INTRODUCTION
Acute myeloid leukemia (AML) is a frequently fatal malignant disease of hematopoietic stem and progenitor cells. The molecular and phenotypic characteristics of AML are highly heterogeneous [1]. It can arise from a series of genetic and epigenetic alterations that disrupt the differentiation, proliferation, and survival of myeloid progenitor cells [2]. The incidence of AML peaks in early childhood and late adulthood [3]. Although the survival rate among younger AML patients has improved, the prognosis of older patients is still poor.
Long non-coding RNAs (lncRNAs) are a heterogeneous class of RNAs greater than 200 nucleotides in length [4]. Many studies have indicated lncRNAs are important for cell cycle control [5], survival [6], migration [7], and metabolism [8]. LncRNAs participate in multiple networks that control cellular differentiation and development [9], and alterations in lncRNA expression/regulation have been associated with many diseases including cancer [10]. Recently, more and more studies have shown that lncRNAs are deregulated in a wide variety of cancers [11, 12]. Several studies have assessed the roles of lncRNAs such as ANRIL, lncRNA-P21, MEG3, Dleu2, HOTAIRM1, EGO, and lncRNA-a7 in leukemia. The results highlight the importance of investigating lncRNAs as diagnostic, prognostic, and therapeutic targets [13]. The maternally expressed gene 3 (MEG3) gene, located on chromosome 14q32, encodes a myelocyte-related lncRNA that has been implicated in several human malignancies [14]. However, the function of MEG3 has not been elucidated [15]. MEG3 is involved in both physiological and pathological processes. For example, it participates in signaling cascades involved in cell proliferation and differentiation, survival, and angiogenesis. Dysregulation of MEG3 has been associated with several types of cancer [16]. Previous studies have indicated that loss of MEG3 expression in cancer can result from hypermethylation of the MEG3 promoter as well as the intergenic germline-derived differentially methylated region [16–18]. Intriguingly, hypermethylation of the MEG3 promoter has been observed in approximately 50% of patients with myelodysplastic syndrome (MDS) and AML [19]. These results were confirmed in an independent analysis of 40 AML patient samples [20]. Hypermethylation of the MEG3 promoter was correlated with decreased overall survival and is a prognostic marker in myeloid malignancies [19]. Thus, aberrant methylation of the MEG3 promoter may promote AML progression [19, 21]. However, the mechanisms underlying hypermethylation of the MEG3 promoter in AML are unclear.
Tet methylcytosine dioxygenase 2 (TET2) is a putative tumor suppressor gene located on chromosome 4q24.1 [22]. Mutations in TET2 have been observed in a variety of myeloid disorders [23]. Subsequent sequencing analysis revealed that TET2 mutations are present in approximately 7%−23% of AML patients [24–26] and in 14%−55% of patients with other myeloid malignancies [23, 24, 27]. Reduced TET2 activity and 5-hydroxymethylcytosine (5hmC) levels were observed in AML, MDS, chronic myelomonocytic leukemia (CMML), lymphoid leukemia, and other patients with hematological malignancies. Thus, TET2 inactivation and MEG3 promoter methylation frequently coexist.
Micro RNAs (miRNAs) regulate many cellular processes including cell proliferation, differentiation, development, apoptosis, metabolism, and hematopoiesis [28]. Interestingly, miRNA 22 (miR-22) negatively regulates TET2 expression, which results in a decrease in 5hmC and an increase in the methylation of the promoters of multiple genes. Here, we investigated the relationship between TET2 inactivation and MEG3 promoter methylation in Hainan patients with AML.
RESULTS
Analysis of MEG3, TET2, miR-22-3p, and miR-22-5p expression, and MEG3 promoter methylation
In Table 1 MEG3 expression was significantly reduced in the AML compared to the control group. TET2, miR-22-3p, and miR-22-5p expression was not significant in either group. Analysis of MEG3 promoter methylation revealed no significant differences in 19 CpG units between the AML and control groups: MEG3_1_CpG_1, MEG3_1_CpG_3.4, MEG3_1_CpG_15, MEG3_2_CpG_2, MEG3_2_CpG_6, MEG3_2_CpG_10, MEG3_3_CpG_4, MEG3_3_CpG_5, MEG3_3_CpG_11, MEG3_4_CpG_9, MEG3_5_CpG_5.6, MEG3_5_CpG_10, MEG3_7_CpG_6, MEG3_7_CpG_7, MEG3_7_CpG_12, MEG3_8_CpG_7, MEG3_8_CpG_9, MEG3_8_CpG_11, and MEG3_8_CpG_13 (Figure 1).
Table 1: Analysis of MEG3, TET2, miR-22-3p, and miR-22-5p expression, and MEG3 promoter methylation
Mean |
SD |
P |
OR |
95% CI |
P |
|||
|---|---|---|---|---|---|---|---|---|
MEG3 2-ΔΔCt |
Control |
2.095 |
3.725 |
0.021 |
1.00 |
|||
Case |
0.765 |
1.156 |
3.80 |
0.66 |
21.97 |
0.136 |
||
TET2 2-ΔΔCt |
Control |
1.259 |
0.751 |
0.214 |
1.00 |
|||
Case |
1.069 |
0.783 |
1.75 |
0.40 |
7.71 |
0.459 |
||
miR-22-3p 2-ΔΔCt |
Control |
3.107 |
6.433 |
0.857 |
1.00 |
|||
Case |
4.600 |
17.543 |
2.16 |
0.38 |
12.30 |
0.385 |
||
miR-22-5p 2-ΔΔCt |
Control |
2.064 |
4.495 |
0.857 |
1.00 |
|||
Case |
4.079 |
8.213 |
0.63 |
0.12 |
3.33 |
0.588 |
||
MEG3_1_CpG_1 |
Control |
0.528 |
0.104 |
0.034 |
1.00 |
|||
Case |
0.644 |
0.121 |
7.19 |
0.77 |
66.89 |
0.083 |
||
MEG3_1_CpG_3.4 |
Control |
0.442 |
0.120 |
0.003 |
1.00 |
|||
Case |
0.630 |
0.176 |
3.09 |
0.49 |
19.62 |
0.232 |
||
MEG3_1_CpG_15 |
Control |
0.528 |
0.104 |
0.034 |
1.00 |
|||
Case |
0.644 |
0.121 |
7.19 |
0.77 |
66.89 |
0.083 |
||
MEG3_2_CpG_2 |
Control |
0.231 |
0.094 |
0.005 |
1.00 |
|||
Case |
0.324 |
0.150 |
10.01 |
0.92 |
108.78 |
0.058 |
||
MEG3_2_CpG_6 |
Control |
0.403 |
0.125 |
0.010 |
1.00 |
|||
Case |
0.430 |
0.111 |
6.12E+09 |
0.00 |
. |
0.999 |
||
MEG3_2_CpG_10 |
Control |
0.432 |
0.126 |
0.006 |
1.00 |
|||
Case |
0.492 |
0.123 |
43.54 |
2.25 |
843.64 |
0.013 |
||
MEG3_3_CpG_4 |
Control |
0.771 |
0.108 |
0.044 |
1.00 |
|||
Case |
0.855 |
0.104 |
4.39 |
0.66 |
28.96 |
0.125 |
||
MEG3_3_CpG_5 |
Control |
0.750 |
0.086 |
0.027 |
1.00 |
|||
Case |
0.825 |
0.090 |
0.04 |
1.21 |
202.945 |
0.035 |
||
MEG3_3_CpG_11 |
Control |
0.755 |
0.137 |
0.031 |
1.00 |
|||
Case |
0.847 |
0.083 |
24.24 |
1.26 |
466.05 |
0.035 |
||
MEG3_4_CpG_9 |
Control |
0.329 |
0.202 |
0.049 |
1.00 |
|||
Case |
0.472 |
0.159 |
5.87 |
0.38 |
91.13 |
0.206 |
||
MEG3_5_CpG_5.6 |
Control |
0.468 |
0.173 |
0.024 |
1.00 |
|||
Case |
0.567 |
0.162 |
5.09 |
0.66 |
39.51 |
0.12 |
||
MEG3_5_CpG_10 |
Control |
0.561 |
0.168 |
0.030 |
1.00 |
|||
Case |
0.722 |
0.194 |
2.79 |
0.44 |
17.62 |
0.276 |
||
MEG3_7_CpG_6 |
Control |
0.293 |
0.187 |
0.016 |
1.00 |
|||
Case |
0.495 |
0.264 |
6.72 |
0.77 |
58.91 |
0.086 |
||
MEG3_7_CpG_7 |
Control |
0.420 |
0.230 |
0.021 |
1.00 |
|||
Case |
0.538 |
0.194 |
21.99 |
1.20 |
401.45 |
0.037 |
||
MEG3_7_CpG_12 |
Control |
0.357 |
0.176 |
0.072 |
1.00 |
|||
Case |
0.383 |
0.153 |
40.16 |
2.00 |
805.15 |
0.016 |
||
MEG3_8_CpG_7 |
Control |
0.308 |
0.078 |
0.048 |
1.00 |
|||
Case |
0.378 |
0.112 |
22.52 |
1.07 |
473.11 |
0.045 |
||
MEG3_8_CpG_9 |
Control |
0.292 |
0.030 |
0.011 |
1.00 |
|||
Case |
0.371 |
0.138 |
9.32 |
0.64 |
134.89 |
0.102 |
||
MEG3_8_CpG_11 |
Control |
0.308 |
0.078 |
0.048 |
1.00 |
|||
Case |
0.378 |
0.112 |
22.52 |
1.07 |
473.11 |
0.045 |
||
MEG3_8_CpG_13 |
Control |
0.315 |
0.057 |
0.008 |
1.00 |
|||
Case |
0.409 |
0.151 |
7.80 |
0.58 |
104.79 |
0.121 |
||
P < 0.05 indicates statistical significance; OR: odds ratio; 95% CI: 95% confidence interval.
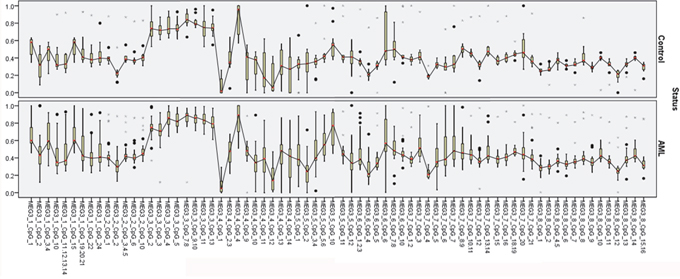
Figure 1: MEG3 expression diagnosis effect analysis.
Analysis of the relationship between MEG3 promoter methylation, and MEG3 and TET2 expression
Spearman’s rank correlation coefficient analysis indicated there was no linear correlation between MEG3 promoter methylation and MEG3 expression. However, a negative correlation between MEG3 promoter methylation and MEG3 expression was observed in the AML group (57 methylation units) (Table 2). Analysis of the relationship between TET2 expression and MEG3 promoter methylation revealed a positive correlation between one CpG unit (MEG3_5_CpG_5.6) and TET2 expression in the control group. A negative correlation between MEG3 promoter methylation (8 CpG units) and TET2 expression was observed in the AML group (Table 3).
Table 2: Spearman’s rank correlation analysis of MEG3 promoter methylation and expression
MEG3 2-ΔΔCt |
Control |
Case |
||
|---|---|---|---|---|
r |
P |
r |
P |
|
MEG3_1_CpG_1 |
0.236 |
0.460 |
-0.632 |
0.002 |
MEG3_1_CpG_2 |
0.049 |
0.880 |
-0.390 |
0.080 |
MEG3_1_CpG_3.4 |
0.275 |
0.388 |
-0.524 |
0.015 |
MEG3_1_CpG_10 |
0.014 |
0.965 |
-0.736 |
0.000 |
MEG3_1_CpG_11.12.13.14 |
-0.057 |
0.861 |
-0.663 |
0.001 |
MEG3_1_CpG_15 |
0.236 |
0.460 |
-0.632 |
0.002 |
MEG3_1_CpG_19.20.21 |
0.056 |
0.862 |
-0.624 |
0.003 |
MEG3_1_CpG_22 |
-0.021 |
0.948 |
-0.339 |
0.133 |
MEG3_1_CpG_24 |
-0.025 |
0.940 |
-0.647 |
0.002 |
MEG3_2_CpG_1 |
0.050 |
0.878 |
-0.492 |
0.028 |
MEG3_2_CpG_2 |
-0.158 |
0.623 |
-0.317 |
0.174 |
MEG3_2_CpG_3.4.5 |
-0.099 |
0.759 |
-0.516 |
0.020 |
MEG3_2_CpG_6 |
0.273 |
0.391 |
-0.070 |
0.769 |
MEG3_2_CpG_10 |
-0.145 |
0.653 |
-0.339 |
0.144 |
MEG3_3_CpG_2 |
0.237 |
0.483 |
-0.306 |
0.217 |
MEG3_3_CpG_3 |
0.032 |
0.926 |
-0.107 |
0.671 |
MEG3_3_CpG_4 |
-0.005 |
0.989 |
-0.149 |
0.555 |
MEG3_3_CpG_5 |
0.146 |
0.668 |
-0.419 |
0.084 |
MEG3_3_CpG_7.8 |
-0.036 |
0.915 |
-0.135 |
0.594 |
MEG3_3_CpG_9.10 |
-0.060 |
0.861 |
-0.020 |
0.938 |
MEG3_3_CpG_11 |
0.224 |
0.508 |
-0.506 |
0.032 |
MEG3_3_CpG_13 |
0.046 |
0.894 |
-0.548 |
0.019 |
MEG3_4_CpG_1 |
-0.125 |
0.685 |
-0.232 |
0.312 |
MEG3_4_CpG_2.3 |
-0.179 |
0.558 |
-0.439 |
0.046 |
MEG3_4_CpG_4 |
0.243 |
0.424 |
0.061 |
0.797 |
MEG3_4_CpG_9 |
-0.081 |
0.802 |
-0.672 |
0.006 |
MEG3_4_CpG_10 |
0.069 |
0.823 |
-0.084 |
0.716 |
MEG3_4_CpG_11 |
-0.144 |
0.656 |
-0.644 |
0.003 |
MEG3_4_CpG_12 |
-0.138 |
0.653 |
-0.369 |
0.100 |
MEG3_4_CpG_13 |
-0.213 |
0.484 |
-0.232 |
0.311 |
MEG3_4_CpG_14 |
-0.124 |
0.685 |
-0.220 |
0.364 |
MEG3_5_CpG_1 |
-0.072 |
0.815 |
-0.491 |
0.024 |
MEG3_5_CpG_2 |
0.637 |
0.026 |
-0.605 |
0.004 |
MEG3_5_CpG_3.4 |
-0.105 |
0.734 |
-0.535 |
0.012 |
MEG3_5_CpG_5.6 |
0.274 |
0.389 |
-0.664 |
0.001 |
MEG3_5_CpG_10 |
0.098 |
0.761 |
-0.492 |
0.032 |
MEG3_5_CpG_11 |
0.290 |
0.336 |
-0.661 |
0.001 |
MEG3_5_CpG_12 |
0.391 |
0.187 |
-0.497 |
0.022 |
MEG3_6_CpG_1.2.3 |
0.198 |
0.538 |
-0.564 |
0.008 |
MEG3_6_CpG_4 |
-0.093 |
0.775 |
-0.609 |
0.003 |
MEG3_6_CpG_5 |
-0.302 |
0.316 |
-0.482 |
0.027 |
MEG3_6_CpG_7.8 |
-0.201 |
0.511 |
-0.662 |
0.001 |
MEG3_7_CpG_3 |
0.274 |
0.415 |
-0.562 |
0.012 |
MEG3_7_CpG_4 |
-0.393 |
0.232 |
-0.592 |
0.008 |
MEG3_7_CpG_5 |
0.156 |
0.648 |
-0.640 |
0.003 |
MEG3_7_CpG_8.9 |
-0.415 |
0.205 |
-0.530 |
0.020 |
MEG3_7_CpG_15 |
-0.111 |
0.746 |
-0.528 |
0.020 |
MEG3_7_CpG_20 |
-0.187 |
0.582 |
-0.486 |
0.035 |
MEG3_8_CpG_1 |
0.290 |
0.387 |
-0.554 |
0.017 |
MEG3_8_CpG_4.5 |
0.602 |
0.038 |
-0.599 |
0.009 |
MEG3_8_CpG_7 |
0.392 |
0.207 |
-0.498 |
0.035 |
MEG3_8_CpG_9 |
0.252 |
0.455 |
-0.734 |
0.001 |
MEG3_8_CpG_10 |
-0.125 |
0.699 |
-0.503 |
0.033 |
MEG3_8_CpG_11 |
0.392 |
0.207 |
-0.498 |
0.035 |
MEG3_8_CpG_13 |
0.004 |
0.991 |
-0.610 |
0.007 |
MEG3_8_CpG_14 |
-0.125 |
0.699 |
-0.503 |
0.033 |
MEG3_8_CpG_15.16 |
0.077 |
0.811 |
-0.476 |
0.046 |
P < 0.05 indicates statistical significance.
Table 3: Spearman’s rank correlation analysis of MEG3 promoter methylation and TET2 expression
TET2 2-ΔΔCt |
Control |
Case |
||
|---|---|---|---|---|
r |
P |
r |
P |
|
MEG3_1_CpG_3.4 |
0.479 |
0.115 |
-0.414 |
0.049 |
MEG3_1_CpG_10 |
0.319 |
0.312 |
-0.459 |
0.028 |
MEG3_1_CpG_11.12.13.14 |
0.371 |
0.235 |
-0.435 |
0.038 |
MEG3_3_CpG_5 |
0.511 |
0.108 |
-0.484 |
0.031 |
MEG3_4_CpG_11 |
0.227 |
0.502 |
-0.448 |
0.042 |
MEG3_5_CpG_5.6 |
0.596 |
0.041 |
-0.458 |
0.032 |
MEG3_5_CpG_11 |
0.392 |
0.208 |
-0.437 |
0.042 |
MEG3_7_CpG_5 |
-0.297 |
0.374 |
-0.486 |
0.025 |
P < 0.05 indicates statistical significance.
We performed multivariable linear regression analysis of the relationship between MEG3 promoter methylation and MEG3 expression in Table 4. After adjusting for sex and age, we identified as association between MEG3 promoter methylation (7 CpG units) and MEG3 expression (P < 0.05). Among the CpG units, linear changes in MEG3 expression were correlated with MEG3_4_CpG_9 (control, B = -21.60, P = 0.01; case, B = -10.56, P < 0.001) and MEG3_5_CpG_2 (control, B = 20.50, P < 0.001; case, B = -6.19, P = 0.02). In Table 5 we also found that six CpG methylation units were correlated with TET2 expression (P < 0.05). There was no significant correlation in the control group but an inverse linear correlation was observed in the case group (B < 0).
Table 4: Multivariable linear regression analysis of MEG3 promoter methylation and expression
MEG3 2-ΔΔCt |
B |
P |
95% CI |
||
|---|---|---|---|---|---|
MEG3_1_CpG_1 |
Control |
3.74 |
0.53 |
-6.40 |
11.49 |
Case |
-8.68 |
0.09 |
-19.17 |
1.81 |
|
MEG3_1_CpG_3.4 |
Control |
3.74 |
0.30 |
-3.96 |
11.44 |
Case |
-6.62 |
0.00 |
-10.28 |
-2.97 |
|
MEG3_4_CpG_9 |
Control |
-21.60 |
0.01 |
-37.04 |
-6.16 |
Case |
-10.56 |
0.00 |
-13.54 |
-7.57 |
|
MEG3_5_CpG_2 |
Control |
20.50 |
0.00 |
12.17 |
28.82 |
Case |
-6.19 |
0.02 |
-11.34 |
-1.05 |
|
MEG3_5_CpG_12 |
Control |
11.27 |
0.18 |
-6.43 |
28.97 |
Case |
-3.97 |
0.05 |
-7.91 |
-0.03 |
|
MEG3_8_CpG_4.5 |
Control |
6.92 |
0.20 |
-4.47 |
18.31 |
Case |
-18.80 |
0.04 |
-36.47 |
-1.14 |
|
MEG3_8_CpG_9 |
Control |
13.78 |
0.15 |
-6.28 |
33.83 |
Case |
-12.68 |
0.02 |
-23.05 |
-2.31 |
|
P < 0.05 indicates statistical significance; 95% CI: 95% confidence interval.
Table 5: Multivariable linear regression analysis of MEG3 promoter methylation and TET2 expression
TET2 2-ΔΔCt |
B |
P |
95% CI |
||
|---|---|---|---|---|---|
MEG3_1_CpG_10 |
Control |
0.040 |
0.593 |
-0.125 |
0.204 |
Case |
-0.152 |
0.040 |
-0.296 |
-0.008 |
|
MEG3_1_CpG_24 |
Control |
0.024 |
0.741 |
-0.136 |
0.184 |
Case |
-0.119 |
0.047 |
-0.236 |
-0.002 |
|
MEG3_2_CpG_2 |
Control |
0.059 |
0.104 |
-0.015 |
0.134 |
Case |
-0.091 |
0.028 |
-0.171 |
-0.012 |
|
MEG3_5_CpG_11 |
Control |
0.024 |
0.351 |
-0.032 |
0.081 |
Case |
-0.057 |
0.001 |
-0.084 |
-0.03 |
|
MEG3_6_CpG_1.2.3 |
Control |
-0.023 |
0.213 |
-0.095 |
0.049 |
Case |
-0.155 |
0.048 |
-0.307 |
-0.002 |
|
MEG3_8_CpG_2 |
Control |
-0.038 |
0.509 |
-0.166 |
0.089 |
Case |
-0.026 |
0.020 |
-0.047 |
-0.005 |
|
P < 0.05 indicates statistical significance; 95% CI: 95% confidence interval.
Analysis of the correlation between TET2 expression and miR-22-3p, miR-22-5p, and MEG3 expression
We did not observe a correlation between miR-22-3p, miR-22-5p, and TET2 expression in either the AML or control group before or after adjustment for age and gender (Table 6). We did observe a positive correlation between TET2 and MEG3 expression in the AML group (Spearman’s rank correlation coefficient, r = 0.634 > 0). However, no significant correlation was detected after adjustment for age and gender. Finally, multivariable linear regression analysis indicated TET2 expression was positively correlated with MEG3 expression in the control group (B = 0.708 > 0).
Table 6: Analysis of the correlation between TET2 expression, and miR-22-3p and miR-22-5p expression
TET2 2-ΔΔCt |
Control |
Case |
||
|---|---|---|---|---|
r |
P |
r |
P |
|
miR-22-3p 2-ΔΔCt |
0.341 |
0.181 |
0.202 |
0.334 |
miR-22-5p 2-ΔΔCt |
0.118 |
0.653 |
0.072 |
0.731 |
P < 0.05 indicates statistical significance.
Analysis of MEG3 expression as a diagnostic test
ROC curve analysis showed MEG3 expression was effective as a diagnostic (area under the curve = 0.713, 95% confidence interval [CI] = 0.554−0.871, P = 0.021) (Figure 2).
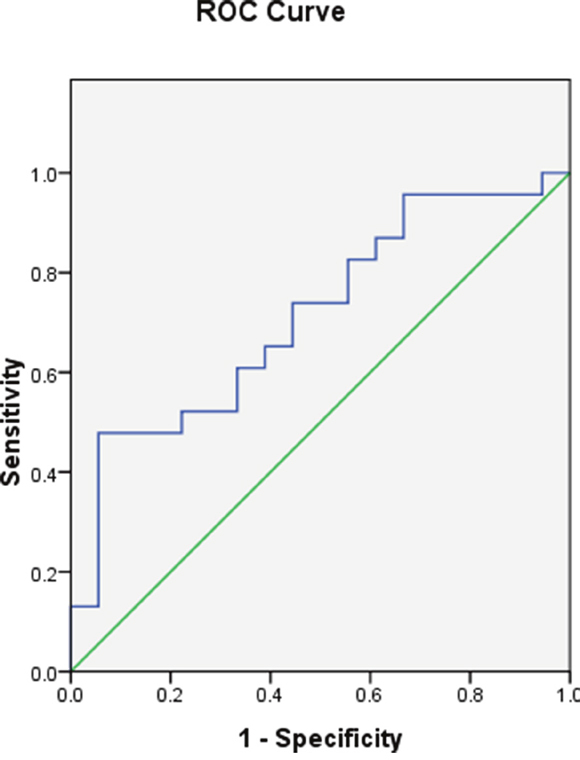
Figure 2: MEG3 promoter methylation in the AML and control groups.
DISCUSSION
Aberrant promoter methylation can result in silencing of gene expression and contribute to the development of leukemia. Changes in DNA methylation state (particularly hypermethylation of tumor suppressor genes) is a diagnostic and prognostic marker in patients with hematological malignancies [29]. Previous studies of the role of DNA methylation in AML have achieved conflicting results. Analysis of epigenetic patterns in AML could enable identification of new patient subgroups and/or provide new prognostic biomarkers. Here, we assessed the relationship between MEG3 promoter methylation and MEG3, TET2, miR-22-3p, and miR-22-5p expression.
MEG3 is a maternally expressed gene on that encodes a lncRNA with a length of 1.6 kb [30, 31]. The functions of MEG3 have not yet been defined. However, it has been implicated in normal physiological processes as well as tumorigenesis [32]. MEG3 promoter methylation was also correlated with reduced overall survival, and could serve as a prognostic marker in myeloid malignancies [15]. Promoter methylation is not always disease-related. It also occurs under normal conditions and is important for the expression of growth factors and their receptors, cytokines, and various other molecules during normal myeloid development [21]. Promoter hypermethylation and aberrant silencing of genes involved in cell adhesion, cell cycle regulation, and tumor suppression has been observed in hematological malignancies such as MDS and AML. These alterations are thought to occurs at approximately the same frequency as mutations [17].
Reduced MEG3 expression has been observed in tumor tissue. For example, MEG3 expression was significantly lower in non-functional pituitary adenoma compared to normal tissue [14, 33]. Reduced MEG3 expression has also been observed in breast, cervical, colon, liver, lung, and prostate cancer cell lines [14, 34]. We observed reduced MEG3 expression in the AML compared to the control group. Because MEG3 enhances the activity of the tumor suppressor P53, down-regulation of MEG3 expression may promote cancer progression. Indeed, down-regulation of MEG3 expression has been observed in approximately 50% of AML patients and is mediated by promoter hypermethylation.
Altered DNA methylation is an important mechanism underlying tumor development and progression [35, 36]. TET2 catalyzes the oxidation of 5-methylcytosine (5mC) to 5hmC, and decreased TET2 activity can result in an altered methylation pattern (e.g. promoter hypermethylation) [37]. TET2 inactivation can cause impaired DNA demethylation and ultimately promote AML development. TET2 inactivation promotes hematological malignancies [38]. The TET enzyme catalyzes the oxidation of 5mC to 5hmC, resulting in active DNA demethylation. The TET family of proteins includes TET1, TET2, and TET3. TET2 inactivation is the most common alteration in hematological malignancies. TET2 activity and 5hmC levels were shown to be reduced in AML, MDS, CMML, lymphocytic leukemia, and other hematological malignancies [38]. Mutations in TET2 and inactivation through methylation have been observed in AML patients, and impact the complete remission rate and disease-free survival. Thus, TET2 inactivation may promote the development and progression of a variety of hematological malignancies including AML [39]. We observed differences in the methylation level of the MEG3 promoter between the AML and control groups. Additionally, we observed a negative correlation between MEG3 promoter methylation and MEG3 expression. Finally, we determined that MEG3 promoter methylation was negatively correlated with TET2 expression.
TET2 expression is negatively regulated by miR-22, which reduces the expression of 5hmC and enhancing the methylation of multiple genes [40]. However, we did not detect an association between miR-22 and TET2 expression in AML. Our data demonstrate inactivation of TET2 and hypermethylation of the MEG3 promoter in AML. We hypothesize that TET2 inactivation causes hypermethylation of the MEG2 promoter based on the negative correlation between TET2 expression and MEG3 promoter methylation. TET2 expression and MEG3 promoter hypermethylation may serve as prognostic markers in AML and lead to new targeted therapeutics.
MATERIALS AND METHODS
Patients and samples
Bone marrow samples were obtained from 29 patients with AML (diagnosed according to the French-American-British criteria [41]) who were treated at the People’s Hospital of Hainan Province between February 2014 and August 2015. The control population consisted of 20 healthy volunteers. The protocol was approved by the People’s Hospital of Hainan Province. Written informed consent was obtained from all patients.
Quantitative real-time PCR
Total RNA was extracted from frozen tissue samples or cells using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s protocol. A total of 1 μg of RNA was reverse transcribed using the TIANScript RT Kit (TIANGEN, Beijing, China). Quantitative real-time PCR was performed using the BIO-RAD iQ5 Real-Time System (BIO-RAD, Hercules, CA, USA) and SYBR Green (TIANGEN) as a double-stranded DNA-specific dye. We performed the cDNA synthesis using a Thermo Script RT-qPCR System (Invitrogen). Target genes were amplified with primers designed using the Primer Premier Version 5.0 software. The following protocol was used for real-time PCR: 95°C for 2 min followed by 40 cycles at 95°C for 15 sec, and then 60°C for 1 min. Standard curves were generated for each assay to produce a linear plot of threshold cycle (Ct) against log (dilution). Target gene expression was quantified using the standard curve method. Data are presented as relative Ct values (n = 6). MEG3 and TET2 expression was normalized to GAPDH, while miR-22-3p and miR-22-5p levels were normalized to U6 snRNA. The relative levels of MEG3, TET2, miR-22-3p, and miR-22-5p were calculated using the 2-ΔΔCt method [(Ct, HOTAIR - Ct, GAPDH, U6) - (Ct, HOTAIR - Ct, GAPDH, U6) control].
DNA extraction and bisulfite modification
DNA was extracted from bone marrow tissue collected into EDTA-containing tubes using a Qiagen DNA Extraction kit. Bisulfite treatment was performed using the EZ DNA Methylation kit (Zymo Research, Irvine, CA, USA) and the manufacturer’s protocol. Quantification of DNA methylation was performed using the Sequenom MassARRAY platform and the EpiTYPER software (Sequenom, San Diego, CA, USA). This platform contained 125 CpG sites. There were 8 CpG units that resulted from cleavage after T, and each unit included single or multiple CpG sites. Using the Mass Cleave assay (Sequenom), we quantitatively assessed the levels of DNA methylation at single CpG units consisting of at least one CpG dinucleotide. Sequenom MassARRAY primers were designed to cover all possible alternative CpG cleavage sites using the Methyl Primer Express software v1.0. Amplicons were designed using the Sequenom Epityper software v.1.0. The PCR conditions were the following: 94°C for 5 min, 94°C for 30 s, 64.6°C for 30 s (annealing), 72°C for 1 min (elongation), and 72°C for 7 min.
Statistical analysis
Statistical analyses were performed using the SPSS 17.0 software (SPSS, Chicago, IL, USA). Demographic and clinical data are reported as the mean, median, or a proportion. The data were analyzed using Student’s t-tests or one-way analysis of variance, and a P value < 0.05 was considered statistically significant. Mann-Whitney U tests were performed using the GraphPad Prism 5 software. Spearman’s correlation was used to assess differences in methylation levels. Finally, receiver operating characteristic (ROC) curves were used to evaluate MEG3 promoter methylation as a diagnostic marker for AML.
ACKNOWLEDGMENTS
We wish to thank all of the patients who participated in this study.
CONFLICTS OF INTEREST
The authors declare that there are no conflicts of interest.
FUNDING
This work was supported by the National Natural Science Foundation of China (No. 81460031).
REFERENCES
1. Link DC. Molecular genetics of AML. Best practice & research Clinical haematology. 2012; 25:409-414.
2. Marchesi V. Genetics: the AML mutational landscape. Nature reviews Clinical oncology. 2013; 10:305.
3. Dohner H, Estey EH, Amadori S, Appelbaum FR, Buchner T, Burnett AK, Dombret H, Fenaux P, Grimwade D, Larson RA, Lo-Coco F, Naoe T, Niederwieser D, Ossenkoppele GJ, Sanz MA, Sierra J, et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood. 2010; 115:453-474.
4. Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, Huarte M, Zuk O, Carey BW, Cassady JP, Cabili MN, Jaenisch R, Mikkelsen TS, Jacks T, Hacohen N, Bernstein BE, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009; 458:223-227.
5. Meola N, Pizzo M, Alfano G, Surace EM, Banfi S. The long noncoding RNA Vax2os1 controls the cell cycle progression of photoreceptor progenitors in the mouse retina. Rna. 2012; 18:111-123.
6. Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD, Horlings HM, Shah N, Umbricht C, Wang P, Wang Y, Kong B, Langerod A, Borresen-Dale AL, Kim SK, van de Vijver M, et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nature genetics. 2011; 43:621-629.
7. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, Li R, West RB, van de Vijver MJ, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464:1071-1076.
8. Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Science signaling. 2010; 3:ra8.
9. Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends in cell biology. 2011; 21:354-361.
10. Hung T, Chang HY. Long noncoding RNA in genome regulation: prospects and mechanisms. RNA biology. 2010; 7:582-585.
11. Gao W, Chan JY, Wong TS. Long non-coding RNA deregulation in tongue squamous cell carcinoma. BioMed research international. 2014; 2014:405860.
12. Quagliata L, Matter MS, Piscuoglio S, Arabi L, Ruiz C, Procino A, Kovac M, Moretti F, Makowska Z, Boldanova T, Andersen JB, Hammerle M, Tornillo L, Heim MH, Diederichs S, Cillo C, et al. Long noncoding RNA HOTTIP/HOXA13 expression is associated with disease progression and predicts outcome in hepatocellular carcinoma patients. Hepatology. 2014; 59:911-923.
13. Heuston EF, Lemon KT, Arceci RJ. The Beginning of the Road for Non-Coding RNAs in Normal Hematopoiesis and Hematologic Malignancies. Frontiers in genetics. 2011; 2:94.
14. Zhang X, Zhou Y, Mehta KR, Danila DC, Scolavino S, Johnson SR, Klibanski A. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. The Journal of clinical endocrinology and metabolism. 2003; 88:5119-5126.
15. Benetatos L, Vartholomatos G, Hatzimichael E. MEG3 imprinted gene contribution in tumorigenesis. International journal of cancer. 2011; 129:773-779.
16. Astuti D, Latif F, Wagner K, Gentle D, Cooper WN, Catchpoole D, Grundy R, Ferguson-Smith AC, Maher ER. Epigenetic alteration at the DLK1-GTL2 imprinted domain in human neoplasia: analysis of neuroblastoma, phaeochromocytoma and Wilms' tumour. British journal of cancer. 2005; 92:1574-1580.
17. Zhao J, Dahle D, Zhou Y, Zhang X, Klibanski A. Hypermethylation of the promoter region is associated with the loss of MEG3 gene expression in human pituitary tumors. The Journal of clinical endocrinology and metabolism. 2005; 90:2179-2186.
18. Pantoja C, de Los Rios L, Matheu A, Antequera F, Serrano M. Inactivation of imprinted genes induced by cellular stress and tumorigenesis. Cancer research. 2005; 65:26-33.
19. Benetatos L, Hatzimichael E, Dasoula A, Dranitsaris G, Tsiara S, Syrrou M, Georgiou I, Bourantas KL. CpG methylation analysis of the MEG3 and SNRPN imprinted genes in acute myeloid leukemia and myelodysplastic syndromes. Leukemia research. 2010; 34:148-153.
20. Khoury H, Suarez-Saiz F, Wu S, Minden MD. An upstream insulator regulates DLK1 imprinting in AML. Blood. 2010; 115:2260-2263.
21. Benetatos L, Dasoula A, Hatzimichael E, Georgiou I, Syrrou M, Bourantas KL. Promoter hypermethylation of the MEG3 (DLK1/MEG3) imprinted gene in multiple myeloma. Clinical lymphoma & myeloma. 2008; 8:171-175.
22. Langemeijer SM, Kuiper RP, Berends M, Knops R, Aslanyan MG, Massop M, Stevens-Linders E, van Hoogen P, van Kessel AG, Raymakers RA, Kamping EJ, Verhoef GE, Verburgh E, Hagemeijer A, Vandenberghe P, de Witte T, et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nature genetics. 2009; 41:838-842.
23. Delhommeau F, Dupont S, Della Valle V, James C, Trannoy S, Masse A, Kosmider O, Le Couedic JP, Robert F, Alberdi A, Lecluse Y, Plo I, Dreyfus FJ, Marzac C, Casadevall N, Lacombe C, et al. Mutation in TET2 in myeloid cancers. The New England journal of medicine. 2009; 360:2289-2301.
24. Abdel-Wahab O, Mullally A, Hedvat C, Garcia-Manero G, Patel J, Wadleigh M, Malinge S, Yao J, Kilpivaara O, Bhat R, Huberman K, Thomas S, Dolgalev I, Heguy A, Paietta E, Le Beau MM, et al. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood. 2009; 114:144-147.
25. Nibourel O, Kosmider O, Cheok M, Boissel N, Renneville A, Philippe N, Dombret H, Dreyfus F, Quesnel B, Geffroy S, Quentin S, Roche-Lestienne C, Cayuela JM, Roumier C, Fenaux P, Vainchenker W, et al. Incidence and prognostic value of TET2 alterations in de novo acute myeloid leukemia achieving complete remission. Blood. 2010; 116:1132-1135.
26. Metzeler KH, Maharry K, Radmacher MD, Mrozek K, Margeson D, Becker H, Curfman J, Holland KB, Schwind S, Whitman SP, Wu YZ, Blum W, Powell BL, Carter TH, Wetzler M, Moore JO, et al. TET2 mutations improve the new European LeukemiaNet risk classification of acute myeloid leukemia: a Cancer and Leukemia Group B study. Journal of clinical oncology. 2011; 29:1373-1381.
27. Kim M, Yahng SA, Kwon A, Park J, Jeon YW, Yoon JH, Shin SH, Lee SE, Cho BS, Eom KS, Lee S, Min CK, Kim HJ, Cho SG, Kim DW, Lee JW, et al. Mutation in TET2 or TP53 predicts poor survival in patients with myelodysplastic syndrome receiving hypomethylating treatment or stem cell transplantation. Bone marrow transplantation. 2015; 50:1132-1134.
28. Gangaraju VK, Lin H. MicroRNAs: key regulators of stem cells. Nature reviews Molecular cell biology. 2009; 10:116-125.
29. Babaei H, Mohammadi M, Salehi R. DNA methylation analysis of secreted frizzled-related protein 2 gene for the early detection of colorectal cancer in fecal DNA. Nigerian medical journal. 2016; 57:242-245.
30. Miyoshi N, Wagatsuma H, Wakana S, Shiroishi T, Nomura M, Aisaka K, Kohda T, Surani MA, Kaneko-Ishino T, Ishino F. Identification of an imprinted gene, Meg3/Gtl2 and its human homologue MEG3, first mapped on mouse distal chromosome 12 and human chromosome 14q. Genes to cells. 2000; 5:211-220.
31. Schuster-Gossler K, Bilinski P, Sado T, Ferguson-Smith A, Gossler A. The mouse Gtl2 gene is differentially expressed during embryonic development, encodes multiple alternatively spliced transcripts, and may act as an RNA. Developmental dynamics. 1998; 212:214-228.
32. Braconi C, Kogure T, Valeri N, Huang N, Nuovo G, Costinean S, Negrini M, Miotto E, Croce CM, Patel T. microRNA-29 can regulate expression of the long non-coding RNA gene MEG3 in hepatocellular cancer. Oncogene. 2011; 30:4750-4756.
33. Cao X, Zhuang S, Hu Y, Xi L, Deng L, Sheng H, Shen W. Associations between polymorphisms of long non-coding RNA MEG3 and risk of colorectal cancer in Chinese. Oncotarget. 2016; 7:19054-19059. doi: 10.18632/oncotarget.7764.
34. Zhang J, Yao T, Wang Y, Yu J, Liu Y, Lin Z. Long noncoding RNA MEG3 is downregulated in cervical cancer and affects cell proliferation and apoptosis by regulating miR-21. Cancer biology & therapy. 2016; 17:104-113.
35. Dawson MA, Kouzarides T. Cancer epigenetics: from mechanism to therapy. Cell. 2012; 150:12-27.
36. Jin B, Li Y, Robertson KD. DNA methylation: superior or subordinate in the epigenetic hierarchy? Genes Cancer. 2011; 2:607-17. doi: 10.1177/1947601910393957.
37. Tahiliani M, Koh KP, Shen Y, Pastor WA, Bandukwala H, Brudno Y, Agarwal S, Iyer LM, Liu DR, Aravind L, Rao A. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science. 2009; 324:930-935.
38. de Oliveira FM, Miguel CE, Lucena-Araujo AR, de Lima AS, Falcao RP, Rego EM. FISH analysis for TET2 deletion in a cohort of 362 Brazilian myeloid malignancies: correlation with karyotype abnormalities. Medical oncology. 2013; 30:483.
39. Zawlik I, Vaccarella S, Kita D, Mittelbronn M, Franceschi S, Ohgaki H. Promoter methylation and polymorphisms of the MGMT gene in glioblastomas: a population-based study. Neuroepidemiology. 2009; 32:21-29.
40. Song SJ, Ito K, Ala U, Kats L, Webster K, Sun SM, Jongen-Lavrencic M, Manova-Todorova K, Teruya-Feldstein J, Avigan DE, Delwel R, Pandolfi PP. The oncogenic microRNA miR-22 targets the TET2 tumor suppressor to promote hematopoietic stem cell self-renewal and transformation. Cell stem cell. 2013; 13:87-101.
41. Bennett JM, Catovsky D, Daniel MT, Flandrin G, Galton DA, Gralnick HR, Sultan C. Proposed revised criteria for the classification of acute myeloid leukemia. A report of the French-American-British Cooperative Group. Annals of Internal Medicine. 1985; 103:620-625.

