INTRODUCTION
The natural occurred antisense transcripts (Nats), widely transcribed in human and animals, usually interacted with the sense RNA via their complemented regions [1, 2] and involved in various biological processes [3, 4], development and diseases [5], suggesting critical roles of antisense transcripts in mammalian gene expression. In eukaryotes, the double transcribed gene was first found in mouse in 1986 [6], subsequently, they were also identified in avian [7]. Currently, Nats have been found existed in diverse organisms, such as mouse [2], chicken [8], pig [9], sheep [10] and human [1]. A lot of Nats have been reported playing important roles in human diseases and mammalian development, such as Tsix [4], DHRS4-AS [3] and AS-IL1α [11]. However, until now only four gene’s Nats have been studied in detail in avian and they were chicken bFGF [7], IGF-II [12], Collagen a1 (I) [13] and the programmed cell death 2 [14].
Growth hormone receptor played key roles in human and animal growth. Both human laron type dwarfism [15] and sex linked dwarf chicken [16] were caused by the mutation of GHR gene. Chicken GHR gene, located in chromosome Z, comprised 8 exons and was highly expressed in liver. Growth hormone (GH), combined with GHR in liver and formed GH-GHR-IGFs signal pathway, affected animal development [17]. Previous studies revealed that chicken GHR gene possessed several kinds of sense transcripts by alternatively using different 5′UTR and other ways [17, 18].
In the present study, we identified and characterized a tail to tail Nat of chicken GHR gene, GHR-AS (GHR antisense transcript), in chicken liver. GHR-AS is a 4,337 bp spliced and polyadenylated RNA transcribed from 3′UTR of the GHR gene in the antisense direction that affects the local epigenetic status of GHR gene promoter. GHR-AS could protect GHR mRNA (GHR sense transcript, GHR-S) by forming double strand RNAs via exon 8 of GHR gene. The results will be beneficial in revealing complex regulation network of GHR gene.
RESULTS
Identification of a naturally occurring antisense transcript overlapping chicken GHR gene
Antisense transcripts were found extensively expressed in chicken liver in our previous study by digital gene expression sequencing of RNA [8]. In addition to the known expressed antisense transcripts, a lot of novel Nats were also detected in our analysis, including GHR-AS. Based on the obtained tags of RNA sequencing, by performing 5′ and 3′ RACE, we cloned a 4,337 bp antisense transcript of GHR, designated GHR-AS, consisting of three exons with a 3′ polyadenylation tail. 5′UTR, exon 1, 2 and 3 of GHR-AS overlaps with the 3′UTR, exon 8, exon 7and exon 6 of GHR in an antisense type, respectively (Figure 1). The sequence of GHR-AS from chicken liver has been submitted to GenBank (accession nos.KX268230). Based on the analysis results of the Coding Potential Calculator (http://cpc.cbi.pku.edu.cn/) and ORF-Finder (http://www.ncbi.nlm.nih.gov/gorf/gorf.html), GHR-AS was classified as “noncoding” RNA with a coding potential score of -1.0185 (Supplementary Figure S1).
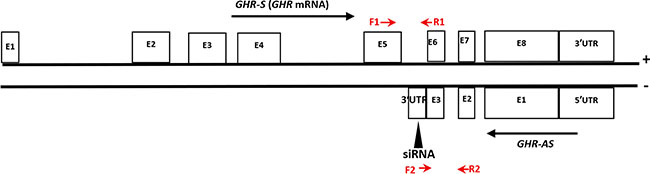
Figure 1: The relative positions of chicken GHR-AS and GHR-S transcripts in DNA. The exons (E) of GHR-S (GHR mRNA) were noted according to the GHR sequence characteristic deposited in Genbank (accession nos.NC_006127.4). Position of the siRNA oligonucleotides was indicated by black triangle. The qPCR primers of GHR-S (F1 and R1) and GHR-AS (F2 and R2) were labeled with red arrows.
To verify that GHR-AS was an antisense RNA transcribed from the opposite strand of GHR gene in chicken liver, by designing biotin-labeled sense or antisense probe, we conducted northern blot. Results indicated that the hybrid signals can be detected at about 4.5 kb position with the sense probe complemented with the GHR-AS sequence, and the GHR mRNA (GHR-S) also can be detected by the antisense probe (Supplementary Figure S2).
GHR-AS and GHR-S exhibit similar expression trends within the chicken development
We examined a panel of RNAs from female chicken tissues at 11 w of age for the presence of GHR-AS transcripts by quantitative RT-PCR. Chicken GHR-AS transcripts were expressed in all tissues examined, however, there were significant differences in their expression levels. GHR-AS were higher in the lung, pituitary and liver than in the other tissues, and it was the lowest in thigh muscle (Figure 2). GHR-S played important roles in liver and can form GH-GHR-IGFs signal pathway to affect animal growth [19]. Subsequently, we mainly analyzed the temporal characteristics of GHR-AS and GHR-S in chicken livers and leg muscles. The GHR-AS was expressed more lowly than the GHR-S in the two tissues at the indicated points (Figure 3). And the trends of GHR-AS expression displayed similarities to those of GHR-S in the two tissues. In liver, both GHR-AS and GHR-S ascended with the development of chicken from E10 to 3 w of age, however they were descendant in leg muscles (Figure 3).
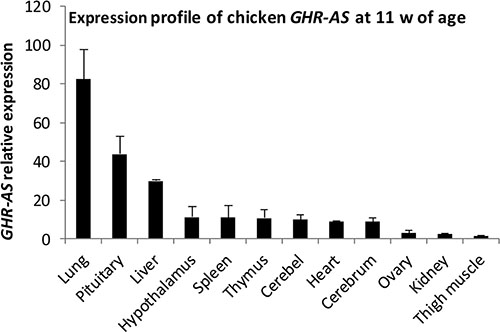
Figure 2: Characterization of the expression profiles of chicken GHR-AS.
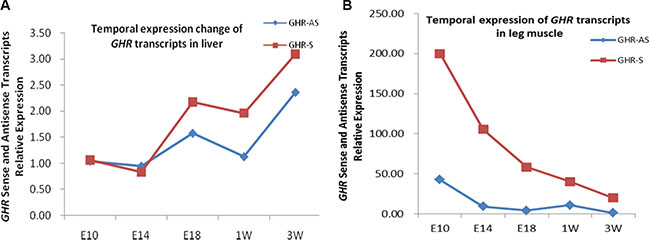
Figure 3: The temporal expression of chicken GHR-AS in liver and leg muscle.
GHR-AS regulates GHR-S in chicken LMH cells
To check whether the antisense RNA plays roles in GHR-S expression, we analyzed the changes of GHR-S, after silencing of GHR-AS using transcript-specific siRNA that specifically targeted 3′UTR of GHR-AS (Figure 1). We firstly measured GHR-AS levels in the chicken LMH cells treated with siGHR-AS. Results indicated that siGHR-AS caused visible reductions in GHR-AS levels in LMH cell lines. The GHR-S RNA expression decreased significantly after the knockdown of GHR-AS (Figure 4). To examine whether knockdown of GHR-AS affected expression of the nearby ZNF131 (zinc finger protein 131) gene, we analyzed RNA levels in parallel. Results indicated that knockdown of GHR-AS had no effect on ZNF131 gene expression at the transcriptional level (Figure 4), suggesting that GHR-AS specifically regulates the GHR-S.
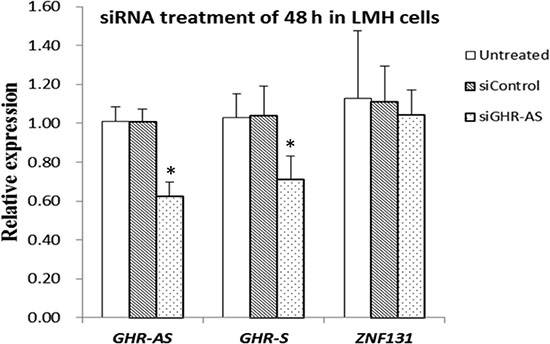
Figure 4: GHR-AS regulates GHR-S in LMH cells. qPCR analysis of the RNA levels of the GHR-S after siRNA treatment for 48 h, with the GAPDH gene as an internal control. Error bars represent the SEs of three independent experiments.
Silencing GHR-AS accompanied with the change of histone modifications in the GHR-S promoter
To identify a possible mechanism of GHR-AS mediated transcriptional inhibition, we analyzed histone modification status in the promoter regions of the GHR gene using ChIP assays. We firstly performed western blot to detect the specificity of H3K9me2 antibody and then conducted ChIP assays after knockdown of GHR-AS in LMH cells. Inactive gene indicator, H3K9me2, was increased in the GHR-S promoter regions in LMH cells (Figure 5A). These results indicated that silencing of GHR-AS could change the chromatin status from an open and active state to a closed and inactive state, thereby decreasing GHR-S expression. We further examined whether knockdown of GHR-AS affected the LMH cell cycles. Results demonstrated that more cells were blocked in G1 phase compared with the control group (Figure 5B).
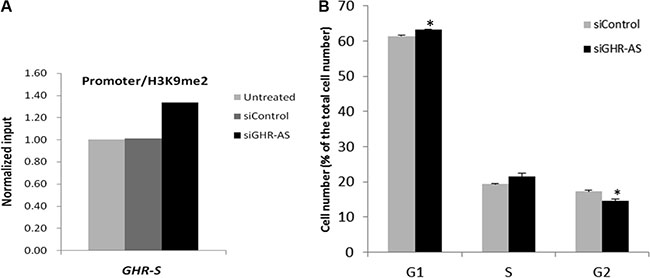
Figure 5: Silencing GHR-AS in LMH cell induced an increase of H3K9me2 in the promoter region of GHR gene and cell cycle arrest. (A) ChIP analysis of H3K9me2 enrichment in the GHR gene promoters. (B) Cell cycle analysis after silencing GHR-AS.
GHR-AS and GHR-S can form an RNA duplex via the last exon of GHR gene in vivo and in vitro
Having manifested the existence of GHR sense and antisense transcripts, it was of interest to know whether GHR-AS and GHR-S can form RNA duplex in vivo and in vitro. This question was examined by treating total RNA obtained from chicken liver and LMH cells with the single-strand specific nucleases RNase A. The remaining RNA was analyzed by RT-PCR using gene-specific primers which located in different overlapping regions of the both transcripts. Only primers within the exon 8 of GHR gene generated a PCR product (Figure 6C) while primers within the exon 6 and 7 of GHR gene did not (Figure 6B). Therefore, the overlapping part, exon 8 of GHR gene, of both transcripts was protected from degradation. In contrast, GHR mRNAs that is not protected by GHR antisense RNAs were totally degraded.
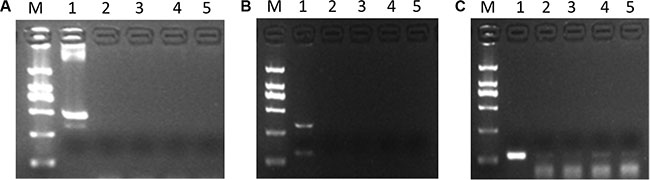
Figure 6: GHR-AS and GHR-S can form double strand RNAs at the last exon of GHR gene. (A) DNA contamination detection, using β-actin gene, demonstrated the RNAs were DNA-free. 1: positive control using DNA as template for PCR; 2,3: cDNA obtained from RNA of LMH cells; 4,5: cDNA obtained from RNA of chicken liver. (B) RT-PCR with gene specific primers, located in exon6 and 7 of GHR gene, suggested no production detected. 1: positive control using cDNA as template for PCR; 2,3: cDNA obtained from LMH cells RNA treated with RNase A; 4,5: cDNA obtained from chicken liver RNA treated with RNase A. (C) The GHR-S and GHR-AS RNA hybrids were detected by RT-PCR with gene specific primers, located in exon8 of GHR gene. The templates used were the same to (B).
DNA methylation affects both GHR-AS and GHR-S transcription
To determine whether the expression of GHR-AS and GHR-S was related to DNA methylation, we treated LMH cells with 5-AZA-d (C) (Sigma), the inhibitor of DNA methyltransferase, at the concentration of 0 μM, 0.1 μM, 1 μM and 10 μM for 48 h. Quantitative RT-PCR results indicated that both GHR-AS and GHR-S increased with the concentration ascending of 5-AZA-d(C) (Figure 7). Both GHR-AS and GHR-S increased at least 2 folds when adding 10 μM of 5-AZA-d(C) in DMEM which indicated that their expression were regulated by DNA methylation. Bioinformatic analysis indicated that CpG islands were rich in the predicted promoter of GHR-S but no CpG island was found in the promoter of GHR-AS (Supplementary Figure S3). The CpG islands were located in the regions spanning −67,909 to −66,733 bp of GHR relative to the A of the initiation codon (accession nos.NC_006127.4), and they were predicted using the Browser (http://www.urogene.org/cgi-bin/methprimer/methprimer.cgi). Further, we conducted DNA methylation analysis on the promoter of GHR-S in normal and dwarf WRR female chicken livers at 7 w of age. The CpG sites of the GHR-S gene promoter were highly methylated in normal WRR chicken livers, with the lower GHR-S and GHR-AS expressions while in dwarf WRR chicken, they were lowly methylated, with higher GHR-S and GHR-AS expressions (Table 1, Figure 8).
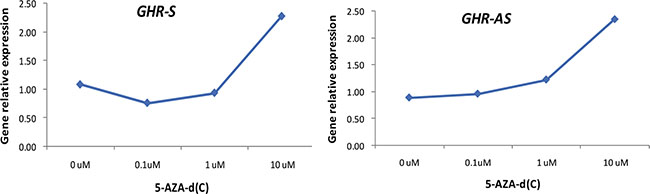
Figure 7: Both GHR-AS and GHR-S increased with the concentration ascending of 5-AZA-d(C) in LMH cells.
Table 1: DNA methylation comparison analyses on the promoter of GHR-S between normal and dwarf WRR female chicken livers
Chicken |
individual (1) |
individual (2) |
individual (3) |
X ± std |
P |
|---|---|---|---|---|---|
Normal WRR |
56.31% |
57.41% |
50.52% |
54.73% ± 3.71% |
0.0075 |
Dwarf WRR |
23.20% |
19.81% |
22.80% |
21.93% ± 1.86% |
The percentage represents the methylated CpG sites out of the total CpG number in the promoter of GHR-S.
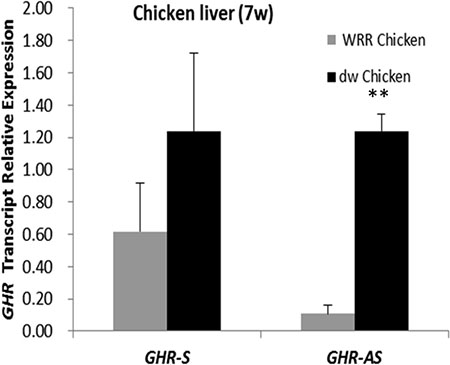
Figure 8: GHR-S and GHR-AS were highly expressed in dwarf chicken liver than in normal one WRR Chicken: normal chicken; dw Chicken: dwarf chicken.
DISCUSSION
GHR, expressed highly in the liver, was important in human and animal growth. In this study, we characterized GHR-AS, the Nats of chicken GHR gene, which were highly expressed in chicken lung, pituitary and liver at 11 w of age. According to the previous report [20], GHR-AS overlapped with GHR-S in a tail to tail manner. The transcription start site of GHR-AS resided in the 3′UTR of GHR-S while the AATAA motif and 3′polyadenylation tail of GHR-AS were located in the intron 5 of GHR gene. Generally, mRNAs were formed by splicing the exons and adding an extra polyadenylation tail. But GHR-AS was different from that, and it used the polyadenylation sequences from the intron 5 of GHR gene. Coincidentally, an alternative splicing of GHR sense transcript used the polyadenylation sequences from the exon of GHR gene as its tail to form a 0.7 kb transcript [18]. The GHR-AS and the 0.7 kb GHR-S transcripts, expressed in liver simultaneously, transcribed from the opposite directions and stopped at the site before they met. It may be related with RNA polymerase II complexes collide, known as “transcriptional interference” [21].
Chicken GHR gene was about 80 kb, with 7 polyadenylation sequences located in the DNA region, and possessed multiply transcripts. Both previous studies and this study indicated that chicken GHR gene can generate various transcripts via different ways, such as by using the alternative 5′UTR [17], different terminal signals [18] and transcribing from different directions. The function and significance of gene introns were controversial problems. Chicken GHR-AS used the intron 5 of GHR gene as its 3′UTR to end its transcription, which reminded us that the intron was functional sequences but not the junk DNA and it can be transcribed and used by organism when needed. In this study the intron 5 helped GHR gene to generate GHR-AS transcript. Hence, this study added a new case for explaining that the intron was functional sequence.
Most sense and antisense RNA pairs showed positive correlations in various tissues [22, 23]. But other studies showed that sense and antisense RNA pairs can be expressed in opposite trends [3, 4, 24, 25]. In chicken liver and leg muscle, GHR-AS and GHR-S exhibited positive correlations, and espectially in dwarf chicken they were simultanously increased compared with the normal one. The GHR-AS was expressed ascendently in liver with the chicken development but was expressed desendently in leg muscle that implied that GHR-AS may play different roles in different tissues. It may act activate roles in myoblast fiber formation in ealier chicken embyro development for its higher expression in E10, the important time for the myoblast fiber formation, than in 3 w of age [26].
Until now, most of studies suggested that antisense RNA can regulate sense RNA by the three ways. Firstly, sense and antisense transcripts may potentially interact through their complementary regions, resulting in double-stranded RNA structures that promote stability, transport, and/or translation of the sense transcript [27]. Secondly, antisense RNA can mediate the sense RNA suppressive or active by epigenetic regulation [28]. Thirdly, antisense RNA protects and transports the sense RNA from cell nucleus to the cytoplasm [29]. In this study, interfering GHR-AS caused GHR-S decreased, accompanied with the inactive gene indicator, H3K9me2, increasing in the GHR-S promoter regions in LMH cells. It indicated that GHR-AS could regulate GHR-S by modifying the histone epigenetic marker and hence affected the cell cycles. It needs to further study whether H3K4me2 can be affected by GHR-AS or not.
Several reports showed that the sense/antisense RNA duplex protected the sense RNA via their complementary sequences [30–32]. RNaseA experiment indicated that GHR-AS and GHR-S can form double strand RNAs at the last exon of GHR gene in vivo and in vitro which hinted they could act on each other through the region. In addition to the exon, the 5′UTR of GHR-AS was complemented with the 3′UTR of GHR-S. Our group has proved that chicken GHR-S 3′UTR was the target of Let-7b [33]. Consequently, the 3′UTR of GHR-S may be the competitive binding site for GHR-AS and Let-7b if the double strand GHR-S/AS RNAs can be formed in 3′UTR of GHR-S. Further, many long noncoding RNAs have been confirmed regulating gene mRNA stability via competing for binding to miRNA [34].
DNA methylation is generally considered as repressive epigenetic modification and is critical for the control of gene expression in animals. In this study, both GHR-S and GHR-AS were risen with the 5-AZA-d(C) concentration increasing, which indicated that the levels of GHR-S and GHR-AS can be affected by DNA methylation. Antisense transcripts have been proposed to cause DNA methylation. Tsix and AS1DHRS4, long antisense transcripts, induce DNA methylation at the promoter of the sense transcript Xist and DHRS4L2, respectively [3, 4]. Moreover, some antisense transcripts have coordinated expression with its counterpart sense gene promoter methylation [22]. In this study, methylation of GHR-S promoter was significantly higher in normal chicken liver, coupled with lower GHR-AS levels, while the situation was the opposite in dwarf chicken, exhibiting lower DNA methylation status in GHR-S promoter and higher GHR-AS expression. This implies that GHR-S and GHR-AS transcripts are not regulated independently but are regulated sufficiently to give simultaneous activation or inactivation of the gene pair. A cis-acting co-regulatory mechanism could exist that is working in a coordinated manner between sense and antisense transcription.
Many genes, such as BDNF [5], XIST [4] and DHRS4 [3], were bidirectionally transcribed in various species which implied that the transcribing directions of gene were conserved and the Nats play key roles in gene regulation and biological process. In human and animals, GHR genes are highly conserved in structure and function. GH exerts growth-promoting and metabolic effects in target tissues by binding to the transmembrane GHR and triggering enhanced GHR association with, and activation of, the cytoplasmic tyrosine kinase JAK2 [35, 36]. By checking website (http://www.ncbi.nlm.nih.gov/gene), we found the DNA region containing mouse GHR gene also exhibited transcribing in both directions. The mouse noncoding transcript Gm41254 showed an opposite direction to GHR mRNA (Figure 9). It suggested that GHR gene may be transcribed in double directions in various species and GHR-AS may act an important role in mammalian and avian growth.

Figure 9: The direction of mouse GHR gene and the non-coding transcript Gm41254. The figure was cited from website (http://www.ncbi.nlm.nih.gov/gene) and the arrows represent the directions of transcripts from different genes.
MATERIALS AND METHODS
Ethics statement
The Animal Care Committee of South China Agricultural University (Guangzhou, People’s Republic of China) approved this study (approval number SCAU#0017). The animals involved in this study were humanely sacrificed as necessary to ameliorate suffering.
Animals and samples
Animals
Recessive White Rock chickens, provided by Guangdong Nanhai Poultry Company (Guangdong, China), were used in this study. Two strains, including normal WRR and dwarf WRR chickens, were used to compare DNA methylation and the expression of the two transcripts. Dwarf chickens had a 1,773 bp deletion mutation at the last exon and in the 3′UTR of GHR gene. The weight of dwarf chickens was about 30% less than that of normal chickens. The two strains were fed under the same conditions to 7 w of age.
RNA extraction and cloning chicken GHR-AS by RACE technology
Total RNA was extracted from chicken liver at 7 w of age using the Trizol reagent (Invitrogen, USA) and was tested by agarose electrophoresis. Rapid amplification of the 3′ or 5′cDNA ends (RACE) was conducted using the RACE kit (Clontech, Japan) as described by the manufacturer, followed by nested PCRs of the cDNA copies. The PCR products were cloned and sequenced in the forward and reverse directions. All RACE primers were listed in Table 2.
Table 2: Primers used in this study
Application |
Primer name |
Primer sequence |
Tm (°C) |
|---|---|---|---|
3′RACE |
GHR-AS3(GSP1) |
GGGTCAATCCCTTTAATCTTT |
52 |
GHR-AS3(NGSP1) |
CAACAACTAAGAACCAGGGAAA |
||
5′RACE |
GHR-AS5(GSP1) |
TCCTCCTGTGCCAGTTCC |
52 |
GHR-AS5(NGSP1) |
GCGTGTTCAGGAGCAAAGCT |
||
GHR-AS full length PCR |
GHR-AS-F |
TTTTTTTTTTTTTTTTTT |
52 |
GHR-AS-R |
GTATGAAGAGTCCCAACCAAC |
||
GHR-AS quantitative PCR |
qGHR-AS-F |
TTGCTAATGTTTCTGTTCTGTG |
56.3 |
qGHR-AS-R |
GGGTCAATCCCTTTAATCTTT |
||
GHR-S quantitative PCR |
qGHR-S-F |
AGTCCGATCAAGACAACGTAC’ |
56.3 |
qGHR-S-R |
CTAAGAACCAGGGAAACTCG |
||
ZNF131 quantitative PCR |
ZNF131-F |
ATGTCCAAACTGCCACG |
56.3 |
ZNF131-R |
CACGCTGTTACAAACCTGA |
||
DNA contamination detection |
β-actin-F |
TCATTGTGCTAGGTGCCA |
50–60 |
β-actin-R |
CCTCTTCCAGCCATCTTT |
||
Internal control in qPCR analysis |
GAPDH-F |
TCCTCCACCTTTGATGCG |
50–60 |
GAPDH-R |
GTGCCTGGCTCACTCCTT |
||
Overlapping region(1) detection |
OL1-F |
TTTCCCTGGTTCTTAGTTGTTG |
55 |
OL1-R |
GGGTCAATCCCTTTAATCTTT |
||
Overlapping region(2) detection |
OL2-F |
GCGTGTTCAGGAGCAAAGCT |
60 |
OL2-R |
TGGGACAGGCATTTCCATACTT |
||
ChIP-qPCR |
GHR-S promoter-V1-F |
GCCTATAGCTGTCGCCTA |
60 |
GHR-Spromoter-V1-R |
GGAGAGCACTGTCTGATG |
||
DNA methylation analysis |
GHR- S promoter-BSP-F1 |
GAGGGCGGTCGTTGTTCG |
60 |
GHR- S promoter-BSP-R1 |
TTCTCCGCAACGCCCGCTCGTC |
||
GHR- S promoter-BSP-F2 |
GTGATATTTAAGTAAAATAAATTGT GGGAT |
66–56 (−0.5°C /cycle) |
|
GHR- S promoter-BSP-R2 |
ACTCRACRCATTCCTAAAAACRCR ACTAACC |
Northern blot analysis
Northern blotting was employed to identify the expression of GHR-AS RNA in the chicken livers. The sense and antisense RNA probes were designed to minimize non-specific hybridization against mRNAs following homology searches using the Basic Local Alignment Tool (BLAST; http://blast.ncbi.nlm.nih.gov/). The selected probe sequence for the detection of GHR-AS and GHR-S RNA were listed in Table 3. The sense and antisense probes were synthesized and labeled with biotin (Sangon Biotech Company, China). The total RNA, obtained from chicken livers, was DNA removed and transferred to a nylon membrane and UV-crosslinked. The membranes were pre-hybridized with alkaline-denatured salmon sperm DNA and then probed with biotin-labeled sense or antisense probe. The membranes were washed with wash buffer A for 10 min at 25°C and then washed twice with wash buffer B for 20 min at 60°C (Beyotime Biotechnology, China). Signals were visualized on a phosphorimager machine.
Table 3: The single strand probes, biotin labeled, used for Northern blot to detect GHR-Sand GHR-AS
Application |
Name |
Probe sequence |
|---|---|---|
GHR-S detection |
GHR-antisense probe |
5′-TCAATCCCTTTAATCTTTGGAACTGGCACAGGAGGAAAAATCA GCATTTTTAACCTTGGCTGTTTAGACAACAGGATTAAGATCGCTGT TACGGCCAGCCCACACACT -3′ |
GHR-AS detection |
GHR-sense probe |
5′-GCGTGTTCAGGAGCAAAGCTGTAACGAGGACACTTACTTCACC ACAGAAAGCCTTACCACTACCGGTATCAATCTTGGAGCTTCAATG GCAGAAACCCCAAGTATGG-3′ |
Reverse transcription and quantitative PCR
Reverse transcription reaction was performed using PrimeScript RT reagent Kit (Perfect Real Time) (Takara , Otsu, Japan) as described in the manufacturer’s protocol. Real time PCR was conducted in anBio-rad CFX96 Real-Time Detection system (Bio-rad, Hercules, CA, USA) using KAPA SYBR FAST qPCR Kit (KAPA Biosystems, Wobrun, MA, USA). All primers were listed in Table 2. Gene expression was normalized with GAPDH.
Cell lines and culture conditions
We cultured chicken LMH cells, gifted from China Harbin Veterinary Research Institute, in high-glucose Dulbecco’s modified Eagle’s medium (Gibco, Grand Island, NY, USA) with 10% (v/v) fetal bovine serum (Hyclone, Logan, UT, USA) and 0.2% penicillin/streptomycin (Invitrogen, Carlsbad, CA, USA).
RNA interference
LMH cells were transfected with 50 nM of siRNAs targeting GHR-AS using lip3000 kit (Invitrogen, USA) according to the manufacturer’s direction. Total RNA was harvested 48 h later for Real-time quantitative PCR analysis. The siRNA sequence was listed in Table 4.
Table 4: siRNA targeting chicken GHR-AS
siRNA name |
Sequence (5′–3′) |
|
|---|---|---|
Sense |
Antisense |
|
siRNA-GHR-AS (siGHR-AS) |
GCAUCUCUGUUGCAUCUUUTT |
AAAGAUGCAACAGAGAUGCTT |
Negative control (siControl) |
UUCUCCGAACGUGUCACGUTT |
ACGUGACACGUUCGGAGAATT |
Cell cycle distribution analysis
In order to determine the effects of GHR-AS on the cell cycle distribution, the cells were collected 48 h after siRNA transfection. The cells were washed twice with ice cold PBS and fixed with 70% ice-cold ethanol overnight at −20°C until further processing. After incubation in 50 μg/mL propidium iodide (Sigma Life Science, St. Louis, MO, USA) containing 10 μg/mL RNase A (Takara, Otsu, Japan) and 0.2% (v/v) Triton X-100 (Sigma) for 30 min at 4 °C, the cells were analyzed using a FACSAriaII flow cytometer (BD Biosciences, San Jose, CA, USA) and ModFit Lt 4.1 software (Verity Software House, Topsham, ME, USA).
Characterization of double-stranded RNA
Total RNA was treated with DNase I (Takara, Otsu, Japan) in DNase I buffer according to the manufacturer’s instructions until no traces of DNA could be detected by PCR analysis. DNA-free total RNA was digested with RNase A (Takara, Otsu, Japan) at a terminal concentration of 20 ng/μL at 37°C for 1 h according to the manufacturer’s instructions. Following the RNase A protection assay, we used RT-PCR to detect duplex formation by gene-specific primers located in different overlapping regions (the primers are listed in Table 1).
Chromatin immunoprecipitation (ChIP) assays for histone 3 lysine 9 methylation
Standard ChIP assays were performed using the Pierce™ AgaroseChIP Kit (Thermo Fisher Scientific, USA) according to the manufacturer’s instructions. LMH cells were harvested after transfected with siRNAs for 48 h to reach 90% confluence. The cross-linking between the nuclear protein and genomic DNA was performed in the growth medium with 1% formaldehyde supplementation at room temperature for 10 min. The harvested cells were lysed with Lysis Buffer I. Chromatin was digested with MNase for 15 min at 37°C and incubated with dimethylated H3K9 antibodies (Abcam, Cambridge, UK) or normal rabbit IgG overnight at 4°C. The antibody-chromatin complex beads were captured by Protein A/G agarose. DNA was separated from the beads and eluted using IP Elution Buffer for 40 min at 65°C. The DNA was further purified using a DNA Clean-up Column. Immunoprecipitated DNA was analyzed using real-time PCR, with gene-specific primers for GHR-S promoter, normalized by input DNA.
Bisulfite sequencing for DNA methylation analysis
Genomic DNA of chicken livers was extracted with a DNA tissue kit (Omega, USA), and the bisulfite conversion reaction was performed using an EpiTect Bisulfite kit (Qiagen, Germany) according to the manufacturer’s instructions. PCR amplification of bisulfate-treated DNA was respectively carried out with methylated site primers (listed in Table 1). The amplified products were cloned and sequenced. For each PCR product, more than eight clones were sequenced to analyze the GHR gene methylation status in the promoter region.
Statistical analysis
Data were processed using the statistical software package SAS 9.1.3 (SAS Institute Inc., NC) and expressed as the mean ± SE. Variance analysis was completed using a GLM procedure. P < 0.05 was considered a significant difference between the groups.
ACKNOWLEDGMENTS AND FUNDING
The authors would like to thank Prof. Yunfeng Wang and Yan Zhao (Harbin Veterinary Research Institute, Chinese Academy of Agricultural Science, Heilongjiang, China) for donating us LMH cells. This work was supported by the Chinese Postdoctoral Science Foundation (2013T60808), the National Natural Science Foundation (31301958, 31672412), the Guangdong Provincial Natural Science Foundation (S2013010013382) and the China Agriculture Research System (CARS-42-G05).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Chen J, Sun M, Kent WJ, Huang X, Xie H, Wang W, Zhou G, Shi RZ, Rowley JD. Over 20% of human transcripts might form sense-antisense pairs. Nucleic Acids Res. 2004; 32:4812–20.
2. Li JT, Zhang Y, Kong L, Liu QR, Wei L. Trans-natural antisense transcripts including noncoding RNAs in 10 species: implications for expression regulation. Nucleic Acids Res. 2008; 36:4833–44.
3. Li Q, Su Z, Xu X, Liu G, Song X, Wang R, Sui X, Liu T, Chang X, Huang D. AS1DHRS4, a head-to-head natural antisense transcript, silences the DHRS4 gene cluster in cis and trans. Proc Natl Acad Sci USA. 2012; 109:14110–5.
4. Do JT, Han DW, Gentile L, Sobek-Klocke I, Stehling M, Scholer HR. Enhanced reprogramming of Xist by induced upregulation of Tsix and Dnmt3a. Stem Cells. 2008; 26:2821–31.
5. Modarresi F, Faghihi MA, Lopez-Toledano MA, Fatemi RP, Magistri M, Brothers SP, van der Brug MP, Wahlestedt C. Inhibition of natural antisense transcripts in vivo results in gene-specific transcriptional upregulation. Nat Biotechnol. 2012; 30:453–9.
6. Williams T, Fried M. A mouse locus at which transcription from both DNA strands produces mRNAs complementary at their 3′ ends. Nature. 1986; 322:275–9.
7. Zuniga Mejia Borja A, Meijers C, Zeller R. Expression of alternatively spliced bFGF first coding exons and antisense mRNAs during chicken embryogenesis. Dev Biol. 1993; 157:110–8.
8. Zhang L, Li Y, Xie X, Xu H, Xu Z, Ma J, Li B, Lin S, Nie Q, Luo Q, Zhang X. A Systematic Analysis on mRNA and MicroRNA Expression in Runting and Stunting Chickens. PLoS One. 2015; 10:e0127342.
9. Chen C, Wei R, Qiao R, Ren J, Yang H, Liu C, Huang L. A genome-wide investigation of expression characteristics of natural antisense transcripts in liver and muscle samples of pigs. PLoS One. 2012; 7:e52433.
10. Yue Y, Guo T, Liu J, Guo J, Yuan C, Feng R, Niu C, Sun X, Yang B. Exploring Differentially Expressed Genes and Natural Antisense Transcripts in Sheep (Ovis aries) Skin with Different Wool Fiber Diameters by Digital Gene Expression Profiling. PLoS One. 2015; 10:e0129249.
11. Chan J, Atianand M, Jiang Z, Carpenter S, Aiello D, Elling R, Fitzgerald KA, Caffrey DR. Cutting Edge: A Natural Antisense Transcript, AS-IL1alpha, Controls Inducible Transcription of the Proinflammatory Cytokine IL-1alpha. J Immunol. 2015; 195:1359–63.
12. Taylor ER, Seleiro EA, Brickell PM. Identification of antisense transcripts of the chicken insulin-like growth factor-II gene. J Mol Endocrinol. 1991; 7:145–54.
13. Watanabe T, Totoki Y, Toyoda A, Kaneda M, Kuramochi-Miyagawa S, Obata Y, Chiba H, Kohara Y, Kono T, Nakano T, Surani MA, Sakaki Y, et al. Endogenous siRNAs from naturally formed dsRNAs regulate transcripts in mouse oocytes. Nature. 2008; 453:539–43.
14. Mihola O, Forejt J, Trachtulec Z. Conserved alternative and antisense transcripts at the programmed cell death 2 locus. BMC Genomics. 2007; 8:20.
15. Godowski PJ, Leung DW, Meacham LR, Galgani JP, Hellmiss R, Keret R, Rotwein PS, Parks JS, Laron Z, Wood WI. Characterization of the human growth hormone receptor gene and demonstration of a partial gene deletion in two patients with Laron-type dwarfism. Proc Natl Acad Sci USA. 1989; 86:8083–7.
16. Agarwal SK, Cogburn LA, Burnside J. Dysfunctional growth hormone receptor in a strain of sex-linked dwarf chicken: evidence for a mutation in the intracellular domain. J Endocrinol. 1994; 142:427–34.
17. Lau JS, Yip CW, Law KM, Leung FC. Cloning and characterization of chicken growth hormone binding protein (cGHBP). Domest Anim Endocrinol. 2007; 33:107–21.
18. Oldham ER, Bingham B, Baumbach WR. A functional polyadenylation signal is embedded in the coding region of chicken growth hormone receptor RNA. Mol Endocrinol. 1993; 7:1379–90.
19. Mavalli MD, DiGirolamo DJ, Fan Y, Riddle RC, Campbell KS, van Groen T, Frank SJ, Sperling MA, Esser KA, Bamman MM, Clemens TL. Distinct growth hormone receptor signaling modes regulate skeletal muscle development and insulin sensitivity in mice. J Clin Invest. 2010; 120:4007–20.
20. Lapidot, M. Pilpel Y. Genome-wide natural antisense transcription: coupling its regulation to its different regulatory mechanisms. EMBO Rep. 2006; 7:1216–22.
21. Zghidi-Abouzid O, Merendino L, Buhr F, Malik Ghulam M, Lerbs-Mache S. Characterization of plastid psbT sense and antisense RNAs. Nucleic Acids Res. 2011; 39:5379–87.
22. Watanabe Y, Numata K, Murata S, Osada Y, Saito R, Nakaoka H, Yamamoto N, Watanabe K, Kato H, Abe K, Kiyosawa H. Genome-wide analysis of expression modes and DNA methylation status at sense-antisense transcript loci in mouse. Genomics. 2010; 96:333–41.
23. Su WY, Li JT, Cui Y, Hong J, Du W, Wang YC, Lin YW, Xiong H, Wang JL, Kong X, Gao QY, Wei LP, Fang JY. Bidirectional regulation between WDR83 and its natural antisense transcript DHPS in gastric cancer. Cell Res. 2012; 22:1374–89.
24. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, Chang HY. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007; 129:1311–23.
25. Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell. 2008; 32:232–46.
26. Luo W, Wu H, Ye Y, Li Z, Hao S, Kong L, Zheng X, Lin S, Nie Q, Zhang X. The transient expression of miR-203 and its inhibiting effects on skeletal muscle cell proliferation and differentiation. Cell Death Dis. 2014; 5:e1347.
27. Mahmoudi S, Henriksson S, Corcoran M, Mendez-Vidal C, Wiman KG, Farnebo M. Wrap53, a natural p53 antisense transcript required for p53 induction upon DNA damage. Mol Cell. 2009; 33:462–71.
28. Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP, Cui H. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature. 2008; 451:202–6.
29. Johnsson P, Ackley A, Vidarsdottir L, Lui WO, Corcoran M, Grander D, Morris KV. A pseudogene long-noncoding-RNA network regulates PTEN transcription and translation in human cells. Nat Struct Mol Biol. 2013; 20:440–6.
30. Matsui K, Nishizawa M, Ozaki T, Kimura T, Hashimoto I, Yamada M, Kaibori M, Kamiyama Y, Ito S, Okumura T. Natural antisense transcript stabilizes inducible nitric oxide synthase messenger RNA in rat hepatocytes. Hepatology. 2008; 47:686–97.
31. Farnebo M. Wrap53, a novel regulator of p53. Cell Cycle. 2009; 8:2343–6.
32. Beltran M, Puig I, Pena C, Garcia JM, Alvarez AB, Pena R, Bonilla F, de Herreros AG. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008; 22:756–69.
33. Lin S, Li H, Mu H, Luo W, Li Y, Jia X, Wang S, Jia X, Nie Q, Li Y, Zhang X. Let-7b regulates the expression of the growth hormone receptor gene in deletion-type dwarf chickens. BMC Genomics. 2012; 13:306.
34. Wang ZD, Shen LP, Chang C, Zhang XQ, Chen ZM, Li L, Chen H, Zhou PK. Long noncoding RNA lnc-RI is a new regulator of mitosis via targeting miRNA-210–3p to release PLK1 mRNA activity. Sci Rep. 2016; 6:25385.
35. Zhang Y, Jiang J, Kopchick JJ, Frank SJ. Disulfide linkage of growth hormone (GH) receptors (GHR) reflects GH-induced GHR dimerization. Association of JAK2 with the GHR is enhanced by receptor dimerization. J Biol Chem. 1999; 274:33072–84.
36. Argetsinger LS, Campbell GS, Yang X, Witthuhn BA, Silvennoinen O, Ihle JN, Carter-Su C. Identification of JAK2 as a growth hormone receptor-associated tyrosine kinase. Cell. 1993; 74:237–44.

