INTRODUCTION
Hepatocellular carcinoma (HCC) poses a substantial threat to public health, particularly in less developed regions [1]. More than 50% of HCC cases occur in China [2]. It has been estimated that 75–85% of HCC cases in China are attributable to chronic HBV infection. However, only a minority of chronic HBV carriers eventually develop HCC [3], thus highlighting the importance of genetic susceptibility in HBV-related HCC.
Different genotypes of HBV are defined when the entire genome displays a sequence divergence of > 8%. HBV genotypes have distinct geographic distributions, and the incidence of HCC also varies in different regions of the world [4]. HBV genotypes have been found to differ in terms of clinical liver diseases and disease outcomes [5, 6]. However, because of small sample sizes, low success rates of HBV typing and different study designs, the previously reported effects of HBV genotypes on the outcomes of persistent HBV infections have varied greatly.
Zinc ribbon domain-containing 1 (ZNRD1) is a zinc finger-related protein cloned from human leukocyte antigen (HLA) [7]. HLA is an important control gene for immune characteristics, and previous data have demonstrated that HLA polymorphisms exhibit certain correlations with the outcome of HBV infections and the development of HCC [8, 9]. Additionally, ZNRD1 has been analyzed as a host cellular factor that influences virus replication [10]. Interestingly, as a transcription-associated gene, ZNRD1 has recently been found to be a novel negative modifier in carcinogenesis [11, 12]. These findings highlight the importance of ZNRD1 in the development of HBV-related HCC.
ZNRD1 is located on chromosome 6p21.3. There is a long non-coding RNA (lncRNA) ZNRD1-AS1 in the upstream region of ZNRD1. Using bioinformatic analyses, we have previously determined that three SNPs (rs3757328, rs6940552 and rs9261204) in ZNRD1-AS1 may be expression quantitative trait loci (eQTLs) for ZNRD1 (http://www.regulomedb.org) [13, 14]. The significant associations of the three ZNRD1 eQTLs SNPs in ZNRD1-AS1 with the risks of both chronic HBV infection and HCC have been tested in our previous study. Our previous in vitro experiments have also demonstrated that ZNRD1 knockdown inhibits the expression of HBV mRNA and promotes the proliferation of HepG2.2.15 cells [15]. Hence, we hypothesized that ZNRD1 eQTL SNPs may influence the HCC risks associated with HBV genotypes. To test this hypothesis, we performed a case-control study to evaluate the effects of the HBV genotype, ZNRD1 eQTL SNPs and their interactions on HCC risk.
RESULTS
The demographic characteristics of the 1,507 HBV-related HCC patients and the 1,560 persistent HBV carriers have been summarized previously [16]. There were no significant differences in the distributions of age or gender between the HCC patients and the persistent HBV carriers (P = 0.835 and 0.687, respectively). The HBV genotypes B, C, BC (coinfection), and D were identified in the subjects of this study through nested multiplex PCR and sequencing, as previously described [16].
The genotype distributions of the SNPs rs3757328, rs6940552 and rs9261204 in the HCC cases and the persistent HBV carriers are described in Table 1. The observed genotype frequencies of the three SNPs in the HBV persistent carriers were all in Hardy-Weinberg equilibrium (P = 0.721 for rs3757328, P = 0.723 for rs6940552 and P = 0.971 for rs9261204). The logistic regression analysis with an additive genetic model indicated that the variant alleles rs6940552 and rs9261204 increased the host’s HCC risk compared with the persistent HBV carriers (adjusted OR = 1.16, 95% CI = 1.03-1.32 for rs6940552; adjusted OR =1.20, 95% CI = 1.06-1.35 for rs9261204). Moreover, a borderline significant association was observed between rs3757328 and the HCC risk (adjusted OR = 1.11, 95% CI = 0.96-1.28; Table 1).
Table 1: Associations between three SNPs and HBV-related HCC susceptibility
Genotype |
HCC patients |
HBV persistent carriers |
OR (95% CI) |
P |
|---|---|---|---|---|
(n = 1507) |
(n = 1560) |
|||
N (%) |
N (%) |
|||
rs3757328 |
||||
G/G |
1038 (71.9) |
1146 (73.8) |
1.00 |
|
G/A |
362 (25.1) |
375 (24.1) |
1.07 (0.90-1.26) |
|
A/A |
43 (3.0) |
33 (2.1) |
1.44 (0.91-2.29) |
|
Dominant |
1.10 (0.93-1.29) |
0.262 |
||
Recessive |
1.42 (0.90-2.25) |
0.134 |
||
Additive |
1.11 (0.96-1.28) |
0.150 |
||
rs6940552 |
||||
G/G |
958 (64.2) |
1048 (67.5) |
1.00 |
|
G/A |
461 (30.9) |
453 (29.2) |
1.11 (0.95-1.30) |
|
A/A |
73 (4.9) |
52 (3.4) |
1.54 (1.07-2.22) |
|
Dominant |
1.16 (1.00-1.35) |
0.055 |
||
Recessive |
1.49 (1.03-2.14) |
0.031 |
||
Additive |
1.16 (1.03-1.32) |
0.018 |
||
rs9261204 |
||||
A/A |
853 (58.3) |
976 (62.9) |
1.00 |
|
G/A |
521 (35.6) |
510 (32.8) |
1.17 (1.00-1.36) |
|
G/G |
88 (6.0) |
67 (4.3) |
1.51 (1.08-2.10) |
|
Dominant |
1.21 (1.04-1.40) |
0.011 |
||
Recessive |
1.42 (1.03-1.97) |
0.034 |
||
Additive |
1.20 (1.06-1.35) |
0.004 |
||
Combined genotypes (unfavorable genotypes carried) a |
||||
0 |
784 (55.5) |
971 (62.8) |
1.00 |
|
1-3 |
553 (39.2) |
514 (33.3) |
1.34 (1.15-1.56) |
<0.001 |
4-6 |
75 (5.3) |
61 (4.0) |
1.52 (1.07-2.16) |
0.019 |
Note: The logistic regression analyses were adjusted for age and gender.
a The combined genotypes were trichotomized according to the unfavorable genotypes carried (rs3757328 AA, rs6940552 AA and rs9261204 GG were considered to be unfavorable genotypes). 0 indicates the presence of no unfavorable genotypes, 1–3 indicating the presence of 1–3 unfavorable genotypes, and 4-6 indicates the presence of 4-6 unfavorable genotypes.
Next, we estimated the cumulative effects of the three SNPs on HBV-related HCC susceptibility by adding the numbers of variant alleles. As summarized in Table 1, the HCC risk significantly and dose-dependently increased with the number of variant alleles (adjusted OR =1.34, 95% CI =1.15-1.56 for one to three variant alleles and adjusted OR =1.52, 95% CI = 1.07-2.16 for four to six variant alleles; P for trend <0.001), compared with the wild-type (WT) subjects who were homozygous for the three SNPs (Table 1).
The combined effects of the three SNPs on HCC susceptibility were also evaluated by stratifying on the basis of age, gender and HBV genotype (Table 2). Consequently, similar association strengths were found in most of the subgroups (P > 0.05 for heterogeneity test). Interestingly, a stronger combined effect of the three SNPs on HCC risk was observed among the non-B groups (adjusted OR = 1.26, 95% CI = 1.05-1.50) compared with the B-related groups (adjusted OR = 0.89, 95% CI = 0.69-1.15; P = 0.029 for the heterogeneity test). Further interactive analysis detected a significant multiplicative interaction between the variant alleles and the HBV genotypes on HCC susceptibility (P = 0.030; Table 3). Crossover analysis suggested that the non-B groups with 0 alleles (i.e., rs3757328-G, rs6940552-G and rs9261204-A), 1–3 alleles, and 4–6 alleles were associated with significantly increased risks (adjusted OR = 7.02, 95% CI = 5.66-8.72, P < 0.001; adjusted OR = 9.17, 95% CI = 7.20-11.66, P < 0.001; and adjusted OR = 10.17, 95% CI = 6.33-16.34, P < 0.001, respectively) of chronic HBV infection, as compared with the most common combination (i.e., the B-related groups with “0” alleles; Table 3, Figure 1).
Table 2: Stratified analyses of the combined variant alleles and HCC susceptibility
Variables |
HCC susceptibility (0 / 1-3 / 4-6) |
|||
|---|---|---|---|---|
HCC patients |
HBV persistent carriers |
OR (95% CI) |
P* |
|
Age |
||||
≤53 |
476/351/42 |
510/313/34 |
1.04 (0.86-1.25) |
0.252 |
>53 |
308/202/33 |
461/201/27 |
1.23 (0.99-1.53) |
|
Gender |
||||
Male |
644/440/54 |
770/426/49 |
1.06 (0.91-1.25) |
0.133 |
Female |
140/113/21 |
201/88/12 |
1.39 (1.01-1.90) |
|
HBV Genotype |
||||
B-related† |
174/88/3 |
645/317/33 |
0.89 (0.69-1.15) |
0.029 |
Non-B¶ |
594/446/72 |
317/183/27 |
1.26 (1.05-1.50) |
|
Note: The logistic regression analyses were adjusted for age, gender and HBV genotype (excluding the stratified factor in each stratum) in the additive genetic model.
*P-value for the heterogeneity test.
†B-related genotypes including B and BC.
¶Non-B genotypes including C and D.
Table 3: Crossover analysis of the combined variant allele-HBV genotype interactions on HCC susceptibility
Variables |
HBV genotype |
HCC patients |
HBV persistent carriers |
OR (95% CI) |
P |
|---|---|---|---|---|---|
0 |
B-related† |
174 (12.6) |
645 (42.4) |
1 |
|
1-3 |
B-related† |
88 (6.4) |
317 (20.8) |
1.02 (0.76-1.36) |
0.916 |
4-6 |
B-related† |
3 (0.2) |
33 (2.2) |
0.34 (0.10-1.11) |
0.731 |
0 |
Non-B¶ |
594 (43.1) |
317 (20.8) |
7.02 (5.66-8.72) |
<0.001 |
1-3 |
Non-B¶ |
446 (32.4) |
183 (12.0) |
9.17 (7.20-11.66) |
<0.001 |
4-6 |
Non-B¶ |
72 (5.2) |
27 (1.8) |
10.17 (6.33-16.34) |
<0.001 |
Interaction |
aP = 0.030 |
||||
Note: The logistic regression analyses were adjusted for age and gender.
aP value for the multiplicative interaction.
†B-related genotypes including B and BC.
¶Non-B genotypes including C and D.
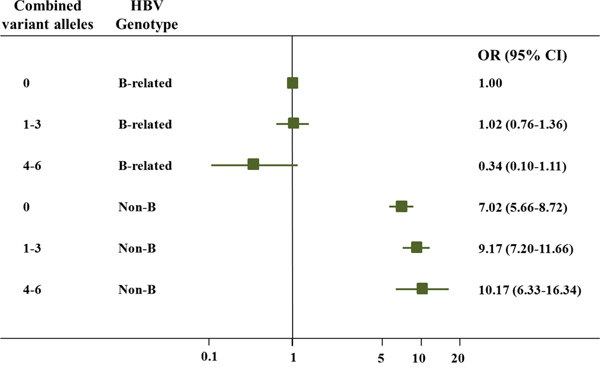
Figure 1: Crossover analysis of the effect of the three SNPs-HBV genotype interactions on HCC susceptibility. The non-B groups with 0 alleles (i.e., rs3757328-G, rs6940552-G and rs9261204-A), 1–3 alleles, and 4–6 alleles were associated with significantly increased risks (adjusted OR = 7.02, 95% CI = 5.66-8.72, P < 0.001; adjusted OR = 9.17, 95% CI = 7.20-11.66, P < 0.001; and adjusted OR = 10.17, 95% CI = 6.33-16.34, P < 0.001, respectively) of chronic HBV infection, as compared with the most common combination (i.e., the B-related groups with “0” alleles).
DISCUSSION
The development of HCC is a multistage process, and most HCCs arise from chronic hepatitis induced by HBV infection, particularly in China. The natural histories of hepatitis B virus infection are not uniform and are affected by several that include the HBV genotype [17]. Studies have found that genotype C is an independent risk factor for HCC development [18–20]. Viral genotype may be an independent and direct cause of hepatocarcinogenesis, which is the result of continuous necroinflammation [21]. In a previous study, we have also identified a significant association between the C genotype and the risk of HCC [16].
Previously, Cao et al have conducted a case-control study (1342 healthy controls, 327 HBV surface antigen (HBsAg) seroclearance subjects, and 1531 HCC patients) to assess the effects of HLA-DQ genetic polymorphisms, HBV genotypes, HBV mutations and their interactions on the risks of HCC. The results indicate that the rs9275319 variant genotype (GG) is significantly associated with an increased frequency of preS1 start codon mutations, which are HCC-risk mutations, in genotype C [22]. Recently, our group has detected significant interactions between HLA-DQ/DR rs9272105 and both of the HBV genotypes (P < 0.05 for each), as enabled by a large sample size and a high detection rate [16]. However, additional genes with potential interactions with HBV sequence variations need to be identified. Here, we detected significant interactions between the ZRND1 eQTL SNP and the HBV genotypes and found that these interactions were associated with HCC risk in a large sample.
Previous study had evidenced that ZNRD1, in close proximity to the HLA-A locus, was cloned from the human MHC class I region [23]. Our previous study has varified the significant association between two HLA-DQ/DR SNPs found by former GWAS studies and HCC risk (OR = 1.31, 95%CI = 1.18-1.45 for rs9272105; OR = 0.66, 95%CI = 0.56-0.78 for rs9275319) [16]. As a clone from MHC class I region, ZNRD1 eQTLs also appear the similar association with HCC risk. Furthermore, a series of studies have demonstrated that ZNRD1 variation affects host susceptibility to virus acquisition [10, 24]. In our previous work, we have found that the down-regulation of ZNRD1 reduced the expression level of HBV, which was a major risk factor for the progression of HBV infection; this finding was consistent with the protective roles of the variant alleles of the three SNPs on persistent HBV infection. Interestingly, the opposite findings have been reported in relation to tumorigenesis after persistent HBV infection [15]. Several studies have demonstrated that the up-regulation of ZNRD1 may inhibit DNA damage and enhance DNA repair capacity, which has been identified as a novel negative modifier in carcinogenesis [11, 12]. As we all kown, genes in the HLA region on chromosome 6 (such as HLA-DP and HLA-DQ, which were expressed as surface antigens and determine the immune response to virus infection) were associated with HBV infection and progression [25]. Which is more evidence that ZNRD1 and HLA play alike roles in cancer risk and virus infection.
Recently, a large number of lncRNAs have been identified by next-generation sequencing-based analyses in studies of complex diseases, and research on lncRNA genetic variants and biological function has been gaining increasing interest and emphasis in the literature [26, 27]. The dysregulation of multiple lncRNAs has been reported in HCCs, including HEIH and HULC [28, 29]. Moreover, our previous study has demonstrated that the genetic variants of ZNRD1-AS1 may exhibit different effects on HBV infection and HCC development [15]. The SNPs in lncRNAs may regulate the expressions of localized lncRNAs and subsequently affect the expression or function of nearby genes. Our current study further revealed that ZNRD1 eQTL SNPs may influence the HCC risks associated with the HBV genotypes.
The effect of ZNRD1 eQTLs was particularly evident in non-B genotype HBV-infected patients. Compared with HBV genotype B, genotype C was more prone to cause chronic HBV infection/inflammation and HCC [6, 30]. Our previous study also showed the similar conclusions. Results in Table 4 showed that the three eQTLs was associated with an increased risk of HCC. In addition, its interaction with HBV non-B genotype was greater associated with HCC risk (Table 3). In brief, the three eQTLs in ZNRD1-AS1 might predispose the HBV-infected patients to dysregulation of ZNRD1 which affect viral replication and interaction with HBV genotype, thus contributing to HBV-induced hepatocarcinogenesis.
Table 4: Primers and probes used in TaqMan allelic discrimination
Polymorphism |
Sequence (5’-3’) |
|
|---|---|---|
rs3757328 |
Primer |
F: TTTCTTGACTACTGCTAGCCTCACTT |
R: GGTGGTGGAACAGAGGAGCTT |
||
Probe |
G: FAM-TCTGGCAGGAGTCGA-MGB |
|
A: HEX-TCTGGCAGAAGTC-MGB |
||
rs6940552 |
Primer |
F: TACATAGCTAGAAGCAGCATCTATAATCC |
R: GCTTTAGAGTGTCATTGGTATGAACAG |
||
Probe |
G: FAM-TCACATAGGAATCACTG-MGB |
|
A: HEX-TCTCACATAAGAATCAC-MGB |
||
rs9261204 |
Primer |
F: TCCTTGCTCTGCTCTGCATTAT |
R: TGGTCTTTTAGTGGATGTTTTTGG |
||
Probe |
G: FAM-ATTTATTGGGACAGTCGTA-MGB |
|
A: HEX-ATGGGTAAGATTTATTGAGAC-MGB |
In summary, this large case-control study revealed that multiplicative interactions of ZNRD1-AS1 SNPs with HBV genotypes significantly affect HCC risk and suggested that HBV genotypes and ZNRD1-AS1 SNPs (rs3757328, rs6940552 and rs9261204) may serve as susceptibility biomarkers for the risk of HBV-related HCC. Additional well-designed prospective studies are warranted to validate and extend our findings.
MATERIALS AND METHODS
Patient samples
The ethical committees of all of the institutions involved in this study approved this case-control study. The subject enrollment has previously been described [16]. Briefly, the HCC cases were recruited from January 2006 to May 2014 at the First Affiliated Hospital of Nanjing Medical University (Nanjing, China), the Nantong Tumor Hospital (Nantong, China) and the Qidong Liver Cancer Institute (Qidong, China) in central and southern Jiangsu Province, China. The diagnosis of HCC was confirmed by a pathological examination and/or an alpha-fetoprotein elevation (> 400 ng/ml) combined with an imaging examination. HCV-related HCC patients were excluded. Ultimately, 1,507 HBV-related HCC patients agreed to participate in the study. Controls who were recognized as being persistent HBV carriers from 2009 to 2010 and were also from three cities in central and southern Jiangsu Province (48,417 subjects from Zhangjiagang, 43,563 subjects from Taixing and 57,192 subjects from Danyang) were recruited. Chronic HBV carriers were defined as subjects who were positive for both hepatitis B surface antigen (HBsAg) and antibodies against the hepatitis B core antigen (anti-HBc) but were negative for the HCV antibody (anti-HCV) at their second visit. Approximately 2,475 (5.11%), 3,413 (7.83%) and 5,587 (9.77%) chronic HBV carriers were identified from Zhangjiagang, Taixing and Danyang, respectively. In total, 1,560 cancer-free controls were randomly selected from these three cities and frequency-matched to the cases on the basis of age and sex. The demographic information of these selected controls, including age and gender, was collected via face-to-face interviews.
Serological testing
HBsAg, anti-HB, anti-HBc, and anti-HCV antibodies were detected in the subjects’ sera with enzyme-linked immunosorbent assays (Kehua Bio-Engineering Co., Ltd., Shanghai, China) according to the manufacturer’s instructions, as described previously [31].
Genotyping of the ZNRD1-AS1 SNPs
Genomic DNA was extracted from leukocyte pellets by traditional proteinase K digestion, phenol-chloroform extraction and ethanol precipitation. The SNPs rs3757328, rs6940552 and rs9261204 were genotyped using a TaqMan allelic discrimination assay on an ABI 7900 system (Applied Biosystems, La Jolla, CA). Information about the primers and probes is displayed in Table 4. The genotyping assays were performed blindly, and two blank (i.e., water) controls in each 384-well plate were used for quality control. More than 10% of the samples were randomly selected for repeat analyses, which yielded a 100% concordance rate. The genotyping success rates for these three SNPs were all above 98%.
Identification of the HBV genotypes
We extracted HBV DNA from 200 μl HBV-positive serum samples with High Pure Viral Nucleic Acid kits (Roche Diagnostics GmbH, Mannheim, Germany) following the manufacturer’s instructions. We used an improved nested multiplex PCR to identify the HBV genotypes. P1-S and P1-AS primers were used for the amplifications of the first round of the HBV genotypes A, B, C and D. Moreover, genotype primers were used for the amplifications of the A, B, C and D genotypes. The primers used are detailed in our previous study [16]. All genotyping assays were performed blindly with respect to the subjects’ case or control status, and two blank (i.e., water) controls in each 96-well plate were used for quality control. More than 10% of the samples were randomly selected for confirmation by DNA sequencing. The success rates of the determinations of the HBV genotypes and subgenotypes for each area were all above 96%.
Statistical analysis
The associations of the HBV genotypes with the SNP genotypes and HCC risk were estimated by computing the odds ratios (ORs) and their 95% confidence intervals (CIs) based on logistic regression analyses. The heterogeneities of the associations among the subgroups were assessed with the χ2-based Q test. All of the statistical analyses were performed with R software (version 2.13.0; the R Foundation for Statistical Computing), and P ≤ 0.05 from two-sided tests was considered to be statistically significant.
ACKNOWLEDGMENTS
We thank the ENCODE Projects for making the data public.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
GRANT SUPPORT
This work was supported in part by the National Natural Science Foundation of China (81302107), the Cheung Kong Scholars Programme of China, the National Science Foundation for Distinguished Young Scholars of China (81225020), Jiangsu Specially-Appointed Professor project, National Program for Support of Top-notch Young Professionals from the Organization Department of the CPC Central Committee, National Natural Science Foundation of China (81502861), National Hepatitis Prevention and Treatment Foundation TianQing Hepatitis Study Foundation (TQGB20140221) and the Priority Academic Program for the Development of Jiangsu Higher Education Institutions (Public Health and Preventive Medicine).
REFERENCES
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J and Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87-108.
2. Li GJ, Harrison TJ, Yang JY, Chen QY, Wang XY and Fang ZL. Combined core promoter mutations and pre-S deletion of HBV may not increase the risk of HCC: a geographical epidemiological study in Guangxi, China. Liver Int. 2013; 33:936-943.
3. Yang HI, Lu SN, Liaw YF, You SL, Sun CA, Wang LY, Hsiao CK, Chen PJ, Chen DS and Chen CJ. Hepatitis B e antigen and the risk of hepatocellular carcinoma. The New England journal of medicine. 2002; 347:168-174.
4. Anzola M. Hepatocellular carcinoma: role of hepatitis B and hepatitis C viruses proteins in hepatocarcinogenesis. Journal of viral hepatitis. 2004; 11:383-393.
5. Chan HL, Tse CH, Mo F, Koh J, Wong VW, Wong GL, Lam Chan S, Yeo W, Sung JJ and Mok TS. High viral load and hepatitis B virus subgenotype ce are associated with increased risk of hepatocellular carcinoma. J Clin Oncol. 2008; 26:177-182.
6. Zhang HW, Yin JH, Li YT, Li CZ, Ren H, Gu CY, Wu HY, Liang XS, Zhang P, Zhao JF, Tan XJ, Lu W, Schaefer S and Cao GW. Risk factors for acute hepatitis B and its progression to chronic hepatitis in Shanghai, China. Gut. 2008; 57:1713-1720.
7. Hong L, Zhao Y, Han Y, Guo W, Jin H, Qiao T, Che Z and Fan D. Mechanisms of growth arrest by zinc ribbon domain-containing 1 in gastric cancer cells. Carcinogenesis. 2007; 28:1622-1628.
8. Hu Z, Liu Y, Zhai X, Dai J, Jin G, Wang L, Zhu L, Yang Y, Liu J, Chu M, Wen J, Xie K, Du G, Wang Q, Zhou Y, Cao M, et al. New loci associated with chronic hepatitis B virus infection in Han Chinese. Nature genetics. 2013; 45:1499-1503.
9. Zhang H, Zhai Y, Hu Z, Wu C, Qian J, Jia W, Ma F, Huang W, Yu L, Yue W, Wang Z, Li P, Zhang Y, Liang R, Wei Z, Cui Y, et al. Genome-wide association study identifies 1p36.22 as a new susceptibility locus for hepatocellular carcinoma in chronic hepatitis B virus carriers. Nature genetics. 2010; 42:755-758.
10. Ballana E, Senserrich J, Pauls E, Faner R, Mercader JM, Uyttebroeck F, Palou E, Mena MP, Grau E, Clotet B, Ruiz L, Telenti A, Ciuffi A and Este JA. ZNRD1 (zinc ribbon domain-containing 1) is a host cellular factor that influences HIV-1 replication and disease progression. Clin Infect Dis. 2010; 50:1022-1032.
11. Hong L, Zhang Y, Liu N, Liu C, Zhi M, Pan Y, Lan M, Sun L and Fan D. Suppression of the cell proliferation in stomach cancer cells by the ZNRD1 gene. Biochemical and biophysical research communications. 2004; 321:611-616.
12. Zhang YM, Zhao YQ, Pan YL, Shi YQ, Jin XH, Yi H and Fan DM. Effect of ZNRD1 gene antisense RNA on drug resistant gastric cancer cells. World journal of gastroenterology. 2003; 9:894-898.
13. Veyrieras JB, Kudaravalli S, Kim SY, Dermitzakis ET, Gilad Y, Stephens M and Pritchard JK. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS genetics. 2008; 4:e1000214.
14. Stranger BE, Nica AC, Forrest MS, Dimas A, Bird CP, Beazley C, Ingle CE, Dunning M, Flicek P, Koller D, Montgomery S, Tavare S, Deloukas P and Dermitzakis ET. Population genomics of human gene expression. Nature genetics. 2007; 39:1217-1224.
15. Wen J, Liu Y, Liu J, Liu L, Song C, Han J, Zhu L, Wang C, Chen J, Zhai X, Shen H and Hu Z. Expression quantitative trait loci in long non-coding RNA ZNRD1-AS1 influence both HBV infection and hepatocellular carcinoma development. Molecular carcinogenesis. 2015; 54:1275-1282.
16. Wen J, Song C, Jiang D, Jin T, Dai J, Zhu L, An J, Liu Y, Ma S, Qin N, Liang C, Chen J, Jiang Y, Yang L, Liu J, Liu L, et al. Hepatitis B virus genotype, mutations, human leukocyte antigen polymorphisms and their interactions in hepatocellular carcinoma: a multi-centre case-control study. Scientific reports. 2015; 5:16489.
17. Ozaras R, Inanc Balkan I, Yemisen M and Tabak F. Epidemiology of HBV subgenotypes D. Clinics and research in hepatology and gastroenterology. 2015; 39:28-37.
18. Kao JH, Chen PJ, Lai MY and Chen DS. Hepatitis B genotypes correlate with clinical outcomes in patients with chronic hepatitis B. Gastroenterology. 2000; 118:554-559.
19. Fujie H, Moriya K, Shintani Y, Yotsuyanagi H, Iino S and Koike K. Hepatitis B virus genotypes and hepatocellular carcinoma in Japan. Gastroenterology. 2001; 120:1564-1565.
20. Chan HL, Hui AY, Wong ML, Tse AM, Hung LC, Wong VW and Sung JJ. Genotype C hepatitis B virus infection is associated with an increased risk of hepatocellular carcinoma. Gut. 2004; 53:1494-1498.
21. Ohkubo K, Kato Y, Ichikawa T, Kajiya Y, Takeda Y, Higashi S, Hamasaki K, Nakao K, Nakata K and Eguchi K. Viral load is a significant prognostic factor for hepatitis B virus-associated hepatocellular carcinoma. Cancer. 2002; 94:2663-2668.
22. Zhang Q, Yin J, Zhang Y, Deng Y, Ji X, Du Y, Pu R, Han Y, Zhao J, Han X, Zhang H and Cao G. HLA-DP polymorphisms affect the outcomes of chronic hepatitis B virus infections, possibly through interacting with viral mutations. Journal of virology. 2013; 87:12176-12186.
23. Fan W, Wang Z, Kyzysztof F, Prange C and Lennon G. A new zinc ribbon gene (ZNRD1) is cloned from the human MHC class I region. Genomics. 2000; 63:139-141.
24. An P, Goedert JJ, Donfield S, Buchbinder S, Kirk GD, Detels R and Winkler CA. Regulatory variation in HIV-1 dependency factor ZNRD1 associates with host resistance to HIV-1 acquisition. The Journal of infectious diseases. 2014; 210:1539-1548.
25. Xi-Lin Z, Te D, Jun-Hong L, Liang-Ping L, Xin-Hui G, Ji-Rong G, Chun-Yan G, Zhuo L, Ying L and Hui L. Analysis of HLA-DQB1 gene polymorphisms in asymptomatic HBV carriers and chronic hepatitis B patients in the Chinese Han population. International journal of immunogenetics. 2006; 33:249-254.
26. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, Ruan Y, Lassmann T, Carninci P, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome research. 2012; 22:1775-1789.
27. Li P, Ruan X, Yang L, Kiesewetter K, Zhao Y, Luo H, Chen Y, Gucek M, Zhu J and Cao H. A liver-enriched long non-coding RNA, lncLSTR, regulates systemic lipid metabolism in mice. Cell metabolism. 2015; 21:455-467.
28. Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, Shang JL, Gao CF, Zhang FR, Wang F and Sun SH. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology. 2011; 54:1679-1689.
29. Yang X, Xie X, Xiao YF, Xie R, Hu CJ, Tang B, Li BS and Yang SM. The emergence of long non-coding RNAs in the tumorigenesis of hepatocellular carcinoma. Cancer letters. 2015; 360:119-124.
30. Yang HI, Yeh SH, Chen PJ, Iloeje UH, Jen CL, Su J, Wang LY, Lu SN, You SL, Chen DS, Liaw YF and Chen CJ. Associations between hepatitis B virus genotype and mutants and the risk of hepatocellular carcinoma. Journal of the National Cancer Institute. 2008; 100:1134-1143.
31. Hu L, Zhai X, Liu J, Chu M, Pan S, Jiang J, Zhang Y, Wang H, Chen J, Shen H and Hu Z. Genetic variants in human leukocyte antigen/DP-DQ influence both hepatitis B virus clearance and hepatocellular carcinoma development. Hepatology. 2012; 55:1426-1431.

