INTRODUCTION
Non-small-cell lung cancer (NSCLC), which accounts for approximately 85% of lung cancers, has been largely identified by oncogenic driver mutations with potential opportunities for targeted therapies [1, 2]. Recently, activation of the anaplastic lymphoma kinase (ALK) gene in lung cancer by fusion to echinoderm microtubule-associated protein-like 4 (EML4) or other gene partners (such as BIRC6 [3], TFG [4], KIF5B [5] and KLC1 [6]) has been identified as oncogenic events [7]. Clinical studies have shown that locally advanced or metastatic NSCLC patients harboring ALK gene rearrangement are highly sensitive to Crizotinib, which is a small molecular inhibitor of ALK tyrosine kinase [8, 9]. In addition, NSCLC with sensitive epidermal growth factor receptor (EGFR) mutations are well responded to tyrosine kinase inhibitors (TKI) and these patients will have a longer progression-free survival (PFS) than the patients whose tumors do not contain EGFR mutations [1, 10].
Approximately 60% of patients with NSCLC are diagnosed at a late stage for the first time [11]. These patients are not suitable for the resection of the primary tumor, and the only pathologic material guiding systemic therapy should be small biopsy or cytological specimens. Recent studies have demonstrated that cytological specimens, including fine-needle aspiration (FNA), fibreoptic bronchoscopic (FOB) and pleural effusion (PLE), are suitable for the molecular testing [12–16]. Although FISH (fluorescent in situ hybridization) is currently the gold standard method to detect ALK gene rearrangement approved by FDA, its application on cytological specimens remains a worth area of investigation. In this study, we investigate the use of ALK FISH and EGFR, KRAS mutational testing on cytological specimens and to evaluate PFS of the patients who received targeted therapies.
RESULTS
Specimen and patient characteristics
Demographic and clinicopathologic features were summarized in Table 1. Of the 137 patients enrolled in the study, 54 (39.4%, of 137) were male and 83 (60.6%, of 137) were female. The mean age at diagnosis was 58.8 years (range: 27.0 - 85.0 years) and the median age was 59.0 years. Cytological specimens (n = 137) included FNAs (n = 91), FOBs (n = 5) and PLEs (n = 41). Of these, 126 (92.0%, of 137) were diagnosed as ADC, 3 (2.2%, of 137) as SCC, 1 (0.7%) as adenosquamous carcinoma and 7 (5.1%, of 137) as NSCLC, not otherwise specified.
Table 1: Demographic and clinicopathologic features of the study patients
Characteristics |
No. |
Percent |
|---|---|---|
Sex |
||
Male |
54 |
39.4% |
Female |
83 |
60.6% |
Sample type |
||
FNA |
91 |
66.4% |
FOB |
5 |
3.6% |
PLE |
41 |
30% |
Histologic type |
||
ADC |
126 |
92.0% |
SCC |
3 |
2.2% |
Adenosquamous |
1 |
0.7% |
NSCLC, NOS |
7 |
5.1% |
Age |
||
Mean (SD) |
58.8 ± 12.1 |
|
Median |
59.0 |
|
Range |
27.0–85.0 |
|
ALK FISH |
||
Positive |
16 |
11.7% |
Negative |
121 |
88.3% |
EGFR mutation status |
||
Sensitive mutation |
53 |
39.5% |
Exon 20 mutation |
4 |
3.0% |
Complex mutation with exon 19 deletion + T790M |
1 |
0.7% |
Complex mutation with L858R + T790M |
2 |
1.5% |
Negative |
74 |
55.3% |
Not tested |
3 |
NA |
KRAS mutation status |
||
Positive |
8 |
6.0% |
Negative |
126 |
94.0% |
Not tested |
3 |
NA |
Abbreviations: ALK = anaplastic lymphoma kinase; FISH = fluorescence in situ hybridization; FNA = fine needle aspiration; FOB = fibreoptic bronchoscopic; PLE = pleural effusion; ADC = adenocarcinoma; SCC = squamous cell carcinoma; NSCLC, NOS = non-small cell lung cancer, not otherwise specified.
ALK FISH analysis
Of the 137 NSCLCs analyzed for ALK FISH, 16 (11.7%, of 137) were detected to harbor ALK rearrangement (FISH positive) and 121 (88.3%, of 137) were FISH negative. The ALK FISH positive cases of the cytological samples were 12 (75.0%, of 16) FNAs, 1 (6.2%, of 16) FOB and 3 (18.8%, of 16) PLE (Supplementary Table S1). The FNA samples showed highest FISH positive rate (13.2%, 12/91) among three groups, although this did not demonstrate a statistically significant difference (P = 0.32). On FISH examination (Figure 1), split pattern was observed in 14 cases (87.5%) and unbalanced rearrangement, characterized by a loss of the 5’ probe, was shown in 2 cases (12.5%).
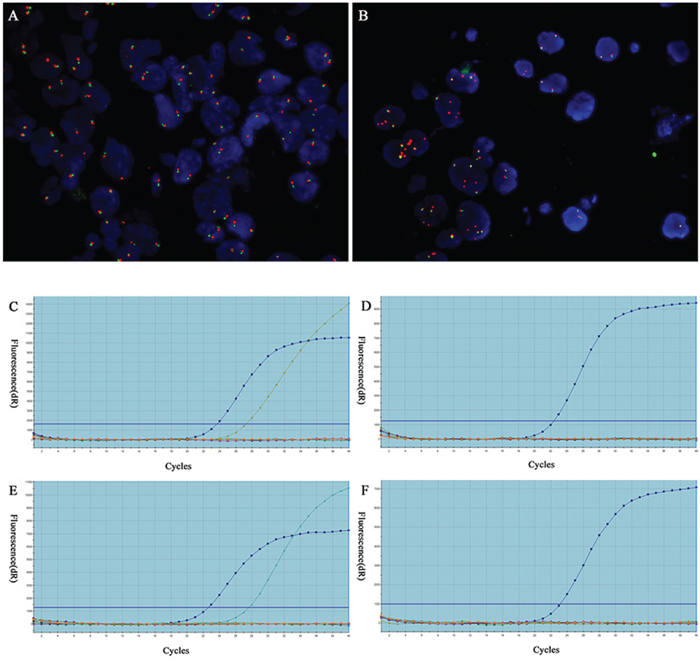
Figure 1: Detection of ALK fusion by FISH and EGFR/KRAS mutations in cytological specimens by qRT-PCR. A. Representative image of FISH negative case showing intact two fused signals per nucleus. B. Representative image of FISH carried out with Vysis LSI ALK Dual color Break-Apart FISH probes detected ALK fusion as split red and green signals. Original magnification ×1000. C-D. Detection of EGFR and EGFR L858R mutation and negative case. E-F. KRAS p.G12D mutation and negative case.
The ALK FISH positive cases included 6 men and 10 women and there was no significant difference in gender distribution between FISH positive and negative cases (P = 0.87). The mean age at diagnosis for FISH positive cases was 52.7 ± 11.8 years, which was much younger than that of FISH negative cases (P < 0.05). FISH positive cases were all defined as ADC histologic subtype. In addition, there was one FISH positive case also demonstrated an EGFR L858R mutation (Table 2).
Table 2: Clinicopathologic characteristics of ALK FISH positive and negative cytology cases
Characteristics |
ALK FISH positive (n = 16) |
ALK FISH negative (n = 121) |
P value |
|---|---|---|---|
Sex |
0.87† |
||
Male |
6 (37.5%) |
48 (39.7%) |
|
Female |
10 (62.5%) |
73 (60.3%) |
|
Histologic type |
NA |
||
ADC |
16 (100%) |
110 (90.9%) |
|
SCC |
0 |
3 (2.5%) |
|
Adenosquamous |
0 |
1 (0.8%) |
|
NSCLC, NOS |
0 |
7 (5.8%) |
|
Age |
0.03‡ |
||
Mean (SD) |
16 |
59.6 ± 11.9 |
|
Median |
53.5 |
60.0 |
|
Range |
27.0 – 73.0 |
30.0 – 85.0 |
|
EGFR |
0.002† |
||
Positive |
1 (6.7%) |
59 (49.6%) |
|
Negative |
14 (93.3%) |
60 (50.4%) |
|
KRAS |
0.42§ |
||
Positive |
0 |
8 (6.7%) |
|
Negative |
15 (100%) |
111 (93.3%) |
Abbreviations: ALK = anaplastic lymphoma kinase; FISH = fluorescence in situ hybridization; ADC = adenocarcinoma; SCC = squamous cell carcinoma; NSCLC, NOS = non-small cell lung cancer, not otherwise specified.
† Two-sided χ2 test
‡ Two-sided Kruskal Wallis test
§ Fischer’s exact test
EGFR and KRAS mutation status
Among 134 NSCLCs tested, 60 (44.8%, of 134) cases carried EGFR mutations, which was 53 (39.5%, of 134) sensitive mutations and 4 (3.0%, of 134) exon 20 mutations with S768I. There was one case demonstrated complex mutation with exon 19 deletion and T790M and the other two cases showed complex mutation with L858R and T790M. Among them, two patients carried acquired T790M mutation after TKI treatment and one patient carried primary coexisting mutations of T790M and L858R. In addition, four patients carried primary exon 20 S768I mutations and did not receive targeted therapies. The mutated EGFR cases of the cytological samples were 36 (60.0%, of 60) FNAs, 1 (1.7%, of 60) FOB and 23 (38.3%, of 60) PLEs (Supplementary Table S2). The PLE samples showed highest EGFR mutation rate (58.9%, 23/39) among three groups, although this did not demonstrate a statistically significant difference (P = 0.41). Mutations of KRAS occurred in 8 (6.0%, of 134) cases and this was mutually exclusive from EGFR mutation. In addition, compared with male patients, female patients were more likely to carry EGFR mutations (55.6% vs 28.3%, P = 0.002) (Supplementary Table S3).
Outcomes of targeted therapies
Of the 9 patients who received crizotinib treatment, there were 5 men and 4 women with mean age of 58.6 years (Table 3). The median follow-up duration was 9 months (range: 6–15 months). At the end of follow up, 8 (88.9%) patients were still receiving crizotinib treatment and there was one person stopping the treatment because of disease progression.
Table 3: Progression-free survival of crizotinib treatment ALK FISH positive patients
Case no. |
Age |
Sex |
Type of samples |
PFS (months) |
Status |
|---|---|---|---|---|---|
1 |
55 |
Female |
PLE |
8 |
PR |
2 |
48 |
Male |
FNA |
5 |
PD |
3 |
72 |
Male |
FNA |
12 |
PR |
4 |
42 |
Female |
FNA |
3 |
PR |
5 |
46 |
Female |
FNA |
5 |
PR |
6 |
51 |
Male |
FNA |
6 |
PR |
7 |
27 |
Male |
FNA |
5 |
PR |
8 |
62 |
Female |
PLE |
4 |
PR |
9 |
55 |
Male |
PLE |
4 |
PR |
Abbreviations: ALK = anaplastic lymphoma kinase; FISH = fluorescence in situ hybridization; FNA = fine needle aspiration; PLE = pleural effusion; PFS = progression-free survival; PR = partial response; PD = progressive disease.
We performed targeted next-generation sequencing (NGS) in one case who did not response well to crizotinib treatment. Results confirmed the existence of EML4-ALK transloction, and the relative abundance of EML4-ALK fusion was 25.3%. In addition, there were a nonsense mutation (c.991 C > T, p.Q331) in exon 9 of P53 and a deletion of CDKN2A. The mutation in P53 could result in a stop codon and lead to loss of P53 function as a transcription factor. [17]
There were 22 patients with EGFR sensitive mutations who received gefitinib or erlotinib treatment. The median PFS was 16.0 months (95% confidence interval, 12.9 - 19.1) (Figure 2). Of them, 8 patients (36.4%) exhibited disease progression after 3 months due to acquired resistance to EGFR-TKI treatment. At the data cutoff point, 14 patients were still taking TKI treatment and tumor regression was observed in these patients.
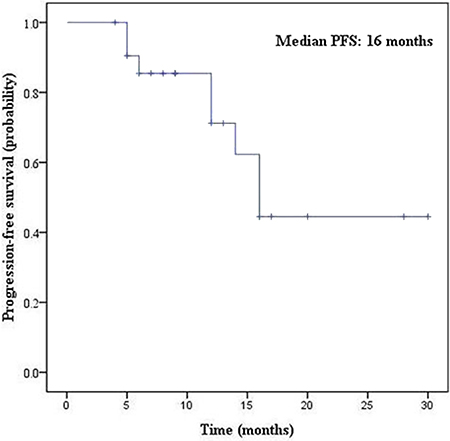
Figure 2: Kaplan-Meier survival curves of PFS for EGFR-TKI treatment.
DISCUSSION
The diagnosis of advanced-stage NSCLC is usually based on a small amount of cytological specimens, consequently, the use of ALK FISH and EGFR, KRAS mutational testing on cytological specimens is gradually becoming a necessity in the routine molecular pathological diagnosis. Although FISH is considered to be the gold standard for detection of ALK rearrangements on histological tissues approved by FDA, its application on cytological specimens remains a worth area of investigation. In this study, for the first time, we demonstrated the possibility of using ALK FISH and EGFR, KRAS mutational testing on cytological specimens in a large cohort of Chinese patients and correlate the analysis results with PFS in patients with targeted therapies.
Targeted therapies are mostly effective in patients with locally advanced or metastatic disease, thus, molecular analysis on cytological samples rather than on resected specimens would be preferred on patients with late stage tumors. The data in previous studies demonstrated that the performance of cytology based EGFR and KRAS mutational analysis was at least as effective as histology based analysis [18–20]. In this study, various cytological specimens including FNA, FOB and PLE were all submitted for EGFR/KRAS mutational analysis and 97.8% (134/137) of samples were suitable for test. The other three samples not suitable for analysis were due to inadequate cell numbers and this prompted us to use more sensitive methods, such as digital PCR and NGS, to test EGFR mutations. Recently, a large cohort of mutational analysis on histological specimens was performed in 5125 Chinese patients and revealed that 36.2% and 8.4% patients with NSCLC carried EGFR and KRAS mutations respectively [21]. Cytological specimens of our study demonstrated slightly higher EGFR mutational rates (44.8% vs 36.2%) than that of histological specimens in above study and this could partly be attributed to the patients undertaken cytological mutational analysis often underwent a late stage disease.
Previous studies had rarely evaluated the EGFR-TKI treatment responses on patients with sensitive EGFR mutations tested on cytological specimens, although the outcome of the patients treated with TKI was important to confirm the predictive value of cytological mutational analysis. In our study, fifty-seven patients with EGFR mutations demonstrated an ADC subtype and this population would largely benefit from targeted therapies. Twenty-two patients with ADC subtype and EGFR sensitive mutations received TKI treatment with a median PFS of 16 months and all the patients were observed with tumor regression after 3-months treatment. This was in accordance with previous studies, which demonstrated that the median PFS interval of 16.2 months was observed in 11 EGFR mutated patients who treated with gefitinib [22]. In addition, Maria D. Lozano also observed patients with EGFR mutations had a 12.3-months median PFS [23]. These results indicated that EGFR mutational testing on cytological samples was adequate for patient selection with EGFR-TKI treatment. In addition to guide the targeted therapy for patients with unresected tumors, another possible advantage of cytological mutational analysis was to identify genetic shifts of EGFR acquired mutations during TKI treatment.
It is currently accepted that FISH is the most appropriate method to detect ALK rearrangements and to guide crizotinib treatment. Only a few reports have assessed the adequacy of cytological specimens for ALK FISH analysis, thus, its application on cytological specimens still needs further studies. Weiya Wang, et al analyzed a cohort of 58 patients for ALK rearrangements using pleural effusion cell blocks and found 10.3% positive cases by ALK FISH [16]. Agnese Proietti, et al demonstrated 4.4% cases with ALK FISH rearrangements in both cell blocks and small biopsies [24]. In addition, MJ. Neat, et al compared FISH and IHC methods to screen for ALK status in endobronchial ultrasound (EBUS)-transnronchial needle aspiration (TBNA) derived cytological specimens and found that FISH was superior to IHC for the detection of ALK rearrangement in these cytological samples [25]. To best of our knowledge, this is the largest report of ALK detection in cytological specimens in Chinese patients and first described the outcome of patients with ALK FISH positive who received crizotinib treatment. In this study, we detected 11.7% ALK rearrangement in 137 cases of various cytological specimens, including FNA, FOB and PLE, which was significantly higher than the rate of 6.8% previously reported in a large Chinese consecutive case series [26]. Our results were in accordance with previous report that 12.7% of ALK rearrangements were observed in malignant pleural effusion cell blocks from patients with advanced NSCLC [16]. The increased percentage of positive cases was likely attributed to the advanced stage of disease among patients with cytological specimens. Of the nine patients who received crizotinib treatment, one patient exhibited disease progression after 5 months and eight patients were still receiving crizotinib treatment at the end of cutoff point. We performed targeted next-generation sequencing in one case who did not response well to Crizotinib treatment and found that there was a nonsense mutation in exon 9 of P53 which could lead to loss of P53 function. Previous studies have indicated that P53 alterations might accelerate cancer development and could also lead to shorter survival in patients with P53 mutated NSCLC [27]. Our findings first demonstrated the original P53 mutation in ALK positive patients and this mutation of P53 may correlate to the shorter PFS.
In summary, we conclude thatALK FISH and EGFR, KRAS mutational analysis on cytological specimens are sensitive methods for screening advanced stage NSCLC patients who are adequate for targeted treatment. Cytological specimens including FNA, FOB and PLE all provide feasible and effective material for the molecular analysis and further studies are required to validate the application in the routine molecular pathological diagnosis.
MATERIALS AND METHODS
Patients
One hundred and thirty-seven patients with advanced NSCLC were enrolled in this study from September 2013 to June 2015. All these cytological samples, including FNA, FPB and PLE, were tested for ALK fusion by FISH and 134 were performed with EGFR and KRAS mutational testing because 3 samples were not adequate for DNA extraction. Imaging data were independently reviewed by authors to evaluate their treatment responses according to the Response Evaluation Criteria in Solid Tumors (RECIST) version 1.1. PFS was calculated from the date of initiating TKI treatment to a radiologic or clinical observation of disease progression. The study protocol was approved by the Institute Review Board of the Cancer Hospital, Chinese Academy of Medical Sciences (CAMS), Beijing, China. The methods were carried out in accordance with the approved guidelines. Each participant signed an Institutional Review Board approved informed consent in accordance with current guidelines.
Specimen preparation
The cytological specimen preparations were conducted according to a standard specimen processing protocol in our laboratory. The cases included FNA specimens obtained under image guidance with a cytopathologist present for adequacy assessment and pleural fluid specimens obtained by thoracentesis. The percentage of tumor cells more than 5% or over 500 tumor cells were used for DNA extraction and 100 tumor cells for ALK FISH analysis. The algorithm used for molecular testing in our study was depicted in Supplementary Figure S1. Each case had air-dried slides stained with Diff-Quik (DQ stain, Protocol Hema 3; Fisher Scientific, Kalamazoo, MI) and additional slides fixed in 95% alcohol for Papanicolaou staining. Fluid specimens also had a ThinPrep (Hologic, Marlborough, MA) slide prepared.
Fluorescence in situ hybridization
FISH analysis was conducted as previously described [28]. Briefly, FISH analysis was performed using the Vysis LSI ALK Dual color, Break Apart Rearrangement Probe (Abbott/Vysis, Abbott Park, IL, USA). Samples were considered to be FISH positive if more than 15% of the scored tumour cells had split one or both ALK 5’ and 3’ probe signals or had isolated 3’ signals. Slides were evaluated independently by two experts blind to the patient’s history and histological findings.
EGFR and KRAS mutational testing
EGFR and KRAS mutational testing was conducted as previously described [29]. Briefly, mutational testing was performed using the HumanEGFR or KRAS Mutation Qualitative Detection Kit (Beijing ACCB Biotech Ltd., China), which applies quantitative real-time PCR (qRT-PCR) platform combining amplification refractory mutation system (ARMS) primers and TaqMan probes. The sensitive mutations were defined as p.G719S/C/A, p.L858R, p.L861Q and insertions in exon 19. The hotspot mutations in KRAS gene were within codon 12 and 13 including p.G12C/V/S/R/D/A and p.G13D. The assay was carried out according to the manufacturer’s protocol using the Stratagene Mx3000P real-time PCR system (Agilent technologies Inc, USA). Presence or absence of mutations was assessed from the fluorescence amplification curve.
Targeted next-generation sequencing
For one patient who did not response well to crizotinib treatment, targeted next-generation sequencing was performed. Genomic DNA was profiled by using a capture-based targeted sequencing panel (Burning Rock Biotech, Guangzhou, People’s Republic of China). In brief, human genomic regions of 271 kb, including all exons in 56 genes and selected introns in ALK, RET and ROS1 for the detection of translocation events, were captured by using 120-bp probes and were sequenced (Supplementary Table S4). The concentration of the DNA samples was measured with the Qubit dsDNA assay. Fragments of 200 to 400-bp sizes were selected with beads (Agencourt AMPure XP kit; Beckman-Coulter, Brea, CA), followed by hybridization with the capture probes baits, hybrid selection with magnetic beads, and PCR amplification. A bioanalyzer high-sensitivity DNA assay was then used to assess the quality and size range. Available indexed samples were then sequenced on a Nextseq (Illumina, San Diego, CA) with pair-end reads. Sequence data were analyzed by GATK 3.2 (https://www.broadinstitute.org/gatk/) and DNA translocation analysis was performed by using both Tophat2 (https://ccb.jhu.edu/software/tophat/index.shtml) and Factera 1.4.3 ((http://factera.stanford.edu) [30].
Statistical analysis
Differences of patient characteristics and clinicopathologic factors in the two-dimensional cross-comparison were evaluated statistically by Pearson’s χ2-test orFischer’s exact test. Statistical tests were two-sided, and P < 0.05 was considered significant. Estimation of PFS was calculated using the Kaplan–Meier method. Statistics were carried out using SPSS software (version 16.0 of SPSS, Chicago, IL, USA).
ACKNOWLEDGMENTS
We thank all study participants of the Department of Pathology for their contributions to this project.
CONFLICTS OF INTEREST
The authors declare no competing financial interests.
GRANT SUPPORT
This work was supported by a grant from Youth Backbone Program (to Jianming Ying) of Cancer Hospital, CAMS, Beijing, China).
Authors’ contributions
Conceived and designed the experiments: Jianming Ying. Performed the experiments: Wenbin Li, Lei Guo, Tian Qiu, Yun Ling and Jian Cao, Huiqin Guo, Huan Zhao,. Analyzed the data: Wenbin Li and Zhihui Zhang. Contributed reagents/materials/analysis tools: Lin Li. Wrote the paper: Wenbin Li and Zhihui Zhang.
REFERENCES
1. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, Harris PL, Haserlat SM, Supko JG, Haluska FG, Louis DN, Christiani DC, Settleman J and Haber DA. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004; 350:2129–2139.
2. Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, Herman P, Kaye FJ, Lindeman N, Boggon TJ, Naoki K, Sasaki H, Fujii Y, Eck MJ, Sellers WR, Johnson BE, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004; 304:1497–1500.
3. Shan L, Jiang P, Xu F, Zhang W, Guo L, Wu J, Zeng Y, Jiao Y and Ying J. BIRC6-ALK, a Novel Fusion Gene in ALK Break-Apart FISH-Negative Lung Adenocarcinoma, Responds to Crizotinib. J Thorac Oncol. 2015; 10:e37–39.
4. Rikova K, Guo A, Zeng Q, Possemato A, Yu J, Haack H, Nardone J, Lee K, Reeves C, Li Y, Hu Y, Tan Z, Stokes M, Sullivan L, Mitchell J, Wetzel R, et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell. 2007; 131:1190–1203.
5. Takeuchi K, Choi YL, Togashi Y, Soda M, Hatano S, Inamura K, Takada S, Ueno T, Yamashita Y, Satoh Y, Okumura S, Nakagawa K, Ishikawa Y and Mano H. KIF5B-ALK, a novel fusion oncokinase identified by an immunohistochemistry-based diagnostic system for ALK-positive lung cancer. Clin Cancer Res. 2009; 15:3143–3149.
6. Togashi Y, Soda M, Sakata S, Sugawara E, Hatano S, Asaka R, Nakajima T, Mano H and Takeuchi K. KLC1-ALK: a novel fusion in lung cancer identified using a formalin-fixed paraffin-embedded tissue only. PLoS One. 2012; 7:e31323.
7. Soda M, Choi YL, Enomoto M, Takada S, Yamashita Y, Ishikawa S, Fujiwara S, Watanabe H, Kurashina K, Hatanaka H, Bando M, Ohno S, Ishikawa Y, Aburatani H, Niki T, Sohara Y, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007; 448:561–566.
8. Shaw AT, Kim DW, Nakagawa K, Seto T, Crino L, Ahn MJ, De Pas T, Besse B, Solomon BJ, Blackhall F, Wu YL, Thomas M, O'Byrne KJ, Moro-Sibilot D, Camidge DR, Mok T, et al. Crizotinib versus chemotherapy in advanced ALK-positive lung cancer. N Engl J Med. 2013; 368:2385–2394.
9. Camidge DR, Bang YJ, Kwak EL, Iafrate AJ, Varella-Garcia M, Fox SB, Riely GJ, Solomon B, Ou SH, Kim DW, Salgia R, Fidias P, Engelman JA, Gandhi L, Janne PA, Costa DB, et al. Activity and safety of crizotinib in patients with ALK-positive non-small-cell lung cancer: updated results from a phase 1 study. Lancet Oncol. 2012; 13:1011–1019.
10. Rosell R, Moran T, Queralt C, Porta R, Cardenal F, Camps C, Majem M, Lopez-Vivanco G, Isla D, Provencio M, Insa A, Massuti B, Gonzalez-Larriba JL, Paz-Ares L, Bover I, Garcia-Campelo R, et al. Screening for epidermal growth factor receptor mutations in lung cancer. N Engl J Med. 2009; 361:958–967.
11. Azzoli CG, Temin S, Aliff T, Baker S, Jr., Brahmer J, Johnson DH, Laskin JL, Masters G, Milton D, Nordquist L, Pao W, Pfister DG, Piantadosi S, Schiller JH, Smith R, Smith TJ, et al. 2011 Focused update of 2009 american society of clinical oncology clinical practice guideline update on chemotherapy for stage iv non-small-cell lung cancer. J Clin Oncol. 2011; 29:3825–3831.
12. Ma ES, Ng WK and Wong CL. EGFR gene mutation study in cytology specimens. Acta Cytol. 2012; 56:661–668.
13. Betz BL, Roh MH, Weigelin HC, Placido JB, Schmidt LA, Farmen S, Arenberg DA, Kalemkerian GP and Knoepp SM. The application of molecular diagnostic studies interrogating EGFR and KRAS mutations to stained cytologic smears of lung carcinoma. Am J Clin Pathol. 2011; 136:564–571.
14. Malapelle U, de Rosa N, Rocco D, Bellevicine C, Crispino C, Illiano A, Piantedosi FV, Nappi O and Troncone G. EGFR and KRAS mutations detection on lung cancer liquid-based cytology: a pilot study. J Clin Pathol. 2012; 65:87–91.
15. Bravaccini S, Tumedei MM, Ulivi P, Zoli W, Calistri D, Candoli P, Amadori D and Puccetti M. ALK translocation detection in non-small cell lung cancer cytological samples obtained by TBNA or EBUS-TBNA. Cytopathology. 2015. doi: 10.1111/cyt.12237. PMID: 25757141.
16. Wang W, Tang Y, Li J, Jiang L, Jiang Y and Su X. Detection of ALK rearrangements in malignant pleural effusion cell blocks from patients with advanced non-small cell lung cancer: a comparison of Ventana immunohistochemistry and fluorescence in situ hybridization. Cancer Cytopathol. 2015; 123:117–122.
17. Yamaguchi F, Kugawa S, Tateno H, Kokubu F and Fukuchi K. Analysis of EGFR, KRAS and P53 mutations in lung cancer using cells in the curette lavage fluid obtained by bronchoscopy. Lung Cancer. 2012; 78:201–206.
18. Mitiushkina NV, Iyevleva AG, Poltoratskiy AN, Ivantsov AO, Togo AV, Polyakov IS, Orlov SV, Matsko DE, Novik VI and Imyanitov EN. Detection of EGFR mutations and EML4-ALK rearrangements in lung adenocarcinomas using archived cytological slides. Cancer Cytopathol. 2013; 121:370–376.
19. Bruno P, Mariotta S, Ricci A, Duranti E, Scozzi D, Noto A, Mancini R, Giarnieri E and Giovagnoli MR. Reliability of direct sequencing of EGFR: comparison between cytological and histological samples from the same patient. Anticancer Res. 2011; 31:4207–4210.
20. Pang B, Dettmer M, Ong CW, Dhewar AN, Gupta S, Lim GL, Nga ME, Seet JE, Qasim A, Chin TM, Soo R, Soong R and Salto-Tellez M. The positive impact of cytological specimens for EGFR mutation testing in non-small cell lung cancer: a single South East Asian laboratory's analysis of 670 cases. Cytopathology. 2012; 23:229–236.
21. Li S, Li L, Zhu Y, Huang C, Qin Y, Liu H, Ren-Heidenreich L, Shi B, Ren H, Chu X, Kang J, Wang W, Xu J, Tang K, Yang H, Zheng Y, et al. Coexistence of EGFR with KRAS, or BRAF, or PIK3CA somatic mutations in lung cancer: a comprehensive mutation profiling from 5125 Chinese cohorts. Br J Cancer. 2014; 110:2812–2820.
22. Oshita F, Matsukuma S, Yoshihara M, Sakuma Y, Ohgane N, Kameda Y, Saito H, Yamada K, Tsuchiya E and Miyagi Y. Novel heteroduplex method using small cytology specimens with a remarkably high success rate for analysing EGFR gene mutations with a significant correlation to gefitinib efficacy in non-small-cell lung cancer. Br J Cancer. 2006; 95:1070–1075.
23. Lozano MD, Zulueta JJ, Echeveste JI, Gurpide A, Seijo LM, Martin-Algarra S, Del Barrio A, Pio R, Idoate MA, Labiano T and Perez-Gracia JL. Assessment of epidermal growth factor receptor and K-ras mutation status in cytological stained smears of non-small cell lung cancer patients: correlation with clinical outcomes. Oncologist. 2011; 16:877–885.
24. Proietti A, Ali G, Pelliccioni S, Lupi C, Sensi E, Boldrini L, Servadio A, Chella A, Ribechini A, Cappuzzo F, Miccoli M and Fontanini G. Anaplastic lymphoma kinase gene rearrangements in cytological samples of non-small cell lung cancer: comparison with histological assessment. Cancer Cytopathol. 2014; 122:445–453.
25. Neat MJ, Foot NJ, Hicks A, Breen R, Wilkins B, McLean E and Santis G. ALK rearrangements in EBUS-derived transbronchial needle aspiration cytology in lung cancer. Cytopathology. 2013; 24:356–364.
26. Wang WY, Liang DN, Yao WQ, Wu WL, Li JN, Chen M, Liao DY, Zhang M and Li GD. Immunohistochemical screening and fluorescence in situ hybridization confirmation of ALK translocation in lung adenocarcinoma and its clinicopathological significance: a single-center large-scale investigation of Chinese patients. Hum Pathol. 2014; 45:1414–1422.
27. Molina-Vila MA, Bertran-Alamillo J, Gasco A, Mayo-de-las-Casas C, Sanchez-Ronco M, Pujantell-Pastor L, Bonanno L, Favaretto AG, Cardona AF, Vergnenegre A, Majem M, Massuti B, Moran T, Carcereny E, Viteri S and Rosell R. Nondisruptive p53 mutations are associated with shorter survival in patients with advanced non-small cell lung cancer. Clin Cancer Res. 2014; 20:4647–4659.
28. Ying J, Guo L, Qiu T, Shan L, Ling Y, Liu X and Lu N. Diagnostic value of a novel fully automated immunochemistry assay for detection of ALK rearrangement in primary lung adenocarcinoma. Ann Oncol. 2013; 24:2589–2593.
29. Shan L, Wang Z, Guo L, Sun H, Qiu T, Ling Y, Li W, Li L, Liu X, Zheng B, Lu N and Ying J. Concurrence of EGFR amplification and sensitizing mutations indicate a better survival benefit from EGFR-TKI therapy in lung adenocarcinoma patients. Lung Cancer. 2015; 89:337–342.
30. Frampton GM, Fichtenholtz A, Otto GA, Wang K, Downing SR, He J, Schnall-Levin M, White J, Sanford EM, An P, Sun J, Juhn F, Brennan K, Iwanik K, Maillet A, Buell J, et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol. 2013; 31:1023–1031.

