Introduction
Cutaneous melanoma, arising from abnormal proliferation of melanocytes in the epidermis, is one of the most aggressive forms of skin cancer [1]. Tumor metastasis to distant organs is responsible for the majority of melanoma-related death; only 14% of metastatic melanoma patients survive for 5 years [2]. The treatment of metastatic melanoma has been notably improved by recent development of the specific MAP-Kinase (BRAFV600E, MEK) inhibitors and the immune checkpoint antibodies (anti-CTLA-4, anti-PD1/PDL1) [3-6]. Both therapies have shown survival benefit for patients with metastatic melanoma, however, both regimens have their own limitations [7]. Patients with metastatic melanoma still face a very poor prognosis: with a median survival of well under one year [8].
Melanoma has long been recognized as a highly heterogeneous disease [9-10]. It has been shown that approximately 50% of patients develop metastases within 15 years after treatment of the primary melanoma, and the occurrence rate of metastasis is 15% in patients with a thin (<1 mm) melanoma after their initial treatment [11]. However, the currently used AJCC (American Joint Committee of Cancer) staging system, which is based on clinical and histological parameters, although highly useful as a general guideline for prognostication, cannot precisely define such metastasis/mortality risks in many cases. For example, sentinel node biopsy has been recommended by AJCC as an important prognostic factor in early stage melanoma. But one third of patients diagnosed with metastatic melanoma do not present with regional lymph node involvement, and therefore are not detected by sentinel lymph node biopsy [12]. Some other factors that are not mentioned in AJCC staging system have potential to stratify metastatic risk. For example, a very recent study demonstrated that melanogenesis (the biochemical process to produce melanin by melanocytes) correlates with melanoma clinical outcome and promotes tumor progression [13-15]. Development of a prognostic assay that could triage high risk primary melanomas will be highly valuable for melanoma management and beneficial for melanoma patients.
To metastasize, tumor cells need to gain a series of biological capabilities to achieve invasion, distant growth, extravasation and colonization [16]. Acquisition of these capabilities requires certain genetic and epigenetic events occurring in the tumor cells. Therefore, the heterogeneity of tumors, including metastatic potential, is dictated by underlying genetic and molecular alterations [17]. In this regard, molecular or genetic expression profiles and potential biomarkers could be utilized to evaluate the metastatic risk of primary melanoma and predict patient survival [18-23]. However, previous studies on melanoma indicated that individual biomarkers are of narrow statistical significance and are unlikely to be widely adopted [23], so combinations of two or more biomarkers have gained increasing importance. Gould-Rothberg et al. found a tissue microarray and genetic-algorithm based five-marker prognostic model for stage II melanoma patients [24]; while Meyer et al. demonstrated a seven-marker signature to predict clinical outcome in malignant melanoma [25]. So far, none of these combinative biomarkers have become standard use in the clinical setting. These prognostic models still need to be validated in further prospective tests or clinical trials.
In this present study, we retrieved the melanoma tissue microarray clinical data for seven previously reported independent prognostic biomarkers [26-32], performed statistical analysis, and identified a molecular signature (loss of both KAI1 and p27) that defined a metastatic-like-subgroup (ML) among patients with primary melanomas, in both the training dataset and in a validation dataset. Our data showed that the ML subgroup had a poor 5-year melanoma-specific survival compared to the non-metastatic like (NML) subgroup (P < 0.001, Log-rank test). Additionally, multivariate Cox regression analysis revealed that KAI1-/p27- is an independent prognostic factor in primary melanomas, showing a stronger correlation with patient survival than when used as individual markers.
Results
KAI1 and p27 differentiate metastatic from primary melanomas
First of all, logistic regression analysis was performed to identify the signature proteins that most efficiently discriminated metastatic from primary melanomas. Seven previously reported independent prognostic biomarkers were selected based on the correlation with metastatic tumors [27, 35-40] (Table 1). Representative staining of BRAF, Dicer, Fbw7, KAI1, MMP2, p27 and Tip60 is shown in Figure 1. As shown in the images, cytoplasmic staining was observed for BRAF, MMP2, Dicer, KAI1, and Tip60, whereas nuclear staining was observed for Fbw7 and p27.
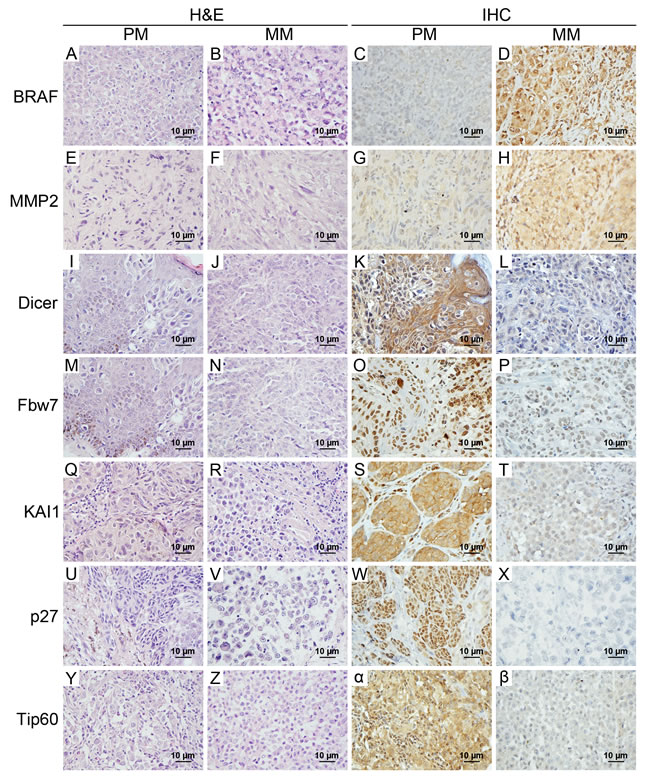
Figure 1: Representative H&E and immunohistochemical staining of 7 candidate biomarkers in primary melanomas and metastatic melanomas. Cytoplasmic staining was investigated for BRAF, MMP2, Dicer, KAI1, and Tip60, and nuclear staining was observed for Fbw7 and p27. Metastatic melanomas had overexpression of BRAF and MMP2, but did not express/ had low levels of Dicer, Fbw7, KAI1, p27 and Tip60 compared with primary melanomas. Magnification used × 400 for H & E and immunohistochemical staining. H & E, hematoxylin and eosin; IHC, immunohistochemistry; PM, primary melanoma; MM, metastatic melanoma.
Table 1: Seven selected candidate biomarkers. |
||
Marker |
Full name |
Function |
BRAF |
V-raf murine sarcoma viral oncogene homolog B |
Proto-oncogene |
Dicer |
Dicer 1, ribonuclease type III |
Endoribonuclease |
Fbw7 |
F-box and WD repeat domain containing 7 |
E3 ubiquitin protein ligase |
KAI1 |
CD82 molecule |
Metastasis suppressor |
MMP2 |
Matrix metallopeptidase 2 |
Matrix metalloproteinase |
P27 |
Cyclin-Dependent Kinase Inhibitor 1B |
Tumor suppressor |
Tip60 |
Tat-interactive protein 60 kDa |
Regulates metastasis suppressor |
Univariate analysis showed that BRAF, Dicer, KAI1, P27 and Tip60 were differentially expressed in metastatic melanoma as compared to primary melanoma. Multiple logistic regression analyses revealed four biomarkers that were significant: BRAF (P = 0.037), Tip60 (P = 0.029), p27 (P = 9.677E-5) and KAI1 (P = 6.321E-5); KAI1 and p27 were the two most significant biomarkers (Table 2). KAI1 and p27 were expressed in 95.24% and 78.10% of metastatic tumors, and in 66.21% and 50.34% of primary melanomas, respectively. This difference between primary melanoma and metastatic melanoma expression was highly significant (P < 0.001), indicating that loss of KAI1 or p27 may represent a relatively robust feature of more advanced melanomas. Representative KAI1 and p27 staining patterns in primary melanomas are shown in Figure 2. Furthermore, we ran the same analysis on the combination of KAI1 and p27, and found loss of both proteins better differentiated metastatic from primary melanomas, suggesting that double loss of both KAI1 and p27 can serve as a potential signature for metastatic like (ML) primary melanomas.
Table 2: Expression of seven biomarkers in primary versus metastatic melanomas |
||||||
Biomarker |
Primary melanoma No. (%) |
Metastatic melanoma No. (%) |
P value |
|||
- |
+ |
- |
+ |
Univariate |
Multivariate |
|
BRAF |
79 (54.48) |
66 (45.52) |
36 (34.29) |
69 (65.71) |
0.002 |
0.037 |
Dicer |
10 (6.90) |
135 (93.10) |
23 (21.90) |
82 (78.10) |
0.001 |
0.053 |
Fbw7 |
22 (15.17) |
123 (84.83) |
21 (20.00) |
84 (80.00) |
0.318 |
0.666 |
KAI1 |
96 (66.21) |
49 (33.79) |
100 (95.24) |
5 (4.76) |
4.000E-8 |
6.321E-5 |
MMP2 |
102 (70.34) |
43 (29.66) |
63 (60.00) |
42 (40.00) |
0.089 |
0.166 |
P27 |
73(50.34) |
72(49.66) |
82(78.10) |
23(21.90) |
8.130E-6 |
9.677E-5 |
Tip60 |
30(20.69) |
115(79.31) |
46(43.81) |
59(56.19) |
8.768E-6 |
0.029 |
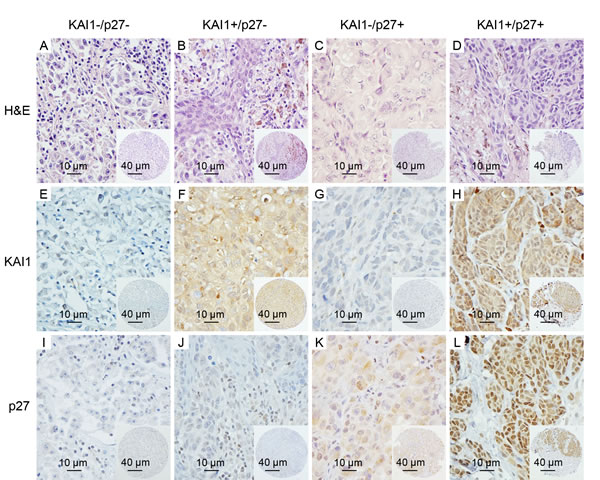
Figure 2: Representative images of KAI1 and p27 staining in subgroups of primary melanomas. A.-D. the H & E staining images of representative cores are shown for each subgroup; E. and I., absent/minimal cytoplasmic expression of KAI1 and nuclear expression of p27 (KAI1-/p27-) were seen in 48 of the 145 cases; F.-H., J.-L., moderate or high cytoplasmic expression of KAI1 or nuclear expression of p27 (KAI1+/p27- or KAI1-/p27+), or expression of both (KAI1+/p27+), are shown in 97 of 145 primary melanoma patients. (magnification: × 400); insets show corresponding tissue microarray cores (magnification: × 100).
The subgroup of primary melanoma with KAI1-/p27- signature had poor survival in the training cohort
To evaluate whether a KAI1-/p27- signature can distinguish high risk primary melanomas, and examine the prognostic value of this combined marker, we separated 145 primary melanomas into two subgroups: a metastasis-like group (ML) of 48 patients with negative expression of both KAI1 and p27 (33.10%), and a non-metastasis-like group (NML) of 97 patients with positive expression of either KAI1 or p27 or both (66.90%) (Table 3). Representative images of the expression patterns are shown in Figure 2. Kaplan-Meier survival curves showed that melanoma-specific survival is significantly reduced for patients in the metastasis-like group (52.10%) compared to those in the non-metastasis-like subgroup (76.30%, P = 0.002, Figure 3A).
Table 3: Subgroups of primary melanoma in training cohort, validation cohort and combined cohort |
|||
Population |
Subgroup |
Expression |
Case no. (%) |
Training cohort (145 cases) |
ML |
KAI1-/ P27- |
48 (33.10) |
NML |
Either KAI1 or p27 is negative |
97 (66.90) |
|
Validation cohort (92 cases) |
ML |
KAI1- P27- |
34 (37.00) |
NML |
Either KAI1 or p27 is negative |
58 (63.00) |
|
Combined cohort (237 cases) |
ML |
KAI1- P27- |
82 (34.60) |
NML |
Either KAI1 or p27 is negative |
155 (65.40) |
|
ML, metastasis-like; NML, non-metastasis-like
The ML subgroup of primary melanoma had poor survival in both the independent validation cohort and the combined cohort
To validate our results, 92 additional independent primary melanoma patients were classified into NML and ML subgroups and analysed (Table 3). Survival analysis showed that the NML group had significantly better disease-specific survival (87.9% in NML versus 70.6% in ML, P = 0.030, Figure 3B). In addition, the same trend was observed in the combined patient’s cohort: 79.8% of NML subgroup patients can survive for 5 years, while this number drops to 57.9% in ML subgroup (P = 3.00E-4, Figure 3C, Table 3).
Table 4: Clinicopathological characteristics of NML and ML subgroups in combination cohort of patients with primary melanoma. |
|||
Variable |
ML (N = 82) |
NML (N = 155) |
P value |
Age |
63.42 ± 1.97 |
59.48 ± 1.53 |
0.123 |
Sex |
0.194 |
||
Male |
48(58.54) |
77(49.68) |
|
Female |
34(41.46) |
78(50.32) |
|
Thickness |
4.94 ± 0.73 |
2.66 ± 0.24 |
0.001 |
Ulceration (%) |
0.117 |
||
Absent |
57(69.51) |
122(78.71) |
|
Present |
25(30.49) |
33(21.29) |
|
Subtype (%) |
0.830 |
||
AL |
4(4.88) |
6(3.87) |
0.714 |
LM |
8(9.76) |
18(11.61) |
0.663 |
N |
20(24.39) |
28(18.06) |
0.249 |
SS |
27(32.93) |
62(40.00) |
0.285 |
Other |
7(8.54) |
13(8.39) |
0.969 |
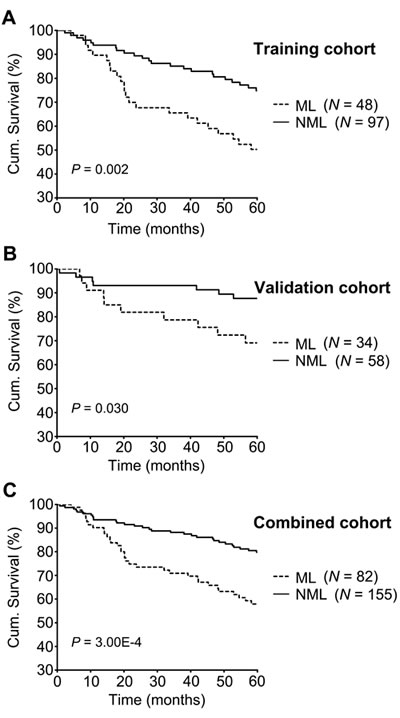
Figure 3: 5-year melanoma-specific survival analyses in training, validation and combined cohorts. Melanoma-specific survival in different subgroups of primary melanomas was analyzed by the Kaplan-Meier survival model. Patients in the ML subgroup showed poorer prognosis than patients in the NML subgroup in the training cohort A. poor survival for patients in the ML subgroup was found in the validation cohort B. and combination cohort C. Log-rank P value is indicated in the graphs.
The KAI1-/p27- signature is an independent factor for primary melanoma survival
Univariate Cox proportional hazards regression analysis showed that the ML subgroup had significantly worse disease-specific survival, with a hazard ratio of 2.421 (95% CI: 1.481-3.972; P = 4.649E-4) (Table 5), indicating that the risk of dying in the ML subgroup was 2.421 times greater compared to that of the NML subgroup. Multivariate Cox proportional hazards regression model adjusted for age, gender, thickness, ulceration were assessed (Tables 6, 7, 8). In the multivariable Cox model, the KAI1-/p27- signature was found to be of independent prognostic significance for melanoma-specific survival (P = 0.009). More important, compared to KAI1 and p27 as an individual prognostic marker, the KAI1-/p27- signature is more closely associated with melanoma patient survival: the P value is 0.009 for KAI1-/p27-, compared to 0.025, 0.264 for KAI1 and p27, respectively (Tables 6, 7, 8).
Table 5: Univariate Cox proportional hazard regression analysis in combined cohort (237 primary melanoma patients). |
|||
Hazard ratio |
95% CI |
P value |
|
Age: > 60 vs ≤ 60 y |
2.372 |
1.392 - 4.041 |
0.001 |
Gender: male vs female |
0.980 |
0.601 - 1.615 |
0.937 |
Thickness: > 2 vs ≤ 2 mm |
4.883 |
2.692 - 8.863 |
1.791E-7 |
Ulceration: + vs - |
4.354 |
2.652 - 7.168 |
6.510E-9 |
KAI1: - vs + |
2.996 |
1.602 - 5.621 |
0.001 |
p27: - vs + |
1.690 |
1.020 - 2.815 |
0.043 |
KAI1/p27: ML vs NML |
2.421 |
1.481 - 3.972 |
4.649E-4 |
CI: confidence interval; ML: metastasis-like subgroup, KAI1-/p27-; NML: non-metastasis-like subgroup: KAI1-/p27+ or KAI1+/p27- or KAI1+/p27+.
Table 6: Multivariate Cox proportional hazard regression analysis in combined cohort (237 primary melanoma patients) for KAI1 as a single biomarker. |
|||
Hazard Ratio |
95% CI |
P value |
|
Age: > 60 vs ≤ 60 y |
1.271 |
0.727-2.224 |
0.400 |
Gender: male vs female |
0.985 |
0.609-1.692 |
0.954 |
Thickness: > 2 vs ≤ 2 mm |
3.256 |
1.728-6.137 |
2.609E-4 |
Ulceration: + vs - |
2.616 |
1.535-4.459 |
4.082E-4 |
KAI1: - vs + |
2.083 |
1.096-3.953 |
0.025 |
CI: confidence interval.
Table 7: Multivariate Cox proportional hazard regression analysis in combined cohort (237 primary melanoma patients) for p27 as a single biomarker. |
|||
Hazard Ratio |
95% CI |
P value |
|
Age: > 60 vs ≤ 60 y |
1.265 |
0.725-2.205 |
0.408 |
Gender: male vs female |
0.915 |
0.550-1.522 |
0.732 |
Thickness: > 2 vs ≤ 2 mm |
3.114 |
1.647-5.888 |
3.158E-4 |
Ulceration: + vs - |
2.900 |
1.689-4.979 |
1.129E-4 |
p27: - vs + |
1.345 |
0.798-2.278 |
0.264 |
CI: confidence interval.
Loss of KAI1 and p27 in primary melanoma was correlated with thickness
It is noteworthy that ML subgroup exhibited a relatively high hazard ratio for 5-year melanoma-specific survival (HR = 1.957; 95% CI = 1.182-3.236), as compared to tumor thickness (HR = 3.114; 95% CI = 1.647-5.888) when analyzed together, suggesting a potential interdependence between KAI1- /p27- signature and tumor thickness. To investigate the clinical phenotype in the ML subgroup of primary melanoma patients, we compared the relevant patient clinicopathologic characteristics and histological features between ML and NML subgroups in the combination cohort. The associations of the KAI1-/p27- signature with other patient and tumor characteristics are shown in Table 4. There was no difference in age, gender, clinical subtype and ulceration between ML and NML subgroups. However, we found that the ML subgroup was characterized by greater mean thickness compared to the NML subgroup (4.94 vs. 2.66 mm; P = 0.001, Table 4), suggesting that the ML subgroup has more aggressive characteristics.
Table 8: Multivariate Cox proportional hazard regression analysis in combined cohort (237 primary melanoma patients) for combined KAI1-/p27- signature. |
|||
Hazard Ratio |
95% CI |
P value |
|
| Age: > 60 vs ≤ 60 y | 1.277 | 0.727-2.244 | 0.394 |
| Gender: male vs female | 0.894 | 0.523-1.486 | 0.665 |
| Thickness: > 2 vs ≤ 2 mm | 3.114 | 1.647-5.888 | 4.757E-4 |
| Ulceration: + vs - | 3.005 | 1.760-5.130 | 5.525E-5 |
| KAI1/p27: ML vs NML | 1.957 | 1.182-3.236 | 0.009 |
CI: confidence interval; ML: metastasis-like subgroup, KAI1-/p27-; NML: non-metastasis-like subgroup: KAI1-/p27+ or KAI1+/p27- or KAI1+/p27+.
Discussion
Since Fidler first demonstrated the heterogeneity of mouse melanoma cells with respect to metastatic potential in 1973 [41], several genome-wide high-throughput studies have described gene expression signatures to predict metastasis of primary melanoma patients. However, the gene-sets identified have shown minimal overlap between various studies, and lack the convenience and simplicity necessary for clinical application [42].
In the present study, we performed statistical analysis on a patient cohort with large sample size, and identified the metastasis-like subgroup of primary melanomas in order to develop a clinically effective classification model. We demonstrated that simultaneous loss of both KAI1 and p27 was a novel molecular feature associated with metastasis, discriminating between primary and metastatic melanomas, identifying a metastasis like subgroup (ML subgroup) within primary melanoma patients, and constituting a strong prognostic marker for poor survival in patients with primary melanoma.
Initially, our data demonstrated that KAI1- and p27- were the two markers among 7 previously identified independent markers that showed the most significant difference in expression level between metastatic melanomas (95.24% vs. 78.10%) and primary melanomas (66.21% vs. 50.34%, P < 10E-4) (Table 2). This result was consistent with the previous finding that highly aggressive tumors showed the lowest KAI1 and p27 expression levels [43-44]. Our study extends the previous work in that the combination of KAI1 and p27 loss is a metastatic feature and significantly differentiates metastatic melanomas from primary melanomas (P < 0.001).
Tumor metastasis suppressor KAI1 has previously been shown to interfere with multiple steps of the metastatic cascade, including proliferation, invasion, and migration [45]. Also the inverse association of KAI1 expression with formation of metastasis was reported [46]. Studies from our group have found dramatic decrease of KAI1 protein expression in human melanomas using tissue microarray technology, and KAI1 expression was negatively correlated with patient outcome [32]. The prognostic value and the role of p27 in cancer metastasis have been intensely reported [47-48]. However, no studies have examined the combination of KAI1 and p27, two important proteins for metastatic disease, to identify high-risk primary tumors.
Using the metastatic signature (loss of both KAI1- and p27-), a metastasis-like subgroup was identified accounting for 33.10% of primary melanomas. Tumors identified within the same subgroup were more likely to present with similar clinical features. Kaplan-Meier survival analysis showed that the patients in the ML subgroup have a worse 5-year survival rate compared with patients in the NML subgroup (52.1% vs 76.3%). More importantly, multivariate Cox regression analysis revealed that KAI1-/p27- is an independent prognostic factor in primary melanomas, showing a stronger correlation with patient survival than when used as individual markers. Additionally, we found that melanomas in the ML subgroup exhibited greater thickness compared to the NML subgroup (4.94 vs. 2.66 mm; P = 0.001, Table 4). However, we did not detect significant association between KAI1-/p27- signature and other important clinical parameters, such as ulceration (P = 0.117, Table 4) and subtype (P = 0.83, Table 4). Superficial spreading and nodular subtypes are responsible for most melanoma related-mortality, and the nodular subtype is the most aggressive subtype [49], yet we did not observe significant difference in the distribution of specific subtypes in ML and NML subgroups, which may due to the relatively small case number in each subtype group. Because of a lack of recorded information, we were unable to conduct analyses on other clinicopathological parameters, such as mitotic activity, tumor-infiltrating lymphocyte response, regression, lymph vascular invasion, etc.
Taken together, our results suggest that loss of KAI1 and p27 leads to enhanced metastatic potential for primary melanomas, and that restoring expression or function of KAI1 and p27 can be a potential strategy for melanoma therapy. The KAI1-/p27- signature could potentially identify a distinct subgroup of primary melanoma patients that need to be monitored more closely and treated more aggressively. This study presents a new alternative way of subgroup discrimination which will hopefully facilitate the search for subgroup-specific therapeutic targets and the development of personalized medicine for primary melanoma patients in the metastasis-like subgroup.
Materials and Methods
Ethics statement
Our study on archival melanoma biopsies was approved by the Clinical Research Ethics Board of the University of British Columbia. The experiments were performed in accordance with the Declaration of Helsinki guidelines.
Study patients and tissue microarray
The construction of melanoma tissue microarrays (TMAs) and corresponding clinical database have been described previously [30, 33]. The protocol was approved by the Institutional Review Board at the University of British Columbia. Due to core loss, 250 cases could be evaluated for all 7 markers in a training cohort, including 145 primary melanomas and 105 metastatic melanomas. Further, 92 additional cases of primary melanomas were evaluated for both KAI1 and p27. These formed the independent validation cohort used for confirmation analysis of the KAI1-/p27- signature.
Immunohistochemistry and intensity assessment
The immunohistochemistry staining was performed as described previously [33]. Briefly, Primary rabbit polyclonal anti-BRAF antibody (1:100 dilution; Sigma, St. Louis, MO, USA), rabbit polyclonal anti-Dicer (1:100 dilution; Sigma), mouse monoclonal anti-Fbw7 antibody (1:50 dilution; Abcam, Cambridge, MA), mouse monoclonal anti-KAI1 (1:100 dilution, Novus Biologicals, Littleton, CO, USA), mouse monoclonal anti-MMP2 (1:50 dilution; Biolegend, San Diego, CA, USA), mouse monoclonal anti-p27 (1:100 dilution; Santa Cruz Biotechnology, Santa Cruz, CA, USA), rabbit polyclonal anti-Tip60 (1:50 dilution; Millipore, Temecula, CA, USA) and the biotin-labeled secondary antibody (Dako Diagnostics, Glostrup, Denmark) were used. The technical negative control used for immunohistochemistry included the use of PBS instead of primary antibody, with all other conditions kept the same. Briefly, the staining intensity was scored using the following scale: no staining (0), weak (1), moderate (2), and strong (3). The percentage of positive cells was scored into 4 categories: 1 (0–25%), 2 (26–50%), 3 (51–75%), and 4 (76–100%). The staining intensity and percentage of positive cells were evaluated in a blinded manner by three independent observers (including two dermatologists) simultaneously, and a consensus score was reached for each core. Immunoreactive score (IRS) was used to determine the level of staining by multiplying the scores of staining intensity and the percentage of positive cells. Since cytoplasmic expression of Dicer was correlated with survival in our previous studies, only cytoplasmic Dicer scores were used in this present study. For each biomarker, x-tile software (version 3.6.1) was used to determine the optimized cut-off points, by selecting the maximal χ2 values of the log-rank test for survival between two groups [34].
Statistical analysis
Univariate and multivariate logistic regression analysis was performed to determine the discriminant biomarker between metastatic and primary melanomas. The best two markers were selected by lower P values; they were KAI1 and p27 in this study. The Kaplan–Meier survival analysis and log-rank test were used to evaluate the effects of KAI1/-p27- signature on the 5-year melanoma-specific survival. Univariate and multivariate Cox proportional hazards regression analysis were performed to estimate the crude hazard ratios (HRs) or adjusted HRs and their 95% confidence intervals (CI). A P value of <0.05 was considered to indicate statistical significance. All statistical analyses were carried out using the SPSS version 16.0 software (SPSS, Chicago, IL, USA).
Acknowledgments and funding
The authors thank Dr. Ladan Fazil and Scott Kwong for assistance in imaging, Dr. Gang Li for assistance in construction of melanoma tissue microarrays, Dr. Seyed Mehdi Jafarnejad for permission of reusing Dicer TMA data. This work is supported by grants from the Canadian Institutes of Health Research (CIHR; MOP-93810, MOP-110974, and CCI-117958), Canadian Cancer Society Research Institute (CCSRI), Cancer Research Society (CRS) and the Canadian Dermatology Foundation (CDF).
Conflicts of Interest
Dr. Kevin McElwee is the Chief Scientific Officer of Replicel Life Science Inc. This company has no role in this work and is not involved in melanoma research or treatment development.
References
1. Houghton AN and Polsky D. Focus on melanoma. Cancer Cell. 2002; 2:275-278.
2. Miller AJ and Mihm MC, Jr. Melanoma. N Engl J Med. 2006; 355:51-65.
3. Chapman PB, Hauschild A, Robert C, Haanen JB, Ascierto P, Larkin J, Dummer R, Garbe C, Testori A, Maio M, Hogg D, Lorigan P, Lebbe C, et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. The New England journal of medicine. 2011; 364:2507-2516.
4. Hauschild A, Grob JJ, Demidov LV, Jouary T, Gutzmer R, Millward M, Rutkowski P, Blank CU, Miller WH, Jr., Kaempgen E, Martin-Algarra S, Karaszewska B, Mauch C, et al. Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial. Lancet. 2012; 380:358-365.
5. Flaherty KT, Robert C, Hersey P, Nathan P, Garbe C, Milhem M, Demidov LV, Hassel JC, Rutkowski P, Mohr P, Dummer R, Trefzer U, Larkin JM, et al. Improved survival with MEK inhibition in BRAF-mutated melanoma. N Engl J Med. 2012; 367:107-114.
6. Hodi FS, O’Day SJ, McDermott DF, Weber RW, Sosman JA, Haanen JB, Gonzalez R, Robert C, Schadendorf D, Hassel JC, Akerley W, van den Eertwegh AJ, Lutzky J, et al. Improved survival with ipilimumab in patients with metastatic melanoma. N Engl J Med. 2010; 363:711-723.
7. Slominski AT and Carlson JA. Melanoma resistance: a bright future for academicians and a challenge for patient advocates. Mayo Clin Proc. 2014; 89:429-433.
8. Eggermont AM, Spatz A and Robert C. Cutaneous melanoma. Lancet. 2014; 383:816-827.
9. Meguerditchian AN, Asubonteng K, Young C, Lema B, Wilding G and Kane JM, 3rd. Thick primary melanoma has a heterogeneous tumor biology: an institutional series. World J Surg Oncol. 2011; 9:40.
10. Gimotty PA, Elder DE, Fraker DL, Botbyl J, Sellers K, Elenitsas R, Ming ME, Schuchter L, Spitz FR, Czerniecki BJ and Guerry D. Identification of high-risk patients among those diagnosed with thin cutaneous melanomas. J Clin Oncol. 2007; 25:1129-1134.
11. Kalady MF, White RR, Johnson JL, Tyler DS and Seigler HF. Thin melanomas: predictive lethal characteristics from a 30-year clinical experience. Ann Surg. 2003; 238(4):528-535; discussion 535-527.
12. Carlson JA, Slominski A, Linette GP, Mihm MC, Jr. and Ross JS. Biomarkers in melanoma: staging, prognosis and detection of early metastases. Expert Rev Mol Diagn. 2003; 3:303-330.
13. Brozyna AA, Jozwicki W, Carlson JA and Slominski AT. Melanogenesis affects overall and disease-free survival in patients with stage III and IV melanoma. Hum Pathol. 2013; 44:2071-2074.
14. Slominski A, Tobin DJ, Shibahara S and Wortsman J. Melanin pigmentation in mammalian skin and its hormonal regulation. Physiol Rev. 2004; 84:1155-1228.
15. Slominski A, Zmijewski MA and Pawelek J. L-tyrosine and L-dihydroxyphenylalanine as hormone-like regulators of melanocyte functions. Pigment Cell Melanoma Res. 2012; 25:14-27.
16. Hanahan D and Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011; 144:646-674.
17. Scott KL, Nogueira C, Heffernan TP, van Doorn R, Dhakal S, Hanna JA, Min C, Jaskelioff M, Xiao Y, Wu CJ, Cameron LA, Perry SR, Zeid R, et al. Proinvasion metastasis drivers in early-stage melanoma are oncogenes. Cancer Cell. 2011; 20:92-103.
18. Kabbarah O, Nogueira C, Feng B, Nazarian RM, Bosenberg M, Wu M, Scott KL, Kwong LN, Xiao Y, Cordon-Cardo C, Granter SR, Ramaswamy S, Golub T, et al. Integrative genome comparison of primary and metastatic melanomas. PLoS One. 2010; 5:e10770.
19. Hoek KS, Schlegel NC, Brafford P, Sucker A, Ugurel S, Kumar R, Weber BL, Nathanson KL, Phillips DJ, Herlyn M, Schadendorf D and Dummer R. Metastatic potential of melanomas defined by specific gene expression profiles with no BRAF signature. Pigment Cell Res. 2006; 19:290-302.
20. Riker AI, Enkemann SA, Fodstad O, Liu S, Ren S, Morris C, Xi Y, Howell P, Metge B, Samant RS, Shevde LA, Li W, Eschrich S, et al. The gene expression profiles of primary and metastatic melanoma yields a transition point of tumor progression and metastasis. BMC Med Genomics. 2008; 1:13.
21. Xu L, Shen SS, Hoshida Y, Subramanian A, Ross K, Brunet JP, Wagner SN, Ramaswamy S, Mesirov JP and Hynes RO. Gene expression changes in an animal melanoma model correlate with aggressiveness of human melanoma metastases. Mol Cancer Res. 2008; 6:760-769.
22. Tang L, Zhang W, Su B and Yu B. Long Noncoding RNA HOTAIR Is Associated with Motility, Invasion, and Metastatic Potential of Metastatic Melanoma. Biomed Res Int. 2013; 2013:251098.
23. Gould Rothberg BE, Bracken MB and Rimm DL. Tissue biomarkers for prognosis in cutaneous melanoma: a systematic review and meta-analysis. Journal of the National Cancer Institute. 2009; 101:452-474.
24. Gould Rothberg BE, Berger AJ, Molinaro AM, Subtil A, Krauthammer MO, Camp RL, Bradley WR, Ariyan S, Kluger HM and Rimm DL. Melanoma prognostic model using tissue microarrays and genetic algorithms. J Clin Oncol. 2009; 27:5772-5780.
25. Meyer S, Fuchs TJ, Bosserhoff AK, Hofstadter F, Pauer A, Roth V, Buhmann JM, Moll I, Anagnostou N, Brandner JM, Ikenberg K, Moch H, Landthaler M, et al. A seven-marker signature and clinical outcome in malignant melanoma: a large-scale tissue-microarray study with two independent patient cohorts. PLoS One. 2012; 7:e38222.
26. Chen G, Cheng Y, Zhang Z, Martinka M and Li G. Prognostic significance of cytoplasmic p27 expression in human melanoma. Cancer epidemiology, biomarkers & prevention : a publication of the American Association for Cancer Research, cosponsored by the American Society of Preventive Oncology. 2011; 20:2212-2221.
27. Chen G, Cheng Y, Tang Y, Martinka M and Li G. Role of Tip60 in Human Melanoma Cell Migration, Metastasis, and Patient Survival. J Invest Dermatol. 2012.
28. Jafarnejad SM, Ardekani GS, Ghaffari M, Martinka M and Li G. Sox4-mediated Dicer expression is critical for suppression of melanoma cell invasion. Oncogene. 2012.
29. Rotte A, Martinka M and Li G. MMP2 expression is a prognostic marker for primary melanoma patients. Cellular oncology. 2012; 35:207-216.
30. Cheng Y, Chen G, Martinka M, Ho V and Li G. Prognostic significance of Fbw7 in human melanoma and its role in cell migration. The Journal of investigative dermatology. 2013; 133:1794-1802.
31. Ardekani GS, Jafarnejad SM, Khosravi S, Martinka M, Ho V and Li G. Disease progression and patient survival are significantly influenced by BRAF protein expression in primary melanoma. Brit J Dermatol. 2013; 169:320-328.
32. Tang Y, Cheng YB, Martinka M, Ong CJ and Li G. Prognostic significance of KAI1/CD82 in human melanoma and its role in cell migration and invasion through the regulation of ING4. Carcinogenesis. 2014; 35:86-95.
33. Jafarnejad SM, Wani AA, Martinka M and Li G. Prognostic significance of Sox4 expression in human cutaneous melanoma and its role in cell migration and invasion. Am J Pathol. 2010; 177:2741-2752.
34. Camp RL, Dolled-Filhart M and Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clinical cancer research : an official journal of the American Association for Cancer Research. 2004; 10:7252-7259.
35. Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA, O’Dwyer PJ, Lee RJ, Grippo JF, Nolop K and Chapman PB. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med. 2010; 363:809-819.
36. Martello G, Rosato A, Ferrari F, Manfrin A, Cordenonsi M, Dupont S, Enzo E, Guzzardo V, Rondina M, Spruce T, Parenti AR, Daidone MG, Bicciato S, et al. A MicroRNA targeting dicer for metastasis control. Cell. 2010; 141:1195-1207.
37. Cheng Y and Li G. Role of the ubiquitin ligase Fbw7 in cancer progression. Cancer Metastasis Rev. 2012; 31:75-87.
38. Tsai YC and Weissman AM. Dissecting the diverse functions of the metastasis suppressor CD82/KAI1. FEBS Lett. 2011; 585:3166-3173.
39. Deryugina EI and Quigley JP. Matrix metalloproteinases and tumor metastasis. Cancer metastasis reviews. 2006; 25:9-34.
40. Wander SA. p27 and Metastatic Progression: Molecular Mechanisms Underlying Bone Metastasis. Open Access Dissertations. 2011; Paper 690.
41. Fidler IJ. Selection of successive tumour lines for metastasis. Nat New Biol. 1973; 242:148-149.
42. Gschaider M, Neumann F, Peters B, Lenz F, Cibena M, Goiser M, Wolf I, Wenzel J, Mauch C, Schreiner W and Wagner SN. An attempt at a molecular prediction of metastasis in patients with primary cutaneous melanoma. PLoS One. 2012; 7:e49865.
43. Wander SA, Zhao D and Slingerland JM. p27: a barometer of signaling deregulation and potential predictor of response to targeted therapies. Clinical cancer research : an official journal of the American Association for Cancer Research. 2011; 17:12-18.
44. Lijovic M, Somers G and Frauman AG. KAI1/CD82 protein expression in primary prostate cancer and in BPH associated with cancer. Cancer Detect Prev. 2002; 26:69-77.
45. Maloney SC, Fernandes BF, Penteado RC, Antecka E, Bravo-Filho V, Sanft DM and Burnier MN, Jr. Expression of the Metastasis Suppressor KAI1 in Uveal Melanoma. J Ophthalmol. 2013; 2013:683963.
46. Romanska HM and Berditchevski F. Tetraspanins in human epithelial malignancies. J Pathol. 2011; 223:4-14.
47. Hoshino D, Koshikawa N and Seiki M. A p27(kip1)-binding protein, p27RF-Rho, promotes cancer metastasis via activation of RhoA and RhoC. J Biol Chem. 2011; 286:3139-3148.
48. Chu IM, Hengst L and Slingerland JM. The Cdk inhibitor p27 in human cancer: prognostic potential and relevance to anticancer therapy. Nature reviews Cancer. 2008; 8:253-267.
49. Shaikh WR, Xiong M and Weinstock MA. The contribution of nodular subtype to melanoma mortality in the United States, 1978 to 2007. Arch Dermatol. 2012; 148:30-36.

