Introduction
KRAS is a member of the ras gene family which encodes small G proteins with intrinsic GTPase activity. GTPase activity leads to protein inactivation and activates downstream effectors involved in multiple pathways including proliferation, differentiation and apoptosis. Point mutations occur in tumors resulting in the loss of intrinsic GTPase activity and consequently in the deregulation of cell proliferation signals [1].
KRAS is the most frequently mutated oncogene in Non-Small-Cell Lung Cancer (NSCLC) [2]. KRAS mutations are present in approximately 20% of lung adenocarcinomas, are more frequent in smokers, while infrequent in squamous cell tumors [3]. KRAS mutations in NSCLC are mainly missense in exon 2, codon 12 and 13, although other rare variants, such as codon 61, are also occasionally detected [4].
Although the ras gene was discovered almost thirty years ago, the role of KRAS mutations as prognostic and predictive markers in NSCLC cancer is still contentious [5, 6]. The available meta-analyses suggest that patients with wild-type KRAS have a better prognosis. On the other hand, the predictive role of KRAS mutations is uncertain caused by evidence mainly based on retrospective series with contradicting results likely due to patients selection bias, and therefore to the lack of proper planned randomized trials [7-11].
In addition, it seems that different types of KRAS mutations, according to the replaced bases, have a different role in carcinogenesis and drug response [12-15].
The aim of the study was to investigate, in terms of overall survival (OS) and progression free survival (PFS), the role of KRAS mutations in advanced EGFR wild-type NSCLC patients treated with first-line platinum-based chemotherapy.
Results
Between October 12, 2007 and March 13, 2012 we collected and genotyped for KRAS and EGFR 540 patients in the TAILOR trial [16]. Of these, 213 patients were not eligible for the present study for various reasons: adjuvant therapy (N = 177), missing data (N = 24), early stages at the time of first-line treatment (N = 6), KRAS status not evaluable (N = 3) and early death (N = 3). Eighty patients with tumor harboring EGFR gene mutations were also excluded. Of the remaining 247 eligible patients, 187 (76.8%) had wild-type KRAS tumor, whereas 60 (24.3%) had a tumor with a mutated KRAS. Nine different types of KRAS mutations were identified and the three most common were G12C (43.3%), G12V (23.3%) and G12D (10.0%) as reported in Table 1. G13 mutation isoforms (G13C and G13D) were seen in 6.7% (N = 4) of all mutated cases.
Table 1: Different type of KRAS mutations
KRAS mutations |
N |
% |
| G12A | 6 | 10.0 |
| G12C | 26 | 43.3 |
| G12D | 6 | 10.0 |
| G12F | 2 | 3.3 |
| G12R | 1 | 1.7 |
| G12S | 1 | 1.7 |
| G12V | 14 | 23.3 |
| G13C | 2 | 3.3 |
| G13D | 2 | 3.3 |
The CONSORT diagram is illustrated in Figure 1 whereas the baseline characteristics of the patients included in the study according to KRAS mutational status are illustrated in Table 2.
KRAS mutational status was associated with tumor histology (P = 0.038) and smoking habit (P = 0.006). The mutated KRAS subgroup of patients had, as expected, a higher percentage of adenocarcinoma histology (85.0% compared to 65.8% for mutated and wild-type respectively) and a lower prevalence of never smoker patients (6.7% compared to 22.5% for mutated and wild-type respectively). All the other characteristics were well balanced between the two groups.
All patients received platinum-doublet chemotherapy in the first-line setting with higher percentage of wild-type KRAS tumor patients receiving gemcitabine (57.1%) as compared to mutated tumor patients (37.9%). The latter received pemetrexed in a higher (50.0%) percentage compared to wild-type (30.4%). Vinorelbine option was less frequent but homogenously administered (12.5% and 12.1% in wild-type and mutated tumor patients respectively).
One-hundred and thirty-five patients were randomized in the main clinical trial. In particular, 52.4% and 56.7% of wild-type and mutated patients respectively were further treated with docetaxel in second-line treatment. On the other hand, 47.6% of wild-type and 43.3% of mutated patients received erlotinib. Among the randomized patients a higher percentage of patients with KRAS mutant tumor did not reach the third-line treatment (76.7%) compared to wild-type (54.3%) (P = 0.028).
Table 2: Patient’s characteristics
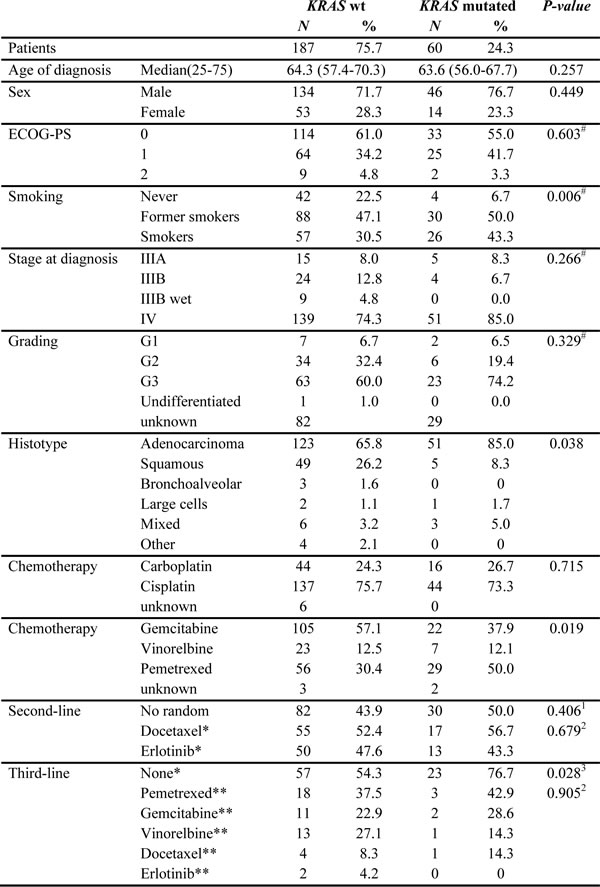
# Chi-square for trend
* percentage calculated on randomized patients
** percentage calculated on patients who received third-line treatment
1 comparison between randomized and not randomized patients
2 comparison among different treatments
3 comparison between treatment performed and no treatment
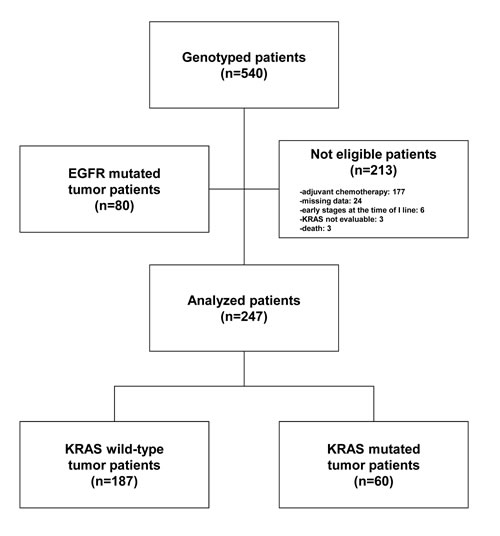
Figure 1: Patient CONSORT diagram.
Survival outcomes
After a median follow-up of 52.5 months (95%Confidence Interval [CI] 42.0-64.7), 225 patients had progressed or died and 202 had died.
Median OS was 14.4 months (95%CI 10.9-19.4) for patients with wild-type KRAS tumor and 10.6 months (95%CI 8.4-12.9) for those with mutant tumor (Figure 2A). The survival of patients with tumor harboring mutated KRAS was significantly lower than in wild-type group (unadjusted Hazard Ratio [HR] = 1.41 95%CI: 1.03-1.94 P = 0.032; adjusted HR = 1.39, 95%CI: 1.00-1.94 P = 0.050). The OS of patients expressing the three most common KRAS mutations, separately analyzed (G12C, G12D and G12V), was not different when compared to wild-type although all mutations had worsening trend (Figure 3A). The 4 patients harboring G13 KRAS mutations showed a median OS of 9.0 months.
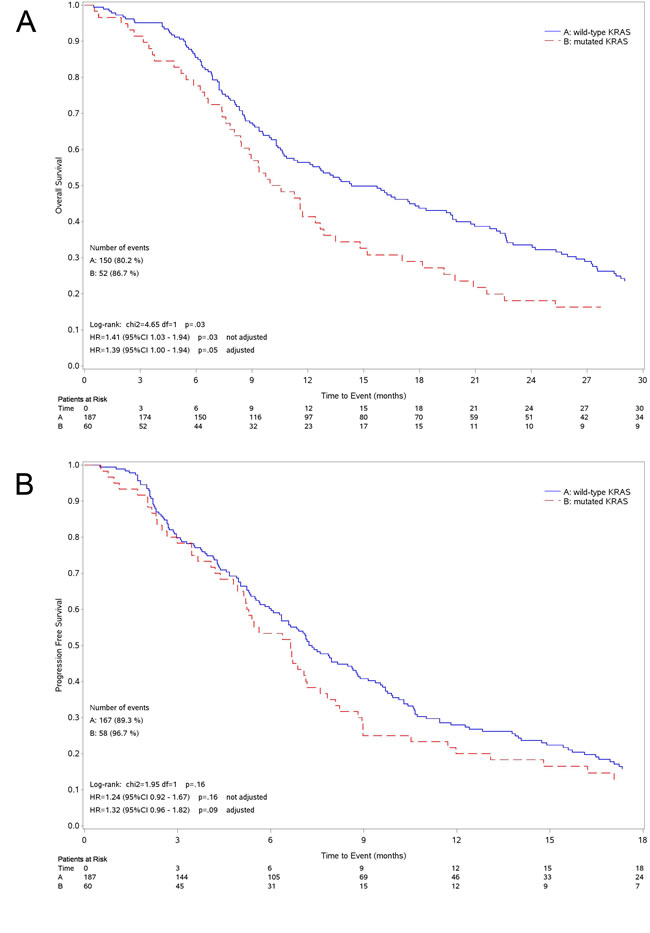
Figure 2: Kaplan-Meier curves for survival. Curves for overall survival A. and progression free survival B. according to the KRAS status.
ECOG-PS (HR = 1.79, 95%CI: 1.41-2.27 P < 0.001), sex (HR = 0.65, 95%CI: 0.47-0.89 P = 0.007) and tumor stage (HR = 1.18, 95%CI: 1.02-1.37 P = 0.024) were clinical factors significantly associated with OS. Risk estimate covariates are reported in Table 2.
Median PFS was 7.2 months (95%CI 6.3-8.8) for patients with wild-type KRAS tumor and 6.6 months (95%CI: 5.1-7.6) for those with mutant tumor (Figure 2B) (unadjusted HR = 1.24, 95%CI: 0.92-1.67 P = 0.164; adjusted HR = 1.32, 95%CI: 0.96-1.82 P = 0.092). As for OS, we could not find any statistical difference among patients with tumor harboring the three more common KRAS mutations, separately analyzed, and those with wild-type tumor (Figure 3B). Patients with G13 KRAS mutant tumor showed a median PFS of 3.9 months.
Tumor histology (HR = 0.70, 95%CI: 0.51-0.97 P = 0.033), ECOG-PS (HR = 1.53, 95%CI: 1.23-1.90 P < .001), and tumor stage (HR = 1.29, 95%CI: 1.11-1.48 P = 0.001) were statistically correlated with PFS. Risk estimate covariates are reported in Table 4.
Table 3: Prognostic evaluation of clinical and histopatological characteristics – Overall Survival
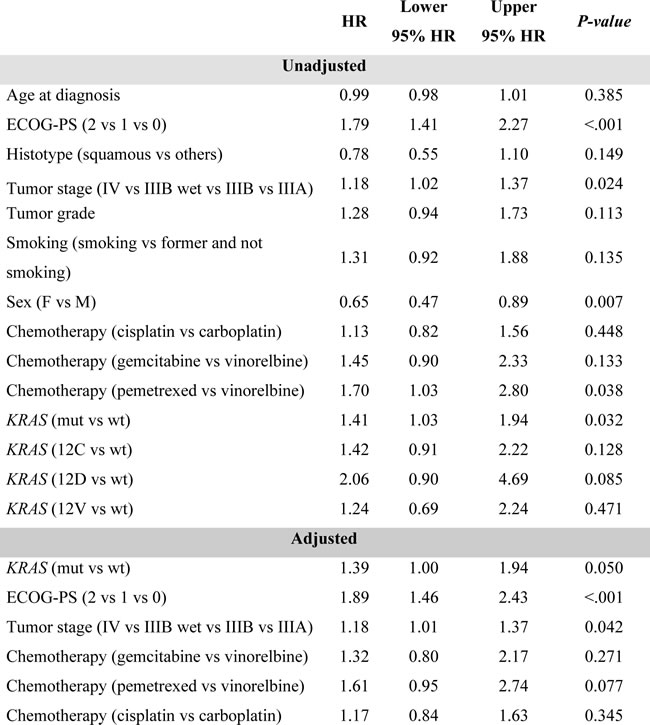
Table 4: Prognostic evaluation of clinical and histopatological characteristics- Progression Free Survival
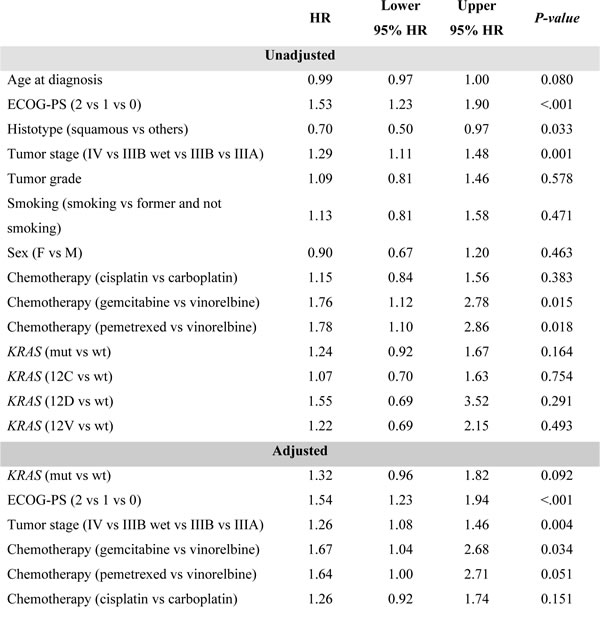
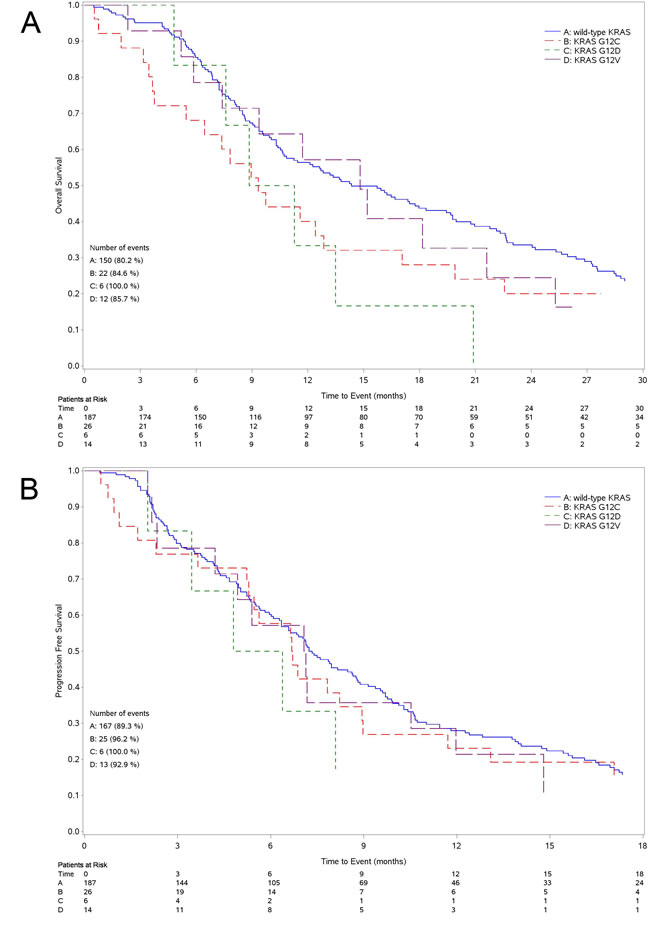
Figure 3: Kaplan Meier curves for survival. Curves for overall survival A. and progression free survival B. according to the main different types of mutations (G12C, G12D, G12V).
Discussion
In the last 20 years many studies, including meta-analyses, with more than 8,000 patients considered have been published analyzing the prognostic and predictive role of KRAS mutations in NSCLC The majority of these results indicated KRAS as a negative prognostic and predictive marker [7-11, 17]. All these data were drawn from uncontrolled series and they included patients with different biological characteristics.
Our study confirms that a small negative prognostic effect of KRAS mutations can be observed in advanced NSCLC patients treated with a platinum-based doublet when EGFR mutant patients are excluded from the analysis. TAILOR results are superimposable to those found by Mascaux et al., the only meta-analysis performed on the role of KRAS in predicting efficacy of chemotherapy [10].
In addition, our epidemiological results are in line with literature: KRAS mutations are strongly correlated with smoking habit and adenocarcinoma histology and are mutually exclusive with EGFR mutations [3].
In a planned subgroup analysis of the main TAILOR study the KRAS mutations effect was not so clearly observed, although we cannot exclude a similar effect to the one presented in this paper with higher number of patients [18]. This difference may lay in the low statistical power of the TAILOR second-line subgroup analysis, or it may depend on a selected population with a better prognosis that can receive a second-line. Considering that a higher percentage of patients with a mutant KRAS tumor did not even underwent third-line treatment compared to wild-type we do not find these results in contrast with those previously reported by our group.
This analysis has some strengths and some limitations. The major strength is that this is the only pre-planned study in which KRAS status was evaluated in all consecutive patients included in the analysis and for whom EGFR mutations were also known. However, it is not possible to discriminate if the slightly worse prognosis of KRAS mutated patients is dependent on the poor prognostic effect of KRAS mutation or on a lesser response to the chemotherapy of these patients.
The treatment choice was done by physicians based on patients and disease characteristics therefore we cannot differentiate the final effect of cisplatin, carboplatin or other chemotherapeutic doublet agents according to KRAS status. On the other hand, the association between KRAS status and type of first-line chemotherapy can be explained by choice of treatment based on tumor histology, which is associated to the KRAS status. Nevertheless, the poorer outcome of mutated KRAS patients cannot be explained by a confounding effect related to second-line treatments given that wild-type and mutated KRAS patients were equally distributed in the two treatment arms in the main trial.
Results of the LACE-Bio pooled analysis, the largest pooled analysis encompassing 1543 patients from four large adjuvant studies (JBR.10, ANITA, CALGB, IALT), showed that there is no difference in terms of prognosis in early stage lung cancer patients with either wild-type or mutated KRAS [19]. Our different results could raise the hypothesis that KRAS mutations may play a different role in early and in advanced disease. This biological hypothesis could corroborate the evidence that, in advanced stages, KRAS is a condition necessary, but not sufficient to explain a more aggressive phenotype. Other additional factor(s) could contribute to this KRAS effect such as DNA repair capability. Advanced KRAS mutated tumors might have a DNA repair imbalance more pronounced than in early stages and our group is working actively on this issue [20, 21].
Due to small sample number, we are not able to elucidate any role for the three single most common KRAS mutations although the different variants seem to confer different OS when compared to the wild-type. Interestingly enough, preliminary preclinical data obtained in our laboratory in isogenic NSCLC cells differing only for the presence of KRAS mutations seem to suggest that the activity of cisplatin is different in cells expressing the different KRAS mutations [12].
Furthermore, the LACE-Bio pooled analysis pointed out that mutations in the codon 13 may confer a worse prognosis than others [19]. Although in TAILOR we have only an handful of G13 mutations, results seem to confirm this hypothesis.
The concept that not all KRAS mutations behave in the same way and that they differently impact on tumor progression has been addressed [12, 14, 15]. This data together with the indications reported in our study may suggest that proper trials need to be planned to define the role of the specific mutations in terms of response to treatment and tumor progression.
In conclusion, although KRAS showed a prognostic effect in first-line platinum-based treatment in advanced NSCLC, this study leads us to conclude that it is not warranted to test KRAS in clinical practice, at least until a specific targeted therapy is available for this group of patients. However, the potential mechanism of resistance to platinum-based therapies of these tumors should be further explored.
materials and Methods
Study design
TAILOR was a not-for-profit multicentre, open label, randomized trial, funded by the Italian Regulatory Agency AIFA and conducted in 52 Italian hospitals, comparing erlotinib versus docetaxel in second-line NSCLC and details have been published elsewhere [16]. Within the TAILOR trial we planned an ancillary study to assess the prognostic value of KRAS mutations in advanced NSCLC patients treated with a first-line platinum containing regimens.
Briefly, tumor samples from registered patients were histologically centrally reclassified according with the 2004 WHO classification. Suitable samples were genotyped in parallel by investigators in two independent laboratories using two different techniques: EGFR by Sanger’s sequencing and restriction fragment length polymorphism whereas KRAS by Sanger sequencing and high-resolution melting analysis. Scorpion/ARMS technique was used for low-material samples. The Italian central authority and ethical review board at each participating Institution approved the protocol. The study complied with the declaration of Helsinki and was done in accordance with good clinical practice guidelines. Trial Registration: clinicaltrials.gov identifier NCT00637910.
Patients and eligibility criteria
Participating centers registered all consecutive patients with NSCLC before or during first-line platinum-based chemotherapy as well as patients recurred after a first-line adjuvant platinum-based chemotherapy. Only those with both EGFR and KRAS status centrally determined were included in the trial. All patients received platinum-based chemotherapy in combination with either vinorelbine, gemcitabine or pemetrexed according to the physician’s choice. Combinations with taxanes and with anti-EGFR agents were not allowed. Patients with EGFR mutations, early stages patients and patients receiving the adjuvant therapy were excluded from this analysis. All patients had an Eastern Cooperative Oncology Group (ECOG) Performance Status (PS) between 0 and 2 and were at least 18 years of age. Exclusion criteria included any evidence of serious co-morbidities that the investigator judged as a contraindication to the participation in the study, pregnancy and breast feeding. Patients were considered former smokers if they smoked more than 100 cigarettes in their life and they stopped this habit for at least one year at the time of diagnosis as used in most of smoking-habits analyses [22, 23]. All patients who were eligible for participation provided written informed consent with all applicable governing regulations before undergoing any study procedure.
Statistical methods
The analysis was planned at the occurrence of 200 events, needed to detect a Hazard Ratio ≥1.60 (mutated KRAS vs. wild-type KRAS) assuming a KRAS mutation frequency of 25% with a statistical power of 80% and two-sided type I error of 5%.
The primary endpoint was OS defined as the time from the day of first-line treatment start to the date of death from any cause. The secondary endpoint was PFS defined as the time from the day of first-line treatment start up to the date of first progression or death from any cause, whichever came first. Patients who had not died or had disease progression at the date of study cutoff were censored at the last available information on status. Time-to-event data were described by the Kaplan-Meier curves. Cox proportional hazards models were used for univariate and multivariate analysis (adjusted for ECOG-PS, stage, type of first-line chemotherapy) to estimate and test demographic characteristics, clinical features, and biologic parameters for their associations with OS and PFS. We also evaluated OS and PFS according to the different subtypes of KRAS mutations.
Moreover we evaluated the association between the status of KRAS and clinical and histopathological characteristics by means of Chi-square test.
Results were expressed as HRs and their 95% confidence intervals and P values for two sided hypothesis test were reported.
All statistical analyses were carried out using SAS version 9.2 (SAS Institute, Cary, NC).
Funding
This work was supported by grants from the Agenzia Italiana del Farmaco (FARM6F5JER), Fondazione CARIPLO (2010-0794) and AIRC (IG-12915).
Conflicts of Intrests
The authors have declared no conflicts of interest.
References
1. Pylayeva-Gupta Y, Grabocka E and Bar-Sagi D. RAS oncogenes: weaving a tumorigenic web. Nat Rev Cancer. 2011; 11:761-774.
2. Riely GJ, Marks J and Pao W. KRAS mutations in non-small cell lung cancer. Proc Am Thorac Soc. 2009; 6:201-205.
3. Piva S, Ganzinelli M, Garassino MC, Caiola E, Farina G, Broggini M and Marabese M. Across the universe of k-ras mutations in non-small-cell-lung cancer. Curr Pharm Des. 2014; 20:3933-3943.
4. Karachaliou N, Mayo C, Costa C, Magri I, Gimenez-Capitan A, Molina-Vila MA and Rosell R. KRAS mutations in lung cancer. Clin Lung Cancer. 2013; 14:205-214.
5. Loriot Y, Mordant P, Deutsch E, Olaussen KA and Soria JC. Are RAS mutations predictive markers of resistance to standard chemotherapy? Nat Rev Clin Oncol. 2009; 6:528-534.
6. Roberts PJ and Stinchcombe TE. KRAS mutation: should we test for it, and does it matter? J Clin Oncol. 2013; 31:1112-1121.
7. Garassino MC, Borgonovo K, Rossi A, Mancuso A, Martelli O, Tinazzi A, Di Cosimo S, La Verde N, Sburlati P, Bianchi C, Farina G and Torri V. Biological and Clinical Features in Predicting Efficacy of Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitors: A Systematic Review and Meta-analysis. Anticancer Research. 2009; 29:2691-2701.
8. Linardou H, Dahabreh IJ, Kanaloupiti D, Siannis F, Bafaloukos D, Kosmidis P, Papadimitriou CA and Murray S. Assessment of somatic k-RAS mutations as a mechanism associated with resistance to EGFR-targeted agents: a systematic review and meta-analysis of studies in advanced non-small-cell lung cancer and metastatic colorectal cancer. The Lancet Oncology. 2008; 9:962-972.
9. Mao C, Qiu L-X, Liao R-Y, Du F-B, Ding H, Yang W-C, Li J and Chen Q. KRAS mutations and resistance to EGFR-TKIs treatment in patients with non-small cell lung cancer: A meta-analysis of 22 studies. Lung Cancer. 2010; 69:272-278.
10. Mascaux C, Iannino N, Martin B, Paesmans M, Berghmans T, Dusart M, Haller A, Lothaire P, Meert AP, Noel S, Lafitte JJ and Sculier JP. The role of RAS oncogene in survival of patients with lung cancer: a systematic review of the literature with meta-analysis. Br J Cancer. 2005; 92:131-139.
11. Meng D, Yuan M, Li X, Chen L, Yang J, Zhao X, Ma W and Xin J. Prognostic value of K-RAS mutations in patients with non-small cell lung cancer: a systematic review with meta-analysis. Lung Cancer. 2013; 81:1-10.
12. Garassino MC, Marabese M, Rusconi P, Rulli E, Martelli O, Farina G, Scanni A and Broggini M. Different types of K-Ras mutations could affect drug sensitivity and tumour behaviour in non-small-cell lung cancer. Ann Oncol. 2011; 22:235-237.
13. Janakiraman M, Vakiani E, Zeng Z, Pratilas CA, Taylor BS, Chitale D, Halilovic E, Wilson M, Huberman K, Ricarte Filho JC, Persaud Y, Levine DA, Fagin JA, Jhanwar SC, Mariadason JM, Lash A, et al. Genomic and Biological Characterization of Exon 4 KRAS Mutations in Human Cancer. Cancer Res. 2010; 70:5901-5911.
14. Ihle NT, Byers LA, Kim ES, Saintigny P, Lee JJ, Blumenschein GR, Tsao A, Liu S, Larsen JE, Wang J, Diao L, Coombes KR, Chen L, Zhang S, Abdelmelek MF, Tang X, et al. Effect of KRAS Oncogene Substitutions on Protein Behavior: Implications for Signaling and Clinical Outcome. Journal of the National Cancer Institute. 2012; 104:228-239.
15. Miller MS and Miller LD. RAS mutations and oncogenesis: not all RAS mutations are created equally. Frontiers in Genetics. 2012; 2.
16. Garassino MC, Martelli O, Broggini M, Farina G, Veronese S, Rulli E, Bianchi F, Bettini A, Longo F, Moscetti L, Tomirotti M, Marabese M, Ganzinelli M, Lauricella C, Labianca R, Floriani I, et al. Erlotinib versus docetaxel as second-line treatment of patients with advanced non-small-cell lung cancer and wild-type EGFR tumours (TAILOR): a randomised controlled trial. Lancet Oncol. 2013; 14:981-988.
17. Ferte C, Besse B, Dansin E, Parent F, Buisine MP, Copin MC, Penel N and Soria JC. Durable responses to Erlotinib despite KRAS mutations in two patients with metastatic lung adenocarcinoma. Ann Oncol. 2010; 21:1385-1387.
18. Rulli E, Marabese M, Torri V, Farina G, Veronese S, Bettini A, Longo F, Moscetti L, Ganzinelli M, Lauricella C, Copreni E, Labianca R, Martelli O, Marsoni S, Broggini M and Garassino MC. Value of KRAS as prognostic or predictive marker in NSCLC: results from the TAILOR trial. Ann Oncol. 2015; 26:2079-84.
19. Shepherd FA, Domerg C, Hainaut P, Janne PA, Pignon JP, Graziano S, Douillard JY, Brambilla E, Le Chevalier T, Seymour L, Bourredjem A, Le Teuff G, Pirker R, Filipits M, Rosell R, Kratzke R, et al. Pooled analysis of the prognostic and predictive effects of KRAS mutation status and KRAS mutation subtype in early-stage resected non-small-cell lung cancer in four trials of adjuvant chemotherapy. J Clin Oncol. 2013; 31:2173-2181.
20. Vilmar AC, Santoni-Rugiu E and Sorensen JB. ERCC1 and histopathology in advanced NSCLC patients randomized in a large multicenter phase III trial. Ann Oncol. 2010; 21:1817-1824.
21. Postel-Vinay S, Vanhecke E, Olaussen KA, Lord CJ, Ashworth A and Soria JC. The potential of exploiting DNA-repair defects for optimizing lung cancer treatment. Nat Rev Clin Oncol. 2012; 9:144-155.
22. Janne PA, Wang X, Socinski MA, Crawford J, Stinchcombe TE, Gu L, Capelletti M, Edelman MJ, Villalona-Calero MA, Kratzke R, Vokes EE and Miller VA. Randomized phase II trial of erlotinib alone or with carboplatin and paclitaxel in patients who were never or light former smokers with advanced lung adenocarcinoma: CALGB 30406 trial. J Clin Oncol. 2012; 30:2063-2069.
23. Pham D, Kris MG, Riely GJ, Sarkaria IS, McDonough T, Chuai S, Venkatraman ES, Miller VA, Ladanyi M, Pao W, Wilson RK, Singh B and Rusch VW. Use of cigarette-smoking history to estimate the likelihood of mutations in epidermal growth factor receptor gene exons 19 and 21 in lung adenocarcinomas. J Clin Oncol. 2006; 24:1700-1704.

