INTRODUCTION
There is firm evidence that human papillomavirus (HPV) is implicated in the pathogenesis of a subset of patients with head and neck squamous cell carcinoma (HNSCC) who are typically younger and without the traditional risk factors of smoking and alcohol consumption [1–4]. Most of these HPV-associated HNSCCs are found in the oropharyngeal subsite of tonsil and base of tongue [4].
There are approximately 18 oncogenic HPV genotypes which have been implicated in the causation of cancers in humans [5, 6]. The World Health Organization has categorized HPV subtypes into high-risk (HR) and low-risk (LR) group based on their oncogenic potential in causing cervical cancer. In HNSCC, HPV16 is the predominant genotype which has been associated with up to 90% of HPV-associated oropharyngeal squamous cell carcinoma (OPSCC) [7–11]. This data is however predominantly derived from Caucasian populations and the prevalence of HPV in HNSCC among Asian patients is not well characterized.
This study aims to determine the prevalence and characterize the HR-HPV genotypes in HNSCC in Singapore, which constitutes a multiethnic, urban South-East Asian society; and to explore the prognostic implication of HPV detection in patients with HNSCC in this population.
RESULTS
HPV status and HPV genotyping
159 HNSCC patients with sufficient archival tumor specimen were accrual for HPV detection. The distribution of HPV subtypes according to the head and neck subsites is summarized in Table 1 and the representative figure of the HPV-DNA genotypes detection is showed in Figure 1. One hundred forty-four patients (90.6%) were tested positive for HR-HPV using the DNA-PCR based assay. Among the 18 HR-HPV genotypes tested, six HPV genotypes (HPV16, 18, 31, 45, 56 and 68) were identified. HPV45 was the commonest (87.4%), followed by HPV31 (79.2%). HPV16 and HPV18 were found in 5.0% and 15.1% of HNSCC respectively. When we analyzed the specific HR-HPV genotypes according to sites of cancer, only HPV16 was predominantly detected in OPSCC than non-OPSCC sites (p < 0.001). Additionally, multiple co-existing HPV genotypes (2 and more HPV genotypes) were detected in 79.9% of the tumor specimen and they were equally detected across all subsites. In OPSCC, 74.2% of tumor specimens demonstrated 2 or more HPV genotypes and only 19.4% had only one HPV strain detected (4 HPV16-positive, 2 HPV45-positive).
Table 1: Summary of HPV genotypic detection and p16 IHC in our HNSCC cohort
Sites of cancer (N = 159) |
HPV (%) |
HPV 16 |
HPV 18 |
HPV 31 |
HPV 45 |
HPV 56 |
HPV 68 |
P16 |
|---|---|---|---|---|---|---|---|---|
Oral cavity/% (n = 52) |
48 (92.3) |
0 |
9 (17.3) |
44 (84.6) |
48 (92.3) |
6 (11.5) |
26 (50) |
2 (3.8) |
Oropharynx/% (n = 31) |
29 (93.5) |
6 (19.4) |
4 (12.9) |
22 (71.0) |
25 (80.6) |
6 (19.4) |
11 (35.5) |
14 (45.2) |
Larynx/% (n = 52) |
46 (88.5) |
2 (3.8) |
8 (15.4) |
41 (78.8) |
45 (86.5) |
9 (17.3) |
27 (51.9) |
1 (1.9) |
Hypopharynx/% (n = 13) |
11 (84.6) |
0 |
2 (15.4) |
9 (69.2) |
11 (84.6) |
1 (7.7) |
6 (46.2) |
0 |
Others/% (n =11) |
10 (90.9) |
0 |
1 (9.1) |
10 (90.9) |
10 (90.9) |
2 (18.2) |
8 (72.7) |
2 (18.2) |
144 (90.6) |
8 (5.0) |
24 (15.1) |
126 (79.2) |
139 (87.4) |
24 (15.1) |
78 (49.1) |
19 (11.9) |
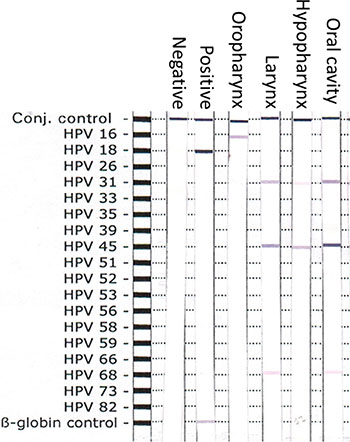

Figure 1: Representative plot of the nitrocellulose strip with HPV genotype-specific probes. Positive control, negative control, Oropharynx SCC, Laryngeal SCC, Hypopharynx SCC, Oral cavity SCC and human β-globin controls were shown.
P16 as a surrogate marker of HPV in comparison to detection of HPV genotyping
There was strong concordance of p16 IHC detection with the presence of HR-HPV detected using the PCR assay across all sites of HNSCC (94.7%) (Table 2). Representative plot of p16-positive and p16-negative tumor is illustrated in Figure 2. Using p16 IHC as a surrogate marker for HR-HPV detection, 12.2% (19 out of 156 cases) of the HNSCC specimen were tested positive with the majority detected in the oropharynx (14 out of 19 cases; 73.7%; p = 0.005). Of the remaining 5 cases with p16-positive tumor, 2 were detected in oral tongue SCC; 1 from laryngeal SCC and the remaining 2 cases were from primary sino-nasal SCC. All p16 IHC-positive cases were HPV-positive except for one patient with OPSCC who was HPV PCR-negative. Using p16 IHC as a reference test for HPV detection, the sensitivity and specificity of HPV-DNA PCR detection were 96%, and 11%, and 93% and 5.9% for HNSCC and OPSCC, respectively.
Table 2: Distribution of P16 and HPV positive tumors according to sites of HNSCC
Sites of cancer |
p16+ve |
HPV+ve/p16 +ve |
HPV-ve/p16+ve |
|---|---|---|---|
Oral cavity (n = 52) |
2 (3.8) |
2 (100) |
0 |
Oropharyngeal (n = 31) |
14 (45.2) |
13 (92.9) |
1 (7.1) |
Laryngeal (n = 50) |
1 (2.0) |
1 (100) |
0 |
Hypopharyngeal (n = 12) |
0 |
0 |
0 |
Others (Sinonasal) (n = 11) |
2 (18.2) |
2 (100) |
0 |
Total =156 |
19 (12.2) |
18 (94.7) |
1 (5.3) |
*3 patients did not have p16 IHC performed (1 with hypopharynx SCC and 2 with laryngeal SCC).
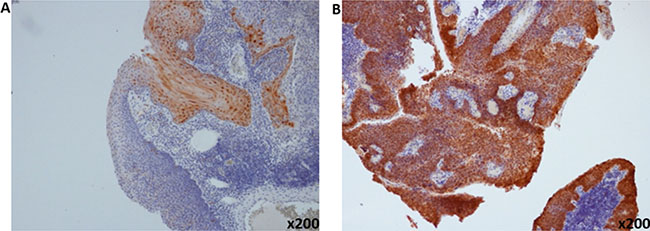
Figure 2: Representative pictures of immunohistochemical staining of p16. (A) P16-negative (B) and p16-positive oropharyngeal squamous cell carcinoma (magnification ×200).
Clinical outcomes in p16 positive and p16 negative OPSCC
Since p16 IHC was predominantly identified in our OPSCC cohort, comparison of clinical features between p16-positive and p16-negative OPSCC patients was performed. There were no statistical differences in age, gender, T- or N classification and overall AJCC stage among patients with p16-positive or p16-negative OPSCC (Table 3).
Table 3: Summary of cohort characteristics in patients with oropharyngeal squamous cell carcinoma
P16+ OPSCC |
P16– OPSCC |
P value (Fisher’s test) |
|
|---|---|---|---|
Age Median = 60.7 |
0.16 |
||
< 20 |
0 |
0 |
|
20–40 |
1 |
1 |
|
40–60 |
7 |
5 |
|
> 60 |
6 |
11 |
|
Age > 60 |
6 |
12 |
|
Age < 60 |
8 |
5 |
|
Gender, |
0.44 |
||
Male |
8 |
13 |
|
Female |
6 |
4 |
|
Smoking, |
0.06 |
||
Yes |
5 |
12 |
|
No |
9 |
4 |
|
Not available (n =1) |
|||
Alcohol |
0.42 |
||
Yes |
3 |
6 |
|
No |
11 |
8 |
|
Not available (n =3) |
|||
Ethnic |
0.41 |
||
Chinese |
12 |
12 |
|
Others |
2 |
5 |
|
T stage |
1.00 |
||
T1 |
2 |
2 |
|
T2 |
8 |
6 |
|
T3 |
1 |
0 |
|
T4 |
3 |
9 |
|
N stage |
0.70 |
||
N0 |
3 |
5 |
|
N1 |
4 |
2 |
|
N2 |
4 |
8 |
|
N3 |
3 |
2 |
|
Definitive treatment |
0.48 |
||
Surgery +/– adjuvant therapy |
7 |
11 |
|
Radiation/ChemoRT |
7 |
6 |
|
Median follow up |
55.8 month |
19.1 month |
|
Overall survival (OS) N3-year OS |
93% |
59% |
0.022 (0.21, 0.05–0.84) |
P16-positive OPSCC patients showed significantly better overall survival (OS) compared to p16-negative patients (92.9% versus 58.8%) (log rank test: p = 0.022, Hazard Ratio = 0.21; 95% Confidence Interval (CI) = 0.05–0.84) (Figure 3). Additionally, patients who are non-smokers and do not consume alcohol demonstrated demonstrated better OS compared to patients with smoking history or who consume alcohol (p = 0.007; p = 0.01) (Figure 4 and Figure 5).
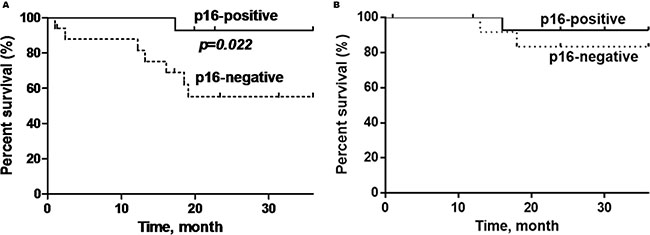
Figure 3: Kaplan-Meier curves showing (A) overall survival and; (B) disease-specific survival of p16-positive and p16-negative patients with oropharyngeal SCC. Black line = p16-positive, dashed line= p16-negative. The p16-positive OPSCC has a significantly better overall survival (p = 0.022), but not disease-specific survival (p = 0.46).
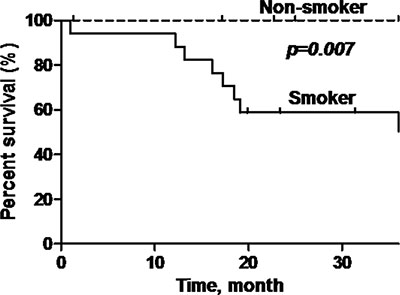
Figure 4: Overall survival of oropharyngeal SCC patients stratified according to smoking status. Black line = smoker, dashed line = non-smoker. Non-smokers have a significantly better overall survival (p = 0.007) compared to smokers.
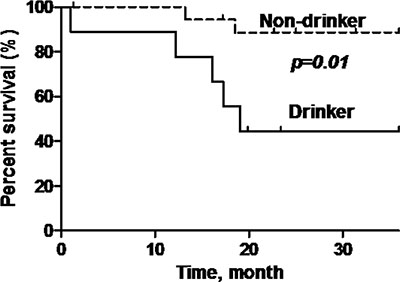
Figure 5: Overall survival of oropharyngeal SCC patients stratified according to alcohol consumption status. Black line = drinker, dashed line = non-drinker. Non-drinker has a significantly better overall survival (p = 0.01) compared to drinker.
DISCUSSION
In this study, we have chosen the digene HPV genotyping RH test kit because it detects a panel of HR-HPV genotypes which is associated with cervical cancer [5, 6, 12]. This RH test kit is designed for routine clinical use and supersedes the previous FDA-approved HR-hybrid capture 2 (HC2) detection system which does not identify specific HR-HPV genotypes. The RH test kit utilizes the HR-HPVGP5+/6+ consensus primer which targets the L1 region of the HPV genome. Using this technique, it has demonstrated high concordance with the previously established HR-hybrid HC2 system [13]. Therefore, this RH test offers the opportunity to detect HR-HPV genotypes in HNSCC and may identify similarities or differences in the HPV genotypes in both cervical cancer and HNSCC, given that HPV is sexually transmitted.
Using this digene HPV genotyping RH test kit, the overall prevalence of HPV-associated HNSCC was 90.6%. Six out of the 18 HR-HPV genotypes were detected (HPV16, 18, 31, 45, 56, 68) in our cohort of HNSCC patients. Among these HPV genotypes, only HPV16, HPV18, HPV31 and HPV45 had been previously reported in HNSCC. The detection of these HR-HPV genotypes is consistent with the epidemiology data of HPV-associated cervical cancer where HPV31 and HPV45 have been reported, but HPV16 and HPV18 remain the predominant genotypes [5, 14–17]. The prevalence of HPV 31 and HPV45 were disproportionate relative to other studies countries since HPV31 was detected in 0.2% of oral cavity SCC, and HPV45 was reported in 0.2% in OPSCC and larynx SCC [18]. This finding cautions the routine use of HPV-DNA PCR based commercial detection kits which may have an inherently high false positive rate. This is especially crucial because HPV-associated OPSCC is associated with a superior clinical outcome compared to HPV-negative counterpart; and dose de-intensification clinical trials are underway to screen HPV-associated OPSCC patients for a less toxic radiotherapy regime. Therefore, using a HPV detection test with a high false positive rate will invariably result in under-treatment in these patients.
The digene RH test kit has a sensitivity limit of detection for HPV31 and HPV45 at 10 and 23 copies/PCR, respectively; compared to the PapilloCheck high-risk HPV kit (Greiner Bio-one, Germany) which has a higher detection level of 50 and 90 copies/PCR. Additionally, the type of primer employed by the different commercial test kits may play a role in the detection rate of HPV. The GP5+/6+ used in our study had a higher sensitivity than the MY09/11 primer set in detecting HPV45 [19]. Lastly, since there was no micro-dissection of the tumour cells in the paraffin blocks, HPV may be detected in the non-cancerous tissue. Nevertheless, the presence of HPV31 and HPV45 in HNSCC is worth exploring further since persistent HPV31 and HPV45 infections have been associated with an increased risk of progressing to cervical intraepithelial neoplasia and cervical cancer [20].
To our knowledge this is the first report of HPV56 and HPV68 being associated in HNSCC. Interestingly, HPV56 and HPV68 have been reported in cervical smears of healthy Singaporean women although these genotypes are not seen in the malignant counterparts in Singapore [14, 15, 17]. Overall, these data suggest that not all HR-HPVs detected are biologically relevant in the causation of these virally-associated cancers, and some may represent biologically inactive co-infection of neoplastic cells [21, 22]. Since the digene RH test kit only targets the L1 region of the HPV genome, it does not have the capability to differentiate integrated versus episomal forms of the virus, and hence may not identify biologically active HPV transcripts. Therefore, some authors have advocated the use of transcriptionally active HPV-E6/E7 mRNA expression as the gold standard of HPV detection [21, 23]. To update, HPV E6/E7-mRNA has been shown to be oncogenic product of HPV16 driven carcinogenesis pathway in HNSCC [24, 25]. The presence of this onco-protein is not well-characterized for the other HPV subtypes [26]. Therefore, it remains to be ascertained if the other HR-HPV genotypes detected in this study have corresponding onco-proteins which may drive or support the carcinogenesis pathway in HNSCC in similar way as HPV16.
Multiple HR-HPV genotypes are typically detected in our HNSCC cohort with HPV31 and HPV45 genotypes being the most prevalent. HPV16 genotype is hardly detected in non-oropharyngeal sites and is primarily detected in OPSCC. This information is compatible with our Western cohorts where HPV16 genotype is the commonest genotype detected in OPSCC. Specific to the Asian context, our findings differ from the HPV genotype expression in China where HPV18 and HPV16 exists in equal proportions among patients with OPSCC [27]. Besides, co-existence of multiple HR-HPVs (HPV31, HPV33, HPV35 and HPV66) with HPV16 has been reported in 23% of HPV16-positive OPSCC [23]. Therefore, geographic location appears to influence the specific HPV genotypes detected in OPSCC and further studies should evaluate the reasons for these genotypic differences in HR-HPV types in OPSCC.
P16 immunohistochemistry (IHC) detection is a useful surrogate marker for HPV-related oncogenic pathway as it represents active HPV oncogenic activation. In the HPV-driven oncogenic pathway, HPV integrates into the host genome and increases HPV-E6 and E7 gene transcription. E7 onco-protein specifically results in phosphorylation of the retinoblastoma tumor suppressor which releases E2F transcription factor. This in turn increases p16 expression [28–31]. Therefore, p16 IHC expression may result in a lower sensitivity but high specificity in OPSCC as compared to HPV-DNA PCR based assay because not all HPV infection results in downstream transcription of p16 over-expression. Our data showed that 45.2% of OPSCCs were p16-positive which was compatible with another Asian cohort which reported a prevalence of 30.7% [32]. These two Asian cohorts demonstrated a lower prevalence rate of p16-positive OPSCC as compared to our Western counterparts.
Additionally, using p16 IHC as a surrogate in OPSCC allows easy clinical translations as it is much easier to perform and less time consuming compared to HPV-DNA PCR assay. As with other studies, p16-positive OPSCC patients have consistently demonstrated improved clinical outcomes compared to patients with p16-negative OPSCC [33]. This observation is similar to our cohort of OPSCC where p16-positive OPSCC patients demonstrate superior overall survival. However, given the relative small numbers of OPSCC in Singapore, we were unable to demonstrate a superior disease-specific survival among patients with p16-positive OPSCC. Last but not least, the history of smoking and alcohol consumption did adversely affect the survival outcomes in these patients with OPSCC.
One limitation in our study was the relatively small sample size of OPSCC. Therefore, we are unable to correlate the presence of HPV16 with p16 IHC expression. The correlation between HPV16 and p16 was 42.8% (6/14) in our study; as compared to more than 80% reported in other studies [34, 35]. Furthermore, we were unable to further examine the viral RNA or protein, such as E6 in our samples due to degradation of RNA from formalinized paraffin tissue sample. Majority of the commercially available E6-RNA detection kits are designed for fresh tissue samples and we will be prospectively collecting these OPSCC tissues to validate the detection of HPV in our local cohort. Lastly, it has been suggested that consensus HPV primer may not be suitable for HNSCC where HPV16 is the predominant HPV genotype detected due to possible cross reactivity with other HPV genotypes [36, 37]. It is possible that the detection of other HR-HPVs in HNSCC may be due to extreme sensitivity of the digene test kit which may not distinguish HPV16 from HPV31 and HPV18 from HPV45; since these paired HPV genotypes are genetically closely related [38]. Be that as it may, we believe that our results represent the first complete characterization of HR-HPVs in HNSCC using a commonly used HPV-DNA PCR test kit designed for cervical cancer.
In conclusion, our HNSCC cohort from an urban, multi-ethnic, South East Asian society, demonstrated a high prevalance of HR-HPVs (HPV16, 18, 31, 45, 56, 68) using a commercial PCR-based test kit. HPV16 and P16 IHC expression were predominantly detected in the oropharyngeal subsite. Patients with p16-positive oropharyyngeal HNSCC exhibited a superior overall survival compared to those with p16-negative HNSCC; making p16 IHC a convenient surrogate marker for HPV detection in HNSCC. Routine clinical use of commercial HPV DNA based test kits should not be utilized given the high percentage of false positive results.
MATERIALS AND METHODS
Patients
Tissue blocks of patients diagnosed with primary HNSCC were retrieved from the archives of the Department of Pathology at the National University Hospital Singapore. These tissue blocks were verified to be squamous cell carcinoma by one experienced pathologist (FP) and the DNA was extracted from the tissue blocks using the QIAamp DNA Formalin-Fixed, Paraffin-Embedded (FFPE) tissue kit as per manufacturer protocol (Qiagen Inc., Germany). One hundred and fifty-nine tissue blocks contained adequate DNA and were accrued for HPV detection. HPV was detected using a Polymerase Chain Reaction (PCR)-based amplification assay (Qiagen digene HPV Genotyping RH test amplification kit, Germany) followed by a reverse hybridization procedure (Qiagen digene HPV Genotyping RH test detection kit, Germany). P16 was detected using immunohistochemical staining of the tissue slides (Roche Diagnostics GmbH, Germany). Approval by our institution review board was obtained prior to the commencement of the study.
The demographics, clinical of pathological parameters of these patients were analyzed. Clinical outcomes included overall survival, disease-specific survival and recurrence in correlation of outcomes with HPV status and p16 was performed.
P16 immunohistochemistry and scoring
For the immunohistochemistry analysis, an anti-human p16INK4a monoclonal antibody (clone: E6H4, Roche Diagnostics GmbH, Germany) was used. In brief, tissue sections were cut into 3-μm slices and deparaffinized. Antigen retrieval was done with pH9 CC1 buffer (Dako, Denmark) for 36 mins at 100°C. The tissues sections were incubated with peroxidase blocking reagent for 30 mins and then with 10% goat serum for 60 mins. The tissues were then stained with anti-p16INK4a antibody at the dilution 1:100 for 32 mins at 37°C followed by HRP Polymer anti-mouse Envision (Dako, Denmark) system for 30 mins. Slides were visualized using Ventana Ultra View DAB kit with DAB (Dako, Denmark) for 8 mins. Tissues sections were counterstained with hematoxylin for 2 mins. The specimens were considered to be p16 positive when at least 80% of the tumour cells showed both cytoplasmic and nuclear p16INK4a staining.
Genomic DNA extraction
Genomic DNA was extracted using Qiagen QiaAmp DNA FFPE kit (Qiagen Inc., Germany). Briefly, 5–10 μm thick tissue sections were deparaffinized with xylene and residual xylene were extracted by ethanol. The tissue was then lysed and digested at 56°C followed by 90°C for 1 hr. Cells were lysed and DNA was extracted using DNA column. DNA was then eluted and stored at −20°C until further testing.
Digene HPV genotyping RH test- polymerase chain reaction (PCR) and viral detection
In this study, the digene RH test was used to detect 18 carcinogenic HR-HPVs. These 18 HR-HPVs are HPV 16, 18, 26, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68, 73 and 82. The test comprised of an amplification and detection phase. Briefly, 10 μl of the extracted DNA was mixed with 25 mM MgCl2 and the commercial master mix solution containing GP5+/6+ primer sets and an internal control primer set. L1-segment PCR amplification was carried out with cycle parameters as follows; initial activation step, 94°C for 5 mins; 3-cycling steps comprising denaturation at 94°C for 20 sec; annealing at 38°C for 30 sec; extension at 71°C for 80 sec. This 3-cycling-step was repeated 40 times using PE Applied Biosystem GeneAMP PCR system 9700 machine (Applied Biosystems, USA).
After GP5+/6+ PCR amplification, detection was carried out according to the kit protocol. In brief, 10 μl of the amplicon was denatured and hybridized with the specific immobilized probes on the nitrocellulose strip in parallel lines, including an internal control- human β-globin amplicon. The nitrocellulose strip was then incubated with a conjugate and a substrate and produced a color band for visualization and identification of the specific HPV strain.
Clinical correlation statistical analysis
The statistical association between HPV-DNA PCR positivity, p16 expression and clinico-pathological was evaluated using Chi-square Fisher’s exact test (GraphPad Prism version 6 San Diego, USA). Clinical outcome measures include overall survival and disease specific survival. Overall survival was defined as time of diagnosis to death from any cause or last follow-up and disease-specific survival was defined as the time to death from cancer or treatment related complications. The survival outcomes were estimated using Kaplan-Meier and survival curve were compared using Mantel-cox log-rank tests (GraphPad Prism version 6). All calculated p values < 0.05 were considered statistically significant.
ACKNOWLEDGMENTS
Dr Lim Chwee Ming acknowledges the funding support from the National Medical Research Council Singapore Transitional Award and National University of Singapore Clinician Scientist Leadership Grant for this work.
CONFLICT OF INTEREST
None.
FINANCIAL SUPPORT
None.
REFERENCES
1. Haraf DJ, Nodzenski E, Brachman D, Mick R, Montag A, Graves D, Vokes EE, Weichselbaum RR. Human papilloma virus and p53 in head and neck cancer: clinical correlates and survival. Clin Cancer Res. 1996; 2:755–762.
2. Gillison ML, Koch WM, Capone RB, Spafford M, Westra WH, Wu L, Zahurak ML, Daniel RW, Viglione M, Symer DE, Shah KV, Sidransky D. Evidence for a causal association between human papillomavirus and a subset of head and neck cancers. J Natl Cancer Inst. 2000; 92:709–720.
3. Chaturvedi AK, Engels EA, Pfeiffer RM, Hernandez BY, Xiao W, Kim E, Jiang B, Goodman MT, Sibug-Saber M, Cozen W, Liu L, Lynch CF, Wentzensen N, et al. Human papillomavirus and rising oropharyngeal cancer incidence in the United States. J Clin Oncol. 2011; 29:4294–4301.
4. Deschler DG, Richmon JD, Khariwala SS, Ferris RL, Wang MB. The “new” head and neck cancer patient-young, nonsmoker, nondrinker, and HPV positive: evaluation. Otolaryngol Head Neck Surg. 2014; 151:375–380.
5. Munoz N, Bosch FX, de Sanjose S, Herrero R, Castellsague X, Shah KV, Snijders PJ, Meijer CJ. International Agency for Research on Cancer Multicenter Cervical Cancer Study G. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N Engl J Med. 2003; 348:518–527.
6. Ault KA. Human papillomavirus vaccines and the potential for cross-protection between related HPV types. Gynecol Oncol. 2007; 107:S31–33.
7. Singhi AD, Westra WH. Comparison of human papillomavirus in situ hybridization and p16 immunohistochemistry in the detection of human papillomavirus-associated head and neck cancer based on a prospective clinical experience. Cancer. 2010; 116:2166–2173.
8. Nasman A, Attner P, Hammarstedt L, Du J, Eriksson M, Giraud G, Ahrlund-Richter S, Marklund L, Romanitan M, Lindquist D, Ramqvist T, Lindholm J, Sparen P, et al. Incidence of human papillomavirus (HPV) positive tonsillar carcinoma in Stockholm, Sweden: an epidemic of viral-induced carcinoma? Int J Cancer. 2009; 125:362–366.
9. Giuliano AR, Tortolero-Luna G, Ferrer E, Burchell AN, de Sanjose S, Kjaer SK, Munoz N, Schiffman M, Bosch FX. Epidemiology of human papillomavirus infection in men, cancers other than cervical and benign conditions. Vaccine. 2008; 26:K17–28.
10. Olthof NC, Straetmans JM, Snoeck R, Ramaekers FC, Kremer B, Speel EJ. Next-generation treatment strategies for human papillomavirus-related head and neck squamous cell carcinoma: where do we go? Rev Med Virol. 2012; 22:88–105.
11. Howard JD, Chung CH. Biology of human papillomavirus-related oropharyngeal cancer. Semin Radiat Oncol. 2012; 22:187–193.
12. Burd EM. Human papillomavirus and cervical cancer. Clin Microbiol Rev. 2003; 16:1–17.
13. Godinez JM, Tous S, Baixeras N, Moreno-Crespi J, Alejo M, Lejeune M, Bravo IG, Bosch FX, de Sanjose S. Performance of the digene LQ, RH and PS HPVs genotyping systems on clinical samples and comparison with HC2 and PCR-based Linear Array. Infect Agent Cancer. 2011; 6:23.
14. Quek SC, Lim BK, Domingo E, Soon R, Park JS, Vu TN, Tay EH, Le QT, Kim YT, Vu BQ, Cao NT, Limson G, Pham VT, et al. Human papillomavirus type distribution in invasive cervical cancer and high-grade cervical intraepithelial neoplasia across 5 countries in Asia. Int J Gynecol Cancer. 2013; 23:148–156.
15. Tay SK, Oon LL. Prevalence of cervical human papillomavirus infection in healthy women is related to sexual behaviours and educational level: a cross-sectional study. Int J STD AIDS. 2014; 25:1013–1021.
16. Tay SK, Tay YK. The prevalence and significance of high-risk human papillomavirus DNA test in southern Malaysia and Singapore. Aust N Z J Obstet Gynaecol. 2009; 49:323–327.
17. Chan R, Khoo L, Ho TH, Koh CF, Lee IW, Yam KL, Chandra D, Pang M, Chow V. A comparative study of cervical cytology, colposcopy and PCR for HPV in female sex workers in Singapore. Int J STD AIDS. 2001; 12:159–163.
18. Kreimer AR, Clifford GM, Boyle P, Franceschi S. Human papillomavirus types in head and neck squamous cell carcinomas worldwide: a systematic review. Cancer Epidemiol Biomarkers Prev. 2005; 14:467–475.
19. Depuydt CE, Boulet GA, Horvath CA, Benoy IH, Vereecken AJ, Bogers JJ. Comparison of MY09/11 consensus PCR and type-specific PCRs in the detection of oncogenic HPV types. J Cell Mol Med. 2007; 11:881–891.
20. Schettino MT, Ammaturo FP, Grimaldi E, Legnante A, Marcello A, Donnarumma G, Colacurci N, Torella M. Persistent papillomavirus type-31 and type-45 infections predict the progression to squamous intraepithelial lesion. Taiwan J Obstet Gynecol. 2014; 53:494–497.
21. Smeets SJ, Hesselink AT, Speel EJ, Haesevoets A, Snijders PJ, Pawlita M, Meijer CJ, Braakhuis BJ, Leemans CR, Brakenhoff RH. A novel algorithm for reliable detection of human papillomavirus in paraffin embedded head and neck cancer specimen. Int J Cancer. 2007; 121:2465–2472.
22. van Houten VM, Snijders PJ, van den Brekel MW, Kummer JA, Meijer CJ, van Leeuwen B, Denkers F, Smeele LE, Snow GB, Brakenhoff RH. Biological evidence that human papillomaviruses are etiologically involved in a subgroup of head and neck squamous cell carcinomas. Int J Cancer. 2001; 93:232–235.
23. Anantharaman D, Gheit T, Waterboer T, Abedi-Ardekani B, Carreira C, McKay-Chopin S, Gaborieau V, Marron M, Lagiou P, Ahrens W, Holcatova I, Merletti F, Kjaerheim K, et al. Human papillomavirus infections and upper aero-digestive tract cancers: the ARCAGE study. J Natl Cancer Inst. 2013; 105:536–545.
24. Bishop JA, Ma XJ, Wang H, Luo Y, Illei PB, Begum S, Taube JM, Koch WM, Westra WH. Detection of transcriptionally active high-risk HPV in patients with head and neck squamous cell carcinoma as visualized by a novel E6/E7 mRNA in situ hybridization method. Am J Surg Pathol. 2012; 36:1874–1882.
25. Deng Z, Hasegawa M, Kiyuna A, Matayoshi S, Uehara T, Agena S, Yamashita Y, Ogawa K, Maeda H, Suzuki M. Viral load, physical status, and E6/E7 mRNA expression of human papillomavirus in head and neck squamous cell carcinoma. Head Neck. 2013; 35:800–808.
26. Michaud DS, Langevin SM, Eliot M, Nelson HH, Pawlita M, McClean MD, Kelsey KT. High-risk HPV types and head and neck cancer. Int J Cancer. 2014; 135:1653–1661.
27. Mehanna H, Franklin N, Compton N, Robinson M, Powell N, Biswas-Baldwin N, Paleri V, Hartley A, Fresco L, Al-Booz H, Junor E, El-Hariry I, Roberts S, et al. Geographic variation in human papillomavirus-related oropharyngeal cancer: Data from four multinational randomized trials. Head Neck. 2016.
28. Kroupis C VN. Human papilloma virus (HPV) molecular diagnostics. Clin Chem Lab Med. 2011; 49:1783–1799.
29. Munger K, Basile JR, Duensing S, Eichten A, Gonzalez SL, Grace M, Zacny VL. Biological activities and molecular targets of the human papillomavirus E7 oncoprotein. Oncogene. 2001; 20:7888–7898.
30. Munger K, Howley PM. Human papillomavirus immortalization and transformation functions. Virus Res. 2002; 89:213–228.
31. Hammarstedt L, Lindquist D, Dahlstrand H, Romanitan M, Dahlgren LO, Joneberg J, Creson N, Lindholm J, Ye W, Dalianis T, Munck-Wikland E. Human papillomavirus as a risk factor for the increase in incidence of tonsillar cancer. Int J Cancer. 2006; 119:2620–2623.
32. Ang SH, Haaland B, Acharyya S, Thu MM, Krisna SS, Hwang SG, Tan PH, Ng QS, Tan DS, Tai WM, Tan EH, Lim WT, Ang MK. Interactions between clinical factors, p16, and cyclin-D1 expression and survival outcomes in oropharyngeal and hypopharyngeal squamous cell carcinoma. Head Neck. 2015; 37:1650–1659.
33. Fischer CA, Zlobec I, Green E, Probst S, Storck C, Lugli A, Tornillo L, Wolfensberger M, Terracciano LM. Is the improved prognosis of p16 positive oropharyngeal squamous cell carcinoma dependent of the treatment modality? Int J Cancer. 2010; 126:1256–1262.
34. Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-Tan PF, Westra WH, Chung CH, Jordan RC, Lu C, Kim H, Axelrod R, Silverman CC, et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med. 2010; 363:24–35.
35. Reimers N, Kasper HU, Weissenborn SJ, Stutzer H, Preuss SF, Hoffmann TK, Speel EJ, Dienes HP, Pfister HJ, Guntinas-Lichius O, Klussmann JP. Combined analysis of HPV-DNA, p16 and EGFR expression to predict prognosis in oropharyngeal cancer. Int J Cancer. 2007; 120:1731–1738.
36. Schache AG, Liloglou T, Risk JM, Filia A, Jones TM, Sheard J, Woolgar JA, Helliwell TR, Triantafyllou A, Robinson M, Sloan P, Harvey-Woodworth C, Sisson D, et al. Evaluation of human papilloma virus diagnostic testing in oropharyngeal squamous cell carcinoma: sensitivity, specificity, and prognostic discrimination. Clin Cancer Res. 2011; 17:6262–6271.
37. Lajer CB, von Buchwald C. The role of human papillomavirus in head and neck cancer. APMIS. 2010; 118:510–519.
38. Wentzensen N, Sun C, Ghosh A, Kinney W, Mirabello L, Wacholder S, Shaber R, LaMere B, Clarke M, Lorincz AT, Castle PE, Schiffman M, Burk RD. Methylation of HPV18, HPV31, and HPV45 genomes and cervical intraepithelial neoplasia grade 3. J Natl Cancer Inst. 2012; 104:1738–1749.

