INTRODUCTION
Schizophrenia is a chronic, severely debilitating mental disorder. Approximately 1% of the population suffers the torment of schizophrenia [1], and it is reportedly the 8th leading cause of disability among people aged 15 to 44 years, worldwide [2]. The exact pathogenesis of schizophrenia remains unknown and, despite large numbers of trials of potential therapies, the efficacy of pharmacological treatments is poor for many schizophrenia patients [3]. Although the gender ratio among patients is nearly equal, females tend to have a later onset than males, and an earlier onset in females almost associated with a family history of schizophrenia [4]. Lichtenstein et al estimated the heritability of schizophrenia to be 64% [5].
A variety of studies have suggested that genetic factors play a key role in the onset and development of schizophrenia. In genome-wide association studies (GWAS), for example, around 30 loci have been identified as being associated with schizophrenia [6]. In addition, an association between ANK3 and schizophrenia has been detected in several different ethnic groups. The rs10761482 variant is reportedly associated with schizophrenia risk in Norwegian subjects [7], but not in subjects from Germany [8] or southern China [9]. On the other hand, the rs10994336 variant is associated with bipolar disorder in Malays [10] and southern Chinese [9], whereas rs10994336 was found not to be associated with schizophrenia in cohorts of German and Nordic subjects [8, 11].
The associations between ANK3 polymorphisms and schizophrenia risk have not been investigated in the northern Chinese Han population. In present study, therefore, we selected 13 ANK3 SNPs whose association with schizophrenia or bipolar disorder has been previously investigated [7, 10–21] and evaluated whether these polymorphisms are associated with schizophrenia risk in a northern Chinese Han population.
RESULTS
A total of 499 schizophrenia cases and 500 unrelated healthy controls were enrolled in our study. The demographic data for the cases and controls are shown in Table 1. The mean ages were 67.11 ± 9.24 years and 64.31 ± 8.91 years for the case and control groups, respectively. As shown in Table 2, 13 SNPs were analyzed, and two (rs10761482 and rs3808942) displayed significant deviation from the Hardy-Weinberg equilibrium (p < 0.05); We used the Pearson Chi-squared test to compare the different allele frequency distributions between the cases and controls and found that rs958852 was significantly associated with a 0.75-fold reduction in the risk of schizophrenia (95%CI 0.58–0.97; p = 0.025) (Table 2).
Table 1: Characteristics of cases and controls in this study n (%)
Variable |
Case |
Control |
pvalue |
|---|---|---|---|
(n = 499) |
(n = 500) |
||
Sex |
< 0.001a |
||
Male |
263 ( 52.7) |
192 (38.4) |
|
Female |
236 (47.3) |
308 (61.6) |
|
Age, year (mean ± SD) |
36.68 ± 13.25 |
50.35 ± 7.76 |
< 0.001b |
p ≤ 0.05 indicates statistical significance.
aTwo-sided Chi-squared test.
bIndependent samples t test.
Table 2: Candidate SNPs examined in ANK3 (10q21.2) in cases and controls and odds ratio estimates for schizophrenia
SNP No. |
Alleles A/B |
MAF (case) |
MAF (control) |
pavalue for HWE test |
OR(95% CI) |
pbadj. |
|
|---|---|---|---|---|---|---|---|
rs12761450 |
T/G |
0.048 |
0.055 |
0.053 |
0.87 |
0.58–1.29 |
0.485 |
rs10761482 |
T/C |
0.273 |
0.260 |
0.007* |
1.07 |
0.87–1.30 |
0.526 |
rs3808942 |
C/T |
0.230 |
0.232 |
0.032* |
0.99 |
0.81–1.22 |
0.935 |
rs10994336 |
T/C |
0.229 |
0.232 |
0.102 |
0.99 |
0.80–1.21 |
0.893 |
rs10994338 |
A/G |
0.227 |
0.231 |
0.102 |
0.98 |
0.79–1.20 |
0.831 |
rs4948418 |
T/C |
0.230 |
0.232 |
0.102 |
0.99 |
0.81–1.22 |
0.935 |
rs10994359 |
C/T |
0.334 |
0.328 |
0.362 |
1.03 |
0.85–1.24 |
0.788 |
rs10994397 |
T/C |
0.329 |
0.320 |
0.682 |
1.04 |
0.86–1.25 |
0.679 |
rs1938526 |
G/A |
0.341 |
0.320 |
0.679 |
1.10 |
0.91–1.33 |
0.308 |
rs10994415 |
C/T |
0.349 |
0.320 |
0.838 |
1.14 |
0.94–1.37 |
0.174 |
rs958852 |
T/A |
0.123 |
0.158 |
0.313 |
0.75 |
0.58–0.97 |
0.025* |
rs16915157 |
T/C |
0.375 |
0.346 |
0.490 |
1.13 |
0.94–1.36 |
0.181 |
rs1837950 |
T/C |
0.416 |
0.415 |
0.927 |
1.00 |
0.84–1.20 |
0.970 |
SNP single nucleotide polymorphism, HWE Hardy-Weinberg equilibrium, OR odds ratio, 95% CI 95% confidence interval, MAF minor allele frequency
*p ≤ 0.05 indicates statistical significance
ap was calculated by exact test
b p was calculated by Pearson Chi-squared test
We also assumed that the minor SNP allele was a greater risk factor than the wild-type allele (Table 3). We used three genetic models (dominant, recessive and additive) to analyze the associations between the SNPs and the schizophrenia risk. We found that rs10994336, rs10994338 and rs4948418 were associated with schizophrenia risk in the recessive model (OR 2.00; 95%CI 1.01–3,94; p = 0.047, OR 1.99; 95%CI 1.01–3.93; p = 0.047 and OR 2.00; 95%CI 1.01–3,94; p = 0.047, respectively). In addition, rs958852 was associated with schizophrenia risk in the dominant (OR 0.69; 95%CI 0.49–0.97; p = 0.031) and additive (OR 0.73; 95%CI 0.54–0.99; p = 0.044) models.
Table 3: Logistic regression analysis of the association between SNPs and schizophrenia risk n (%)
SNP No. |
Minor |
Dominant model |
Recessive model |
Additive model |
||||||
|---|---|---|---|---|---|---|---|---|---|---|
allele |
OR |
95% CI |
Pa |
OR |
95% CI |
Pa |
OR |
95% CI |
Pa |
|
rs10994336 |
T |
0.91 |
0.68–1.23 |
0.555 |
2.00 |
1.01–3.94 |
0.047* |
1.03 |
0.80–1.33 |
0.799 |
rs10994338 |
A |
0.90 |
0.67–1.21 |
0.487 |
1.99 |
1.01–3.93 |
0.047* |
1.02 |
0.80–1.31 |
0.868 |
rs4948418 |
T |
0.92 |
0.68–1.24 |
0.577 |
2.00 |
1.01–3.94 |
0.047* |
1.04 |
0.81–1.33 |
0.779 |
rs958852 |
T |
0.69 |
0.49–0.97 |
0.031* |
0.87 |
0.28–2.66 |
0.802 |
0.73 |
0.54–0.99 |
0.044* |
*p ≤ 0.05 indicates statistical significance
ap values were calculated by unconditional logistic regression adjusted for age and gender
Table 4: Comparisons of genotypes of some special SNPs and their associations with schizophrenia risk
SNP ID |
Test |
OR |
95% CI |
p |
|---|---|---|---|---|
rs10994336 |
TT/CC |
1.86 |
0.93–3.70 |
0.080 |
TC/CC |
0.83 |
0.60–1.13 |
0.234 |
|
(TT+TC)/CC |
– |
– – |
0.069 |
|
rs10994338 |
AA/GG |
1.84 |
0.92–3.67 |
0.084 |
AG/GG |
0.81 |
0.59–1.11 |
0.193 |
|
(AA+AG)/GG |
– |
– – |
0.060 |
|
rs4948418 |
TT/CC |
1.86 |
0.93–3.71 |
0.079 |
TC/CC |
0.83 |
0.61–1.14 |
0.248 |
|
(TT+TC)/CC |
– |
– |
0.071 |
|
rs958852 |
TT/AA |
0.79 |
0.26–2.43 |
0.678 |
TA/AA |
0.68 |
0.48–0.97 |
0.031* |
|
(TT+TA)/AA |
– |
– – |
0.094 |
*p ≤ 0.05 indicates statistical significance.
p values were calculated from Wald’s test adjusted by age and gender.
Comparisons of the SNP genotypes and their associations with the risk of schizophrenia are shown in Table 4. We found that genotype “TA” of rs958852 in ANK3 was associated with a 0.68-fold lower risk of schizophrenia than genotype “AA” at the 5% level (95%CI 0.48–0.97; p = 0.031).
We next used the genotype data from all the subjects to do an LD analysis. Two blocks were detected based on haplotype analysis. One block included four SNPs (rs10994336, rs10994338, rs4948418 and rs10994359) with D’ = 0.95, and the other included three SNPs (rs10994415, rs958852 and rs16915157) with D’ = 0.98 (Figure 1). The associations between the ANK3 haplotypes and the schizophrenia risk are listed in Table 5. We found the “TTC” haplotype was associated with a reduced risk of schizophrenia (OR 0.73; 95%CI 0.54–0.99; p = 0.044).
Lastly, we looked for interactions among the 13 SNPs tested with schizophrenia risk stratified based on gender (Supplementary Table S1). We found that none of the analyzed SNPs correlated with gender had an impact on the association with schizophrenia risk (data was shown in supplementary).
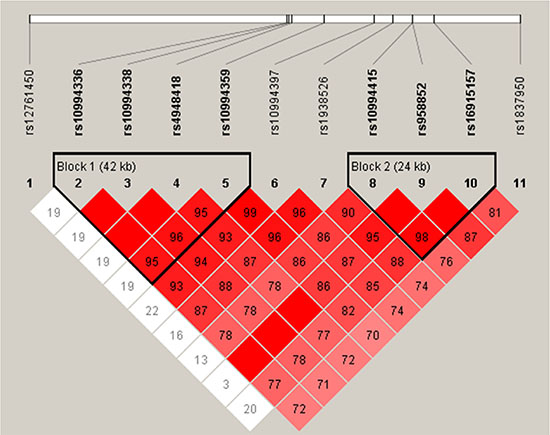
Figure 1: Haplotype block map for all the ANK3 SNPs. Linkage disequilibrium (LD) plots containing eleven SNPs from 10q21.2 of ANK3. Red squares indicate statistically significant associations between a pair of SNPs, as measured by D’; darker shades of red indicate higher D’ values.
Table 5: The haplotypes of three SNPs(rs10994415, rs958852 and rs16915157) and risk of schizophrenia(adjusted by age and gender)
Haplotype |
Freq(case) |
Freq(control) |
Chi-square |
pa |
OR |
95% CI |
pb |
|
|---|---|---|---|---|---|---|---|---|
CAT |
0.347 |
0.317 |
1.986 |
0.159 |
1.11 |
0.89 |
1.39 |
0.347 |
TAT |
0.028 |
0.029 |
0.016 |
0.899 |
1.02 |
0.55 |
1.89 |
0.962 |
TTC |
0.123 |
0.158 |
4.992 |
0.025* |
0.73 |
0.54 |
0.99 |
0.044* |
TAC |
0.500 |
0.493 |
0.098 |
0.754 |
1.06 |
0.86 |
1.30 |
0.585 |
* p ≤ 0.05 indicates statistical significance.
a p value from were calculated from two-sided Chi-squared test.
b p values were calculated by unconditional logistic regression adjusted for age and gender.
DISCUSSION
In this case-control study, we investigated associations between genetic variations in ANK3 and schizophrenia risk in a relatively large sample of individuals. Four susceptibility loci from ANK3 (rs10994336, rs10994338, rs4948418 and rs958852) were significantly associated with schizophrenia risk in a northern Chinese Han population for the first time. We also found that the “TTC” haplotype of ANK3 is associated with a 0.73-fold reduction in the risk of schizophrenia.
Many earlier studies focused on gene-related mechanisms or other potential causes for schizophrenia. Okazaki et al [22] suggested that an imbalance in the immune system contributes to the pathophysiology of schizophrenia. Brown et al reported that cardiovascular disease is the most important natural cause of death among schizophrenia patients [23, 24]. Hjelm et al suggested that the genetic etiology of schizophrenia must be strikingly complex, as they were unable to identify any special gene associated with schizophrenia [25]. Several studies have reported that ANK3 plays an important role in the pathogenesis of schizophrenia. ANK3 is reportedly a neurodevelopmental gene [26] that encodes ankyrin-G (ANKG), which is important for the stability of the neuronal membrane [27]. Notably, the neurodevelopment hypothesis has become the mainstream hypothesis of the etiology of schizophrenia [28]. What’s more, ANKG associates with Nav1.5 and recruits the channel to the myocyte membrane, and impaired Nav1.5 function caused by low ANKG levels leads to sinus node dysfunction, conduction defects, and ventricular arrhythmia [29]. This is consistent with the earlier report that cardiovascular disease is a frequent cause of death among schizophrenia patients. At the same time, ANK3 was found to be associated with bipolar disorder in the large collaborative GWAS [30], and Cassidy et al suggested that schizophrenia and bipolar disorder share common pathogenic pathways [13], as evidenced by the genetic overlap between schizophrenia and bipolar disorder [31].
Rs10994336 was identified as a significant susceptibility locus for bipolar disorder in study of 4,387 cases and 6,209 controls [32], and in a mate analysis [33]. One UK study tried to investigate whether the SNP loci significant in bipolar disorder are also significant in schizophrenia, but they failed to demonstrate a significant association between rs10994336 and schizophrenia risk. In the present study, we observed that rs10994336 is associated with an increased risk of schizophrenia in a northern Chinese Han population. We suppose the difference between our findings and those of the UK study reflects the difference in the ethnic groups.
Associations between rs10994338 and rs4948418 and bipolar disorder have also been previously reported [33, 34]. In present study, we found that rs10994338 and rs4948418 were associated with an increased schizophrenia risk in the northern Chinese Han population. We also found a new significant SNP, rs958852, which appears to reduce the risk of schizophrenia. To our knowledge, this is the first report of an association between ANK3 rs10994336, rs10994338, rs4948418 and rs958852 and schizophrenia risk in a northern Chinese Han population. Our findings suggest ANK3 may play an important role in schizophrenia. It is anticipated that future studies will clarify the precise contribution of ANK3 to schizophrenia, which may offer a new strategies for the prevention and treatment of schizophrenia.
Although there are important discoveries revealed in our study, there are also some potential limitations. First, we selected SNPs with p values higher than 5% in the HWE test. This approach may exclude some SNPs reported to be significantly associated with schizophrenia in previous studies. Second, our study lacks biological function experiments, which will be crucial for elucidating the role of ANK3 in schizophrenia.
MATERIALS AND METHODS
Study participants
We recruited a total of 499 (263 males, 236 females) northern Han Chinese patients diagnosed with schizophrenia between 2011 and 2015. All the subjects were treated at the Xi’an Mental Health Center. All patients were diagnosed and histologically confirmed to suffer from schizophrenia based on the criteria from the DSM-IV (Diagnostic and Statistical Manual of Mental Disorders, the fourth version). The patients had no history of cancer, nephropathy, infection, or other related disease. In addition, to serve as a control sample, 500 (192 male, 308 female) healthy unrelated individuals were recruited from among the Han Chinese living in and around Xi’an city. All of the chosen controls were from the Medical Center in the Affiliated Hospital of Tibet University for Nationalities.
Clinical data and demographic
The study protocol was approved by the Human Research Committee of the Xi’an Mental Health Center for Approval of Research Involving Human Subjects. Informed consent was obtained from all subjects. All subjects were interviewed by a nurse who collected detailed information about gender, age, region, ethnicity, smoking status, education status, working environment, pressures experienced, family history, and other lifestyle factors.
SNP selection and genotyping
In this case-control study, we selected 13 SNPs in ANK3, each with a minor allele frequency (MAF) higher than 5% in the Han Chinese population. We used the GoldMag extraction method to extract genomic DNA from whole blood (GoldMag Co. Ltd, Xi’an, China) [35]. Spectrometry (DU530UV/VIS spectrophotometer, Beckman Instruments, Fullerton, CA, USA) was used to measure the DNA concentration. Multiplexed SNP MassEXTEND assays were designed using Sequenom MassARRAY Assay Design 3.0 software, and the Sequenom MassARRAY RS1000 recommended by the manufacturer was utilized to perform the SNP genotyping [36]. Genotype data were managed and analyzed using Sequenom Typer 4.0 software [36–38].
Statistical analyses
Statistical analyses were performed using the SPSS version 16.0 statistical package (SPSS, Chicago, IL, USA) and Microsoft Excel. All p values were two-sided, and values of p ≤ 0.05 were considered significant. The genotype frequencies for each SNP in the control subjects were checked using the Hardy-Weinberg equilibrium (HWE). Chi-squared test or Fisher’s exact test was used to calculate the allele and genotype frequencies among cases and controls. Associations between the genotypes and schizophrenia risk were estimated by computing odds ratios (ORs) and 95% confidence intervals (CIs) evaluated in three genetic models (dominant, recessive, additive model) using unconditional logistic regression adjusted for age and gender. We determined p values for trend by entering the variable as a single term in the model (i.e., one degree of-freedom) and testing using Wald’s test. Finally, the Haploview software package (version 4.2) and SHEsis software platform (http://analysis.bio-x.cn/myAnalysis.php) were used for estimate the pairwise linkage disequilibrium (LD), haplotype construction, and genetic association at polymorphism loci.
CONCLUSIONS
In sum, we confirmed a positive association between ANK3 and schizophrenia and also demonstrated that four ANK3 variants (rs10994336, rs10994338, rs4948418 and rs958852) are associated with schizophrenia in a northern Chinese Han population. Further evidence will be needed to confirm this interaction in a wider population, and to determine the role of ANK3 in the development of schizophrenia.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Whiteford HA, Degenhardt L, Rehm J, Baxter AJ, Ferrari AJ, Erskine HE, Charlson FJ, Norman RE, Flaxman AD, Johns N, Burstein R, Murray CJL, Vos T. Global burden of disease attributable to mental and substance use disorders: findings from the Global Burden of Disease Study 2010. The Lancet. 2013; 382:1575–86. doi: 10.1016/s0140-6736(13)61611-6.
2. Rossler W, Salize HJ, van Os J, Riecher-Rossler A. Size of burden of schizophrenia and psychotic disorders. Eur Neuropsychopharmacol. 2005; 15:399–409. doi: 10.1016/j.euroneuro.2005.04.009.
3. Lieberman JA, Stroup TS, McEvoy JP, Swartz MS, Rosenheck RA, Perkins DO, Keefe RS, Davis SM, Davis CE, Lebowitz BD, Severe J, Hsiao JK. Effectiveness of antipsychotic drugs in patients with chronic schizophrenia. N Engl J Med. 2005; 353:1209–23. doi: 10.1056/NEJMoa051688.
4. Loranger AW. Sex difference in age at onset of schizophrenia. Arch Gen Psychiatry. 1984; 41:157–61. doi:
5. Lichtenstein P, Yip BH, Björk C, Pawitan Y, Cannon TD, Sullivan PF, Hultman CM. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. The Lancet. 2009; 373:234–9. doi: 10.1016/s0140-6736(09)60072-6.
6. Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014; 511:421–7. doi: 10.1038/nature13595.
7. Athanasiu L, Mattingsdal M, Kahler AK, Brown A, Gustafsson O, Agartz I, Giegling I, Muglia P, Cichon S, Rietschel M, Pietilainen OP, Peltonen L, Bramon E, et al. Gene variants associated with schizophrenia in a Norwegian genome-wide study are replicated in a large European cohort. J Psychiatr Res. 2010; 44:748–53. doi: 10.1016/j.jpsychires.2010.02.002.
8. Gella A, Segura M, Durany N, Pfuhlmann B, Stober G, Gawlik M. Is Ankyrin a genetic risk factor for psychiatric phenotypes? BMC Psychiatry. 2011; 11:103. doi: 10.1186/1471-244x-11-103.
9. Zhang C, Cai J, Zhang J, Li Z, Guo Z, Zhang X, Lu W, Zhang Y, Yuan A, Yu S, Fang Y. Genetic modulation of working memory deficits by ankyrin 3 gene in schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry. 2014; 50:110–5. doi: 10.1016/j.pnpbp.2013.12.010.
10. Lim CH, Zain SM, Reynolds GP, Zain MA, Roffeei SN, Zainal NZ, Kanagasundram S, Mohamed Z. Genetic association of LMAN2L gene in schizophrenia and bipolar disorder and its interaction with ANK3 gene polymorphism. Prog Neuropsychopharmacol Biol Psychiatry. 2014; 54:157–62. doi: 10.1016/j.pnpbp.2014.05.017.
11. Tesli M, Koefoed P, Athanasiu L, Mattingsdal M, Gustafsson O, Agartz I, Rimol LM, Brown A, Wirgenes KV, Smorr LL, Kahler AK, Werge T, Mors O, et al. Association analysis of ANK3 gene variants in nordic bipolar disorder and schizophrenia case-control samples. Am J Med Genet B Neuropsychiatr Genet. 2011; 156b:969–74. doi: 10.1002/ajmg.b.31244.
12. Bergen SE, O'Dushlaine CT, Ripke S, Lee PH, Ruderfer DM, Akterin S, Moran JL, Chambert KD, Handsaker RE, Backlund L, Osby U, McCarroll S, Landen M, et al. Genome-wide association study in a Swedish population yields support for greater CNV and MHC involvement in schizophrenia compared with bipolar disorder. Mol Psychiatry. 2012; 17:880–6. doi: 10.1038/mp.2012.73.
13. Cassidy C, Buchy L, Bodnar M, Dell'elce J, Choudhry Z, Fathalli F, Sengupta S, Fox R, Malla A, Lepage M, Iyer S, Joober R. Association of a risk allele of ANK3 with cognitive performance and cortical thickness in patients with first-episode psychosis. J Psychiatry Neurosci. 2014; 39:31–9. doi: 10.1503/jpn.120242.
14. Chen DT, Jiang X, Akula N, Shugart YY, Wendland JR, Steele CJ, Kassem L, Park JH, Chatterjee N, Jamain S, Cheng A, Leboyer M, Muglia P, et al. Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol Psychiatry. 2013; 18:195–205. doi: 10.1038/mp.2011.157.
15. Chouraki V, De Bruijn RF, Chapuis J, Bis JC, Reitz C, Schraen S, Ibrahim-Verbaas CA, Grenier-Boley B, Delay C, Rogers R, Demiautte F, Mounier A, Fitzpatrick AL, et al. A genome-wide association meta-analysis of plasma Abeta peptides concentrations in the elderly. Mol Psychiatry. 2014; 19:1326–35. doi: 10.1038/mp.2013.185.
16. Grupe A, Li Y, Rowland C, Nowotny P, Hinrichs AL, Smemo S, Kauwe JS, Maxwell TJ, Cherny S, Doil L, Tacey K, van Luchene R, Myers A, et al. A scan of chromosome 10 identifies a novel locus showing strong association with late-onset Alzheimer disease. Am J Hum Genet. 2006; 78:78–88. doi: 10.1086/498851.
17. Hwang SJ, Yang Q, Meigs JB, Pearce EN, Fox CS. A genome-wide association for kidney function and endocrine-related traits in the NHLBI's Framingham Heart Study. BMC Med Genet. 2007; 8:S10. doi: 10.1186/1471-2350-8-s1-s10.
18. Liu Y, Blackwood DH, Caesar S, de Geus EJ, Farmer A, Ferreira MA, Ferrier IN, Fraser C, Gordon-Smith K, Green EK, Grozeva D, Gurling HM, Hamshere ML, et al. Meta-analysis of genome-wide association data of bipolar disorder and major depressive disorder. Mol Psychiatry. 2011; 16:2–4. doi: 10.1038/mp.2009.107.
19. Muhleisen TW, Leber M, Schulze TG, Strohmaier J, Degenhardt F, Treutlein J, Mattheisen M, Forstner AJ, Schumacher J, Breuer R, Meier S, Herms S, Hoffmann P, et al. Genome-wide association study reveals two new risk loci for bipolar disorder. Nat Commun. 2014; 5:3339. doi: 10.1038/ncomms4339.
20. Schizophrenia Psychiatric Genome-Wide Association Study C. Genome-wide association study identifies five new schizophrenia loci. Nat Genet. 2011; 43:969–76. doi: 10.1038/ng.940.
21. Yuan A, Yi Z, Wang Q, Sun J, Li Z, Du Y, Zhang C, Yu T, Fan J, Li H, Yu S. ANK3 as a risk gene for schizophrenia: new data in Han Chinese and meta analysis. Am J Med Genet B Neuropsychiatr Genet. 2012; 159b:997–1005. doi: 10.1002/ajmg.b.32112.
22. Okazaki S, Watanabe Y, Hishimoto A, Sasada T, Mouri K, Shiroiwa K, Eguchi N, Ratta-Apha W, Otsuka I, Nunokawa A, Kaneko N, Shibuya M, Someya T, et al. Association analysis of putative cis-acting polymorphisms of interleukin-19 gene with schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry. 2014; 50:151–6. doi: 10.1016/j.pnpbp.2013.12.006.
23. Brown S, Kim M, Mitchell C, Inskip H. Twenty-five year mortality of a community cohort with schizophrenia. Br J Psychiatry. 2010; 196:116–21. doi: 10.1192/bjp.bp.109.067512.
24. Hennekens CH. Increasing global burden of cardiovascular disease in general populations and patients with schizophrenia. J Clin Psychiatry. 2007; 68:4–7. doi:
25. Hjelm BE, Rollins B, Mamdani F, Lauterborn JC, Kirov G, Lynch G, Gall CM, Sequeira A, Vawter MP. Evidence of Mitochondrial Dysfunction within the Complex Genetic Etiology of Schizophrenia. Mol Neuropsychiatry. 2015; 1:201–19. doi: 10.1159/000441252.
26. Talkowski ME, Rosenfeld JA, Blumenthal I, Pillalamarri V, Chiang C, Heilbut A, Ernst C, Hanscom C, Rossin E, Lindgren AM, Pereira S, Ruderfer D, Kirby A, et al. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell. 2012; 149:525–37. doi: 10.1016/j.cell.2012.03.028.
27. Bennett V, Otto E, Kunimoto M, Kordeli E, Lambert S. Diversity of ankyrins in the brain. Biochem Soc Trans. 1991; 19:1034–9. doi:
28. Savonenko AV, Melnikova T, Laird FM, Stewart KA, Price DL, Wong PC. Alteration of BACE1-dependent NRG1/ErbB4 signaling and schizophrenia-like phenotypes in BACE1-null mice. Proc Natl Acad Sci U S A. 2008; 105:5585–90. doi: 10.1073/pnas.0710373105.
29. Makara MA, Curran J, Little SC, Musa H, Polina I, Smith SA, Wright PJ, Unudurthi SD, Snyder J, Bennett V, Hund TJ, Mohler PJ. Ankyrin-G Coordinates Intercalated Disc Signaling Platform to Regulate Cardiac Excitability In Vivo. Circulation Research. 2014; 115:929–38. doi: 10.1161/circresaha.115.305154.
30. Ferreira MA, O'Donovan MC, Meng YA, Jones IR, Ruderfer DM, Jones L, Fan J, Kirov G, Perlis RH, Green EK, Smoller JW, Grozeva D, Stone J, et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat Genet. 2008; 40:1056–8. doi: 10.1038/ng.209.
31. Williams HJ, Norton N, Dwyer S, Moskvina V, Nikolov I, Carroll L, Georgieva L, Williams NM, Morris DW, Quinn EM, Giegling I, Ikeda M, Wood J, et al. Fine mapping of ZNF804A and genome-wide significant evidence for its involvement in schizophrenia and bipolar disorder. Mol Psychiatry. 2011; 16:429–41. doi: 10.1038/mp.2010.36.
32. Ferreira MA, O'Donovan MC, Meng YA, Jones IR, Ruderfer DM, Jones L, Fan J, Kirov G, Perlis RH, Green EK, Smoller JW, Grozeva D, Stone J, et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat Genet. 2008; 40:1056–8. doi: 10.1038/ng.209.
33. Liu Y, Blackwood DH, Caesar S, de Geus EJ, Farmer A, Ferreira MA, Ferrier IN, Fraser C, Gordon-Smith K, Green EK, Grozeva D, Gurling HM, Hamshere ML, et al. Meta-analysis of genome-wide association data of bipolar disorder and major depressive disorder. Mol Psychiatry. 2011; 16:2–4. doi: 10.1038/mp.2009.107.
34. Fiorentino A, O'Brien NL, Locke DP, McQuillin A, Jarram A, Anjorin A, Kandaswamy R, Curtis D, Blizard RA, Gurling HM. Analysis of ANK3 and CACNA1C variants identified in bipolar disorder whole genome sequence data. Bipolar Disord. 2014; 16:583–91. doi: 10.1111/bdi.12203.
35. Kochl S, Niederstatter H, Parson W. DNA extraction and quantitation of forensic samples using the phenol-chloroform method and real-time PCR. Methods Mol Biol. 2005; 297:13–30. doi:
36. Gabriel S, Ziaugra L, Tabbaa D. SNP genotyping using the Sequenom MassARRAY iPLEX platform. Curr Protoc Hum Genet. 2009; Chapter 2: Unit 2.12. doi: 10.1002/0471142905.hg0212s60.
37. Thomas RK, Baker AC, Debiasi RM, Winckler W, Laframboise T, Lin WM, Wang M, Feng W, Zander T, MacConaill L, Lee JC, Nicoletti R, Hatton C, et al. High-throughput oncogene mutation profiling in human cancer. Nat Genet. 2007; 39:347–51. doi: 10.1038/ng1975.
38. Yunus Z, Liu L, Wang H, Zhang L, Li X, Geng T, Kang L, Jin T, Chen C. Genetic polymorphisms of pharmacogenomic VIP variants in the Kyrgyz population from northwest China. Gene. 2013; 529:88–93. doi: 10.1016/j.gene.2013.07.078.

