INTRODUCTION
Hepatocellular carcinoma (HCC) is a common malignant tumor of digestive system, and a complex disease caused by interactions between multiple genetic and environmental factors [1, 2]. There are about 800,000 new cases of HCC around the world each year, of which more than 50% occurs in China [3, 4]. Guangxi is one region with the highest incidence of HCC in China, where HCC is the leading cause of tumor-related mortality. Because of insidious onset and high malignancy, most HCC patients are diagnosed in advanced stage with poor prognosis. Although comprehensive treatment based on surgery has improved the curative effect of HCC greatly, clinical cure rate and long-term survival rate have no significant improvement. Recently, studies on the causes, prognosis markers, molecular targets and carcinogenesis mechanisms of HCC are increasing widely [5–8], but the exact molecular mechanism is still not well understood. Therefore, it is very crucial to find the molecular markers which can be used in the risk and prognosis evaluation for the prevention and treatment of HCC.
Single nucleotide polymorphism (SNP) is the third generation of molecular marker and one of the most common genetic variations in human. Studies show that SNP can not only be used as a genetic marker locating disease gene, some SNP itself can also directly lead to the occurrence of diseases [9, 10]. Thus SNP has crucial function and application in disease risk assessment, early diagnosis, prevention, treatment and drug development [11–13]. Mismatch repair (MMR) system is a DNA repair system with a high fidelity. It mainly repairs base mismatch and insertion/deletion ring produced in the synthesis of DNA to maintain the stability of the whole genome [14–16]. MutL homolog 1 (MLH1) plays a key role in the MMR system. MLH1 recognizes and repairs the mismatched bases in the process of DNA replication, and also recruits other mismatch repair proteins to the mismatch site to correct DNA replication error [17, 18]. Studies found that SNPs of specific DNA repair genes could affect the expression level and activity of enzymes and individual DNA damage repair efficiency. Repair gene defects may lead to genetic instability and cancer occurrence, suggesting that individual differences in cancer risk was related to polymorphisms of specific repair genes [19–21]. It has been reported MLH1 polymorphisms had a correlation with the pathogenesis of colorectal cancer, lung cancer, breast cancer, prostate cancer, head and neck squamous cell carcinoma, oral squamous cell carcinoma [19–27]. However, there is only a few researches about the relationship between MLH1 polymorphism and HCC [28], and the results still need further verification due to race, nation, region and individual differences.
In order to clarify genetic variations of MLH1 in HCC, this study aims to screen potential MLH1 SNPs in a case-control study from a HCC population in Guangxi. Four MLH1 SNPs, rs1800734, rs10849, rs3774343 and rs1540354, that minor allele frequency (MAF) was more than 0.05, were selected from the National Institute of Environmental Health Sciences (NIEHS) database. Through the investigation and collection of demographic and clinical information from the study population, the relationships between these MLH1 SNPs with risk and prognosis of HCC were analyzed. Finally, we found that rs1800734 was a new predictor for poor prognosis of HCC patients.
RESULTS
General demographic characteristics of study population and Hardy-Weinberg equilibrium test results
The general demographic characteristics of study population were shown in Table 1. The case and control group were significantly different at alcohol intake, smoking, HBV infection and family history (P < 0.001). However, there was no difference about gender, age or nation between the two groups (P > 0.05).
Table 1: Distributions of general demographic characteristics and environmental risk factors in the controls and cases
Characteristics |
Controls (n = 1036) |
Cases (n = 1036) |
χ2 |
P |
|---|---|---|---|---|
Age (years) |
||||
< 50 |
497 |
532 |
2.365 |
0.124 |
≥ 50 |
539 |
504 |
||
Gender |
||||
Male |
896 |
896 |
0.000 |
1.000 |
Female |
140 |
140 |
||
Nation |
||||
Han |
736 |
715 |
1.048 |
0.592 |
Zhuang |
286 |
305 |
||
Others |
14 |
16 |
||
Smoking |
||||
No |
879 |
674 |
108.03 |
< 0.001 |
Yes |
157 |
362 |
||
Alcohol intake |
||||
No |
895 |
689 |
113.75 |
< 0.001 |
Yes |
141 |
347 |
||
HBV infection |
||||
No |
953 |
176 |
1174.97 |
< 0.001 |
Yes |
83 |
860 |
||
Family history |
||||
No |
1022 |
975 |
30.56 |
< 0.001 |
Yes |
14 |
61 |
Bold values indicate significance.
Haploview 4.2 test showed genotype distribution of the four MLH1 SNPs, rs1800734, rs10849, rs3774343, and rs1540354 accorded with the HWE equilibrium in the control group (Supplementary Table 1).
The relationship between genotype distribution of MLH1 SNPs and HCC risk
Multivariate logistic regression analysis showed that MLH1 SNP, rs1800734 had a significant difference between the case and control group (Table 2, P < 0.05). After adjusted for age, gender, smoking, alcohol intake and HBV infection, compared with the AA genotype, the ORs (95% CI) of AG, GG and AG/GG genotype with HCC risk were 1.217 (1.074~1.536), 1.745 (1.301~2.591) and 1.291 (1.126~1.687), respectively. The other three MLH1 SNPs, rs10849, rs3774343 and rs1540354 had no statistically significant differences between the case and control group (P > 0.05).
Table 2: The genotype frequencies of MLH1 polymorphisms and HCC risk
Genotypes |
Cases (%) |
Controls (%) |
Frequencies in HapMap project (%) |
OR (95% CI)a |
OR (95% CI)b |
|---|---|---|---|---|---|
rs1800734 |
|||||
AA |
393 (37.16) |
338 (32.63) |
30.23 |
1.000 |
1.000 |
AG |
522 (50.39) |
529 (51.06) |
44.19 |
1.178 (0.975~1.424) |
1.217 (1.074~1.536) |
GG |
121 (12.45) |
169 (16.31) |
25.58 |
1.624 (1.234~2.138) |
1.745 (1.301~2.591) |
AG/GG |
643 (62.84) |
675 (67.37) |
69.77 |
1.221 (1.018~1.463) |
1.291 (1.126~1.687) |
A |
1308 (63.13) |
1205 (58.16) |
52.33 |
- |
- |
G |
764 (36.87) |
867 (41.84) |
47.67 |
- |
- |
rs10849 |
|||||
AA |
7 (0.67) |
3 (0.29) |
0 |
1.000 |
1.000 |
AG |
151 (14.58) |
162 (15.64) |
0.19 |
2.503 (0.636~9.857) |
2.625 (0.651~11.236) |
GG |
878 (84.75) |
871 (84.07) |
0.81 |
2.315 (0.597~8.890) |
2.431 (0.605~9.873) |
AG/GG |
1029 (99.33) |
1033 (99.71) |
100.00 |
2.342 (0.604~9.083) |
2.526 (0.612~10.585) |
A |
165 (7.96) |
168 (8.11) |
9.30 |
- |
- |
G |
1907 (92.04) |
1904 (91.89) |
90.70 |
- |
- |
rs3774343 |
|||||
CC |
8 (0.77) |
2 (0.19) |
0 |
1.000 |
1.000 |
CT |
143 (13.80) |
125 (12.06) |
11.63 |
3.497 (0.729~16.772) |
3.624 (0.756~19.245) |
TT |
885 (85.43) |
909 (87.75) |
88.37 |
4.108 (0.870~19.401) |
4.162 (0.923~21.523) |
CT/TT |
1028 (90.23) |
1034 (99.81) |
100.00 |
4.023 (0.852~18.992) |
4.101 (0.891~20.317) |
C |
159 (7.67) |
129 (6.23) |
5.81 |
- |
- |
T |
1913 (92.33) |
1943 (93.77) |
94.19 |
- |
- |
rs1540354 |
|||||
AA |
96 (9.27) |
81 (7.82) |
13.95 |
1.000 |
1.000 |
AT |
489 (47.20) |
447 (43.15) |
39.53 |
1.083 (0.785~1.495) |
1.125 (0.813~1.542) |
TT |
451 (43.53) |
508 (49.03) |
46.51 |
1.335 (0.968~1.842) |
1.363 (0.971~1.876) |
AT/TT |
940 (90.73) |
955 (92.18) |
86.04 |
1.204 (0.884~1.640) |
1.237 (0.896~1.679) |
A |
681 (32.87) |
609 (29.39) |
33.72 |
- |
- |
T |
1391 (67.13) |
1463 (70.61) |
66.28 |
- |
- |
a: OR (95% CI) not adjusted; b: OR (95% CI) adjusted by logistic regression for age, gender, nation, smoking, alcohol intake, HBV infection, and HCC family history. Bold values indicate significance.
Gene-environment and SNP-SNP interaction
Logistic regression model analysis showed that rs1800734, rs10849 and rs3774343 had interactions with such environment factors, HBV infection, alcohol intake and smoking in the pathogenesis of HCC (Table 3, P < 0.05).
Table 3: Gene-environment interaction
Factors |
β |
S.E. |
Wald χ2 |
OR (95% CI)a |
P |
|---|---|---|---|---|---|
rs1800734 × Smoking |
0.263 |
0.116 |
6.354 |
1.195(1.040~2.270) |
0.012 |
rs1800734 × Alcohol intake |
0.358 |
0.103 |
14.660 |
1.294(1.134~2.132) |
0.000 |
rs1800734 × HBV infection |
2.132 |
0.127 |
735.759 |
8.685(6.136~13.246) |
0.000 |
rs1800734 × Family history |
0.210 |
0.235 |
1.862 |
1.105(0.748~1.982) |
0.176 |
rs10849 × Smoking |
0.253 |
0.109 |
6.102 |
1.183(1.036~2.131) |
0.014 |
rs10849 × Alcohol intake |
0.342 |
0.121 |
11.638 |
1.217(1.097~2.061) |
0.000 |
rs10849 × HBV infection |
1.532 |
0.127 |
657.842 |
5.685(4.136~9.246) |
0.000 |
rs10849 × Family history |
0.198 |
0.213 |
1.721 |
1.079(0.676~1.837) |
0.225 |
rs3774343 × Smoking |
0.232 |
0.123 |
5.657 |
1.152(1.027~1.846) |
0.021 |
rs3774343 × Alcohol intake |
0.326 |
0.113 |
9.597 |
1.198(1.071~1.956) |
0.007 |
rs3774343 × HBV infection |
1.392 |
0.115 |
548.985 |
4.913(3.694~8.635) |
0.000 |
rs3774343 × Family history |
0.191 |
0.261 |
1.678 |
1.024(0.651~1.736) |
0.292 |
rs1540354 × Smoking |
0.245 |
0.129 |
6.047 |
1.197(1.065~1.976) |
0.019 |
rs1540354 × Alcohol intake |
0.317 |
0.124 |
10.436 |
1.209(1.087~2.012) |
0.011 |
rs1540354 × HBV infection |
1.516 |
0.117 |
632.761 |
5.215(3.956~8.957) |
0.000 |
rs1540354 × Family history |
0.211 |
0.221 |
1.702 |
1.105(0.672~1.894) |
0.258 |
a: OR (95% CI) adjusted by logistic regression for age, gender, nations, smoking, alcohol intake, HBV infection, and HCC family history. Bold values indicate significance.
In addition, as shown in Supplementary Table 2, rs1800734 and rs10849, rs1800734 and rs3774343, rs1800734 and rs1540354 had SNP-SNP interactions in the pathogenesis of HCC (P < 0.05), and these interactions could increase HCC risk.
Associations between MLH1 polymorphisms with clinical-pathological characteristics and the prognosis of HCC patients
As shown in Table 4, MLH1 SNP, rs1800734 was correlated with tumor size, staging and AFP level of HCC patients (P < 0.05), while rs10849, rs3774343 and rs1540354 had no association with these clinical-pathological characteristics of HCC patients.
Table 4: The associations between the MLH1 polymorphisms and clinical characteristics of HCC patients
Variables |
AA |
AG/GG |
OR (95% CI)a |
OR (95% CI)b |
|---|---|---|---|---|
rs1800734 |
||||
Tumor size |
||||
< 5 cm |
311 (29.17) |
455 (70.83) |
1.000 |
1.000 |
≥ 5 cm |
82 (34.56) |
188 (63.44) |
1.567 (1.165~2.109) |
1.671 (1.215~2.252) |
Tumor number |
||||
solitary |
332 (30.58) |
514 (69.42) |
1.000 |
1.000 |
multiple |
61 (29.91) |
129 (70.09) |
1.366 (0.978~1.908) |
1.379 (0.988~1.926) |
TNM staging |
||||
T1 + T2 |
337 (31.30) |
518 (68.70) |
1.000 |
1.000 |
T3 + T4 |
56 (26.14) |
125 (74.86) |
1.452 (1.030~2.048) |
1.545 (1.092~2.185) |
AFP level (ng/ml) |
||||
< 400 |
152 (37.16) |
206 (62.84) |
1.000 |
1.000 |
≥ 400 |
241 (29.45) |
437 (70.55) |
1.338 (1.030~1.739) |
1.463 (1.126~1.903) |
Lymphatic metastasis |
||||
No |
343 (29.02) |
535 (70.98) |
1.000 |
1.000 |
Yes |
50 (40.15) |
107 (59.85) |
1.372 (0.955~1.971) |
1.433 (0.996~2.061) |
rs10849 |
||||
Tumor size |
||||
< 5 cm |
5 (4.64) |
761 (95.36) |
1.000 |
1.000 |
≥ 5 cm |
2 (7.32) |
268 (92.68) |
0.880 (0.170~4.565) |
0.913 (0.215~5.294) |
Tumor number |
||||
solitary |
6 (5.50) |
840 (94.50) |
1.000 |
1.000 |
multiple |
1 (2.56) |
189 (97.44) |
1.350 (0.162~11.279) |
1.401 (0.168~11.706) |
TNM staging |
||||
T1+T2 |
6 (5.73) |
849 (94.27) |
1.000 |
1.000 |
T3+T4 |
1 (1.96) |
180 (98.04) |
1.272 (0.152~10.631) |
1.315 (0.163~11.129) |
AFP level (ng/ml) |
||||
< 400 |
3 (6.89) |
355 (93.10) |
1.000 |
1.000 |
≥ 400 |
4 (4.74) |
674 (95.26) |
1.424 (0.317~6.397) |
1.87 (0.331~6.681) |
Lymphatic metastasis |
||||
No |
7 (5.24) |
871 (94.76) |
1.000 |
1.000 |
Yes |
0 (38.22) |
157 (61.78) |
0.000 |
0.000 |
Variables |
CC |
CT/TT |
OR (95% CI)a |
OR (95% CI)b |
|---|---|---|---|---|
rs3774343 |
||||
Tumor size |
||||
< 5 cm |
6 (3.23) |
760 (96.77) |
1.000 |
1.000 |
≥ 5 cm |
2 (2.44) |
268 (97.56) |
1.058 (0.212~5.273) |
1.117 (0.224~5.569) |
Tumor number |
||||
solitary |
7 (3.09) |
839 (96.91) |
1.000 |
1.000 |
multiple |
1 (2.56) |
189 (97.44) |
1.577 (0.193~12.893) |
1.635 (0.206~13.614) |
TNM staging |
||||
T1 + T2 |
7 (3.58) |
848 (96.42) |
1.000 |
1.000 |
T3 + T4 |
1 (3.92) |
180 (96.08) |
1.486 (0.182~12.151) |
1.537 (0.196~13.513) |
AFP level (ng/ml) |
||||
< 400 |
3 (1.02) |
355 (98.98) |
1.000 |
1.000 |
≥ 400 |
5 (3.88) |
673 (96.12) |
1.137 (0.270~4.787) |
1.198 (0.305~5.477) |
Lymphatic metastasis |
||||
No |
8 (3.50) |
870 (96.50) |
1.000 |
1.000 |
Yes |
0 (4.55) |
157 (95.45) |
0.000 |
0.000 |
Variables |
AA |
AT/TT |
OR (95% CI)a |
OR (95% CI)b |
|---|---|---|---|---|
rs1540354 |
||||
Tumor size |
||||
< 5 cm |
75 (67.34) |
691 (32.66) |
1.000 |
1.000 |
≥ 5 cm |
21 (67.07) |
249 (32.93) |
1.287 (0.777~2.133) |
1.369 (0.819~2.288) |
Tumor number |
||||
solitary |
81 (69.76) |
765 (30.24) |
1.000 |
1.000 |
multiple |
15 (66.66) |
175 (33.34) |
1.235 (0.695~2.195) |
1.324 (0.733~2.389) |
TNM staging |
||||
T1 + T2 |
77 (70.25) |
778 (29.75) |
1.000 |
1.000 |
T3 + T4 |
19 (64.00) |
162 (36.00) |
0.844 (0.497~1.433) |
0.875 (0.518~1.532) |
AFP level (ng/ml) |
||||
< 400 |
36 (69.39) |
322 (30.61) |
1.000 |
1.000 |
≥ 400 |
60 (69.40) |
618 (30.60) |
1.152 (0.746~1.778) |
1.216 (0.792~1.865) |
Lymphatic metastasis |
||||
No |
86 (68.31) |
792 (31.69) |
1.000 |
1.000 |
Yes |
10 (75.00) |
148 (25.00) |
1.607 (0.816~3.166) |
1.786 (0.879~3.627) |
a: OR (95% CI) not adjusted; b: OR (95% CI) adjusted by logistic regression for age, gender, nation, smoking, alcohol intake, HBV infection, and HCC family history.
At the end of the follow-up, there were 37.84% (165/436) of HCC patients died. We found that the prognosis of HCC had a correlation with these clinical-pathological characteristics of patients, such as tumor size, number, staging, AFP level and lymph node metastasis (P < 0.05, Table 5), but was not associated with age, gender, HBV infection or family history (P > 0.05).
Table 5: The associations between general demographic and clinical characteristics with the prognosis of HCC patients
Variables |
Cases |
Survivors |
MST |
Log-rank P |
HR (95% CI) |
|---|---|---|---|---|---|
N = 436 |
N = 271 |
Months |
|||
Age (years) |
|||||
< 50 |
268 |
172 |
49.0 |
0.650 |
1.000 |
≥ 50 |
168 |
99 |
46.7 |
1.074 (0.788–1.464) |
|
Gender |
|||||
Male |
353 |
222 |
48.7 |
0.723 |
1.000 |
Female |
83 |
49 |
46.4 |
0.934 (0.639–1.365) |
|
HBV infection |
|||||
No |
62 |
38 |
46.4 |
0.951 |
1.000 |
Yes |
374 |
233 |
48.7 |
0.821 (0.617–1.467) |
|
Family history |
|||||
No |
399 |
248 |
48.6 |
0.917 |
1.000 |
Yes |
37 |
23 |
46.7 |
0.971 (0.561–1.681) |
|
Tumor size |
|||||
< 5 cm |
243 |
171 |
51.5 |
0.001 |
1.000 |
≥ 5 cm |
193 |
100 |
44.9 |
1.668 (1.226–2.270) |
|
Tumor number |
|||||
solitary |
288 |
202 |
51.5 |
0.000 |
1.000 |
multiple |
148 |
69 |
43.5 |
1.746 (1.300–2.395) |
|
TNM staging |
|||||
T1 + T2 |
300 |
214 |
56.0 |
0.000 |
1.000 |
T3 + T4 |
136 |
57 |
38.4 |
1.471 (1.053–2.054) |
|
AFP level (ng/ml) |
|||||
< 400 |
169 |
120 |
51.0 |
0.023 |
1.000 |
≥ 400 |
267 |
151 |
45.3 |
1.520 (1.080–2.138) |
|
Lymphatic metastasis |
|||||
No |
381 |
254 |
49.6 |
0.000 |
1.000 |
Yes |
55 |
17 |
32.3 |
2.458 (1.710–3.533) |
Bold values indicate significance.
Moreover, the co-dominant and dominant models of rs1800734 had a significant influence on the prognosis of HCC (Table 6, Figure 1 and Supplementary Figure 1). Compared with the AA genotype, the survival time of HCC patients with AG, GG and AG/GG genotype significantly decreased (P < 0.05). But the recessive model of rs1800734 had no effect on the survival time of HCC patients. The genotypes of the other three MLH1 SNPs, rs10849, rs3774343 and rs1540354 had no effect on the survival time of HCC patients (Figure 1 and Supplementary Figure 1).
Table 6: The associations between MLH1 polymorphisms and the prognosis of HCC patients
SNPs |
Genotypes |
Survivors / Cases |
MST (Months) |
Log-rank P |
HR (95% CI) |
|---|---|---|---|---|---|
(N = 271) / (N = 436) |
|||||
rs1800734 |
|||||
Co-dominant |
AA |
110/163 |
56.0 |
0.000 |
1.553 (1.257–1.920) |
AG |
126/212 |
42.0 |
|||
GG |
35/61 |
45.1 |
|||
Dominant |
AA |
110/163 |
56.0 |
0.000 |
2.207 (1.572–3.100) |
AG/GG |
161/273 |
44.3 |
|||
Recessive |
AG/AA |
236/375 |
48.7 |
0.120 |
1.393 (0.915–2.122) |
GG |
35/61 |
45.1 |
|||
rs10849 |
|||||
Co-dominant |
AA |
1/4 |
22.6 |
0.548 |
0.908 (0.631–1.307) |
AG |
47/72 |
48.6 |
|||
GG |
223/360 |
47.6 |
|||
Dominant |
AA |
1/4 |
22.6 |
0.273 |
0.530 (0.166–1.685) |
AG/GG |
270/432 |
48.6 |
|||
Recessive |
AG/AA |
48/76 |
48.6 |
0.780 |
0.944 (0.628–1.418) |
GG |
223/360 |
47.6 |
|||
rs3774343 |
|||||
Co-dominant |
CC |
2/5 |
23.8 |
0.371 |
1.091 (0.756–1.573) |
CT |
55/81 |
46.7 |
|||
TT |
214/350 |
48.6 |
|||
Dominant |
CC |
2/5 |
23.8 |
0.316 |
0.559 (0.176–1.776) |
CT/TT |
269/431 |
48.6 |
|||
Recessive |
CT/CC |
57/86 |
46.7 |
0.450 |
1.167 (0.781–1.744) |
TT |
214/350 |
48.6 |
|||
rs1540354 |
|||||
Co-dominant |
AA |
21/39 |
41.3 |
0.435 |
0.903 (0.711–1.148) |
AT |
133/208 |
51.2 |
|||
TT |
117/189 |
48.6 |
|||
Dominant |
AA |
21/39 |
41.3 |
0.197 |
0.726 (0.445–1.184) |
AT/TT |
250/397 |
48.7 |
|||
Recessive |
AT/AA |
154/247 |
47.6 |
0.733 |
0.948 (0.696–1.290) |
TT |
117/189 |
48.6 |
|||
Bold values indicate significance.
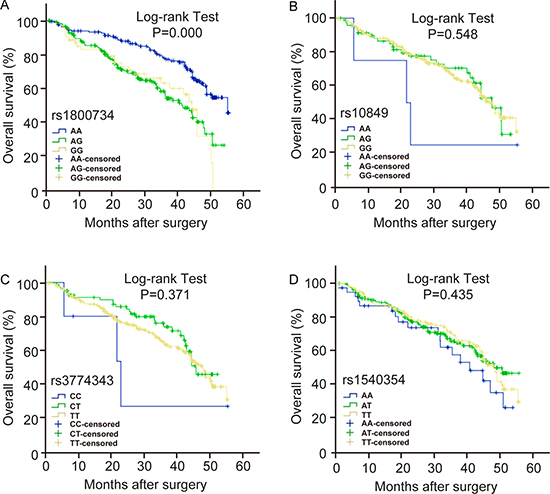
Figure 1: The effect of MLH1 polymorphisms on the prognosis of HCC patients. Kaplan-Meier overall survival curves for HCC patients based on co-dominate genotypes of (A) rs1800734, (B) rs10849, (C) rs3774343, (D) rs1540354. P value is from the log-rank test.
Regression analysis for the survival of HCC patients
We conducted a multivariate regression analysis combined MLH1 polymorphisms, demographic and clinical-pathological characteristics of HCC patients. As shown in Table 7, these factors, such as rs1800734, tumor number, tumor staging and metastasis, were associated with the death risk of HCC patients.
Table 7: COX regression analysis of the prognosis of HCC patients
Variables |
β |
S.E. |
HR |
95% CI |
P |
|---|---|---|---|---|---|
Tumor number |
0.496 |
0.164 |
1.642 |
1.191–2.264 |
0.002 |
TNM staging |
0.691 |
0.162 |
1.995 |
1.452–2.742 |
0.000 |
Metastasis |
1.184 |
0.204 |
3.268 |
2.191–74.875 |
0.000 |
rs1800734 |
1.289 |
0.194 |
3.629 |
2.481–5.310 |
0.000 |
Bold values indicate significance.
DISCUSSION
As the most common malignant tumor in southern Guangxi, HCC is a complex process that multiple genes and environmental factors involved [1, 2]. It is well known that environmental carcinogens cause DNA damage by continuing to attack the genomic DNA. If the damage DNA can’t be repaired timely and effectively, the accumulated damage will lead to an increased genomic instability, resulting in cell apoptosis, deregulated cell proliferation and differentiation, eventually cancer [17, 29, 30]. Recent studies report that MLH1, a mismatch repair gene, plays an important role in carcinogenesis [19, 27, 31, 32]. SNPs of mismatch repair genes are thought to provide useful information for tumor diagnosis, suggesting that SNPs of MLH1 may have potential value for diagnosis of HCC. However, the relationship between MLH1 polymorphisms and HCC has not been reported in Chinese population yet.
This study analyzed the relationship between MLH1 polymorphisms and HCC susceptibility in Guangxi area, where there is a high incidence of HCC every year. Finally, we found that the genotype distribution of rs1800734 had a significant difference between cases and controls. The AG, GG and AG/GG genotype of rs1800734 increased HCC risk compared with the AA genotype. But the other three SNPs of MLH1, rs10849, rs3774343 and rs1540354 were not correlated with HCC risk.
It has been reported that different populations have different genotype distribution of MLH1 polymorphisms [15]. The genotype of MLH1 polymorphisms in European and American population was mainly GG, while our results were in accordance with the genotype in Asians [24, 28, 33]. LO et al. explored the relationship between MLH1 polymorphisms and lung cancer risk in a case-control study and discovered that rs1800734 was closely related to the occurrence of lung cancer [24]. GG genotype of rs1800734 increased the lung cancer risk compared to AA genotype. Due to genetic differences between different regions and populations, we found only rs1800734 had a correlation with the occurrence and development of HCC. The result is consistent with previous study that rs1800734 significantly increased HCC risk [28]. Whether the other MLH1 polymorphisms are associated with HCC needs a prospective and large sample study to be verified.
Previous studies indicated that HBV infection, smoking, alcohol intake and family history of cancer were important environmental risk factors of HCC [6, 34, 35]. Thus we conducted gene-environment interaction analysis between the four MLH1 polymorphisms and these environmental risk factors. The four MLH1 polymorphisms were found to interact with HBV infection, alcohol intake and smoking, and increase the risk of HCC. In addition, rs1800734 had an interaction with SNPs rs10849, rs3774343 and rs1540354, these SNP-SNP interactions also increased HCC risk, suggesting that MLH1 polymorphisms play an important role in the development of HCC. However, we didn’t find any interaction between MLH1 polymorphisms and family history of cancer, which may be due to the small sample size and low test efficiency.
HCC is a highly malignant tumor with a very poor prognosis [36–38]. Accumulated evidences showed that the clinical-pathological features were closely related to the prognosis of tumor, such as tumor size, tumor number, tumor stage, AFP level, lymph node metastasis, tumor thrombus, liver cirrhosis, vascular invasion and migration predicted poor prognosis for HCC [39–42]. This study clarified that rs1800734 was correlated with tumor size, tumor grade and AFP level, indicating that gene polymorphisms may cause carcinogenesis and different clinical-pathological features of patients [39, 42]. Furthermore, we found rs1800734, tumor number, tumor stage and lymph node metastasis were correlated with the prognosis of HCC in a COX regression analysis, and rs1800734 decreased the survival time of HCC patients significantly, suggesting that rs1800734 was a risk factor of HCC prognosis.
In addition, rs1800734 is a transcription factor binding site (TFBS), which can cause individual susceptibility difference by regulating MLH1 activity and the expression of downstream proteins [19, 22]. The relationship between rs1800734 and HCC prognosis may result from its regulation of the binding ability of transcription factors on MLH1, and then lead to expression change of MLH1 in HCC. However, the exact mechanism need further study to prove.
In summary, this is the first report of the relationship between MLH1 polymorphisms with the risk and prognosis of HCC. We found that rs1800734 increased the risk of HCC and was a risk factor for poor prognosis of HCC, which is expected to become a new biomarker of HCC with poor prognosis.
MATERIALS AND METHODS
Ethics statement
The study was approved by the ethics committee of the First Affiliated Hospital of Guilin Medical University. All patients were informed about the aims of sample collection and signed the informed consent in accordance with the ethical guidelines of the hospital.
Study population
1,036 new cases of HCC patients were collected as the case group from Department of Hepatobiliary Surgery, the First Affiliated Hospital of Guilin Medical University and the Guangxi Medical University between July 2009 and June 2015. These HCC patients were diagnosed as HCC by radiological or pathological methods and had not received chemotherapy and radiotherapy before the blood collection. 1,036 cases of non-HCC patients at the same time were selected as the control group from Department of Spinal Bone Marrow Surgery and Hand Trauma Surgery, the First Affiliated Hospital of Guilin Medical University and the Guangxi Medical University, with gender, age and nation matched.
Blood sample collection and investigation
The investigation questionnaire was designed after consulting experts and conducted by trained investigators in a face-to-face way. The questionnaire included general information, such as past history, personal history, family history, smoking and alcohol intake, and clinical data, such as tumor size, number, staging, portal vein tumor thrombus (PVTT), hepatitis virus infection, AFP level, histological grade, and lymph node metastasis. 2 ml peripheral blood was collected from all patients for DNA extraction. The blood DNA was stored at −80°C.
Follow up
475 cases of HCC patients underwent surgical resection between July 2010 and June 2015 were chosen in follow-up by telephone or outpatient review. Since the date of entering the group, they were follow-up every six months until June 2016. The survival time was counted from the first day after operation to the day when patients had metastasis, recurrence, death or the end of the follow-up. At the end of follow-up, 39 patients were lost and 436 patients had complete follow-up data.
The inclusion criteria for follow-up patients: (1) could undergo radical surgery after preoperative blood and imaging examination; (2) were confirmed as HCC by clinical pathology after radical surgery; (3) provided the blood sample; (4) had complete clinical and prognosis information; (5) signed the informed consent. The exclusion criteria for follow-up patients: (1) had not undergone radical surgery; (2) without complete clinical information; (3) refused to provide blood sample; (4) refused to sign the informed consent.
Genotyping
Genotyping was conducted on Applied Biosystems 7500 Fast Real-Time PCR System (ABI, United States) by TaqMan MGB high throughput RT-PCR method and the results were analyzed on 7500 Fast System V1.4.0 SDS software. For quality control, 5% sample DNA was chosen to repeat the genotyping and the concordance rate was 100%. The genotyping was repeated once again when the sample couldn’t be genotyped, and the sample was abandoned if it couldn’t be genotyped in the repeat genotyping.
Statistical analysis
All statistical analysis was conducted using SPSS 19.0 software. Hardy Weinberg equilibrium (HWE) test in the control group was performed using Haploview 4.2. Qualitative and quantitative data were analyzed by χ2 and t test, respectively. SNP-SNP interaction, gene-environment interaction, 95% confidence interval (CI) and odds ratio (OR) were analyzed by binary logistic regression model. The median survival time (MST) was calculated from the date of diagnosis to the date of death or the end of follow-up (June 2016). The overall survival curves were draw by Kaplan-Meier method and the differences between groups were analyzed using log-rank test. Multivariate COX regression was used for calculating hazard ratio (HR) and 95% CI. All tests were two-tailed and P < 0.05 was considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
This work was supported by the National Nature Science Foundation of China (NSFC, 81460515).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Lin H, van den Esschert J, Liu C, van Gulik TM. Systematic review of hepatocellular adenoma in China and other regions. J Gastroenterol Hepatol. 2011; 26:28–35.
2. Nie W, Yan L, Lee YH, Guha C, Kurland IJ, Lu H. Advanced mass spectrometry-based multi-omics technologies for exploring the pathogenesis of hepatocellular carcinoma. Mass Spectrom Rev. 2016; 35:331–349.
3. Chen JG, Zhang SW. Liver cancer epidemic in China: past, present and future. Semin Cancer Biol. 2011; 21:59–69.
4. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108.
5. Alkan C, Coe BP, Eichler EE. Genome structural variation discovery and genotyping. Nat Rev Genet. 2011; 12:363–376.
6. Li J, Xu Y, Long XD, Wang W, Jiao HK, Mei Z, Yin QQ, Ma LN, Zhou AW, Wang LS, Yao M, Xia Q, Chen GQ. Cbx4 governs HIF-1alpha to potentiate angiogenesis of hepatocellular carcinoma by its SUMO E3 ligase activity. Cancer cell. 2014; 25:118–131.
7. Tan C, Liu S, Tan S, Zeng X, Yu H, Li A, Bei C, Qiu X. Polymorphisms in microRNA target sites of forkhead box O genes are associated with hepatocellular carcinoma. PloS one. 2015; 10:e0119210.
8. Xia L, Huang W, Tian D, Zhu H, Qi X, Chen Z, Zhang Y, Hu H, Fan D, Nie Y, Wu K. Overexpression of forkhead box C1 promotes tumor metastasis and indicates poor prognosis in hepatocellular carcinoma. Hepatology. 2013; 57:610–624.
9. Chandrasekaran G, Hwang EC, Kang TW, Kwon D, Park K, Lee JJ, Lakshmanan VK. In Silico Analysis of the deleterious nsSNP’s (missense) in the Homeobox domain of human HOXB13 gene responsible for hereditary prostate cancer. Chem Biol Drug Des. 2017 Jan 10. doi: 10.1111/cbdd.12938. [Epub ahead of print].
10. Haugarvoll K, Johansson S, Rodriguez CE, Boman H, Haukanes BI, Bruland O, Roque F, Jonassen I, Blomqvist M, Telstad W, Mansson JE, Knappskog PM, Bindoff LA. GBA2 Mutations Cause a Marinesco-Sjogren-Like Syndrome: Genetic and Biochemical Studies. PloS one. 2017; 12:e0169309.
11. De Wit P, Pespeni MH, Palumbi SR. SNP genotyping and population genomics from expressed sequences - current advances and future possibilities. Mol Ecol. 2015; 24:2310–2323.
12. Sayyed DR, Nimse SB, Song KS, Sung N, Kim T. Multiplex SNP detection in multiple codons for accurate drug therapy. Chem Commun. 2014; 50:14585–14588.
13. Wills QF, Mead AJ. Application of single-cell genomics in cancer: promise and challenges. Hum Mol Genet. 2015; 24:R74–84.
14. Bodo S, Colas C, Buhard O, Collura A, Tinat J, Lavoine N, Guilloux A, Chalastanis A, Lafitte P, Coulet F, Buisine MP, Ilencikova D, Ruiz-Ponte C, et al. Diagnosis of Constitutional Mismatch Repair-Deficiency Syndrome Based on Microsatellite Instability and Lymphocyte Tolerance to Methylating Agents. Gastroenterology. 2015; 149:1017–1029 e1013.
15. Campbell PT, Curtin K, Ulrich CM, Samowitz WS, Bigler J, Velicer CM, Caan B, Potter JD, Slattery ML. Mismatch repair polymorphisms and risk of colon cancer, tumour microsatellite instability and interactions with lifestyle factors. Gut. 2009; 58:661–667.
16. Scarpa M, Ruffolo C, Canal F, Scarpa M, Basato S, Erroi F, Fiorot A, Dall’Agnese L, Pozza A, Porzionato A, Castagliuolo I, Dei Tos AP, Bassi N, et al. Mismatch repair gene defects in sporadic colorectal cancer enhance immune surveillance. Oncotarget. 2015; 6:43472–43482. doi: 10.18632/oncotarget.6179.
17. Cohen SA, Turner EH, Beightol MB, Jacobson A, Gooley TA, Salipante SJ, Haraldsdottir S, Smith C, Scroggins S, Tait JF, Grady WM, Lin EH, Cohn DE, et al. Frequent PIK3CA Mutations in Colorectal and Endometrial Tumors With 2 or More Somatic Mutations in Mismatch Repair Genes. Gastroenterology. 2016; 151:440–447 e441.
18. Riazy M, Kalloger SE, Sheffield BS, Peixoto RD, Li-Chang HH, Scudamore CH, Renouf DJ, Schaeffer DF. Mismatch repair status may predict response to adjuvant chemotherapy in resectable pancreatic ductal adenocarcinoma. Modern Pathol. 2015; 28:1383–1389.
19. Ma G, Ge Y, Gu D, Du M, Chu H, Chen J, Zhang Z, Wang M. Functional annotation of colorectal cancer susceptibility loci identifies MLH1 rs1800734 associated with MSI patients. Gut. 2016; 65:1227–1228.
20. Nogueira GA, Lourenco GJ, Oliveira CB, Marson FA, Lopes-Aguiar L, Costa EF, Lima TR, Liutti VT, Leal F, Santos VC, Rinck-Junior JA, Lima CS. Association between genetic polymorphisms in DNA mismatch repair-related genes with risk and prognosis of head and neck squamous cell carcinoma. Int J Cancer. 2015; 137:810–818.
21. van der Klift HM, Mensenkamp AR, Drost M, Bik EC, Vos YJ, Gille HJ, Redeker BE, Tiersma Y, Zonneveld JB, Garcia EG, Letteboer TG, Olderode-Berends MJ, van Hest LP, et al. Comprehensive Mutation Analysis of PMS2 in a Large Cohort of Probands Suspected of Lynch Syndrome or Constitutional Mismatch Repair Deficiency Syndrome. Hum Mutat. 2016; 37:1162–1179.
22. Harkness EF, Barrow E, Newton K, Green K, Clancy T, Lalloo F, Hill J, Evans DG. Lynch syndrome caused by MLH1 mutations is associated with an increased risk of breast cancer: a cohort study. J Med Genet. 2015; 52:553–556.
23. Lin LH, Lin MW, Mar K, Lin CS, Ji DD, Lee WP, Lee HS, Cheng MF, Hsia KT. The hMLH1 -93G>A promoter polymorphism is associates with outcomes in oral squamous cell carcinoma patients. Ann Surg Oncol. 2014; 21:4270–4277.
24. Lo YL, Hsiao CF, Jou YS, Chang GC, Tsai YH, Su WC, Chen KY, Chen YM, Huang MS, Hsieh WS, Chen CJ, Hsiung CA. Polymorphisms of MLH1 and MSH2 genes and the risk of lung cancer among never smokers. Lung cancer. 2011; 72:280–286.
25. Niu L, Li S, Liang H, Li H. The hMLH1 -93G>A Polymorphism and Risk of Ovarian Cancer in the Chinese Population. PloS one. 2015; 10:e0135822.
26. Tomlinson IP, Houlston RS, Montgomery GW, Sieber OM, Dunlop MG. Investigation of the effects of DNA repair gene polymorphisms on the risk of colorectal cancer. Mutagenesis. 2012; 27:219–223.
27. Whiffin N, Broderick P, Lubbe SJ, Pittman AM, Penegar S, Chandler I, Houlston RS. MLH1-93G > A is a risk factor for MSI colorectal cancer. Carcinogenesis. 2011; 32:1157–1161.
28. Chen CC, Yang SY, Liu CJ, Lin CL, Liaw YF, Lin SM, Lee SD, Chen PJ, Chen CJ, Yu MW. Association of cytokine and DNA repair gene polymorphisms with hepatitis B-related hepatocellular carcinoma. Int J Epidemiol. 2005; 34:1310–1318.
29. Fukuhara S, Chang I, Mitsui Y, Chiyomaru T, Yamamura S, Majid S, Saini S, Hirata H, Deng G, Gill A, Wong DK, Shiina H, Nonomura N, et al. DNA mismatch repair gene MLH1 induces apoptosis in prostate cancer cells. Oncotarget. 2014; 5:11297–11307. doi: 10.18632/oncotarget.2315.
30. Ma J, Zhang Y, Shen H, Kapesa L, Liu W, Zeng M, Zeng S. Association between mismatch repair gene and irinotecan-based chemotherapy in metastatic colon cancer. Tumor Biol. 2015; 36:9599–9609.
31. Cini G, Carnevali I, Quaia M, Chiaravalli AM, Sala P, Giacomini E, Maestro R, Tibiletti MG, Viel A. Concomitant mutation and epimutation of the MLH1 gene in a Lynch syndrome family. Carcinogenesis. 2015; 36:452–458.
32. Kim JH, Bae JM, Cho NY, Kang GH. Distinct features between MLH1-methylated and unmethylated colorectal carcinomas with the CpG island methylator phenotype: implications in the serrated neoplasia pathway. Oncotarget. 2016; 7:14095–14111. doi: 10.18632/oncotarget.7374.
33. He Y, Xu X, Chen H, Wang J, Xiong W, Xu Y, Liu J. The hMLH1 promoter polymorphisms and cancer susceptibility in Asian populations: a meta-analysis. Gene. 2013; 523:199–204.
34. Ringelhan M, O’Connor T, Protzer U, Heikenwalder M. The direct and indirect roles of HBV in liver cancer: prospective markers for HCC screening and potential therapeutic targets. J Pathol. 2015; 235:355–367.
35. Tan SK, Qiu XQ, Yu HP, Zeng XY, Xiao ZM, Hu L. [Etiologic fraction and interaction of risk factors for primary hepatic carcinoma in Guangxi, China]. [Article in Chinese]. Zhonghua yu fang yi xue za zh. 2008; 42:169–172.
36. Dong QZ, Zhang XF, Zhao Y, Jia HL, Zhou HJ, Dai C, Sun HJ, Qin Y, Zhang WD, Ren N, Ye QH, Qin LX. Osteopontin promoter polymorphisms at locus -443 significantly affect the metastasis and prognosis of human hepatocellular carcinoma. Hepatology. 2013; 57:1024–1034.
37. Uchino K, Tateishi R, Shiina S, Kanda M, Masuzaki R, Kondo Y, Goto T, Omata M, Yoshida H, Koike K. Hepatocellular carcinoma with extrahepatic metastasis: clinical features and prognostic factors. Cancer. 2011; 117:4475–4483.
38. Wang Z, Qu K, Niu W, Lin T, Xu X, Huang Z, Liu S, Liu S, Chang H, Liu Y, Dong X, Liu C, Zhang Y. Glutathione S-transferase P1 gene rs4147581 polymorphism predicts overall survival of patients with hepatocellular carcinoma: evidence from an enlarged study. Tumor Biol. 2016; 37:943–952.
39. Farinati F, Vitale A, Spolverato G, Pawlik TM, Huo TL, Lee YH, Frigo AC, Giacomin A, Giannini EG, Ciccarese F, Piscaglia F, Rapaccini GL, Di Marco M, et al. Development and Validation of a New Prognostic System for Patients with Hepatocellular Carcinoma. Plos Med. 2016; 13:e1002006.
40. Jin J, Niu X, Zou L, Li L, Li S, Han J, Zhang P, Song J, Xiao F. AFP mRNA level in enriched circulating tumor cells from hepatocellular carcinoma patient blood samples is a pivotal predictive marker for metastasis. Cancer Lett. 2016; 378:33–37.
41. Lee JI, Kim JK, Kim do Y, Ahn SH, Park JY, Kim SU, Kim BK, Han KH, Lee KS. Prognosis of hepatocellular carcinoma patients with extrahepatic metastasis and the controllability of intrahepatic lesions. Clin Exp Metastasis. 2014; 31:475–482.
42. Zhu AX, Chen D, He W, Kanai M, Voi M, Chen LT, Daniele B, Furuse J, Kang YK, Poon RT, Vogel A, Chiang DY. Integrative biomarker analyses indicate etiological variations in hepatocellular carcinoma. J Hepatol. 2016; 65:296–304.

