [31]. It is difficult to confirm the sources of autophagosomal membranes because autophagosome formation occurs very rapidly once initiated. It is estimated that autophagosome formation takes only a few minutes in yeast and mammals [11, 24, 32]. How and where this dynamic autophagosome formation occurs is still unknown [8]. The TEM findings of the current study revealed a novel structure that we named the “chrysanthemum flower center” (CFC). The CFC developed from the endoplasmic reticulum (ER), and then isolation membranes (phagophores) arose from the CFC. Phagophore elongate and envelop the cytoplasm to develop autophagosomes. We recently reported the more clear evidence of chrysanthemum flower center in the spermatids of Chinese soft-shelled turtle, because it is better model to study different events of spermiogenesis then the rat. The CFC was more developed in spermatids with compact nuclei than in spermatids with granular nuclei in the turtle [5].
The current work showed that the ER might be a basic source of autophagosome membrane biogenesis. A strong relationship between ER and autophagosome formation sites have been observed in mammalian cell culture [8]. Additionally, 70% of isolation membranes (IM) were associated with the ER (named ER–IM complexes) in Atg4B (Atg4BC74A) mutant cells [33]. In higher eukaryotes, phagophores nucleate at an intricate membranous structure called the omegasome, which was originally identified as being associated with the ER [34]. Our findings are consistent with above studies, while additionally revealing for the first time that the CFC is a novel structure that develops from the ER and that the IM develops from the CFC. Moreover, recent studies suggest that endoplasmic reticulum exit sites (ERES), specialized ER regions where proteins are sorted into the secretory system, are key players in the formation of autophagosomes. Therefore, ERES are a new element that should be integrated into descriptions of autophagosome biogenesis [35]. In yeast, the membrane of the phagophore originates from a structure near the vacuole called the pre-autophagosomal structure (PAS) [36]. We also found several vesicles at the different stages of autophagosomes biogenesis.
In conclusion, the numbers of lysosomes, CFCs, phagophores and autophagosomes increase within male haploid cells as spermiogenesis progresses. Additionally, the endoplasmic reticulum might be a source of autophagosome membrane before phagophore formation via a newly described structure that we refer to as the “chrysanthemum flower center” (CFC). These findings may initiate a new area of focus for understanding the origin of the autophagosome membrane.
MATERIALS AND METHODS
Animals
Six three-month-old male albino rats were purchased from Nanjing, Jiangsu Province, China for this study. Rats were initially weighted (170-200 gm) and were reared for 10 days under a pathogen-free environment. Then, all rats were rendered comatose using intraperitoneally administered sodium pentobarbital (20 mg/animal) and were then sacrificed by cervical dislocation. The testes were collected immediately and fixed to performed different techniques (details below). Sample preparation was conducted according to accepted international standards and was approved by the Ethics Committee for Animal Care and Use by the Science and Technology Agency of Jiangsu Province (SYXK (SU) 2010-0009).
Light microscopy
The testes were fixed in 10% neutral buffered formalin overnight, and then paraffin blocks were prepared. Sectioning was performed at 6μm. The sections were stained with hematoxylin and eosin procedures (Harry’s hematoxylin for 2 min and 1% eosin for 30 sec) for light microscopic analysis using an Olympus microscope (BX53), camera (Olympus DP73, Japan).
Immunohistochemistry (IHC)
Immunohistochemical staining for LC3 and ATG7 was performed according to the manufacturer’s recommendations and as suggested in previous studies [37, 38]. The sections were processed using a standard immunohistochemistry protocol as previously described [39]. Briefly, after deparaffinization, blocking endogenous peroxidase, microwave antigen retrieval, and BSA (bovine serum albumin) blocking, a primary rabbit anti-LC3B polyclonal antibody (ab48394, Abcam, Cambridge, UK) or mouse anti-ATG7 monoclonal antibody (SAB4200304, Sigma Aldrich, St. Louis, USA) was applied for 1 h at room temperature. After washing with PBS, the slides were incubated for 30 min with biotinylated goat anti-rabbit antibody. Peroxidase was visualized with DAB, and the sections were counterstained with hematoxylin. Sections incubated in PBS alone served as negative controls.
Fluorescence microscopy
After deparaffinization, serum blocking and antigen retrieval, the tissue sections of testis were incubated with the primary antibody (LC3) at 4°C overnight. Next, the slides were washed in PBS and corresponding secondary antibodies were added to sections for 2 h at room temperature. After washing again with PBS, then DAPI stain to mark the nuclei. Images were taken immediately using an Olympus microscope (BX53), camera (Olympus DP73), Japan.
Transmission electron microscopy (TEM)
Samples were cut into small pieces and then immersed in 2.5% glutaraldehyde in PBS (4°C, pH 7.4, 0.1 M) overnight. Tissues were rinsed in PBS and then post-fixed for 60 min at room temperature in the same way by using buffered 1% osmium tetroxide (Polysciences Inc. Warrington, PA, USA) and then washed in the buffer. The samples were then dehydrated in ascending concentrations of ethyl alcohol, infiltrated with a propylene oxide-araldite mixture and then embedded in araldite. The blocks were then sectioned using an ultramicrotome (Reichert Jung, Wien, Austria), and the ultrathin sections (50 nm) were mounted on copper-coated grids. The pieces were stained with 1% uranyl acetate and Reynold’s lead citrate for 20 min. Finally, the samples were examined and photographed using a high-resolution digital camera (16 mega pixel) connected to a TEM (Hitachi H-7650, Japan).
Author contributions
Ping Yang and Nisar Ahmed designed the experiments and drafted the manuscript, Wang Linling, Hong Chen, Yasir Waqas, Tengfei Liu, Abdul Haseeb, Nasrullah Bangulzai and Yufei Huang participated in the study design and performed data analysis. Prof.Q. Chen conceived the study and participated in its design and coordination and helped draft the manuscript. All authors read and approved the final manuscript.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (Nos: 31672505 and 31402155) and China Postdoctoral Science Foundation Funded Project (No: 2016M591861), and the Priority Academic Program Development of Jiangsu Higher Education Institutions, PAPD.
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
REFERENCES
1. Zhang L, Han XK, Li MY, Bao HJ, Chen QS. Spermiogenesis in soft-shelled turtle, Pelodiscus sinensis. Anat Rec. 2007; 290:1213-1222.
2. Hermo L, Pelletier R, Cyr DG, Smith CE. Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 1: background to spermatogenesis, spermatogonia, and spermatocytes. Microsc Res Tech. 2010; 73:241-278.
3. Calvel P, Rolland AD, Jégou B, Pineau C. Testicular postgenomics: targeting the regulation of spermatogenesis. Philos Trans R Soc Lond B Biol Sci. 2010; 365:1481-1500.
4. Matzuk MM, Lamb DJ. The biology of infertility: research advances and clinical challenges. Nat Med. 2008; 14:1197-1213.
5. Huang Y, Yang P, Liu T, Chen H, Chu X, Ahmad N, Zhang Q, Li Q, Hu L, Liu Y. Subcellular evidence for biogenesis of autophagosomal membrane during spermiogenesis in vivo. Front Physiol. 2016; 7:470.
6. Wang H, Wan H, Li X, Liu W, Chen Q, Wang Y, Yang L, Tang H, Zhang X, Duan E. Atg7 is required for acrosome biogenesis during spermatogenesis in mice. Cell Res. 2014; 24:852-869.
7. Marzella L, Glaumann H. Autophagy, microautophagy and crinophagy as mechanisms for protein degradation. Lysosomes: Their Role in Protein Breakdown. London: Academic Press, 1987; 319-367.
8. Hamasaki M, Shibutani ST, Yoshimori T. Up-to-date membrane biogenesis in the autophagosome formation. Curr Opin Cell Biol. 2013; 25:455-460.
9. Abada A, Elazar Z. Getting ready for building: signaling and autophagosome biogenesis. EMBO Rep. 2014; 15:839-852.
10. Rubinsztein DC, Shpilka T, Elazar Z. Mechanisms of autophagosome biogenesis. Curr Biol. 2012; 22:29-34.
11. Mizushima N, Yoshimori T, Ohsumi Y. The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol. 2011; 27:107-132.
12. Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell. 2011; 147:728-741.
13. Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature. 2008; 451:1069-1075.
14. Thumm M, Egner R, Koch B, Schlumpberger M, Straub M, Veenhuis M, Wolf D. Isolation of autophagocytosis mutants of Saccharomyces cerevisiae. FEBS Lett. 1994; 349:275-280.
15. Tsukada M, Ohsumi Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 1993; 333:169-174.
16. Feng Y, Yao Z, Klionsky DJ. How to control self-digestion: transcriptional, post-transcriptional, and post-translational regulation of autophagy. Trends Cell Biol. 2015; 25:354-363.
17. Kabeya Y, Mizushima N, Ueno T, Yamamoto A, Kirisako T, Noda T, Kominami E, Ohsumi Y, Yoshimori T. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 2000; 19:5720-5728.
18. Hayashi-Nishino M, Fujita N, Noda T, Yamaguchi A, Yoshimori T, Yamamoto A. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat Cell Biol. 2009; 11:1433-1437.
19. Puri C, Renna M, Bento CF, Moreau K, Rubinsztein DC. Diverse autophagosome membrane sources coalesce in recycling endosomes. Cell. 2013; 154:1285-1299.
20. Zoppino FC, Militello RD, Slavin I, Alvarez C, Colombo MI. Autophagosome formation depends on the small GTPase Rab1 and functional ER exit sites. Traffic. 2010; 11:1246-1261.
21. Graef M, Friedman JR, Graham C, Babu M, Nunnari J. ER exit sites are physical and functional core autophagosome biogenesis components. Mol Biol Cell. 2013; 24:2918-2931.
22. Ge L, Melville D, Zhang M, Schekman R. The ER–Golgi intermediate compartment is a key membrane source for the LC3 lipidation step of autophagosome biogenesis. eLife. 2013; 2:e00947.
23. Bodemann BO, Orvedahl A, Cheng T, Ram RR, Ou YH, Formstecher E, Maiti M, Hazelett CC, Wauson EM, Balakireva M. RalB and the exocyst mediate the cellular starvation response by direct activation of autophagosome assembly. Cell. 2011; 144:253-267.
24. Geng J, Baba M, Nair U, Klionsky DJ. Quantitative analysis of autophagy-related protein stoichiometry by fluorescence microscopy. J Cell Biol. 2008; 182:129-140.
25. Longatti A, Lamb CA, Razi M, Yoshimura S, Barr FA, Tooze SA. TBC1D14 regulates autophagosome formation via Rab11-and ULK1-positive recycling endosomes. J Cell Biol. 2012; 197:659-675.
26. Knævelsrud H, Søreng K, Raiborg C, Håberg K, Rasmuson F, Brech A, Liestøl K, Rusten TE, Stenmark H, Neufeld TP. Membrane remodeling by the PX-BAR protein SNX18 promotes autophagosome formation. J Cell Biol. 2013; 202:331-349.
27. Cheng CY, Mruk DD. The biology of spermatogenesis: the past, present and future. Philos Trans R Soc Lond B Biol Sci. 2010; 365:1459-1463.
28. Ahmed N, Yufei H, Yang P, Muhammad Yasir W, Zhang Q, Liu T, Hong C, Lisi H, Xiaoya C, Chen Q. Cytological study on Sertoli cells and their interactions with germ cells during annual reproductive cycle in turtle. Ecol Evol. 2016, 6:4050-4064.
29. Carr I, Clegg E, Meek G. Sertoli cells as phagocytes: an electron microscopic study. J Anat. 1968; 102:501-506.
30. Shang Y, Wang H, Jia P, Zhao H, Liu C, Liu W, Song Z, Xu Z, Yang L, Wang Y, Li W. Autophagy regulates spermatid differentiation via degradation of PDLIM1. Autophagy. 2016; 12:1575-1592.
31. Shibutani ST, Yoshimori T. A current perspective of autophagosome biogenesis. Cell Res. 2014; 24:58-68.
32. Fujita N, Hayashi-Nishino M, Fukumoto H, Omori H, Yamamoto A, Noda T, Yoshimori T. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol Biol Cell. 2008; 19:4651-4659.
33. Hamasaki M, Yoshimori T. Where do they come from? Insights into autophagosome formation. FEBS Lett. 2010; 584:1296-1301.
34. Carlsson SR, Simonsen A. Membrane dynamics in autophagosome biogenesis. J Cell Sci. 2015; 128:193-205.
35. Sanchez-Wandelmer J, Ktistakis NT, Reggiori F. ERES: sites for autophagosome biogenesis and maturation? J Cell Sci. 2015; 128:185-192.
36. Suzuki K, Ohsumi Y. Current knowledge of the pre-autophagosomal structure (PAS). FEBS Lett. 2010; 584:1280-1286.
37. Obert LA, Sobocinski GP, Bobrowski WF, Metz AL, Rolsma MD, Altrogge DM, Dunstan RW. An immunohistochemical approach to differentiate hepatic lipidosis from hepatic phospholipidosis in rats. Toxicol Pathol. 2007; 35:728-734.
38. Eid N, Ito Y, Maemura K, Otsuki Y. Elevated autophagic sequestration of mitochondria and lipid droplets in steatotic hepatocytes of chronic ethanol-treated rats: an immunohistochemical and electron microscopic study. J Mol Histol. 2013; 44:311-326.
39. Liu T, Chu X, Huang Y, Yang P, Li Q, Hu L, Chen H, Chen Q. Androgen-related sperm storage in oviduct of Chinese soft-shelled turtle in vivo during annual cycle. Sci Rep. 2016; 6:20456.

