The Prevalence of Natural Antisense Transcript (NAT) in Vertebrates
Natural antisense transcript (NAT) is defined as any RNA transcript that is complementary to an endogenous RNA transcript [1]. Our knowledge about NAT began from viral transcripts. In 1969, Bovre K et al found the central b2 region in the coliphage λ genome could produce two opposite oriented mRNA, one originating on the plus strand and the other on the minus strand, which partially overlap with each other [2]. Later, similar transcriptional events were identified in prokaryotes [3] and eukaryotes [4]. Systematic identification of NATs in mammalian cells began in 2002 utilizing a microbead capture double strand cDNA strategy [5]. Genome-wide analyses show that more than 63% of transcripts have antisense (AS) partners, most of which represent non-protein-coding RNAs [6]. In human genome, 22% of the human transcription clusters form sense (S)/antisense (AS) pairs. Through in-depth analysis of the functional elements in the human genome (1% coverage), the Encyclopedia of the DNA element (ENCODE) pilot project discovered that almost all DNA is transcribed into RNA, producing a large number of overlapping transcripts [7]. Several methods are utilized to identify NATs and are reviewed elsewhere [8]. Because NATs are found in low abundance, they are often lost in transcriptome analysis. Analysis of 29 human LongSAGE libraries demonstrated that antisense tag only contributes 8% of the total transcript copies [9]. The percentage of NATs is 2.8% in worm, 11% in yeast and 12% in malaria [10-12]. With the completed sequencing of the human genome, the gene numbers has been revised to 25,000 [13] because more transcripts continue to be discovered, many of which do not correspond to annotated genes, but to intergenic and intronic regions. Most AS transcripts do not have protein-coding capacity [14]. At present, the Unigene database of the National Center for Biotechnology Information indicates 123,200 entries for human and 79,222 for mouse. To date, protein-coding transcripts just account for ~1.5% genome, and the remainder transcribe into a large amount of non-coding RNAs, of which NATs are dominant [13].
Underlying Mechanisms of NAT Function
Most S/AS transcript pairs are conserved in evolution and are concurrently or inversely expressed in a cell suggesting a correlative functional role. Six mechanisms widely dictate how NAT’s function: (a) DNA replication interference, (b) chromatin remodeling, (c) transcriptional interference, (d) RNA masking, (e) double-stranded RNA (dsRNA)-dependent mechanisms and (f) translation interference [15]4. We will discuss the general principles of each NAT mechanism followed by a specific example.
(a) DNA replication interference
DNA replication can be interfered and thereby inhibited by NATs. This was demonstrated elegantly in the E. coli plasmid ColE1. The ColE1 primer transcript (RNA II) binds to the template DNA near replication origin. The plasmid replication is dependent on the primer formation (RNA II). Hybridization of RNA II to DNA is inhibited when a second RNA molecule RNA 1 (NAT RNA) binds to RNA II, which results in the alteration of secondary structure of RNA II [12].
(b) Chromatin Remodeling
Chromatin remodeling is generally defined as any event that alters the nuclease sensitivity of a region of chromatin. These events can occur independently or in concert with other events, such as transcription [16]. S/AS pairs are especially abundant in imprinted loci, suggesting the putative role of non-coding RNA in gene silencing [6]. For example, in leukemia, an inverse relationship was observed between p15 antisense (p15AS) and p15 sense expression. The p15AS induces p15 silencing in cis and in trans through heterochromatin formation in leukemic cells and in mouse ES cells.
(c) Transcriptional interference
Transcriptional interference refers to the direct negative impact of one transcriptional activity on a second transcriptional activity in cis. This happens through 3 kinds of mechanisms: (1) occlusion in which the passage of elongating RNA polymerases (RNAPs) blocks the access to the promoter; (2) collisions between elongating RNAPs, moving in opposite directions, leading to premature termination of transcription; and (3) ‘sitting duck’ interference which refers to the removal of promoter-bound complexes by the passage of RNAPs from the opposing promoter [17]. A non-translated AS RNA (RnaG) transcribes in cis on the complementary strand of icsA, a virulence gene of Shigella flexneri. The icsA and RnaG promoters are in opposite direction and 120 bp apart. In vitro transcription from the strong RnaG promoter (aggressive) dramatically inhibits transcription from the weaker icsA promoter (sensitive) [18].
(d) RNA masking
RNA duplex formation may mask critical regulatory features within transcripts, thereby inhibiting the binding of other trans-acting factors. RNA masking intervenes mRNA splicing, transport, polyadenylation, translation and degradation [19]. Zeb2 NAT involved in epithelial–mesenchymal transition (EMT) uses this mechanism. Efficient translation of Zeb2 does not depend on 5’ cap structure, but on internal ribosome entry site (IRES) embedded in an intron sequence of Zeb2 mRNA. In epithelial cells, Zeb2 mRNA is fully spliced and cannot be translated. During EMT, Zeb2 NAT complements with the 5’ splice site of the intron containing IRES in the 5’ UTR of Zeb2, which prevents the splicing of the intron required for efficient translation and expression of the Zeb2 protein [20].
(e) Double stranded RNA-dependent mechanism
Duplex RNA can be recognized by RNAi machinery. These dsRNAs are rapidly processed into short RNA duplexes of 21 to 28 nucleotides in length, which then guide the recognition and ultimately the cleavage or translational repression of complementary single-stranded RNAs, such as messenger RNAs or viral genomic/anti-genomic RNAs. In a recent experiment in X. Laevis, double stranded Slc34a/antisense Slc34a injected into the nucleus of xenopus oocytes were degraded into short RNAs of ~23 bases within 4 h [21]. Similarly, more than 100 NATs have been processed into siRNA in D. melanogaster [10].
(f) Translation interference
NATs can also prevent transcripts from translating by simply interfering with the translation apparatus. Antisense PU.1 regulates PU.1 expression level via translation interference. The absolute levels of PU.1, a transcription factor required for normal hematopoiesis is critical for specifying cell fate, and, if perturbed, even modest decreases in PU.1 can lead to leukemias and lymphomas. Antisense PU.1 forms complex with three translation factors eIF4A, eEF1A and 4E-BP1. The complex stalls translation between the initiation and elongation steps [22].
NATs Role in Pathology
Although the physiological level of NATs is very low, their levels go up dramatically in disease. For example, β-secretase-1 (BACE1) is a crucial enzyme in Alzheimer’s disease pathophysiology and antisense BACE1 (BACE1-AS) is markedly up-regulated in brain samples from Alzheimer’s disease patients [23]. Similarly, antisense ApoE is 100-fold up-regulated in spinal cord-injured C57BL/6 mice compared with normal levels [24].
Tumor pathology often observes AS RNA level changes and in turn shows aberrant regulation of cognate transcripts. In non-papillary clear-cell renal tumors, the NAT of hypoxia inducible factor α (HIFα) increases 10- to 100-fold [25]. Interestingly, a positive correlation of differential expression was observed for most sense and antisense transcript pairs between normal and malignant breast samples [26]. This suggests that aberrant expression of NATs may provide value as a diagnostic reagent. For neuroblastoma (NB), pediatric brain tumor condition, the ratio of antisense MYCN to MYCN is directly correlated with NB disease stage. In the more advanced NB stages and NBs with MYCN-amplification, relatively more MYCN-AS is present as compared to MYCN. Further, expression of the antisense gene MYCN-AS is speculated to be relevant to the progression of NB [27].
Similar to tumor studies, the relevance of AS RNA levels to progression of cardiovascular disease is unknown. In a recent study, the analysis in peripheral blood mononuclear cells from 1098 patients of coronary artery disease demonstrated positive correlation of AS noncoding RNA in the INK4 locus (ANRIL) expression with atherosclerosis [28]. Our recent study on non-coding RNA identified a natural AS transcript for tyrosine kinase containing immunoglobulin and epidermal growth factor homology domain-1 (tie-1), tie-1AS lncRNA in zebrafish, mouse and humans. In embryonic zebrafish, tie-1AS lncRNA transcript is expressed temporally and spatially in vivo with its native target, the tie-1 coding transcript and in additional locations (ear & brain). This argues for a tie-1 gene role previously unrecognized in the brain and ear function, perhaps in the vasculature of these organs. In fact two recent studies implicate Angiopoietins, ligands for Tie’s, in lymphatic vessel development in ear [29,30]. Since the non-coding RNA for tie-1 is conserved in humans and mice, this paradigm of using non-coding RNA expression pattern to speculate putative novel gene function (simply based on expression alone) of well-studied genes is intriguing.
Mutations in Tie’s have been previously implicated in patients with vascular malformations [31-35]. Our studies showed that the ratio of tie-1 vs. tie-1AS lncRNA is altered in human vascular anomaly samples. The expression level of tie-1AS is much lower than that of tie-1 in human placenta, but it goes up dramatically compared to the expression level of tie-1 in hemangioma samples [36]. We observed similar expression discrepancy between tie-1 and tie-1AS in zebrafish cardiovascular mutant cloche [37] (Figure 1). Interestingly, this data correlates well with previous reported [18] in situ hybridization profile for tie-1 in cloche embryos where the expression is quite low when compared to WT embryos. The vascular anomaly (disease) and the cloche (genetic mutant) results together suggest that when the cognate gene level (tie-1) is low, the cognate tie-1 AS levels are high suggesting an inverse relationship between the tie-1 and the AS transcript. In fact, this interpretation makes sense since we have shown the tie-1AS lncRNA selectively binds tie-1 mRNA in vivo and regulates tie-1 transcript levels [36]. Moreover, the normal expression levels of tie-1AS is a log lower than tie-1 across different embryonic stages in zebrafish.
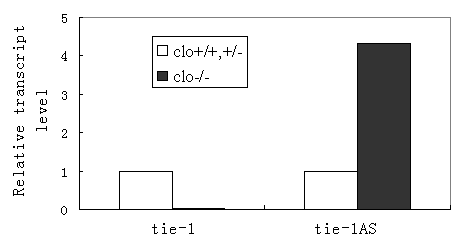
Figure 1. Real time PCR for tie-1 and tie-1AS across Clo+/+ and Clo+/- or Clo-/- embryos. Zebrafish cloche mutant embryos show aberrant level of tie-1AS compared with tie-1.
In most of the examples to date levels of antisense RNA have been correlated to disease. However, studies reporting genetic mutations in the antisense RNA are not common. A genetic mutation that gives rise to pathologic antisense transcript is a 14;18 chromosome translocation observed in most human follicular B-cell lymphomas. This translocation not only creates a bcl-2/IgH hybrid gene, but also gives rise to a bcl-2/IgH AS transcript. The AS transcript is merely present in the t(14;18)-positive, but not in the t(14;18)-negative lymphoid cell lines, and thus appears to be dependent on the bcl-2/IgH fusion [38]. Another recent example is observed in an individual with an inherited form of anemia (α-thalassemia). In this patient, the LUC7L transcription termination sequence is deleted and LUC7L RNA extends into HBA2 locus and causes CpG island methylation of HBA2, a α-globin gene in erythroid cells. The methylation leads to silencing of HBA2 gene resulting in an inherited form of anemia α-thalassemia [39] in this subject.
NATs - Promising Future Applications
NATs hold immense potential for future applications. We will highlight three distinct areas of biology where NATs are expected to make promising contributions. More than 30 years ago, the inhibitory effects of antisense oligonucleotides on translation were observed in vitro. Since then synthetic antisense oligonucleotides have been used extensively to block target gene expression [40]. However, not all sites in mRNA sequences respond to targeting. Only one out of ten randomly selected antisense oligonucleotides exhibit good activity [41]. A study by Van der Krol et al using the petunia as a model to test the effects of antisense gene fragments corresponding to different chromosomal location on flower pigmentation highlights this point. The patterns of pigmentation varied among flowers of different transgenic petunia with sub-genomic fragments of antisense genes being ineffective in establishing a phenotype [42]. NATs are endogenous regulators targeting mRNA, and some of them are processed into endogenous siRNAs [43]. Thereby, NATs are ideal models for designing antisense oligonucleotides.
The second example highlights the role of NATs in a clinical trial setting. In clinical trials, an anticancer agent Gemcitabine can produce cytotoxic synergism with Cisplatin when given to cancer patients. It was proposed that the nucleotide excision repair (NER) process was responsible for the synergism. NER removes cisplatin-DNA adducts by a series of steps including damage recognition, dual incision/excision, repair synthesis and ligation. ERCC1 protein is an important factor in the incision process—the rate-limiting step of the pathway. Antisense therapy targeting the ERCC1 protein was proposed as a target for this approach. As a result, NER activity was downregulated and the cytotoxic synergism between Gemcitabine and cisplatin was abolished [44].
The third example highlights the role of NATs in tumor therapy. The bcl-2/IgH antisense transcript is presumed to be responsible for the overexpressed bcl-2, the root of oncogenicity in follicular lymphomas. Therapeutic ODNs targeting the bcl-2/IgH antisense transcript induced an early strong inhibition of cell growth and a late sudden cell death. The induction is restricted to follicular lymphomas which have t(14;18) chromosomal translocation, but not effective for the cells carrying a normal bcl-2 gene, suggesting tumor selectivity. However, antisense-oriented ODN, complementary to the bcl-2/IgH mRNA is ineffective [45].
Summary
NATs account for majority of non-coding RNAs in the vertebrate genome. Numerous mechanisms are utilized by NATs to regulate gene expression and these mechanisms span over both plant and animal kingdoms. Recently, NATs have been identified in various diseases as well and the regulation that they mediate when lost may in part be responsible for disease progression and pathogenesis. Although most studies identify dysregulation of NATs, very few studies have identified genetic mutations in them. As of now, NATs hold immense clinical value but their application is still at its infancy.
ACKNOWLEDGEMENTS
This work was supported by American Heart Association postdoctoral fellowship to KL, NIH HL090712 to RR, Advancing Healthier Wisconsin and Children’s Research Institute seed funds to RR. We apologize to authors whose work we have not been able to cite due to space limitations.
CONFLICT OF INTEREST STATEMENT
The authors do not have any conflict of interest to declare.
REFERENCES
1. Lavorgna G, Dahary D, Lehner B, et al. In search of antisense. Trends Biochem Sci. 2004; 29: 88-94.
2. Bovre K & Szybalski W. Patterns of convergent and overlapping transcription within the b2 region of coliphage lambda. Virology. 1969; 38: 614-26.
3. Wek RC & Hatfield GW. Nucleotide sequence and in vivo expression of the ilvY and ilvC genes in Escherichia coli K12. Transcription from divergent overlapping promoters. J Biol Chem. 1986; 261: 2441-50.
4. Wong F, Yuh ZT, Schaefer EL, et al. Overlapping transcription units in the transient receptor potential locus of Drosophila melanogaster. Somat Cell Mol Genet. 1987; 13: 661-9.
5. Rosok O & Sioud M. Systematic identification of sense-antisense transcripts in mammalian cells. Nat Biotechnol. 2004; 22: 104-8.
6. Katayama S, Tomaru Y, Kasukawa T, et al. Antisense transcription in the mammalian transcriptome. Science. 2005; 309: 1564-6.
7. ENCODE Project Consortium, Birney E, Stamatoyannopoulos JA, et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature. 2007; 447: 799-816.
8. Faghihi MA & Wahlestedt C. Regulatory roles of natural antisense transcripts. Nat Rev Mol Cell Biol. 2009; 10: 637-43.
9. Ge X, Wu Q, Jung YC, et al. A large quantity of novel human antisense transcripts detected by LongSAGE. Bioinformatics. 2006; 22: 2475-9.
10. Okamura K, Balla S, Martin R, et al. Two distinct mechanisms generate endogenous siRNAs from bidirectional transcription in Drosophila melanogaster. Nat Struct Mol Biol. 2008; 15: 581-90.
11. Faghihi MA, Zhang M, Huang J, et al. Evidence for natural antisense transcript-mediated inhibition of microRNA function. Genome Biol. 2010; 11: R56.
12. Masukata H & Tomizawa J. Control of primer formation for ColE1 plasmid replication: conformational change of the primer transcript. Cell. 1986; 44: 125-36.
13. International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature. 2004; 431: 931-45.
14. Chen J, Sun M, Kent WJ, et al. Over 20% of human transcripts might form sense-antisense pairs. Nucleic Acids Res. 2004; 32: 4812-20.
15. Lapidot M & Pilpel Y. Genome-wide natural antisense transcription: coupling its regulation to its different regulatory mechanisms. EMBO Rep. 2006; 7: 1216-22.
16. Aalfs JD & Kingston RE. What does ‘chromatin remodeling’ mean? Trends Biochem Sci. 2000; 25: 548-55.
17. Shearwin KE, Callen BP & Egan JB. Transcriptional interference--a crash course. Trends Genet. 2005; 21: 339-45.
18. Liao W, Bisgrove BW, Sawyer H, et al. The zebrafish gene cloche acts upstream of a flk-1 homologue to regulate endothelial cell differentiation. Development. 1997; 124: 381-9.
19. Prasanth KV & Spector DL. Eukaryotic regulatory RNAs: an answer to the ‘genome complexity’ conundrum. Genes Dev. 2007; 21: 11-42.
20. Beltran M, Puig I, Peña C, et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008; 22: 756-69.
21. Carlile M, Nalbant P, Preston-Fayers K, et al. Processing of naturally occurring sense/antisense transcripts of the vertebrate Slc34a gene into short RNAs. Physiol Genomics. 2008; 34: 95-100.
22. Ebralidze AK, Guibal FC, Steidl U, et al. PU.1 expression is modulated by the balance of functional sense and antisense RNAs regulated by a shared cis-regulatory element. Genes Dev. 2008; 22: 2085-92.
23. Faghihi MA, Modarresi F, Khalil AM, et al. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat Med. 2008; 14: 723-30.
24. Seitz A, Gourevitch D, Zhang XM, et al. Sense and antisense transcripts of the apolipoprotein E gene in normal and ApoE knockout mice, their expression after spinal cord injury and corresponding human transcripts. Hum Mol Genet. 2005; 14: 2661-70.
25. Reis EM, Nakaya HI, Louro R, et al. Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer. Oncogene. 2004; 23: 6684-92.
26. Grigoriadis A, Oliver GR, Tanney A, et al. Identification of differentially expressed sense and antisense transcript pairs in breast epithelial tissues. BMC Genomics. 2009; 10: 324.
27. Jacobs JF, van Bokhoven H, van Leeuwen FN, et al. Regulation of MYCN expression in human neuroblastoma cells. BMC Cancer. 2009; 9: 239.
28. Holdt LM, Beutner F, Scholz M, et al. ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler Thromb Vasc Biol. 2010; 30: 620-7.
29. Dellinger M, Hunter R, Bernas M, et al. Defective remodeling and maturation of the lymphatic vasculature in Angiopoietin-2 deficient mice. Dev Biol. 2008; 319: 309-20.
30. Kim KE, Cho CH, Kim HZ, et al. In vivo actions of angiopoietins on quiescent and remodeling blood and lymphatic vessels in mouse airways and skin. Arterioscler Thromb Vasc Biol. 2007; 27: 564-70.
31. Limaye N, Wouters V, Uebelhoer M, et al. Somatic mutations in angiopoietin receptor gene TEK cause solitary and multiple sporadic venous malformations. Nat Genet. 2009; 41: 118-24.
32. Vikkula M, Boon LM, Carraway KL 3rd, et al. Vascular dysmorphogenesis caused by an activating mutation in the receptor tyrosine kinase TIE2. Cell. 1996; 87: 1181-90.
33. Wang H, Zhang Y, Toratani S, et al. Transformation of vascular endothelial cells by a point mutation in the Tie2 gene from human intramuscular haemangioma. Oncogene. 2004; 23: 8700-4.
34. Bhandarkar SS, Jaconi M, Fried LE, et al. Fulvene-5 potently inhibits NADPH oxidase 4 and blocks the growth of endothelial tumors in mice. J Clin Invest. 2009; 119: 2359-65.
35. Perry B, Banyard J, McLaughlin ER, et al. AKT1 overexpression in endothelial cells leads to the development of cutaneous vascular malformations in vivo. Arch Dermatol. 2007; 143: 504-6.
36. Li K, Blum Y, Verma A, et al. A noncoding antisense RNA in tie-1 locus regulates tie-1 function in vivo. Blood. 2010; 115: 133-9.
37. Li, K. Unpublished data.
38. Capaccioli S, Quattrone A, Schiavone N, et al. A bcl-2/IgH antisense transcript deregulates bcl-2 gene expression in human follicular lymphoma t(14;18) cell lines. Oncogene. 1996; 13: 105-15.
39. Tufarelli C, Stanley JA, Garrick D, et al. Transcription of antisense RNA leading to gene silencing and methylation as a novel cause of human genetic disease. Nat Genet. 2003; 34: 157-65.
40. Zamecnik PC. History of antisense oligonucleotides. In: Agrawal S, ed. Antisense therapy. Humana Press, Totowa, NJ. 1996; pp. 1-11.
41. Bennett CF & Cowsert LM. Antisense oligonucleotides as a tool for gene functionalization and target validation. Biochim Biophys Acta. 1999; 1489: 19-30.
42. van der Krol AR, Mur LA, de Lange P, et al. Inhibition of flower pigmentation by antisense CHS genes: promoter and minimal sequence requirements for the antisense effect. Plant Mol Biol. 1990; 14: 457-66.
43. Werner A & Swan D. What are natural antisense transcripts good for? Biochem Soc Trans. 2010; 38: 1144-9.
44. Yang LY, Li L, Jiang H, et al. Expression of ERCC1 antisense RNA abrogates gemicitabine-mediated cytotoxic synergism with cisplatin in human colon tumor cells defective in mismatch repair but proficient in nucleotide excision repair. Clin Cancer Res. 2000; 6: 773-81.
45. Morelli S, Delia D, Capaccioli S, et al. The antisense bcl-2-IgH transcript is an optimal target for synthetic oligonucleotides. Proc Natl Acad Sci U S A. 1997; 94: 8150-5.

