INTRODUCTION
Long noncoding RNAs (lncRNAs) are defined as a large class of transcripts that are longer than 200 nt but lack protein-coding ability [1]. According to their genomic locations, lncRNAs can be categorized as follows: large intergenic noncoding RNAs (lincRNAs), intronic lncRNAs, antisense transcripts, bidirectional lncRNAs and pseudogenes [2]. LncRNAs have been shown to be involved in diverse cellular processes including the cell cycle and apoptosis [3–6], development and differentiation [7–9], X chromosome inactivation and gene imprinting [10–12]. Dysregulation of lncRNAs have been implicated in a number of diseases, including neurodegenerative diseases, cardiovascular diseases and cancer [7, 13–15]. Cancer is a group of complex diseases involving abnormal cell growth and tumorigenesis, and increasing evidence indicates that lncRNAs play key roles in cancer cells. However, lncRNA expression is often tissue-specific, and few lncRNAs with a consistent expression pattern in different types of cancer have been reported. One exception is the lncRNA MALAT1, whose expression is deregulated in various cancers. MALAT1 was first shown to be over-expressed in hepatoblastomas compared to hepatocellular carcinomas [16], and later investigations have indicated that MALAT1 is also upregulated in colon, breast, prostate and non-small cell lung cancer (NSCLC) [17–20].
The majority of lncRNAs regulate cellular processes at the epigenetic or transcriptional levels [21–24]. LncRNAs modulate DNA by affecting chromatin state as a tether, scaffold or in other roles while regulating gene transcription through monitoring the activity of specific transcription factors and polymerases. The most celebrated one is lincRNA HOTAIR which not only represses the HOXD locus gene transcription in trans but also regulates chromatin states and epigenetic inheritance by acting as a modular scaffold [21, 25]. However, little is known of the detailed mechanism by which lncRNAs exert their effects at the post-transcriptional level during tumorigenesis. LincRNA-p21 exerts its repression of mRNA translation by interacting with HuR [26], but in general, there are few examples of how lncRNAs influence post-transcriptional processes such as alternative splicing and translation.
Here, we investigated MT1JP, a lncRNA that appears to play a key regulatory role upstream of p53. By examining its expression by microarray analysis, we discovered that MT1JP has a lower expression level in four types of cancerous tissues than in the corresponding normal tissues of liver, lung, colon or gaster. We further demonstrated that MT1JP affects the translational activity of the tumor suppressor p53 through interaction with TIAR, a RNA-binding protein which has not previously been reported to associate with lncRNAs. By acting as a p53 regulator, MT1JP modulates a series of cancer hallmarks in which p53 participates, ranging from cell cycle, apoptosis, proliferation to migration and invasion. Together, our results suggest that MT1JP is a novel tumor suppressor, and broaden our knowledge of the development of human tumors. More generally, its consistent expression pattern in numerous tumor specimens indicates that MT1JP may be a significant biomarker of diagnosis with a potential in cancer therapy.
RESULTS
lncRNA MT1JP has lower expression level in cancer than in normal tissue
To profile the transcriptomes of tumor tissues and investigate the roles of lncRNAs in tumorigenesis, we applied an mRNA + lncRNA custom-designed microarray probing more than 70,000 human transcripts to 76 pairs of cancerous and paracancerous tissue specimens from four types of cancer, including liver, lung, colon and gaster. Bioinformatics analysis showed that 157 lncRNAs were consistently differentially expressed between cancerous and adjacent benign tissue in all types of samples [27]. Among these, we observed one that was down-regulated to less than 50% in all tumor samples compared to the matched normal tissues (Figure 1A), and this transcript has previously been labelled as MT1JP. As an additional test, we determined the abundance of MT1JP in another 29 pairs of tissue samples from liver, colon, lung and gaster cancer by quantitative real time polymerase chain reaction (qRT-PCR). In line with the microarray analysis, we found that MT1JP had considerably lower expression in almost all tumor samples (Figure 1B and Supplementary Figure S1A). Moreover, the RNA-seq data from human body map showed that MT1JP was detected in kinds of tissues including liver, lung and others (Supplementary Figure S1B).
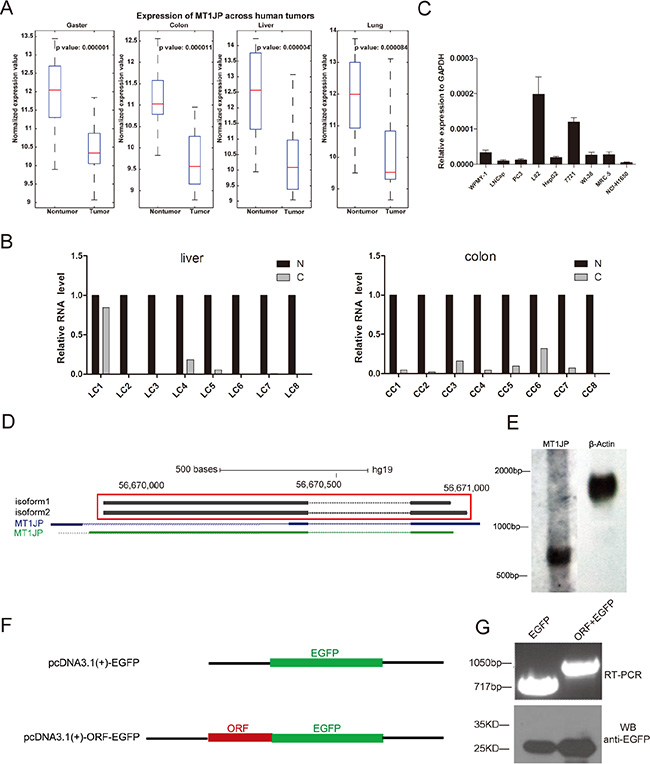
Figure 1: Identification of MT1JP as a tumor-associated lncRNA. (A) Box plots showing MT1JP expression levels in tumor and paired adjacent nontumorous tissues from gaster (n = 20 × 2), colon (n = 20 × 2), liver (n = 16 × 2) and lung (n = 20 × 2). (B) Relative expression of MT1JP in additional 16 pairs of normal (N) and cancer (C) tissue samples from liver and colon were determined by qRT-PCR. (C) MT1JP expression in 9 human cell lines derived from 3 different tissues. (D) Two different isoforms of MT1JP were identified in the L02 cell line by 5′ and 3′ RACE. (E) Northern blot of RNA from the L02 cell line, indicating the MT1JP transcript size. ACTB is used as a control. (F) Left panel: Illustration of plasmids for coding potential verification. Right panel: Analysis of the transfected cells by RT-PCR (top) and Western blot (lower). The fusion transcript of MT1JP ORF and EGFP is detected, but the fusion protein is not.
In order to select one or more cell lines for functional studies of this lncRNA, we examined the MT1JP expression in several cell lines derived from normal or cancer cells of three kinds of tissues (Figure 1C). The results of qRT-PCR analyses showed that MT1JP is expressed at higher levels in non-cancerous than in cancerous cell lines, consistent with the results from the tissue samples. Given that MT1JP had highest expression in the L02 cell line, we conducted the subsequent knock-down experiments in L02 cells, while over-expression experiments were carried out in the HepG2 and SMMC-7721 cell lines, which only express low level of endogenous MT1JP. All three cell lines are derived from liver tissues.
The lncRNA MT1JP resides on chromosome 16 in a cluster consisting of several homologous protein-coding genes of the metallothionein family. The MT1JP locus contains three regions that are highly conserved across a few of vertebrates, and is also annotated with several SNPs, CpG islands and clusters of DNase hypersensitive sites (Supplementary Figure S1C). Using 5′ and 3′ rapid amplification of cDNA ends (RACE) on RNA extracted from the L02 cell line, we showed that the mature MT1JP transcripts comes in two isoforms of 880 nt or 927 nt, both composed of two exons with a poly(A) tail attached (Figure 1D). The length of MT1JP we verified was nearly identical to one of the annotations in UCSC database (Figure 1D). In addition, northern blot analysis also identified MT1JP in the L02 cells, confirming the RACE results (Figure 1E). Since these two isoforms were almost identical, we did not distinguish each other when performing the subsequent experiments.
We then examined the coding potential of MT1JP using both in silico prediction and in vivo experiment. The in silico results obtained with four powerful computational tools for distinguishing coding and noncoding transcripts consistently showed that MT1JP has no ability to code a protein (Supplementary Table S1) [28–31] We nonetheless fused the only open reading frame (ORF) in the MT1JP transcript with EGFP (Figure 1F and Supplementary Figure S1D). By overexpressing the constructs in L02 cells, we found that while fused MT1JP ORF and EGFP could be expressed as a transcript, no fused protein was produced (Figure 1F). These results indicated that MT1JP is a long noncoding RNA which is down-regulated in cancer compared to adjacent normal tissues.
MT1JP modulates the cell cycle
To investigate which pathway MT1JP might be involved in, we conducted a Gene Ontology (GO) analysis of genes whose expression was strongly correlated (r2 > 0.25) to that of MT1JP in the normal and matched cancer tissues. The GO analysis identified significant enrichment for annotations related to the cell cycle (Figure 2A). Therefore, in order to explore the relationship between the cell cycle and the MT1JP expression level, we used the double thymidine method to synchronize the human hepatic cell line L02, and observed a significant increase in MT1JP expression after L02 cells were treated with double thymidine (Figure 2B). Furthermore, RNAi-mediated knockdown of MT1JP in L02 cells were performed with two different siRNA duplexes (Figure 2C). The down-regulation of MT1JP led to an increase in the proportion of cells in S phase and a decrease in G1 phase (Figure 2D). Inversely, overexpression of MT1JP in the liver hepatocellular carcinoma cell line HepG2 by transient transfection (Figure 2E) produced a higher percentage of cells in G1 phase but a lower percentage in S phase, when compared to control cells (Figure 2F), that is, the opposite phenotype of the knockdown experiment. Together, these results indicate that MT1JP has a significant effect on the G1/S check point, and that its ectopic expression can induce cell cycle arrest.
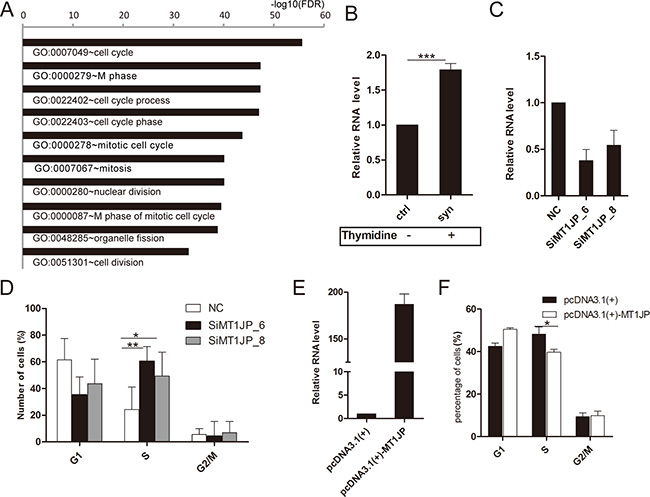
Figure 2: MT1JP acts as a cell cycle regulator. (A) Significantly enriched gene ontology (GO) terms of genes whose expression are correlated with MT1JP in tumor samples. (B) Relative abundance of MT1JP was measured by qRT-PCR before and after treating cell with thymidine. (C) Targeting MT1JP in L02 cells by two independent siRNAs led to depletion of the transcript as measured by qRT-PCR. (D) Effect of MT1JP knock-down on the fraction of L02 cells in G1, S and G2/M cell cycle phases. (E) After transfection of cells with negative control or MT1JP-overexpressed vectors, MT1JP levels were measured as in (B). (F) Relative number of L02 cells in each phase of the cell cycle after treating cell as in (E). Values are the mean and error bars represent standard deviation (± SD) in triplicate experiments. *p < 0.05, **p < 0.01, ***p < 0.001.
MT1JP is a tumor suppressor candidate
Given the significantly differential expression of MT1JP between cancerous and normal tissue samples (Figure 1A), we speculated that this lncRNA might not only regulate the cell cycle but also play important roles in other pathways involving in cell transformation. Toward this end, we performed Gene Set Enrichment Analysis (GSEA) on the above mentioned microarray data in Figure 1A. The GSEA indicated that MT1JP-related genes were enriched for several signatures involved in various kinds of cancers (Supplementary Figure S2).
To address whether MT1JP exert its effects as a tumor suppressor, we first investigated in more detail the role of MT1JP in tumor suppression. The EdU (5-ethynyl-2′-deoxyuridine) incorporation assay was performed to investigate the effects of MT1JP on cell proliferation, and results showed that cells treated with MT1JP-targeting siRNA displayed stronger proliferation ability than cells treated with non-targeting siRNAs and ectopic expression of MT1JP decreased the EdU incorporation (Figure 3A and Supplementary Figure S3A). Next, we examined the impact of MT1JP knockdown on apoptosis in L02 and HepG2 cells by flow cytometry analysis and detecting cleaved-caspase3 which is a key representation of apoptotic cells. In line with our expectation, RNAi of MT1JP decreased the percentage of cells undergoing apoptosis while overexpression of MT1JP increased this percentage (Figure 3B and Supplementary Figure S3B). Collectively, the lncRNA MT1JP plays a potential role in promoting apoptosis to restrain the growth of aberrant cells. Additionally, we applied the transwell assay to examine alterations in L02 cell migration and invasion after knocking down MT1JP, and found that treatment of cells with MT1JP specific siRNAs increased the migration and invasive capacity of L02 cells when compared to ones with wild-type cells (Figure 3C and 3D). These results suggested that MT1JP has an inhibitory effect on proliferation, migration and invasion of L02 cells.
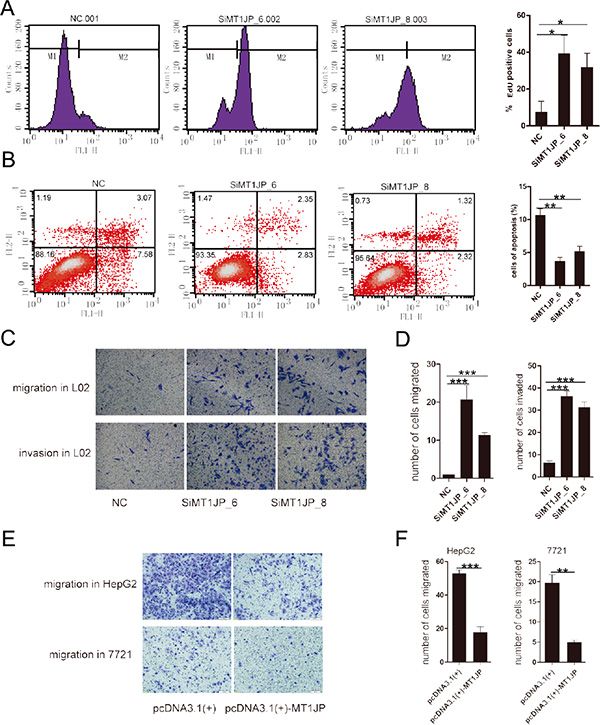
Figure 3: MT1JP is a potential tumor suppressor. (A) BrdU labeling assay indicates that knockdown of MT1JP with siRNAs promotes cell proliferation. (B) Knockdown of MT1JP decreases the percentage of apoptotic cells. The percentage of cells in each quadrant is indicated. (C and D) Transwell assays indicate that MT1JP knockdown increases L02 cell migration and invasion. (E and F) Transwell assays as in (C and D) suggest that overexpression of MT1JP decreases cell migration. All values are means, and error bars represent standard deviation (± SD) of triplicate experiments. **p < 0.01, ***p < 0.001.
To further investigate the effects of MT1JP on tumor inhibition, we constructed a plasmid for overexpression of MT1JP in HepG2 and SMMC-7721 cells, which normally express MT1JP at lower levels than L02 cells. HepG2 overexpressing MT1JP cells were subjected to the transwell assay and their migration ability and invasive capacity were determined. It turned out that ectopic expression of MT1JP in HepG2 and SMMC-7721 cells reduced cellular migration capacitiy (Figure 3E and 3F). These gain-of-function results derived from the low expression cells complemented to the ones from knockdown experiments done in MT1JP high expressed cells, and indicated that MT1JP plays an important role in tumor migration etc. Altogether, these results provide evidence that MT1JP is a modulator of cell cycle, apoptosis, and proliferation, and may play a role as a tumor suppressor.
MT1JP regulates the P53 signaling pathway
To elucidate the underlying molecular mechanisms of MT1JP function in the regulation of cell pathways, microarray analyses were performed to profile the transcriptome of L02 cells upon MT1JP knockdown. Two MT1JP-targeting siRNAs and negative control siRNA were transfected into L02 cells, and the cells were harvested after 24 hour (h), 48 h and 72 h separately. Transcriptome analyses showed major changes in the RNA expression profiles between 24 h and 48 h after transfection, but little changes beyond 48 h. In addition, we also identified 1650 genes (364 up-regulated and 1286 down-regulated) with more than 1.5-fold expression changes in the triplicate siRNA experiments 48 h after knockdown (Figure 4A and Supplementary Table S3). GSEA of these differentially expressed genes indicated enrichment for annotations related to neoplastic transformation, cancer and other tumor associated annotations (Figure 4B), which is consistent with the results as shown in Figures 2 and 3. In addition, the GSEA also showed that MT1JP had an effect on targets of p53, which belongs to the core of cell signaling pathway (Figure 4C). In the KEGG (Kyoto Encyclopedia of Genes and Genomes), p53 is annotated to be involved in apoptosis, cell cycle, angiogenesis, metastasis and other pathways (Figure 4D). To validate the finding that MT1JP regulates targets of p53, we transfected L02 cells with siRNAs targeting MT1JP, and determined the expression levels of several p53 targets by qRT-PCR. The experimental validation is consistent with the results of the GSEA, indicating that MT1JP regulates p53 target genes, for example GADD45a and p21, but has no effect on p27 which is not the member of p53 pathway (Figure 4E).
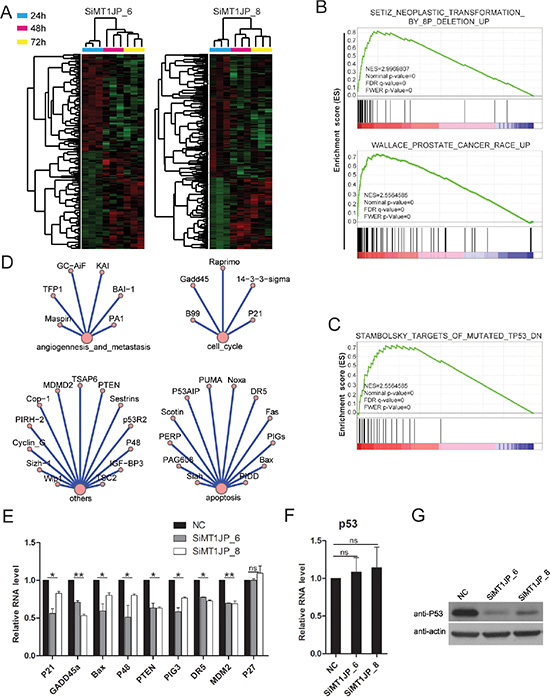
Figure 4: MT1JP regulates the p53 pathway. (A) Heatmap illustration of MT1JP knockdown with two independent siRNAs after 24 h, 48 h and 72 h in L02 cells. Each knockdown has three replicates. (B) GSEA results show that the genes that differentially expressed (1.5-fold) after MT1JP knockdown in L02 cells are enriched for functional annotations related to cancer. (C) GSEA results showing significant enrichment of p53 targets signature for differentially expressed-genes in MT1JP-downregualted L02 cells. (D) TP53-associated pathways annotated in KEGG. (E) Effect of MT1JP knockdown on the expression of p53 target genes (qRT-PCR). (F) Effect of MT1JP knockdown on the p53 mRNA level (qRT-PCR). (G) Western blot analysis of the p53 protein level after MT1JP knockdown. All values are means and error bars represent standard deviation (± SD) in triplicate experiments. ns means not significant.
Given that MT1JP may play a role in regulating the expression of p53 downstream targets, we examined the effect of MT1JP knockdown on p53 expression both at the mRNA and protein levels. Interestingly, MT1JP knockdown resulted in a decrease in the p53 protein level but had no significant effect on its mRNA level (Figure 4F).
All together, these results suggest that MT1JP may facilitate expression of tumor suppressor p53 at the post-transcriptional, but not at the transcriptional level, and in turn affect the downstream genes of p53.
MT1JP is largely cytoplasmic and interacts with TIAR
We next tried to investigate the mechanism by which MT1JP regulates gene expression during cell transformation. We first determined the subcellular distribution of MT1JP by RNA fluorescence in situ hybridization (RNA-FISH), and found that most of the MT1JP transcripts were located in the cytoplasm (Figure 5A). Moreover, we confirmed this observation by quantifying the expression ratio of MT1JP in cytoplasmic and nuclear fractions with qRT-PCR. The results showed that approximately 70% of MT1JP transcripts were found in the cytoplasm (Figure 5B).
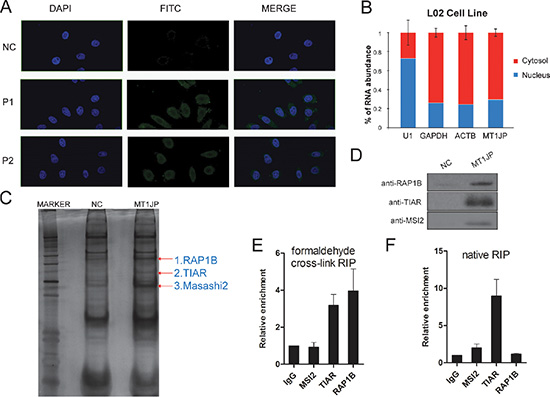
Figure 5: MT1JP is a cytoplasmic lncRNA which interacts with TIAR. (A) RNA fluorescence in situ hybridization (FISH) of MT1JP in L02 cells with two independent LNA probes (P1 and P2) targeting MT1JP. (B) MT1JP subcellular localization is detected through cellular fractionation. The relative abundance in each component is determined by qRT-PCR. (C) RNA pull-down of MT1JP-associated proteins using biotinylated MT1JP or lacZ DNA probes. Isolated proteins were resolved by SDS-PAGE followed by silver staining. The highlighted bands were analyzed by mass spectrometry. (D) Western blot analysis of the specific interaction between candidate proteins and MT1JP with MSI2, TIAR and RAP1B antibody after RNA pull-down as in (B). (E) Association between MT1JP and candidate proteins in formaldehyde cross-link RIP by qRT-PCR. (F) Relative MT1JP enrichment in native RIP (qRT-PCR). The enrichment of MT1JP in F and G is normalized to GAPDH (versus IgG). All values are means and error bars represent standard deviation (± SD) in triplicate experiments.
LncRNAs frequently exert their function through interactions and play roles through their association with proteins [22, 32, 33]. To test whether this might also be the case for MT1JP, we first sought to identify interacting proteins by RNA pull-down assay. Specifically, synthetic biotinylated MT1JP or LacZ antisense probes were incubated with L02 cell lysate, and co-precipitated proteins were then purified with streptavidin magnetic beads, followed by polyacrylamide gel electrophoresis and specifically enriched bands by MT1JP biotinylated probes were further analyzed by mass spectrometry. Three candidates were identified and two of them are Masashi 2 (MSI2) and TIA1-related protein (TIAR), both known to be RNA-binding proteins (Figure 5C), and the third one is Ras-related protein Rap-1b (RAP1B), a protein involved in the Ras pathway [34]. Further analysis by Western blot showed that MSI2, TIAR and RAP1B were pulled down with the MT1JP specific probe, but not with the control lacZ probe (Figure 5D). In addition, we found that the majority of TIAR were located in the cytoplasm by cell nucleocytoplasmic separation, which is similar with MT1JP (Supplementary Figure S4A).
To further validate the specificity of the association between MT1JP and the three identified proteins, we performed both native and formaldehyde cross-linked RNA immunoprecipitation (RIP) with antibodies against RAP1B, TIAR and MSI2. After antibodies had been incubated with the L02 cell lysate, RNA-protein complexes were precipitated with protein A/G beads, and associated RNAs were extracted. We verified the efficacy of the RIP by Western blot (Supplementary Figure S4B), and analyzed the enriched MT1JP content in each RIP sample by qRT-PCR. We observed that when using the formaldehyde cross-linking RIP approach, the enrichment of MT1JP in complexes precipitated with antibodies against TIAR and RAP1B was 3 and 5-fold more, respectively when compared with precipitates using control IgG (Figure 5E). In contrast, when using native RIP approach, in complexes precipitated with antibodies against TIAR and MSI2 the enrichment of MT1JP was 10 and 2-fold more, respectively, when compared with control IgG (Figure 5F). These results implied that TIAR interacts strongly with MT1JP, whereas RAP1B and MSI2 associate more weakly with MT1JP. Collectively, MT1JP interacts specifically with RNA-binding protein TIAR.
MT1JP and TIAR regulates p53 at the post-transcriptional level
Several studies have suggested that a number of RNA-binding proteins are key factors in mRNA translational regulation [35, 36]. The data presented above further suggest that MT1JP interacts directly with TIAR and that MT1JP affects p53 expression, but not at the transcriptional level. Based on these indications, we speculated that MT1JP may form a complex with TIAR which affects the expression of p53 at the translational level. One possibility in such a model is that TIAR also interacts with the p53 mRNA. To test this speculation, we carried out TIAR RIP experiment and detected that the p53 mRNA was precipitated by an antibody against TIAR, whereas the negative control p27 mRNA was not (Figure 6A). To further validate our hypothesis, we treated L02 cell with two specific siRNAs to knockdown TIAR, and observed that down-regulation of TIAR had no effect on the p53 mRNA level (Figure 6B). However, knockdown of TIAR decreased the expression of the p53 protein (Figure 6C). Moreover, p53 protein was reduced when we knocked down MT1JP and TIAR simultaneously, which is consistent with knockdown of MT1JP and TIAR separately (Figure 6D). Nevertheless, the p53 protein could hardly be rescued by overexpression of MT1JP with TIAR repressed (Figure 6E). This result demonstrates that MT1JP regulates p53 protein by associating with TIAR.
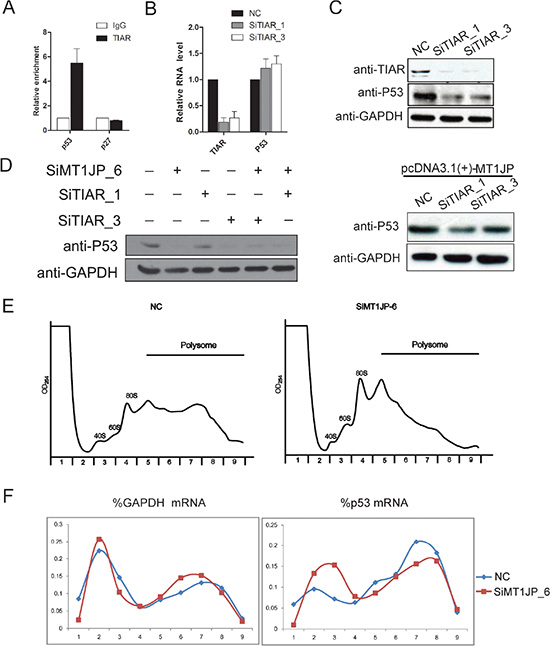
Figure 6: MT1JP and TIAR regulate p53 expression at the post-transcriptional level. (A) Binding of p53 mRNA to TIAR in L02 cells, shown by RNA immunoprecipitation and qRT-PCR normalized with GAPDH (mean ± SD, n = 3, versus IgG). (B) Relative abundance of TIAR and p53 mRNAs was measured by qRT-PCR after TIAR knockdown (mean ± SD, n = 3). (C) Western blot analysis of the p53 protein level after TIAR knockdown. (D) Western blot analysis of the p53 protein level after MT1JP or/and TIAR knockdown. (E) Western blot analysis of the p53 protein level after MT1JP overexpression and TIAR knockdown. (F) Sucrose gradient fractionation of polysomes from L02 cells after MT1JP knockdown. (G) The polysomal distribution of the p53 and GAPDH mRNAs was determined by qRT-PCR analysis of RNA in gradient fractions, and represented as percentage of total RNA in the fractions. Values represent mean of three independent experiments.
To directly test the possibility that MT1JP promotes the translation of p53 mRNA, we utilized polysome fractionation to evaluate the translation status of p53 mRNA in L02 cells. Centrifuged through a 10%–50% sucrose gradient, the lightest components sedimented at the top (Fractions 1 and 2), ribosomal subunit (40S and 60S) and monosomes (80S) in fractions 3 and 4, smaller polysomes in fractions 5–6, and larger polysomes in fractions 7 and 9 (Figure 6E). The translational efficiency of an mRNA is reflected by its distribution within the sucrose gradient. Efficiently translated RNAs are associated with larger polysomes, and inefficiently translated RNAs with smaller polysomes or monosomes. After knockdown of MT1JP, the content of p53 mRNA in larger polysomes was reduced while it increased in smaller polysomes and non-polysomes, whereas the content of GAPDH mRNA in larger polysomes was increased (Figure 6F). Together, these results suggested that altering the MT1JP level affects the polysome distribution of p53 mRNA, thereby influencing p53 translational efficiency.
In addition, we also investigated the relationship between MT1JP and TIAR. Neither the TIAR mRNA nor the protein level was influenced by the knockdown of MT1JP. However, the expression of MT1JP was increased after knockdown of TIAR (Supplementary Figure S4A). TIAR has been reported as a factor controlling RNA turn-over [37], and we therefore treated L02 cells with the RNA polymerase II inhibitor α-amanitin. The de novo synthesis of MT1JP was blocked in L02 cells treated with α-amanitin. The knockdown of TIAR increased the stability of MT1JP when compared to control cells (Supplementary Figure S4B). All these data indicated that TIAR promotes MT1JP decay, thereby affecting the expression level of MT1JP.
DISCUSSION
An increasing number of long noncoding RNAs has been discovered along with the advances in next-generation sequencing technology, however, our knowledge of their biological roles are limited. In this study, we identified a lncRNA MT1JP which appears to play a significant role in the inhibition of tumors. Our data show that MT1JP has considerably lower expression in tumor tissue samples than in matched normal tissue ones, which was observed among samples from gaster, colon, liver and lung. This consistent expression profile of lncRNA among different tissues has only been reported in a few cases. In cell lines from the same organs, the expression profiles of MT1JP were consistent with those in the tissues. Furthermore, knockdown of MT1JP resulted in a range of changes which usually arise during tumor formation, including cell cycle arrest, apoptosis inhibition, proliferation acceleration, and increased invasion and migration, as opposed to the effects of MT1JP overexpression. Down- or up-regulation of MT1JP led to variation in the expression of numerous cancer-associated mRNA molecules. These results suggest that MT1JP is necessary for maintaining the normal life activities of cells, and that it may have a critical function as a tumor suppressor.
Transcription factor p53 is a prominent tumor suppressor, and lncRNAs also have been reported to be involved in the p53 response as master regulators. LincRNA-p21 is transcriptional target of p53, and acts as a repressor in the p53 regulation pathway to induce apoptosis [3], and the lncRNA PANDA is induced by p53 and plays a role in the cell cycle and in the apoptosis pathway [38]. In addition, lncRNA Pint is a bona fide p53 transcriptional effector and serves as a negative modulator of the p53 pathway for proliferation and apoptosis [6]. In contrast to these lncRNAs, MT1JP is not a transcriptional target of p53 (data not shown), but regulates the protein level of p53 through promoting the translation of its mRNA, thereby participating in all p53 pathways. The data presented here thus show that MT1JP acts as an upstream regulator of p53, and add to our understanding of lncRNA-associated gene modulation in the p53 pathway.
Previous studies have shown that a number of RNA-binding proteins can not only enhance but also suppress mRNA translation [39–42]. One such protein is HuR. The HuR protein was found to interact with 3′-UTR of the Fas mRNA to block its translation in liver cancer [43] while upregulation of HuR in human RKO colorectal carcinoma cells led to elevated p53 protein levels in response to ultraviolet light irradiation [44]. The RNA-binding protein TIAR is also a good example. TIAR is reported to have partially repressed global translation in tetracycline-inducible system, whereas a genome-wide profiling showed that TIAR upregulated p53 signaling pathway-related genes selectively [45]. In this study, we have shown that not only the RNA-binding protein TIAR but also the lncRNA MT1JP can upregulate p53 mRNA translation. TIAR interacts strongly with p53 mRNA and MT1JP, however, no direct interaction between MT1JP and p53 mRNA was detected (data not shown), which is explained by the lack of sequence complementarity between these two RNAs. Therefore, the modulation of p53 by MT1JP is TIAR-dependent, this is to say that the translational regulation of p53 occur by a complex of MT1JP and TIAR, in which TIAR interacts with the p53, but where the regulatory effect of MT1JP on p53 translation depends on the presence of TIAR in a fashion yet to be elucidated. Moreover, this is the first report showing TIAR can strongly bind to a lncRNA, and MT1JP is thus the first lncRNA found to act in concert with TIAR. Further experiments are required to confirm the detailed mechanism by which the MT1JP-TIAR complex acts on the p53 mRNA translation. Since MT1JP may have a role as a tumor suppressor in several kinds of cancers, it is likely that the translational regulation by the MT1JP protein complex demonstrated in this study is a widely occurring tumor suppression phenomenon. Even more importantly, the association between MT1JP and RNA-binding protein provides a novel node in the complicated cancer regulatory network.
We observed that in addition to TIAR, MT1JP can also bind to the RAP1B and MSI2 proteins. Given the known functional difference between these three proteins it seems reasonable to assume that MT1JP may play different role in the regulation of other genes or other cellular processes. It is reported that MSI2 can take part in post-transcriptional regulation through associating with specific mRNAs and that it may participate in the modulation of proliferation and maintenance of stem cells in the central nervous system [46, 47], and it has recently demonstrated a potential as a predictive biomarker of HCC prognosis [48]. RAP1B is a member of the RAS-related protein family, and is involved in a number of cellular processes including cell growth, differentiation, migration and invasion [49–51]. Although RAP1B has not been reported as a RNA-binding protein, our data suggest that it may add this feature to its functional repertoire.
In summary, our results suggest that the cytoplasmic lncRNA MT1JP can bind diverse proteins with a potential to affect a variety of cellular processes. In its interaction with the RNA-binding protein TIAR, MJ1JP may function as a tumor suppressor through modulation of the p53 protein expression level and thereby the p53-related pathways (Figure 7).
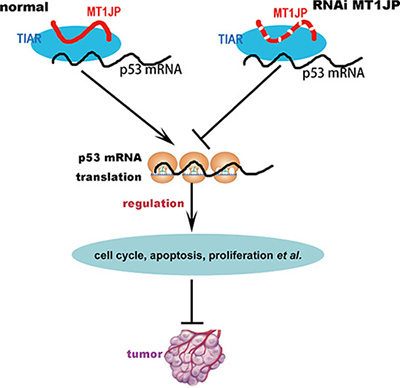
Figure 7: The model of lncRNA MT1JP regulating cell transformation.
MATERIALS AND METHODS
Cell culture
The human cell line L02 were cultured in RPMI 1640 Medium (ATCC Modification) and both SMMC-7721 and HepG2 cells were cultured in DMEM. All media were supplemented with 10% (v/v) FBS.
Northern blotting
Northern blotting of MT1JP was performed according to Xiao et al. [52].
Cell migration and invasion
For knockdown and overexpression of MT1JP, L02, SMMC-7721 or HepG2 cells were transfected with siRNAs or plasmids, trypsinized and counted after 24 h. 50 000 cells were seeded in the upper chamber of Transwell (Corning, #3422) with (for invasion) or without (for migration) BD Matrigel matrix (BD, #356230) coated. The medium in upper chamber has no fetal bovine serum (FBS) while the one in the lower chamber contains 20% FBS. After another 24 h culture at 37°C, 5% CO2, the non-migrated or non-invaded cells in upper chamber were gently removed, while the migrated or invaded cells were stained with 0.1% crystal violet for 20 min, washed with water, air dried and photographed using an Olympus microscope imaging systems. The migrated or invaded cells were counted for differential analysis.
Subcellular fractionation
Cells were fractioned as NE-PER(R) Nuclear and Cytoplasmic Extraction’s instruction (pierce, #78833). The total RNA isolated from each fraction was determined by qRT-PCR. U1 serves as the nuclear control while GAPDH and ACTB as the cytosolic controls.
RNA isolation and RT-qPCR
RNA was isolated with TRIzol Reagent (Invitrogen) according to manufacturer’s instructions, and reverse transcribed using SuperScript® III First-Strand Synthesis System (Invitrogen). QRT-PCR was performed with TransStart Top Green qPCR SuperMix (TransGen) according to the manufacturer’s protocol. All the sequences of primers were in Supplementary Table S2.
Western blotting
Western blotting was performed by standard protocols. Antibodies against the following protein were obtained as indicated: P53 (Senta Cruze), TIAR (Cell Signaling Technology), RAP1B (Proteintech), MSI2 (Abcam), GAPDH (Abnova).
siRNAs, plasmids and cell transfection
siRNAs were designed by using webserver of Integrated DNA Technologies (http://www.idtdna.com). The full length MT1JP DNA sequence, potential ORF sequence and EGFP were cloned into pcDNA3.1(+) with double enzyme digestion reactions. For transfection of siRNAs or plasmids, L02, SMMC-7721 or HepG2 cells were cultured in 12-well or 10 cm plates overnight, and transfected with 20 uM siRNAs or plasmids using the Lipo2000 (Life Technology) kit according to the manufacturer’s recommendations. All the sequences of primers, siRNAs and probes were in Supplementary Table S2.
Silver stain
Silver staining was performed using the SilverQuest™ Silver Staining Kit Dynabeads Protein A (Invitrogen) according to the manufacturer’s recommendations.
RACE
5′ and 3′ RACE were performed using the FirstChoice RLM-RACE Kit (Life Technology) according to the manufacturer’s recommendations. The primers are listed in Supplementary Table S2.
Gene ontology (GO) analysis
The genes that have high correlation of expression with MT1JP in 76 pairs of tissue samples were analyzed to perform functional enrichment at webserver: http://david.abcc.ncifcrf.gov/tools.jsp.
Bioinformatics analysis of microarray data
Feature Extraction v10.7.3.1 (Agilent Technologies, CA) software was used to extract all features of the data obtained from the scanned images. The signal ratio of Cy5 intensity and Cy3 intensity on the same array was subjected to quantile normalization followed by log2-scale transformation. Hierarchical clustering was performed using cluster 3.0 with complete linkage and centered Pearson correlation. The normalized and log2-scaled signal ratios were centered on the median before performing unsupervised hierarchical clustering.
Statistical analysis
Experimental data are represented as the mean ± SD of more than three biologic replicates and Student’s t tests were performed to determine the significance of differences. P-values are indicated as follows: *P < 0.05, **P < 0.01, and ***P < 0.001.
ACKNOWLEDGMENTS
We thank Dr. Geir Skogerbø and Dr. Tengfei Xiao for carefully reading this manuscript and discussing this research. The cell lines are kind gifts from Prof. Jian Zhao (Chinese People’s Liberation Army (PLA) General Hospital). We also thank Prof. Zusen Fan and all members from Chen Lab for discussions about the experiments.
GRANT SUPPORT
This work was supported by grants from Chinese Academy of Science Strategic Project of Leading Science and Technology (XDA01020402) and the National High Technology Research and Development Program (“863” Program) of China (2012AA020402 and 2012AA02A202).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011; 43:904–914.
2. Kung JT, Colognori D, Lee JT. Long noncoding RNAs: past, present, and future. Genetics. 2013; 193:651–669.
3. Huarte M, Guttman M, Feldser D, Garber M, Koziol MJ, Kenzelmann-Broz D, Khalil AM, Zuk O, Amit I, Rabani M, Attardi LD, Regev A, Lander ES, et al. A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell. 2010; 142:409–419.
4. Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci Signal. 2010; 3:ra8.
5. Guttman M, Donaghey J, Carey BW, Garber M, Grenier JK, Munson G, Young G, Lucas AB, Ach R, Bruhn L, Yang X, Amit I, Meissner A, et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature. 2011; 477:295–300.
6. Marin-Bejar O, Marchese FP, Athie A, Sanchez Y, Gonzalez J, Segura V, Huang L, Moreno I, Navarro A, Monzo M, Garcia-Foncillas J, Rinn JL, Guo S, et al. Pint lincRNA connects the p53 pathway with epigenetic silencing by the Polycomb repressive complex 2. Genome Biol. 2013; 14:R104.
7. Klattenhoff CA, Scheuermann JC, Surface LE, Bradley RK, Fields PA, Steinhauser ML, Ding H, Butty VL, Torrey L, Haas S, Abo R, Tabebordbar M, Lee RT, et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell. 2013; 152:570–583.
8. Lin N, Chang KY, Li Z, Gates K, Rana ZA, Dang J, Zhang D, Han T, Yang CS, Cunningham TJ, Head SR, Duester G, Dong PD, et al. An evolutionarily conserved long noncoding RNA TUNA controls pluripotency and neural lineage commitment. Mol Cell. 2014; 53:1005–1019.
9. Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K, Lee CS, Flockhart RJ, Groff AF, Chow J, Johnston D, Kim GE, Spitale RC, et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature. 2013; 493:231–235.
10. Brown CJ, Hendrich BD, Rupert JL, Lafreniere RG, Xing Y, Lawrence J, Willard HF. The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell. 1992; 71:527–542.
11. Mancini-Dinardo D, Steele SJ, Levorse JM, Ingram RS, Tilghman SM. Elongation of the Kcnq1ot1 transcript is required for genomic imprinting of neighboring genes. Genes Dev. 2006; 20:1268–1282.
12. Tian D, Sun S, Lee JT. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell. 2010; 143:390–403.
13. Nishimoto Y, Nakagawa S, Hirose T, Okano HJ, Takao M, Shibata S, Suyama S, Kuwako K, Imai T, Murayama S, Suzuki N, Okano H. The long non-coding RNA nuclear-enriched abundant transcript 1_2 induces paraspeckle formation in the motor neuron during the early phase of amyotrophic lateral sclerosis. Mol Brain. 2013; 6:31.
14. Michalik KM, You X, Manavski Y, Doddaballapur A, Zornig M, Braun T, John D, Ponomareva Y, Chen W, Uchida S, Boon RA, Dimmeler S. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ Res. 2014; 114:1389–1397.
15. Faghihi MA, Modarresi F, Khalil AM, Wood DE, Sahagan BG, Morgan TE, Finch CE, St Laurent G, 3rd, Kenny PJ, Wahlestedt C. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat Med. 2008; 14:723–730.
16. Luo JH, Ren B, Keryanov S, Tseng GC, Rao UN, Monga SP, Strom S, Demetris AJ, Nalesnik M, Yu YP, Ranganathan S, Michalopoulos GK. Transcriptomic and genomic analysis of human hepatocellular carcinomas and hepatoblastomas. Hepatology. 2006; 44:1012–1024.
17. Lin R, Maeda S, Liu C, Karin M, Edgington TS. A large noncoding RNA is a marker for murine hepatocellular carcinomas and a spectrum of human carcinomas. Oncogene. 2007; 26:851–858.
18. Ji P, Diederichs S, Wang W, Boing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E, Thomas M, Berdel WE, Serve H, et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003; 22:8031–8041.
19. Guffanti A, Iacono M, Pelucchi P, Kim N, Solda G, Croft LJ, Taft RJ, Rizzi E, Askarian-Amiri M, Bonnal RJ, Callari M, Mignone F, Pesole G, et al. A transcriptional sketch of a primary human breast cancer by 454 deep sequencing. BMC Genomics. 2009; 10:163.
20. Ren S, Wang F, Shen J, Sun Y, Xu W, Lu J, Wei M, Xu C, Wu C, Zhang Z, Gao X, Liu Z, Hou J, et al. Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 derived miniRNA as a novel plasma-based biomarker for diagnosing prostate cancer. Eur J Cancer. 2013; 49:2949–2959.
21. Tsai MC, Manor O, Wan Y, Mosammaparast N, Wang JK, Lan F, Shi Y, Segal E, Chang HY. Long noncoding RNA as modular scaffold of histone modification complexes. Science. 2010; 329:689–693.
22. Carpenter S, Aiello D, Atianand MK, Ricci EP, Gandhi P, Hall LL, Byron M, Monks B, Henry-Bezy M, Lawrence JB, O’Neill LA, Moore MJ, Caffrey DR, et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science. 2013; 341:789–792.
23. Lai F, Orom UA, Cesaroni M, Beringer M, Taatjes DJ, Blobel GA, Shiekhattar R. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature. 2013; 494:497–501.
24. Fasching L, Kapopoulou A, Sachdeva R, Petri R, Jonsson ME, Manne C, Turelli P, Jern P, Cammas F, Trono D, Jakobsson J. TRIM28 represses transcription of endogenous retroviruses in neural progenitor cells. Cell Rep. 2015; 10:20–28.
25. Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, HY C. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell. 2007; 129:1311–1323.
26. Yoon JH, Abdelmohsen K, Srikantan S, Yang X, Martindale JL, De S, Huarte M, Zhan M, Becker KG, Gorospe M. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012; 47:648–655.
27. Yuan J, Yue H, Zhang M, Luo J, Liu L, Wu W, Xiao T, Chen X, Chen X, Zhang D, Xing R, Tong X, Wu N, et al. Transcriptional profiling analysis and functional prediction of long noncoding RNAs in cancer. Oncotarget. 2016; doi: 10.18632/oncotarget.6993.
28. Lin MF, Jungreis I, Kellis M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics. 2011; 27:i275–282.
29. Sun L, Luo H, Bu D, Zhao G, Yu K, Zhang C, Liu Y, Chen R, Zhao Y. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res. 2013; 41:e166.
30. Wang L, Park HJ, Dasari S, Wang S, Kocher JP, Li W. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res. 2013; 41:e74.
31. Kong L, Zhang Y, Ye ZQ, Liu XQ, Zhao SQ, Wei L, Gao G. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res. 2007; 35:W345–349.
32. Wang Y, He L, Du Y, Zhu P, Huang G, Luo J, Yan X, Ye B, Li C, Xia P, Zhang G, Tian Y, Chen R, et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell. 2015; 16:413–425.
33. McHugh CA, Chen CK, Chow A, Surka CF, Tran C, McDonel P, Pandya-Jones A, Blanco M, Burghard C, Moradian A, Sweredoski MJ, Shishkin AA, Su J, et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature. 2015; 521:232–236.
34. Fischer TH, Gatling MN, Lacal JC, White GC, 2nd. rap1B, a cAMP-dependent protein kinase substrate, associates with the platelet cytoskeleton. J Biol Chem. 1990; 265:19405–19408.
35. Tan LY, Whitfield P, Llorian M, Monzon-Casanova E, Diaz-Munoz MD, Turner M, Smith CW. Generation of functionally distinct isoforms of PTBP3 by alternative splicing and translation initiation. Nucleic Acids Res. 2015; 43:5586–600.
36. Khandjian EW, Corbin F, Woerly S, Rousseau F. The fragile X mental retardation protein is associated with ribosomes. Nat Genet. 1996; 12:91–93.
37. Fechir M, Linker K, Pautz A, Hubrich T, Kleinert H. The RNA binding protein TIAR is involved in the regulation of human iNOS expression. Cell Mol Biol (Noisy-le-grand). 2005; 51:299–305.
38. Hung T, Wang Y, Lin MF, Koegel AK, Kotake Y, Grant GD, Horlings HM, Shah N, Umbricht C, Wang P, Wang Y, Kong B, Langerod A, et al. Extensive and coordinated transcription of noncoding RNAs within cell-cycle promoters. Nat Genet. 2011; 43:621–629.
39. Cathcart AL, Rozovics JM, Semler BL. Cellular mRNA decay protein AUF1 negatively regulates enterovirus and human rhinovirus infections. J Virol. 2013; 87:10423–10434.
40. Ehlers C, Schirmer S, Kehlenbach RH, Hauber J, Chemnitz J. Post-transcriptional regulation of CD83 expression by AUF1 proteins. Nucleic Acids Res. 2013; 41:206–219.
41. Lu M, Pan C, Zhang L, Ding C, Chen F, Wang Q, Wang K, Zhang X. ING4 inhibits the translation of proto-oncogene MYC by interacting with AUF1. FEBS Lett. 2013; 587:1597–1604.
42. Lee KH, Kim SH, Kim HJ, Kim W, Lee HR, Jung Y, Choi JH, Hong KY, Jang SK, Kim KT. AUF1 contributes to Cryptochrome1 mRNA degradation and rhythmic translation. Nucleic Acids Res. 2014; 42:3590–3606.
43. Zhu H, Berkova Z, Mathur R, Sehgal L, Khashab T, Tao RH, Ao X, Feng L, Sabichi AL, Blechacz B, Rashid A, Samaniego F. HuR Suppresses Fas Expression and Correlates with Patient Outcome in Liver Cancer. Mol Cancer Res. 2015; 13:809–18.
44. Mazan-Mamczarz K, Galban S, Lopez de Silanes I, Martindale JL, Atasoy U, Keene JD, Gorospe M. RNA-binding protein HuR enhances p53 translation in response to ultraviolet light irradiation. Proc Natl Acad Sci U S A. 2003; 100:8354–8359.
45. Sanchez-Jimenez C, Ludena MD, Izquierdo JM. T-cell intracellular antigens function as tumor suppressor genes. Cell Death Dis. 2015; 6:e1669.
46. Sakakibara S, Nakamura Y, Satoh H, Okano H. Rna-binding protein Musashi2: developmentally regulated expression in neural precursor cells and subpopulations of neurons in mammalian CNS. J Neurosci. 2001; 21:8091–8107.
47. Okano H, Imai T, Okabe M. Musashi: a translational regulator of cell fate. J Cell Sci. 2002; 115:1355–1359.
48. He L, Zhou X, Qu C, Hu L, Tang Y, Zhang Q, Liang M, Hong J. Musashi2 predicts poor prognosis and invasion in hepatocellular carcinoma by driving epithelial-mesenchymal transition. J Cell Mol Med. 2014; 18:49–58.
49. Malchinkhuu E, Sato K, Maehama T, Ishiuchi S, Yoshimoto Y, Mogi C, Kimura T, Kurose H, Tomura H, Okajima F. Role of Rap1B, tumor suppressor PTEN in the negative regulation of lysophosphatidic acid—induced migration by isoproterenol in glioma cells. Mol Biol Cell. 2009; 20:5156–5165.
50. Guo H, Hu X, Ge S, Qian G, Zhang J. Regulation of RAP1B by miR-139 suppresses human colorectal carcinoma cell proliferation. Int J Biochem Cell Biol. 2012; 44:1465–1472.
51. Zhang M, Zhou S, Zhang L, Zhang J, Cai H, Zhu J, Huang C, Wang J. miR-518b is down-regulated, and involved in cell proliferation and invasion by targeting Rap1b in esophageal squamous cell carcinoma. FEBS Lett. 2012; 586:3508–3521.
52. Xiao T, Liu L, Sun Y, Li H, Luo H, Li T, Wang S, Stephen D, Zhao CR, R C. An Uncharacterized Long Noncoding RNA ADINR Regulates Adipogenesis by Transcriptionally Activating C/EBPα. Stem Cell Reports. 2015; 5:856–65.

