INTRODUCTION
Glioblastomas (GBMs), the most common glioma, account for approximately 70% of astrocytomas and 15% of all intracranial neoplasms [1, 2]. Approximately 90% of GBMs are classified as primary, and such GBMs mainly affect elderly people, with a median survival of approximately 15 months. The remaining 10% of GBMs are classified as secondary. Secondary GBMs develop from World Health Organization (WHO) grade II or III gliomas and predominantly affect younger individuals, with a median survival of 31 months [3].
The type of astrocytic neoplasm is defined through histological analysis. Recently, exomic sequencing has revealed frequent mutations in the isocitrate dehydrogenase 1 (IDH1) gene and its homolog IDH2, in both low- and high-grade gliomas [4, 5]. Almost all IDH1 mutations result in an amino acid substitution at R132. These mutations impair the physiological function of IDH1 in converting isocitrate to α-ketoglutarate (KG) and confer a gain of function in converting α-KG to D-2-hydroxyglutarate (D-2HG), which accumulates in extremely high levels in tumors with IDH1 mutations [5, 6]. Such mutations are predominantly observed in secondary GBMs; therefore, these mutations are typically considered as highly selective molecular markers of secondary disease. In the latest WHO tumor classification of the central nervous system (CNS), GBMs are classified as (1) IDH—wild-type (WT) and (2) IDH-mutant GBMs. However, diagnostic challenges may arise because of the heterogeneity of tumors and partial sampling of lesions. Moreover, not every institute has developed reliable methods to completely evaluate the IDH genotype for every case, particularly for cases that cannot be treated through invasive surgery.
Magnetic resonance imaging (MRI) is the modality of choice for imaging brain tumors. It can provide clear tissue contrasts to help in estimating the malignancy of brain tumors [7, 8]. In addition to tumor grading, several physiological MRI techniques, including diffusion-weighted imaging (DWI), MR spectroscopy (MRS), and perfusion-weighted imaging (PWI), have been developed to more accurately characterize the physiological and molecular features of GBMs [9, 10]. Andronesi and Choi [11, 12] have reported that MRI combined with specialized MRS techniques can detect the in vivo overexpression of D-2HG. Other techniques, including DWI and PWI, were also applied to detect differences between WT and mutant IDH [13]. Tumors with IDH mutations were proposed to have a more heterogeneous imaging microenvironment because of their stepwise gliomagenesis [13]. Nevertheless, the relationship between the IDH status and tissue signals generated by clinically relevant conventional MR sequences has not been well documented.
For quantifying tumor characteristics through imaging, various computer-aided diagnosis (CAD) systems have been developed for classifying tumor types and grades [14–16]. An artificial intelligence classifier can facilitate combining numerous radiomic features to generate a specific model. In particular, differences between tumors with and those without IDH mutations may be subtle and should be explored through the sophisticated integration of various image features. In this study, quantitative morphological, intensity, and textural features were extracted from tumor tissues to determine their IDH status; the performance of both individual feature sets and a combination of the 3 feature sets was evaluated. This quantitative CAD procedure can provide consistent and reliable suggestions for IDH classifications to radiologists.
RESULTS
Considering the IDH genotype as the diagnostic target, the prediction performance of each feature set was determined (Table 1). The global morphological and intensity features exhibited similar performance: accuracy, 51% (20/39); sensitivity, 57% (4/7); and specificity, 50% (16/32) and accuracy, 59% (23/39); sensitivity, 57% (4/7); and specificity, 59% (19/32), respectively. The textural features describing local patterns in tumors yielded an accuracy of 85% (33/39), a sensitivity of 86% (6/7), and a specificity of 84% (27/32), which are significantly higher than the corresponding values for the morphological and intensity features. The agreement level (κ) between the prediction results and biopsy-proven pathology was 0.60. A further experiment for analyzing the combination of the 3 feature sets revealed that only the textural features were selected, with an accuracy of 85%. We selected the features of cluster prominence, cluster shading, maximum probability, difference variance, the information measure of correlation, the inverse difference normalized, and the inverse difference moment.
Table 1: Performance of different image feature sets for predicting IDH mutations
Morphology |
Intensity |
Texture |
Texture |
Texture |
|
|---|---|---|---|---|---|
Accuracy |
51% (20/39) |
59% (23/39) |
85% (33/39) |
0.0016* |
0.0119* |
Sensitivity |
57% (4/7) |
57% (4/7) |
86% (6/7) |
0.2367 |
0.2367 |
Specificity |
50% (16/32) |
59% (19/32) |
84% (27/32) |
0.0034* |
0.0261* |
* p < 0.05 indicates a statistically significant difference.
DISCUSSION
Immunohistochemistry is a standard method for detecting IDH mutations under most clinical scenarios. However, the genetic status within a GBM, including the IDH status, shows intratumoral heterogeneity. Performing biopsies of the different parts of tumors may yield different results regarding the IDH status. Therefore, another noninvasive method to characterize this information is warranted. Radiomic features are typically extracted from pre-existing MRI images stored in a computer; in practice, this is not an expensive and complicated method. This study revealed that computer-aided radiomic features concerning the microenvironmental texture can distinguish the IDH1 status of GBMs. Studies [5, 17–19] have reported that IDH mutations predominantly occur in secondary rather than primary GBMs. These mutations alter the normal enzyme activity, thus reducing the synthesis of α-KG and NADPH, which makes cells more vulnerable to oxidative stress. Furthermore, these mutations impair the function of IDH in converting isocitrate to α-KG and confer a gain of function in converting α-KG to D-2HG, resulting in the overexpression of D-2HG in IDH-mutant tumors. The excessive D-2HG is considered an oncogenic metabolite because it induces epigenetic changes that lead to the aberrant regulation of gene expression and perturbed cellular differentiation, which may contribute to tumorigenesis [20, 21]. Furthermore, D-2HG induces increased levels of hypoxia-inducible factor (HIF) -1α. It is a transcription factor that promotes tumor angiogenesis [22], a key element in GBM formation. Because of the stepwise gliomagenesis pattern of secondary GBMs, tumors with these mutations are considered to have a more heterogeneous microenvironment and imaging presentations [13]. Our results revealed that tumors with IDH mutations tended to have lower values of inverse difference moment features, an indicator of tissue homogeneity [23], which also suggests more heterogeneous intensities in imaging.
Using a radiomic model for predicting IDH mutations provides a connection between intuitive vision and personalized profiling. By applying radiomic image features, tumor characteristics can be quantified without requiring a risky biopsy. In addition, imaging is a routine procedure, with no additional costs. We employed numerous radiomic features in this study for predicting IDH mutations. These features were divided into the categories of morphological, intensity, and textural features, which provided different viewpoints for distinguishing GBMs with and without IDH mutations.
Table 1 shows that the textural features yielded an accuracy of 85% (33/39), a sensitivity of 86% (6/7), and a specificity of 84% (27/32), which are significantly higher (p < 0.05) than the corresponding values for the morphological and intensity features. These results are in agreement with those of a previous study that used textural features to characterize the molecular subtypes of GBM [24]. The difference is that the proposed features and classification method targeted IDH mutations, which are recognized as a critical marker in GBM classification. We expect that more correlations between radiomic features and gene expression will be investigated.
For predicting IDH mutations, we selected the following features: cluster prominence, cluster shading, maximum probability, the information measure of correlation, difference variance, the inverse difference normalized, and the inverse difference moment. The cluster prominence and cluster shading measures indicate whether a lack of symmetry exists in gray-scale distributions. The maximum probability corresponds to the strongest response. The correlation determines the gray-level linear dependence between a pixel and its surrounding neighbors. The difference variance indicates the variance between the co-occurrence probabilities along the x and y axes. The inverse difference moment estimates the homogeneity of the tissue pattern [23]. Regarding the misclassified case in Figure 1, a potential regional feature distinguishing the tissues into numerous regions according to their brightness may be helpful for recognizing the halo observed on the boundary. This type of feature correlated with the regional brightness, and the location can be determined in the future to provide more information than that provided by the existing features.
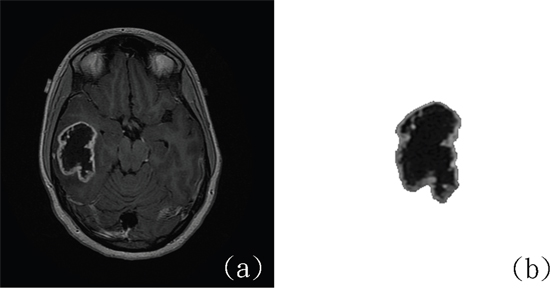
Figure 1: The only IDH-mutant case of GBM misclassified using the CAD system. (a) Original MR image. (b) Delineated tumor area. (http://cancerimagingarchive.net/; “License” and the CC BY license, https://creativecommons.org/licenses/by/3.0/; tumor areas in this figure were extracted from original images.)
A limitation of this preliminary study is that the number of IDH-mutant cases was not comparable with that of IDH-WT cases. Nevertheless, patients in our cohort were enrolled from 4 hospitals, and a balance between the sensitivity (IDH) and specificity (non-IDH) was maintained (86% and 84%, respectively). Future studies should evaluate more cases with respect to the proposed features and CAD system. Another limitation is that we used only contrast-enhanced T1-weighted images (WIs), which might not clearly demonstrate peritumoral edema. However, IDH mutations are associated with angiogenesis activity [22], and the activity of the angiogenesis module within a tumor has been proven to be associated with the degree of contrast enhancement [25, 26]. Therefore, we believe that the measurements of signal intensities on contrast-enhanced T1WIs can be key determinants for differentiating GBMs with and without IDH mutations. However, further investigations on the role of other MRI sequences, such as fluid-attenuated inversion recovery, DWI, PWI, and MRS, are warranted.
MATERIALS AND METHODS
Patient information
The Cancer Genome Atlas and the Cancer Imaging Archive
The MRI data of 32 patients with WT IDH and 7 patients with mutant IDH in GBM were obtained from the Cancer Imaging Archive (TCIA) (http://cancerimagingarchive.net/) of the National Cancer Institute, a portal that contains images of patients in the Cancer Genome Atlas (TCGA) for performing imaging analysis. The materials and data provided by TCGA were used in compliance with all applicable laws, regulations, and policies for patient protection. All necessary approvals, authorizations, participant assurances, informed consent documents, and institutional review board approvals were obtained [27] (Figure 2).
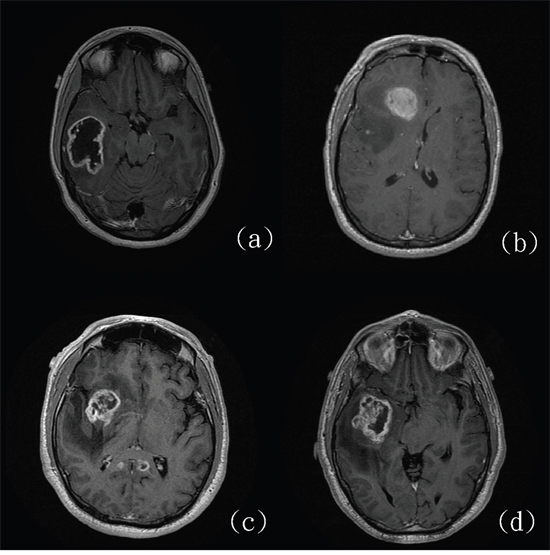
Figure 2: Four glioblastomas with (a and b) and without (c and d) IDH mutations. (http://cancerimagingarchive.net/; “License” and the CC BY license, https://creativecommons.org/licenses/by/3.0/.)
The MR images used in this study were obtained from different institutes. The IDH-WT cases were provided by Henry Ford and Case Western hospitals. The IDH-mutant cases were obtained from Henry Ford Hospital, + Case Western Hospital, Emory University, and Fondazione IRCCS Instituto Neuroligico C. Besta. Among the 291 GBM cases in TCIA, only 15 (5.15%) were found to have the IDH1 mutation. Because 8 of them had no preoperative contrast-enhanced T1WIs, only 7 cases were included in this study. Among them, one case had an IDH1 R132G mutation, whereas the remaining cases had an IDH1 R132H mutation. No case in TCIA had the IDH2 mutation. The IDH-mutant cases in our cohort were enrolled from 4 hospitals in TCIA. To ensure minimum variation in protocols and equipment, the IDH-WT cases were obtained from 2 of the 4 hospitals through consecutive selection from TCGA archive. A total of 40 IDH-WT cases were selected; however, 8 cases were excluded because of incomplete or suboptimal image quality. The demographic information of our cohort is listed in Table 2.
Table 2: Demographic information of the cohort
Age |
Sex |
Tumor |
Tumor |
|
|---|---|---|---|---|
IDH-WT |
62.6 ± 12.5 years |
Female: 9 |
Right: 20 |
Frontal: 11 |
IDH-mutant |
36.5 ± 15.9 years |
Female: 2 |
Right: 3 |
Frontal: 3 |
Tumor delineation
Contrast-enhanced axial T1WIs were used for the image analysis. A board-certified neuroradiologist (K.H., having 13 years of experience), blinded to the molecular information, selected the most representative 2-dimensional image of each tumor. Intensity normalization, which extends the gray-level distribution of each MR image to the whole value range (0–255), was performed to enhance the contrast between the tumor and background tissues for contour delineation. The contour was manually delineated using OsiriX. Image pixels enclosed by the defined tumor contour were used for further feature extraction and analysis (Figure 3).
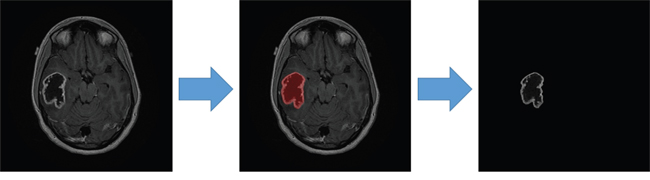
Figure 3: Tumor contour delineation in contrast-enhanced axial T1WIs. (http://cancerimagingarchive.net/; “License” and the CC BY license, https://creativecommons.org/licenses/by/3.0/; tumor areas in this figure were extracted from original images.)
Radiomic features
The latest WHO tumor classification of the CNS describes IDH mutations as a determinant for differentiating GBMs. Whether radiomic features can provide information on IDH mutations is relevant to clinical practice. The features extracted from images can be further combined to develop a prediction model for providing suggestions to and improving the diagnostic accuracy of radiologists. Numerous radiomic features were employed in this study for predicting IDH mutations. Three categories, namely morphological, intensity, and textural features, were examined to interpret the characteristics of GBMs with IDH mutations. Figure 4 illustrates the extraction of radiomic features.
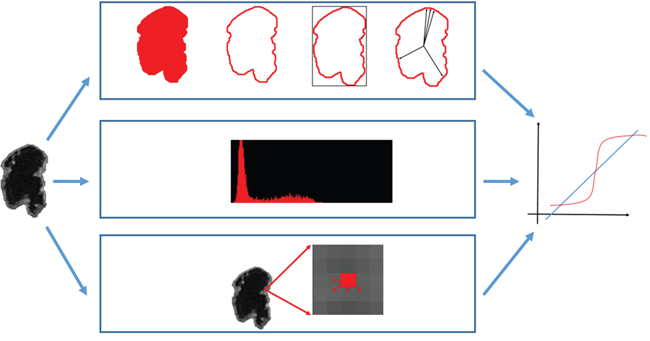
Figure 4: Quantitative features extracted from the tumor area were combined in a logistic regression classifier. (http://cancerimagingarchive.net/; “License” and the CC BY license, https://creativecommons.org/licenses/by/3.0/; tumor areas in this figure were extracted from original images.)
Morphological features
A tumor’s shape may reflect its growth pattern. Therefore, morphological features are widely used in CAD systems to describe the geometric characteristics of tumors, such as the shape and margins. The essential properties, including the area and perimeter, are easily calculated from the delineated tumor contour. Considering the tumor shape (round or irregular), compactness was proposed to correlate to the tumor area and perimeter [28]. The irregularity of the tumor margin can be estimated using the mean and standard deviation of the normalized radial length (NRL) to determine the complexity of the distribution of the boundary pixels [29]. The NRL is defined as the Euclidean distance between the tumor center and boundary pixels normalized by the maximum distance. The effects of tumors on their adjacent normal tissues may vary. Tumor deformation should be evaluated for tissue characterization.
Intensity features
According to the signal intensity characteristics of various tissues in MR images, the brightness distribution reveals the tumor composition. The intensity in contrast-enhanced TIWIs also reveals angiogenesis and blood–tumor barrier abnormalities in brain tumors [9]. By distinguishing the brightness distribution into bins of gray-scale values, a histogram formulates the count of individual values as a probability function. Histogram moments [30, 31] are quantitative metrics for describing probability statistics. Moment features include the first-, second-, third-, and fourth-order central moments of a histogram, namely the mean, variance, skewness, and kurtosis, respectively.
The mean and variance measure the center of the gray-scale value distribution and the extent of increase in these values. Skewness determines the symmetric property of the distribution by describing the balance between 2 sides of a distribution. Kurtosis refers to a single-peak histogram with heavily weighted tails compared with those in a normal distribution.
Textural features
Textural features are widely used in CAD systems to discriminate tumor types [15]. A computational statistical analysis between pixels can reveal the local pattern formed by the correlations between adjacent pixels. The brightness composition, such as whether tissues inside the tumor area are heterogeneous or homogeneous, can be determined from textural features.
The gray-level co-occurrence matrix (GLCM) has been proposed as a promising method for interpreting image textures [32–34]. Various CAD systems use GLCM textures to describe image patterns for tumor classification [35]. The GLCM was established by counting the co-occurrence frequencies of 2 adjacent pixels (i and j) at a distance d and direction θ [32]. A reduced image G with fewer intensity bins facilitated reducing the computational complexity. Parameters including distance d = 1 and 4 directions of θ= 0°, 45°, 90°, and 135° were used individually and in combination. In total, 14 GLCM textural features were used in the experiment:
Autocorrelation = |
(1) |
|
Contrast = |
(2) |
|
Correlation = |
(3) |
|
Cluster prominence = |
(4) |
|
Cluster shading = |
(5) |
|
Dissimilarity = |
(6) |
|
Energy = |
(7) |
|
Entropy = |
(8) |
|
Homogeneity = |
(9) |
|
Difference variance = |
(10) |
|
Difference entropy = |
(11) |
|
Information measure of correlation = |
(12) |
|
Inverse difference normalized = |
; |
(13) |
Inverse difference moment = |
; |
(14) |
where μx,μy,σx, and σy are the mean and standard deviations of the marginal distributions of p(i, j|d, θ):
(15)
(16)
Statistical analysis
The morphological, intensity, and textural image features were implemented in the experiment for tissue characterization. These features completely revealed the global and local appearances of GBMs. The global appearance includes the morphological properties of the tumor shape and overall intensity distribution of tumor tissues, whereas the local texture describes correlations between pixels and their neighbors in different orientations. All features in a category were combined into a feature set in a binary logistic regression classifier for predicting the IDH status. A combination of these features was also examined.
With next-generation sequencing-based molecular profiles as the gold standard, imaging features were evaluated using stepwise backward elimination to explore the most favorable combination of features. The feature set with the lowest error rate was considered the most relevant set. The corresponding prediction model was validated using the leave-one-out method [36] to determine its generalizability. In each validation iteration, an individual case was selected and was used to examine the trained prediction model from the remaining N − 1 cases. Therefore, each tumor was assigned a probability of having an IDH mutation according to its features.
Based on the proven IDH genotype, the performance of the prediction model was evaluated using 3 indices: accuracy, sensitivity, and specificity. The chi-squared test was used to compare the performance indices by using SPSS (Version 16 for Windows; SPSS, Chicago, IL, USA). In addition, the prediction result was compared with the biopsy-proven pathology to obtain the agreement level by using Cohen’s kappa. The resulting κ was −1.0 to 1.0, where a high value indicates high reliability. The agreement was considered slight if κ < 0.20; fair if κ = 0.21–0.40; moderate if κ = 0.41–0.60; substantial if κ = 0.61–0.80; and almost perfect if κ > 0.81.
ACKNOWLEDGMENTS
The authors thank the Ministry of Science and Technology, Taiwan (MOST 104-2218-E-038-004 and MOST 105-2314-B-038-049) and Taipei Medical University (TMU 104-AE1-B04) for financially supporting this study.
CONFLICTS OF INTEREST
The authors declare that they have no financial or personal relationships with other people or organizations that could have inappropriately influenced this study.
REFERENCES
1. Wen PY, Kesari S. Malignant gliomas in adults. N Engl J Med. 2008; 359:492-507.
2. Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007; 114:97-109.
3. Louis DN, Perry A, Reifenberger G, von Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD, Kleihues P, Ellison DW. The 2016 World Health Organization classification of tumors of the central nervous system: a summary. Acta Neuropathol. 2016; 131:803-820.
4. Parsons DW, Jones S, Zhang X, Lin JC, Leary RJ, Angenendt P, Mankoo P, Carter H, Siu IM, Gallia GL, Olivi A, McLendon R, Rasheed BA, et al. An integrated genomic analysis of human glioblastoma multiforme. Science. 2008; 321:1807-1812.
5. Yan H, Parsons DW, Jin G, McLendon R, Rasheed BA, Yuan W, Kos I, Batinic-Haberle I, Jones S, Riggins GJ, Friedman H, Friedman A, Reardon D, et al. IDH1 and IDH2 mutations in gliomas. N Engl J Med. 2009; 360:765-773.
6. Dang L, White DW, Gross S, Bennett BD, Bittinger MA, Driggers EM, Fantin VR, Jang HG, Jin S, Keenan MC, Marks KM, Prins RM, Ward PS, et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature. 2009; 462:739-744.
7. Mahaley MS Jr, Mettlin C, Natarajan N, Laws ER Jr, Peace BB. National survey of patterns of care for brain-tumor patients. J Neurosurg. 1989; 71:826-836.
8. Guzmán-De-Villoria JA, Mateos-Pérez JM, Fernández-García P, Castro E, Desco M. Added value of advanced over conventional magnetic resonance imaging in grading gliomas and other primary brain tumors. Cancer Imaging. 2014; 14:1-10.
9. Jackson A, OʼConnor JP, Parker GJ, Jayson GC. Imaging tumor vascular heterogeneity and angiogenesis using dynamic contrast-enhanced magnetic resonance imaging. Clin Cancer Res. 2007; 13:3449-3459.
10. Jain R, Poisson L, Narang J, Gutman D, Scarpace L, Hwang SN, Holder C, Wintermark M, Colen RR, Kirby J, Freymann J, Brat DJ, Jaffe C, Mikkelsen T. Genomic mapping and survival prediction in glioblastoma: molecular subclassification strengthened by hemodynamic imaging biomarkers. Radiology. 2013; 267:212-220.
11. Andronesi OC, Kim GS, Gerstner E, Batchelor T, Tzika AA, Fantin VR, Vander Heiden MG, Sorensen AG. Detection of 2-hydroxyglutarate in IDH-mutated glioma patients by
12. Choi C, Ganji SK, DeBerardinis RJ, Hatanpaa KJ, Rakheja D, Kovacs Z, Yang XL, Mashimo T, Raisanen JM, Marin-Valencia I, Pascual JM, Madden CJ, Mickey BE, et al. 2-hydroxyglutarate detection by magnetic resonance spectroscopy in IDH-mutated patients with gliomas. Nat Med. 2012; 18:624-629.
13. Lee S, Choi SH, Ryoo I, Yoon TJ, Kim TM, Lee SH, Park CK, Kim JH, Sohn CH, Park SH, Kim IH. Evaluation of the microenvironmental heterogeneity in high-grade gliomas with IDH1/2 gene mutation using histogram analysis of diffusion-weighted imaging and dynamic-susceptibility contrast perfusion imaging. J Neurooncol. 2015; 121:141-150.
14. Hsieh KL, Lo CM, Hsiao CJ. Computer-aided grading of gliomas based on local and global MRI features. Comput Methods Programs Biomed. 2017; 139:31-38.
15. Lo CM, Moon WK, Huang CS, Chen JH, Yang MC, Chang RF. Intensity-invariant texture analysis for classification of bi-rads category 3 breast masses. Ultrasound Med Biol. 2015; 41:2039-2048.
16. Lo CM, Lai YC, Chou YH, Chang RF. Quantitative breast lesion classification based on multichannel distributions in shear-wave imaging. Comput Methods Programs Biomed. 2015; 122:354-361.
17. Balss J, Meyer J, Mueller W, Korshunov A, Hartmann C, von Deimling A. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol. 2008; 116:597-602.
18. Hartmann C, Meyer J, Balss J, Capper D, Mueller W, Christians A, Felsberg J, Wolter M, Mawrin C, Wick W, Weller M, Herold-Mende C, Unterberg A, et al. Type and frequency of IDH1 and IDH2 mutations are related to astrocytic and oligodendroglial differentiation and age: a study of 1,010 diffuse gliomas. Acta Neuropathol. 2009; 118:469-474.
19. Ichimura K, Pearson DM, Kocialkowski S, Backlund LM, Chan R, Jones DT, Collins VP. IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro Oncol. 2009; 11:341-347.
20. Lu C, Ward PS, Kapoor GS, Rohle D, Turcan S, Abdel-Wahab O, Edwards CR, Khanin R, Figueroa ME, Melnick A, Wellen KE, O'Rourke DM, Berger SL, et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature. 2012; 483:474-478.
21. Turcan S, Rohle D, Goenka A, Walsh LA, Fang F, Yilmaz E, Campos C, Fabius AW, Lu C, Ward PS, Thompson CB, Kaufman A, Guryanova O, et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature. 2012; 483:479-483.
22. Pelloski CE, Ballman KV, Furth AF, Zhang L, Lin E, Sulman EP, Bhat K, McDonald JM, Yung WK, Colman H, Woo SY, Heimberger AB, Suki D, et al. Epidermal growth factor receptor variant III status defines clinically distinct subtypes of glioblastoma. J Clin Oncol. 2007; 25:2288-2294.
23. Sharma N, Ray AK, Sharma S, Shukla KK, Pradhan S, Aggarwal LM. Segmentation and classification of medical images using texture-primitive features: application of BAM-type artificial neural network. J Med Phys. 2008; 33:119-126.
24. Yang D, Rao G, Martinez J, Veeraraghavan A, Rao A. Evaluation of tumor-derived MRI-texture features for discrimination of molecular subtypes and prediction of 12-month survival status in glioblastoma. Med Phys. 2015; 42:6725-6735.
25. Diehn M, Nardini C, Wang DS, McGovern S, Jayaraman M, Liang Y, Aldape K, Cha S, Kuo MD. Identification of noninvasive imaging surrogates for brain tumor gene-expression modules. Proc Natl Acad Sci U S A. 2008; 105:5213-5218.
26. Pope WB, Chen JH, Dong J, Carlson MR, Perlina A, Cloughesy TF, Liau LM, Mischel PS, Nghiemphu P, Lai A. Relationship between gene expression and enhancement in glioblastoma multiforme: exploratory DNA microarray analysis. Radiology. 2008; 249:268-277.
27. McLendon R, Friedman A, Bigner D, Van Meir EG, Brat DJ, Mastrogianakis GM, Olson JJ, Mikkelsen T, Lehman N, Aldape K. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature. 2008; 455:1061-1068.
28. Rangayyan RM, Mudigonda NR, Desautels JL. Boundary modelling and shape analysis methods for classification of mammographic masses. Med Biol Eng Comput. 2000; 38:487-496.
29. Nie K, Chen JH, Hon JY, Chu Y, Nalcioglu O, Su MY. Quantitative analysis of lesion morphology and texture features for diagnostic prediction in breast MRI. Acad Radiol. 2008; 15:1513-1525.
30. Baek HJ, Kim HS, Kim N, Choi YJ, Kim YJ. Percent change of perfusion skewness and kurtosis: a potential imaging biomarker for early treatment response in patients with newly diagnosed glioblastomas. Radiology. 2012; 264:834-843.
31. Groeneveld RA, Meeden G. Measuring skewness and kurtosis. Statistician. 1984; 33:391-399.
32. Haralick RM, Shanmugam K, Dinstein IH. Textural features for image classification. IEEE Trans Syst Man Cybern Syst. 1973; SMC-3: 610-621.
33. Clausi DA. An analysis of co-occurrence texture statistics as a function of grey level quantization. Can J Remote Sensing. 2002; 28:45-62.
34. Soh LK, Tsatsoulis C. Texture analysis of SAR sea ice imagery using gray level co-occurrence matrices. IEEE Trans Geosci Remote Sens. 1999; 37:780-795.
35. Lo CM, Chen RT, Chang YC, Yang YW, Hung MJ, Huang CS, Chang RF. Multi-dimensional tumor detection in automated whole breast ultrasound using topographic watershed. IEEE Trans Med Imaging. 2014; 33:1503-1511.
36. Field AP. (2009). Discovering statistics using SPSS, 3rd ed. (Los Angeles: SAGE Publications).

