INTRODUCTION
Breast cancers with overexpression of the human epidermal growth factor 2 (HER2) receptor tyrosine kinase or amplification of HER2 are defined as HER2-positive [1, 2]. These constitute approximately 15%–20% of all breast cancers, and used to be characterized by an aggressive clinical course with a propensity for distant metastases within 5 years of diagnosis [2, 3]. However, the emergence of trastuzumab as a treatment has dramatically improved the outcomes of patients with all stages of HER2-positive breast cancer [4–7]. The combination of HER2-directed agents, including trastuzumab and pertuzumab, and chemotherapy such as docetaxel and paclitaxel was recently approved for use in preoperative and metastatic HER2-positive breast cancer patients [8, 9]. Moreover, the increased use of neoadjuvant HER2-directed agents with chemotherapy has been documented [10].
Following treatment with HER2-directed agents and chemotherapy, some patients achieve a pathological complete response (pCR), and these patients are known to have very long disease-free survival [10–12]. Currently, pCR can only be established after patients have undergone HER2-directed therapy and surgery, and we cannot predict responders in advance or identify responders without surgery. However, if we could predict responders in advance, this would greatly contribute to patient stratification, or personalized treatment. Therefore, a novel marker needs to be developed to identify HER2-positive breast cancer patients who will show pCR in HER2-directed therapy with a high specificity.
To screen and validate a novel marker, robust genome-wide analysis and a large number of samples with high-quality clinical information are essential. For robust genome-wide analysis, we herein focused on DNA methylation. Compared with gene expression, DNA methylation can be analyzed using DNA, which has several advantages over RNA such as resistance to degradation and easy handling [13–15]. In addition, DNA methylation is stably inherited upon cell division, and is capable of predicting that a gene cannot be expressed even if induced. Further, even in specimens with contaminating cells, the DNA methylation level can be corrected using the ratio of cancer cells to total cells. This ratio can be obtained by microscopic examination of thin sections stained with hematoxylin and eosin (H&E) from paraffin-embedded tissue specimens or using DNA methylation markers [16].
In the present study, we used genome-wide methylation analysis to identify genomic regions whose DNA methylation status was associated with pCR to trastuzumab and chemotherapy. In this way, surgery-free treatment may become possible for such patients.
RESULTS
Genome-wide screening of regions with methylation statuses associated with pCR
To identify genomic regions specifically methylated or unmethylated in the initial biopsy specimens of responders (patients with pCR), genome-wide DNA methylation analysis was conducted using samples in the screening set (Table 1), namely: i) 10 samples from responders (pCR samples), ii) 11 samples from non-responders (non-pCR samples), iii) normal breast epithelial cell lines, and iv) an HER2-positive breast cancer cell line.
Table 1: Clinicopathological characteristics of HER2-positive breast cancer patients in the screening and validation sets
Screening set N = 21 |
Validation set N = 46 |
|||
|---|---|---|---|---|
pCR |
Non-pCR |
pCR |
Non-pCR |
|
No. of patients |
10 |
11 |
12 |
34 |
Clinical tumor stage (%) |
||||
cT1 |
0 (0.0) |
2 (18.2) |
1 (8.3) |
1 (2.9) |
cT2 |
8 (80.0) |
8 (72.7) |
9 (75.0) |
24 (70.6) |
cT3 |
1 (10.0) |
1 (9.1) |
2 (16.7) |
7 (20.6) |
cT4 |
1 (10.0) |
0 (0) |
0 (0.0) |
2 (5.9) |
Tumor size (cm) |
2–6.5 (median: 3.5) |
0–7.0 (median: 3.5) |
1.5–7.0 (median: 3.0) |
1.4–10.0 (median: 3.5) |
Clinical nodal stage (%) |
||||
cN0 |
3 (30.0) |
3 (27.3) |
3 (25.0) |
16 (47.1) |
cN1 |
4 (40.0) |
6 (54.5) |
8 (66.7) |
15 (44.1) |
cN2 |
1 (10.0) |
1 (9.1) |
0 (0.0) |
3 (8.8) |
cN3 |
2 (20.0) |
1 (9.1) |
1 (8.3) |
0 (0.0) |
Clinical stage (%) |
||||
cIIA |
3 (30.0) |
5 (45.5) |
4 (33.3) |
15 (44.1) |
cIIB |
3 (30.0) |
3 (27.3) |
6 (50.0) |
10 (29.4) |
cIIIA |
1 (10.0) |
2 (18.1) |
1 (8.3) |
7 (20.6) |
cIIIB |
1 (10.0) |
0 (0.0) |
0 (0.0) |
2 (5.9) |
cIIIC |
2 (20.0) |
1 (9.1) |
1 (8.3) |
0 (0.0) |
Histology (%) |
||||
Invasive ductal carcinoma |
9 (90.0) |
10 (90.9) |
12 (100.0) |
33 (97.1) |
Invasive lobular carcinoma |
0 (0.0) |
1 (9.1) |
0 (0.0) |
1 (2.9) |
Others |
1 (10.0) |
0 (0.0) |
0 (0.0) |
0 (0.0) |
ERa status |
||||
Positive |
2 (20.0) |
8 (72.7) |
3 (25.0) |
21 (61.8) |
Negative |
8 (80.0) |
3 (27.3) |
9 (75.0) |
13 (38.2) |
PgRb status |
||||
Positive |
3 (30.0) |
6 (54.5) |
1 (8.3) |
16 (47.1) |
Negative |
7 (70.0) |
5 (45.5) |
11 (91.7) |
18 (52.9) |
Ki-67 labelling index (%) |
30–60 (median: 45) |
20–80 (median: 40) |
26.5–80 (median: 60) |
10–96.5 (median: 49.3) |
aEstrogen receptor, bProgesterone receptor.
Among the 482,421 CpG sites, 125,856 were found to be unmethylated (β value < 0.2) in two normal breast epithelial cell lines, two non-cancerous breast tissues, and two samples of peripheral leucocytes (Figure 1). Among these CpG sites, 280 were methylated [corrected β value ≥ 0.5] in four or more of the 10 pCR samples and unmethylated in 10 or more of the 11 non-pCR samples. Conversely, 110 CpG sites were methylated in five or more of the 11 non-pCR samples and unmethylated in nine or more of the 10 pCR samples. Finally, by selecting genomic regions containing three or more consecutive CpG sites that were consistently methylated or unmethylated, we isolated eight candidate genomic regions specifically methylated in the pCR samples (Table 2). No genomic regions were specifically methylated in the non-pCR samples.
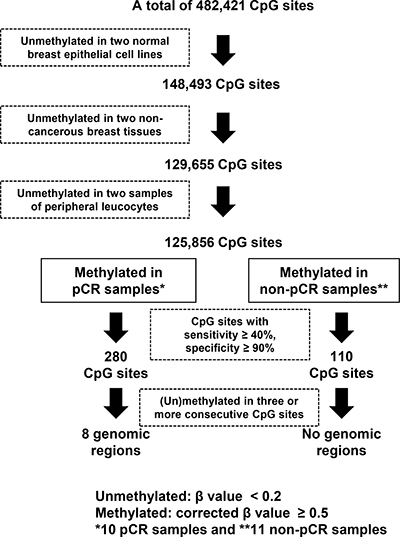
Figure 1: Screening process of genomic regions whose methylation statuses were associated with pCR to trastuzumab and chemotherapy. From the 482,421 CpG sites on the Infinium HumanMethylation450 BeadChip array, those unmethylated in normal breast epithelial cell lines, non-cancerous breast tissues, and peripheral leucocytes were selected. A total of 280 CpG sites were preferentially methylated in responders, and 110 CpG sites were in non-responders. By selecting genomic regions with three or more consecutive differentially methylated CpG sites, eight genomic regions were found to be preferentially methylated in pCR samples. A methylated CpG site was defined when its corrected β value was ≥ 0.5. An unmethylated CpG site was defined when its β value was < 0.2.
Table 2: Candidate genomic regions identified by genome-wide DNA methylation analysis
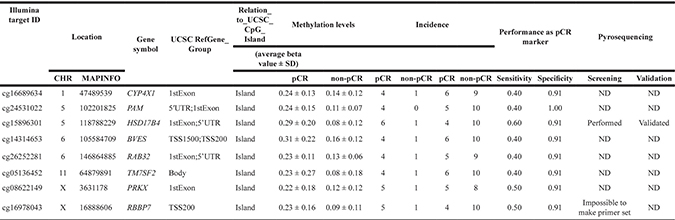
Confirmation of the methylation status of a candidate genomic region
Since a sufficient number of (eight) candidate genomic regions were isolated, we increased the threshold of specificity to 50 %, which three regions (HSD17B4, PRKX, and RBBP7) still met. Further, we focused on methylation changes in 200 bp upstream and downstream of a transcription start site (TSS), which is known to be critical for gene silencing [17], and HSD17B4 and RBBP7 remained as strong candidates. To confirm differential methylation of these two genes, we attempted to design pyrosequencing primers. While it was impossible to design a pyrosequencing primer for RBBP7, we successfully designed a primer for five CpG sites of HSD17B4 (Figure 2A).
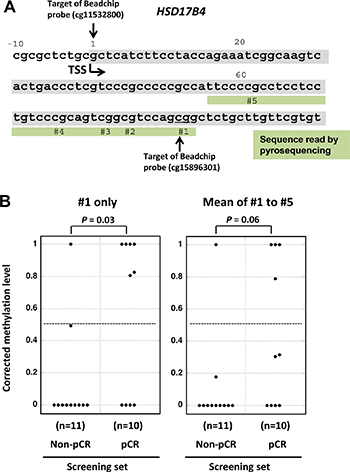
Figure 2: Confirmation of the differential HSD17B4 methylation in the screening set by pyrosequencing. (A) Genomic sequence of HSD17B4 and the genomic region of HSD17B4 analyzed by pyrosequencing are shown. TSS, transcription start site. (B) Corrected methylation levels of the candidate genomic region in the screening set. Methylation levels of the five CpG sites (#1–#5) were measured by pyrosequencing in 10 pCR samples and 11 non-pCR samples, and were corrected by the cancer cell fraction of an individual sample. Corrected methylation levels are shown for CpG #1 (left side) and the mean of five CpG sites (right side). The corrected methylation level of CpG #1 was significantly higher in responders (P = 0.03). The dotted line shows the cutoff value of 0.5 optimized for this screening set and used for validation.
The HSD17B4 methylation levels were measured by pyrosequencing of the 21 samples in the screening set, and, to exclude the effect of contaminating non-cancerous cells in a sample, a corrected methylation level was calculated using the ratio of cancer cells to total cells derived from microscopic observation (cancer cell fraction). The corrected methylation level of one HSD17B4 CpG site was shown to be significantly higher in pCR samples than in non-pCR samples (P = 0.03; Figure 2B, left). The corrected mean methylation level of the five CpG sites tended to be higher in the pCR samples than in the non-pCR samples, but the difference was not significant (P = 0.06; Figure 2B, right). Therefore, we decided to use the CpG position #1 of HSD17B4, and a cutoff value of 0.5 for responders was established so that the Youden index (sensitivity + specificity − 1) would be maximized.
Validation of the association between HSD17B4 methylation and pCR to trastuzumab and chemotherapy
To validate the association between HSD17B4 methylation and pCR to trastuzumab and chemotherapy, the methylation level of CpG site #1 of HSD17B4 was analyzed in an independent sample set (validation set, Table 1), and was then corrected by the cancer cell fraction in a sample. The corrected methylation level of HSD17B4 was found to be significantly higher in pCR samples than in non-pCR samples (P < 0.001; Figure 3). Using the cutoff value of 0.5, which had been prefixed in the screening set, the incidence of HSD17B4 methylation in responders was significantly higher than in non-responders (P = 0.0002). These results established an association between HSD17B4 methylation and the pCR to trastuzumab and chemotherapy. pCR to trastuzumab and chemotherapy was predicted with a sensitivity of 83%, specificity of 79%, and a positive predictive value of 53%.
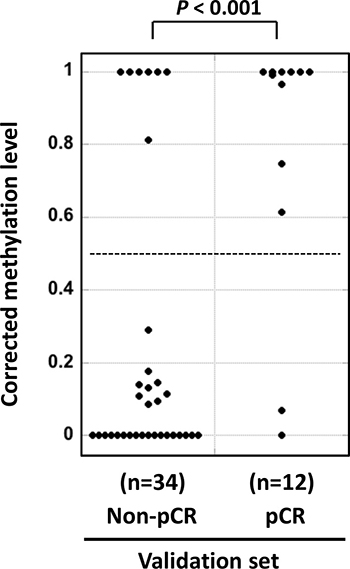
Figure 3: Successful validation of the differential methylation of HSD17B4. Methylation levels of HSD17B4 were measured by pyrosequencing in 12 pCR samples and 34 non-pCR samples in the validation set using the same methodology (pyrosequencing) and threshold established in the screening set. pCR samples had significantly higher corrected methylation levels (P < 0.001), and they were predicted with a sensitivity of 83 % and specificity of 79 %. The horizontal dotted line shows the cutoff value of 0.5, which had been prefixed in the screening set.
Association between HSD17B4 methylation and clinicopathological characteristics
The predictive power of HSD17B4 methylation was examined in comparison with clinicopathological factors. Univariate analyses revealed that ER status and HSD17B4 methylation were significantly associated with pCR to trastuzumab and chemotherapy (Table 3). A multivariate analysis using all clinicopathological factors, including cT, cN, cStage, ER status, PgR status, and Ki-67 labelling index, showed that HSD17B4 methylation was an independent predictive factor for pCR to trastuzumab and chemotherapy (OR 10.0, 95% CI 2.54–39.50, P = 0.001) (Table 4).
Table 3: Association between HSD17B4 methylation and clinicopathological characteristics
Factors |
HSD17B4 |
P valuea |
|
|---|---|---|---|
Methylated |
Unmethylated |
||
Clinical tumor stage |
|||
cT1 |
1 |
3 |
0.76 |
cT2 |
19 |
30 |
|
cT3 |
3 |
8 |
|
cT4 |
1 |
2 |
|
Clinical nodal stage |
|||
cN0 |
9 |
16 |
1 |
cN1 |
12 |
21 |
|
cN2 |
1 |
4 |
|
cN3 |
2 |
2 |
|
Clinical stage |
|||
IIA |
9 |
18 |
1 |
IIB |
9 |
13 |
|
IIIA |
3 |
8 |
|
IIIB |
1 |
2 |
|
IIIC |
2 |
2 |
|
Histology |
|||
Invasive ductal carcinoma |
23 |
41 |
1 |
Others |
1 |
2 |
|
ERb status |
|||
Positive |
7 |
27 |
0.01 |
Negative |
17 |
16 |
|
PgRc status |
|||
Positive |
5 |
21 |
0.04 |
Negative |
19 |
22 |
|
Ki-67 labelling index (%) |
|||
50 ≥ |
15 |
18 |
0.13 |
50 < |
9 |
25 |
|
pCRd |
|||
Positive |
16 |
6 |
2.20E-05 |
Negative |
8 |
37 |
|
aP values were calculated by Fisher’s exact test
bEstrogen receptor,cProgesterone receptor
dPathological complete response
Table 4: Multivariate logistic regression analysis to identify independent factors to determine the response to trastuzumab and chemotherapy
Factors |
No. of patients |
Odds ratio |
95% CI |
P value |
|---|---|---|---|---|
HSD17B4 (methylated vs ummethylated) |
24/43 |
10.02 |
2.54–39.50 |
0.001 |
cT (cT3 + cT4 vs cT1 + cT2) |
14/53 |
0.55 |
0.03–9.32 |
0.681 |
cN (cN1 + cN2 + cN3 vs cN0 ) |
42/25 |
1.79 |
0.38–8.48 |
0.463 |
cStage (III vs II) |
18/49 |
1.28 |
0.09–18.61 |
0.858 |
ER status (negative vs positive) |
33/34 |
6.65 |
0.75–59.32 |
0.090 |
PgR status (negative vs positive) |
41/26 |
0.84 |
0.07–9.61 |
0.890 |
Ki67 labelling index (< 50 vs ≥ 50) |
34/33 |
0.46 |
0.11–1.92 |
0.285 |
To further increase the specificity, we explored combining HSD17B4 methylation with ER status. ER status has been known to be associated with pCR in HER2-positive breast cancer [18], but the above multivariate analysis showed that HSD17B4 methylation provides information independent from the ER status. The independence was confirmed by stratifying the samples into ER+ and ER- tumors, and observing significant associations in both groups (Fisher’s exact test; P = 0.048 for ER+ tumors, and P = 0.005 for ER- tumors) (Supplementary Tables 2 and 3). When a sample with HSD17B4 methylation and negative ER was scored positive, pCR to trastuzumab and chemotherapy was predicted with a sensitivity of 59%, specificity of 91%, and a positive predictive value of 76% (13/17).
DISCUSSION
As a marker to predict pCR to trastuzumab and chemotherapy against HER2-positive breast cancer, HSD17B4 methylation was successfully identified by genome-wide methylation analysis, and was validated by analysis of an independent validation set of samples (P < 0.001). This was achieved by securing the samples for validation in advance, and enabled us to exclude the concern of over-fitting. The sensitivity and specificity in the validation set were 83% and 79%, respectively, after applying the cutoff value established in the screening set. Multivariate analysis showed that HSD17B4 methylation was an independent predictive factor for pCR. These results therefore show that a patient with HSD17B4 methylation is highly probable to achieve pCR with trastuzumab and chemotherapy of a primary HER2-positive breast cancer.
When combined with negative ER status, a high specificity of 91% was achieved with a reasonable sensitivity of 59 %. This shows that HER2-positive/ER-negative breast cancer patients with HSD17B4 methylation are highly likely to achieve pCR. Since pCR is associated with excellent disease-free survival in HER2-positive breast cancer patients with HER2-directed therapy [11, 12], this group of patients might not need surgery. Considering this potential application to surgery-free treatment and serious results due to false-positive diagnosis, we placed emphasis on high specificity, rather than high sensitivity. In addition, to further improve the specificity, we are trying to introduce a molecular, not pathological, marker for cancer cell fractions. The use of such a marker will enable us to assess HSD17B4 methylation levels and cancer cell fractions in the same DNA samples, and accuracy of methylation levels in cancer cells will increase. Even after further improvement of specificity, the usefulness of HSD17B4 methylation should be confirmed by a prospective clinical study with a sufficiently large sample size, and the study must be designed with minimum risk for the participants.
Methylation of the HSD17B4 region, located immediately downstream of the transcription start site and within a CpG island, was associated with its silencing (Supplementary Figure 1), which was in line with a previous study that showed HSD17B4 can be methylation-silenced [19]. However, it remains an issue how HSD17B4 silencing is associated with pCR of HER2-positive breast cancer patients after trastuzumab and chemotherapy. HSD17B4 encodes type 4 17β-hydroxysteroid dehydrogenase. First, this enzyme is involved in the conversion of active 17β-estradiol into inactive estrone [20]. HSD17B4 is specifically down-regulated in breast and ovarian cancers among various human cancers [21], and this suggests that its silencing is associated with sex hormone metabolism and then sensitivity to trastuzumab. Alternatively, the enzyme is involved in β-oxidation of fatty acids and alcohols [22]. In addition, HSD17B4 overexpression has been reported to be associated with aggressive phenotypes in liver and prostate cancers [23, 24]. This suggests that HSD17B4 silencing is associated with sensitivity to trastuzumab via accumulation of a specific fatty acid or other metabolite and might not be related to estrogen metabolism. Further extensive analyses are necessary to clarify the underlying molecular mechanism.
In addition to the potential application to surgery-free treatment of HER2-positive breast cancer patients, unmethylated HSD17B4 may be used to identify patients who will not benefit from trastuzumab treatment but might benefit from lapatinib. This drug is currently used only as a secondary treatment for breast cancer that has progressed after treatment with trastuzumab [25]. Naturally, again, studies with a sufficiently large number of samples from patients who responded to lapatinib and those who did not is essential to explore this important possibility.
Methodologically, CpG site #1 showed better discrimination than the mean of CpG sites #1 to #5. It is known that CpG sites closer to a sequencing primer yield more accurate values than those further away [26]. The accurate measurement of the methylation level of CpG site #1, which is closest to the pyrosequencing primer (Figure 2A), was considered to be the major reason for its good performance. To support this interpretation, the neighboring probe of the bead array (cg11532800) showed a statistically significant difference in the methylation level between the responders and non-responders, as shown in Supplementary Table 1.
In summary, we identified that methylation of the promoter CpG island of HSD17B4 was associated with the pCR of HER2-positive breast cancer to trastuzumab and chemotherapy with a specificity of 79%. When combined with negative ER status, HSD17B4 methylation had a high specificity of 91 %. This high specificity may pave the way for selecting HER2-positive breast cancer patients who would achieve pCR only by HER2-directed therapy and might not need additional surgery, namely surgery-free treatment of breast cancer.
MATERIALS AND METHODS
Tissue samples
A total of 67 tissue samples were collected from the patients enrolled in a neoadjuvant clinical trial, which will be reported elsewhere (Table 1). All the patients in this study were treated with trastuzumab and chemotherapy involving paclitaxel or anthracycline, according to the Japanese guideline of breast cancer treatment [27]. Briefly, the trial aims are: i) to explore a predictive marker of response against HER2-directed therapy using trastuzumab and chemotherapy involving paclitaxel or anthracycline, and ii) to determine whether the clinical response can be optimized by assessing a change in the Ki-67 index during neoadjuvant therapy. For the latter aim, the trial had two arms, but all samples for this study were obtained from patients who received weekly paclitaxel (80 mg/m2 a week) and trastuzumab (a loading dose of 4 mg/kg followed by 2 g/kg a week) for a total of 12 doses. The patients also underwent partial or simple mastectomy, and the response was pathologically assessed. The study protocol was approved by the National Cancer Center Ethics Committee (approval no. 2010–250), and was registered at the UMIN Clinical Trial Registry (Registration no. UMIN000007074) [28]. All patients provided written informed consent.
Two specimens were obtained from each patient by core needle biopsy of a primary tumor before starting neoadjuvant therapy, and fixed using two different methods. One specimen was fixed with 10% neutral buffered formalin for microscopic examination using thin sections stained with H&E, while the other was fixed using the PAXgene Tissue System (Qiagen, Hilden, Germany) and embedded in low-melting paraffin for DNA/RNA extraction using 10 slices of 10-μm sections. A certificated and experienced pathologist (S. F.) conducted the microscopic examination of biopsy specimens to determine the fraction of cancer cells and to select specimens containing a sufficient number of tumor cells for molecular analyses. The pathologist also analyzed surgical specimens to determine the therapeutic response. pCR was defined as no residual tumor cells in surgical specimens.
The 67 HER2-positive breast cancer samples were histologically classified as 64 invasive ductal carcinomas, two invasive lobular carcinomas, and one other histological type according to WHO classification. Thirty-four (50.7%) breast cancer samples were positive for estrogen receptor (ER), and 26 (38.8%) were positive for progesterone receptor (PgR). The samples were divided into a screening set (n = 21) and a validation set (n = 46). Only samples with a tumor cell fraction of 40% or more were used.
Genome-wide DNA methylation analysis
Genome-wide DNA methylation analysis was conducted using an Infinium HumanMethylation450 BeadChip array (Illumina, San Diego, CA), which assessed the degree of methylation of 485,512 probes (482,421 CpG sites and 3,091 non-CpG sites). The methylation level of each probe was obtained as a β value, which ranged from 0 (completely unmethylated) to 1 (completely methylated). Intra-array normalization was performed to exclude probe design biases using the peak-based correction method Beta Mixture Quantile dilation [29]. A corrected β value was obtained from the normalized β value based on the fraction of cancer cells in the sample [corrected β value = 100 × (normalized β value) / (fraction of cancer cells in the sample {%})].
Bisulfite pyrosequencing
Sodium bisulfite modification was performed using 1 μg of genomic DNA as previously described [30]. The modified DNA was suspended in 40 μl of Tris–EDTA buffer, and an aliquot of 2 μl was used for bisulfite pyrosequencing. All pyrosequencing primers were obtained as PyroMark CpG Assays for methylation array validation (Qiagen, Valencia, CA). PCR products labelled with biotin were annealed to 0.2 μM pyrosequencing primers, and pyrosequencing was carried out using the PSQ 96 Pyrosequencing System (Qiagen). Methylation levels were obtained using PSQ Assay Design software (Qiagen).
The measured methylation level was corrected according to the fraction of cancer cells in a sample [corrected methylation level = 100 × (measured methylation level) / (fraction of cancer cells in the sample {%})]. The fraction of cancer cells (%) was obtained by microscopic observation.
Statistical analysis
Fisher’s exact test was used to evaluate the significant difference in relative frequency between two groups. Differences in normalized methylation levels between the responders and non-responders were evaluated by the Mann–Whitney U test. In the univariate analysis, odds ratios (ORs) and 95% confidence intervals (95% CIs) were calculated. Multivariate logistic regression analyses and other statistical analyses were conducted using PASW statistics version 18.0.0 (SPSS Japan Inc., Tokyo, Japan).
Abbreviations
HER2: human epidermal growth factor 2; pCR: pathological complete response; HSD17B4: type 4 17β-hydroxysteroid dehydrogenase; H&E: hematoxylin and eosin; OR: odds ratio; CI: confidence interval; ER: estrogen receptor; PgR: progesterone receptor.
ACKNOWLEDGMENTS
We wish to acknowledge the assistance of Ms. Yuka Nakamura for making paraffin-embedded thin sections for DNA methylation analysis.
CONFLICTS OF INTEREST
The authors have no potential conflicts of interest related to this work.
GRANT SUPPORT
The research described in this report has been funded by the Practical Research for Innovative Cancer Control (grant number; 15ck0106045h0002) from Japan Agency for Medical Research and development, AMED.
REFERENCES
1. Singh JC, Jhaveri K, Esteva FJ. HER2-positive advanced breast cancer: optimizing patient outcomes and opportunities for drug development. Br J Cancer. 2014; 111:1888–1898.
2. Slamon DJ, Clark GM, Wong SG, Levin WJ, Ullrich A, McGuire WL. Human breast cancer: correlation of relapse and survival with amplification of the HER-2/neu oncogene. Science. 1987; 235:177–182.
3. Slamon DJ, Godolphin W, Jones LA, Holt JA, Wong SG, Keith DE, Levin WJ, Stuart SG, Udove J, Ullrich A, Press MF. Studies of the HER-2/neu proto-oncogene in human breast and ovarian cancer. Science. 1989; 244:707–712.
4. Piccart-Gebhart MJ, Procter M, Leyland-Jones B, Goldhirsch A, Untch M, Smith I, Gianni L, Baselga J, Bell R, Jackisch C, Cameron D, Dowsett M, Barrios CH, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005; 353:1659–1672.
5. Romond EH, Perez EA, Bryant J, Suman VJ, Geyer CE Jr, Davidson NE, Tan-Chiu E, Martino S, Paik S, Kaufman PA, Swain SM, Pisansky TM, Fehrenbacher L, et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med. 2005; 353:1673–1684.
6. Slamon D, Eiermann W, Robert N, Pienkowski T, Martin M, Press M, Mackey J, Glaspy J, Chan A, Pawlicki M, Pinter T, Valero V, Liu MC, et al. Adjuvant trastuzumab in HER2-positive breast cancer. N Engl J Med. 2011; 365:1273–1283.
7. Slamon DJ, Leyland-Jones B, Shak S, Fuchs H, Paton V, Bajamonde A, Fleming T, Eiermann W, Wolter J, Pegram M, Baselga J, Norton L. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N Engl J Med. 2001; 344:783–792.
8. Baselga J, Cortes J, Kim SB, Im SA, Hegg R, Im YH, Roman L, Pedrini JL, Pienkowski T, Knott A, Clark E, Benyunes MC, Ross G, et al. Pertuzumab plus trastuzumab plus docetaxel for metastatic breast cancer. N Engl J Med. 2012; 366:109–119.
9. Gianni L, Pienkowski T, Im YH, Roman L, Tseng LM, Liu MC, Lluch A, Staroslawska E, de la Haba-Rodriguez J, Im SA, Pedrini JL, Poirier B, Morandi P, et al. Efficacy and safety of neoadjuvant pertuzumab and trastuzumab in women with locally advanced, inflammatory, or early HER2-positive breast cancer (NeoSphere): a randomised multicentre, open-label, phase 2 trial. Lancet Oncol. 2012; 13:25–32.
10. Dent S, Oyan B, Honig A, Mano M, Howell S. HER2-targeted therapy in breast cancer: a systematic review of neoadjuvant trials. Cancer Treat Rev. 2013; 39:622–631.
11. Gianni L, Eiermann W, Semiglazov V, Manikhas A, Lluch A, Tjulandin S, Zambetti M, Vazquez F, Byakhow M, Lichinitser M, Climent MA, Ciruelos E, Ojeda B, et al. Neoadjuvant chemotherapy with trastuzumab followed by adjuvant trastuzumab versus neoadjuvant chemotherapy alone, in patients with HER2-positive locally advanced breast cancer (the NOAH trial): a randomised controlled superiority trial with a parallel HER2-negative cohort. Lancet. 2010; 375:377–384.
12. Gianni L, Pienkowski T, Im YH, Tseng LM, Liu MC, Lluch A, Staroslawska E, de la Haba-Rodriguez J, Im SA, Pedrini JL, Poirier B, Morandi P, Semiglazov V, et al. 5-year analysis of neoadjuvant pertuzumab and trastuzumab in patients with locally advanced, inflammatory, or early-stage HER2-positive breast cancer (NeoSphere): a multicentre, open-label, phase 2 randomised trial. Lancet Oncol. 2016; 17:791–800.
13. Harada T, Yamamoto E, Yamano HO, Nojima M, Maruyama R, Kumegawa K, Ashida M, Yoshikawa K, Kimura T, Harada E, Takagi R, Tanaka Y, Aoki H, et al. Analysis of DNA methylation in bowel lavage fluid for detection of colorectal cancer. Cancer Prev Res (Phila). 2014; 7:1002–1010.
14. Hoque MO, Prencipe M, Poeta ML, Barbano R, Valori VM, Copetti M, Gallo AP, Brait M, Maiello E, Apicella A, Rossiello R, Zito F, Stefania T, et al. Changes in CpG islands promoter methylation patterns during ductal breast carcinoma progression. Cancer Epidemiol Biomarkers Prev. 2009; 18:2694–2700.
15. Yu F, Jiao Y, Zhu Y, Wang Y, Zhu J, Cui X, Liu Y, He Y, Park EY, Zhang H, Lv X, Ma K, Su F, et al. MicroRNA 34c gene down-regulation via DNA methylation promotes self-renewal and epithelial-mesenchymal transition in breast tumor-initiating cells. J Biol Chem. 2012; 287:465–473.
16. Yamaguchi T, Mukai H, Yamashita S, Fujii S, Ushijima T. Comprehensive DNA Methylation and Extensive Mutation Analyses of HER2-Positive Breast Cancer. Oncology. 2015; 88:377–384.
17. Lin JC, Jeong S, Liang G, Takai D, Fatemi M, Tsai YC, Egger G, Gal-Yam EN, Jones PA. Role of nucleosomal occupancy in the epigenetic silencing of the MLH1 CpG island. Cancer Cell. 2007; 12:432–444.
18. Bhargava R, Dabbs DJ, Beriwal S, Yildiz IA, Badve P, Soran A, Johnson RR, Brufsky AM, Lembersky BC, McGuire KP, Ahrendt GM. Semiquantitative hormone receptor level influences response to trastuzumab-containing neoadjuvant chemotherapy in HER2-positive breast cancer. Mod Pathol. 2011; 24:367–374.
19. Fiegl H, Millinger S, Goebel G, Muller-Holzner E, Marth C, Laird PW, Widschwendter M. Breast cancer DNA methylation profiles in cancer cells and tumor stroma: association with HER-2/neu status in primary breast cancer. Cancer Res. 2006; 66:29–33.
20. Labrie F, Luu-The V, Lin SX, Labrie C, Simard J, Breton R, Belanger A. The key role of 17 beta-hydroxysteroid dehydrogenases in sex steroid biology. Steroids. 1997; 62:148–158.
21. Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, Wilson CJ, Lehar J, Kryukov GV, Sonkin D, Reddy A, Liu M, Murray L, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012; 483:603–607.
22. Adamski J, Jakob FJ. A guide to 17beta-hydroxysteroid dehydrogenases. Mol Cell Endocrinol. 2001; 171:1–4.
23. Rasiah KK, Gardiner-Garden M, Padilla EJ, Moller G, Kench JG, Alles MC, Eggleton SA, Stricker PD, Adamski J, Sutherland RL, Henshall SM, Hayes VM. HSD17B4 overexpression, an independent biomarker of poor patient outcome in prostate cancer. Mol Cell Endocrinol. 2009; 301:89–96.
24. Lu X, Ma P, Shi Y, Yao M, Hou L, Zhang P, Jiang L. NF-kappaB increased expression of 17beta-hydroxysteroid dehydrogenase 4 promotes HepG2 proliferation via inactivating estradiol. Mol Cell Endocrinol. 2015; 401:1–11.
25. Geyer CE, Forster J, Lindquist D, Chan S, Romieu CG, Pienkowski T, Jagiello-Gruszfeld A, Crown J, Chan A, Kaufman B, Skarlos D, Campone M, Davidson N, et al. Lapatinib plus capecitabine for HER2-positive advanced breast cancer. N Engl J Med. 2006; 355:2733–2743.
26. Tost J, Gut IG. DNA methylation analysis by pyrosequencing. Nat Protoc. 2007; 2:2265–2275.
27. Aihara T, Toyama T, Takahashi M, Yamamoto Y, Hara F, Akabane H, Fujisawa T, Ishikawa T, Nagai S, Nakamura R, Tsurutani J, Ito Y, Mukai H. The Japanese Breast Cancer Society Clinical Practice Guideline for systemic treatment of breast cancer, 2015 edition. Breast Cancer. 2016; 23:329–342.
28. Yamaguchi T, Mukai H. Ki-67 index guided selection of preoperative chemotherapy for HER2-positive breast cancer: a randomized phase II trial. Jpn J Clin Oncol. 2012; 42:1211–1214.
29. Teschendorff AE, Marabita F, Lechner M, Bartlett T, Tegner J, Gomez-Cabrero D, Beck S. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics. 2013; 29:189–196.
30. Yoshida T, Yamashita S, Takamura-Enya T, Niwa T, Ando T, Enomoto S, Maekita T, Nakazawa K, Tatematsu M, Ichinose M, Ushijima T. Alu and Satα hypomethylation in Helicobacter pylori-infected gastric mucosae. Int J Cancer. 2011; 128:33–39.

