Introduction
TP53 gene is the most frequently mutated gene (>50%) in human cancer, indicating its crucial role as a tumor suppressor [1]. TP53 gene encodes p53 protein which is considered as “the guardian of the genome” by binding to specific DNA sequences and maintaining genomic stability [2]. P53 protein is involved in cell response to stress signals, activates DNA repair proteins and regulates the production of stem cells [3]. The fundamental role of TP53 is evident in Li-Fraumeni syndrome which is characterized by germline mutations of TP53 and predisposition to aggressive tumors like early-onset breast cancer (25%), soft-tissue or bone sarcomas (35%) and brain tumors. In breast cancer, germline TP53 mutations harbor 5–8% of cases of early-onset (under 30 years old) disease, while up to 85% of women who carry germline TP53 mutations eventually develop breast cancer [4]. On the other hand, somatic TP53 mutations are identified in 37% of all breast cancers and is more frequently mutated in HER-2 positive (72%) and basal-like subtype (80%) [5]. DNA sequencing is considered as the gold-standard for the detection of TP53 mutations that usually harbor the exons 5–8 of the gene [6]. More recently, next generation sequencing has allowed the detection of TP53 mutations outside this restricted region.
There are numerous studies assessing the clinical significance of TP53 mutations in breast cancer. However, the results of these studies are often contradicting. TP53 mutation was associated with worse prognosis in breast cancer patients regardless of the tumor subtype and the type of treatment administered. Initially, p53 IHC expression was associated with an adverse prognosis in small retrospective studies [7–9]. More recent studies evaluated TP53 gene mutations in breast cancer via DNA sequencing [10–13]. P53 mutation was associated with negative estrogen and progesterone receptor (ER/PR) status and increased mortality rate in 859 breast cancer women [10]. TP53 mutations within exons 5 to 8 detected by gene sequencing were related to increased risk of breast cancer-specific death regardless of tumor size, nodal status and hormone receptor expression [11]. Moreover, P53 mutation status has been associated with response to breast cancer treatment. A METABRIC analysis of breast cancer patients that received only endocrine treatment linked TP53 mutations to worse survival [6], while a meta-analysis of 3,476 cases of patients receiving neoadjuvant treatment concluded that TP53 mutation status is a predictor of response to neoadjuvant chemotherapy [14].
The aim of this study is to evaluate the role of somatic TP53 mutations as a potential prognostic marker in Stage I-III breast cancer patients treated in a single center via a case–control study. The identification of TP53 mutations was performed via next-generation sequencing (NGS) in paraffin blocks of the patients enrolled.
Results
Clinicopathological characteristics
Patient characteristics (age, date at diagnosis, menopausal status), histopathological parameters (histological type, grade, ER/PR expression, HER2 expression, ki67, stage) and treatment administered (surgery, chemotherapy, anti-HER2 treatment, hormonotherapy) in cases and controls are summarized in Tables 1 and 2.
Table 1: Clinicopathological characteristics of cases and controls
| Variable Continuous variables | Cases Mean (SD) | Controls Mean (SD) | p-value |
|---|---|---|---|
| Age at diagnosis | 52.35 (11.47) | 49.26 (11.27) | p = 0.362 |
| Menopausal status | p = 0.492 | ||
| Premenopausal | 8 (34.8%) | 12 (52.2%) | |
| Perimenopausal | 4 (17.4%) | 3 (13.0%) | |
| Postmenopausal | 11 (47.8%) | 8 (34.8%) | |
| Surgery | p = 0. 295 | ||
| Yes | 20 (87.0%) | 22 (95.7%) | |
| No | 3 (13.0%) | 1 (4.3%) | |
| Stage at diagnosis | p = 0.802 | ||
| I | 5 (21.7%) | 4 (17.4%) | |
| II | 6 (26.1%) | 8 (34.8%) | |
| III | 12 (52.2%) | 11 (14.8%) | |
| Breast tumor location | p = 0.546 | ||
| Left | 13 (56.5%) | 15 (65.2%) | |
| Right | 10 (43.5%) | 8 (34.8%) | |
| Histology | p = 0.368 | ||
| IDC | 22 (95.7%) | 22 (95.7%) | |
| ILC | 1 (4.3%) | 0 (0%) | |
| Other | 0 (0.0%) | 1 (4.3%) | |
| Hormone status | p = 0.760 | ||
| Positive | 14 (60.9%) | 15 (65.2%) | |
| Negative | 9 (39.1%) | 8 (34.8%) | |
| ER status | p = 0.546 | ||
| Positive | 10 (43.5%) | 8 (34.8%) | |
| Negative | 13 (56.5%) | 15 (65.2%) | |
| PR status | p = 0.552 | ||
| Positive | 12 (52.2%) | 14 (60.9%) | |
| Negative | 11 (47.8%) | 9 (39.1%) | |
| HER2 status | p = 0.760 | ||
| Positive | 9 (39.1%) | 8 (34.8%) | |
| Negative | 14 (60.9%) | 15 (65.2%) | |
| Grade | |||
| G1 | 1 (4.3%) | 0 (0.0%) | p = 0.506 |
| G2 | 5 (21.7%) | 7 (30.4%) | |
| G3 | 17 (73.9%) | 16 (69.6%) | |
| Ki67 | p = 0.681 | ||
| <20% | 3 (13.0%) | 4 (17.4%) | |
| ≥20% | 20 (87.0%) | 19 (82.6%) |
Table 2: Type of treatment administered in cases and controls
| Adjuvant chemotherapy | p = 0.636 | ||
| No | 3 (13.0%) | 2 (8.7%) | |
| Yes | 20 (87.0%) | 21 (91.3%) | |
| Adjuvant radiation | p = 0.743 | ||
| No | 6 (26.1%) | 7 (30.4%) | |
| Yes | 17 (73.9%) | 16 (69.6%) | |
| Anti-Her2 treatment | p = 0.760 | ||
| No | 14 (60.9%) | 15 (65.2%) | |
| Yes | 9 (39.1%) | 8 (34.8%) | |
| Disease progression | |||
| No | 14 (60.9%) | 21 (91.3%) | |
| Yes | 9 (39.1%) | 2 (8.7%) |
Mean age at diagnosis was 52.35 (SD; 11.47) years in cases and 49.26 years (SD; 11.27) years in controls (p = 0.362). Overall, there were not statistically significant differences between cases and controls in terms of age at diagnosis (p = 0.362), menopausal status (p = 0.492), stage at diagnosis (p = 0.802), histology (p = 0.368), hormone status (p = 0.760), HER2 expression (p = 0.760), grade (G1 vs. G2/3; p = 0.506) and Ki67 status (p = 0.681). Patients with stage I–III disease (23; 100%) underwent surgical excision (20; 87%) and adjuvant radiation (17; 73.9%). Invasive ductal carcinoma (IDC) was diagnosed in most of the cases (22; 95.7%). The majority of cases were hormone receptor-positive (14; 60.9%), HER2-negative (14; 60.9%) and characterized by low differentiation (17; 73.9%) and high Ki67 expression (20; 87%).
Genetic polymorphisms of TP53 somatic mutations
Genetic polymorphisms of TP53 pathogenic somatic mutations identified are summarized in Table 3. Of note, the most frequent pathogenic somatic TP53 mutations reported in our patients were c.824G>A p.Cys275Tyr (n = 3) and c.743G>A p.Arg248Gln (n = 2) while the other polymorphisms were detected only once.
Table 3: Genetic polymorphisms of pathogenic somatic TP53 mutations
| TP53 genetic polymorphism | NCBI genomes browser | Type | Clinical significance | Frequency |
|---|---|---|---|---|
| c.614A>G p.Tyr205Cys | Rs1057520007 | Somatic | Pathogenic | 1 |
| c.559+1G>A | Rs1131691042 | Somatic | Pathogenic | 1 |
| c.824G>A p.Cys275Tyr | Rs863224451 | Somatic | Pathogenic | 3 |
| c.488A>G p.Tyr163Cys | Rs148924904 | Somatic | Pathogenic | 1 |
| c.818G>T p.Arg273Leu | Rs28934576 | Somatic | Pathogenic | 1 |
| c.714_715insT p.Asn239Ter | Rs1567549651 | Somatic | Pathogenic | 1 |
| c.536A>G p.His179Arg | Rs1057519991 | Somatic | Pathogenic | 1 |
| c.85_86del p.Asn29GlnfsTer13 | Rs1555526931 | Somatic | Pathogenic | 1 |
| c.586C>T p.Arg196* | Rs397516435 | Somatic | Pathogenic | 1 |
| c.797G>A p.Gly266Glu | Rs193920774 | Somatic | Pathogenic | 1 |
| c.853G>A p.Glu285Lys | Rs112431538 | Somatic | Pathogenic | 1 |
| c.990del p.Gln331Argfs*14 | Rs11575996 | Somatic | Pathogenic | 1 |
| c.742C>T p.Arg248Trp | Rs121912651 | Somatic | Pathogenic | 1 |
| c.722C>T p.Ser241Phe | Rs28934573 | Somatic | Pathogenic | 1 |
| c.743G>A p.Arg248Gln | Rs11540652 | Somatic | Pathogenic | 2 |
| c.638G>T p.Arg213Leu | Rs587778720 | Somatic | Pathogenic | 1 |
| c.455C>T p.Pro152Leu | Rs587782705 | Somatic | Pathogenic | 1 |
| c.817C>T p.Arg273Cys | Rs121913343 | Somatic | Pathogenic | 1 |
| c.681_682insT p.Asp228Ter | Rs1567550002 | Somatic | Pathogenic | 1 |
| c.626_627del p.Arg209LysfsTer6 | Rs1057517840 | Somatic | Pathogenic | 1 |
We examined the presence of other pathogenic mutations along with TP53 mutations. The genomic profile of our cases is summarized in Table 4. Of note, PIK3CA was the most frequent pathogenic mutation detected in somatic TP53-mutant tumors (4/23; 17.4%). Other pathogenic mutations identified included AKT1, PTEN and NRAS mutations. Of note, a number of mutations of unknown significance were frequently reported. The most common mutations of unknown significance (VUS) identified were: ROS1 (10/23), KMT2C (6/23), NF1 (4/23), RET (2/23), NOTCH1 (2/23).
Table 4: Somatic/germline mutations identified in TP53-mutated cases
| Cases | TP53 somatic mutation | TP53 germline mutations | Co-existing pathogenic mutations | Co-existing VUS mutations |
|---|---|---|---|---|
| 1 | c.614A>G p. Tyr205Cys | – | – | – |
| 2 | c.559+1G>A | – | PIK3CA (Glu545Lys) | – |
| 3 | c.824G>A p.Cys275Tyr | – | – | KMT2C (Arg2609Gln), RB1 (Asn663Ser), NOTCH1 (Leu818Pro) |
| 4 | c.488A>G p.Tyr163Cys | – | – | RAD50 (Arg365Gln), TP53 (Leu188Lysfs*59), CCND1 (c.724-2A>C) |
| 5 | c.824G>A p.Cys275Tyr | – | AKT1 (Glu17Lys) | – |
| 6 | c.818G>T p.Arg273Leu | – | PIK3CA (Glu39Lys) | – |
| 7 | c.714_715insT p.Asn239Ter | – | – | ROS1 (Gly2245Ser) ROS1 (Thr145Pro) |
| 8 | c.536A>G p.His179Arg | c.847C>T p.Arg283Cys | – | Somatic: ROS1 (Thr2195Ser) TP53 (Arg283Cys) Germline: PMS2 (His189Pro) |
| 9 | c.85_86del: p.Asn29GlnfsTer13 | – | – | ROS1 (Thr145Pro) BRCA2 (Phe3289Leu) |
| 10 | c.586C>T p.Arg196* | – | – | – |
| 11 | c.797G>A p.Gly266Glu | – | PIK3CA (Glu545Lys), PTEN (Glu285Glyfs*13) | MSH2 (Glu561Lys), NOTCH1 (Glu606Lys) |
| 12 | c.824G>A p.Cys275Tyr | – | – | ROS1 (Thr145Pro), RET (Thr562Ser) |
| 13 | c.853G>A p.Glu285Lys | – | – | ROS1 (Thr145Pro), NF1 (Met102Val) |
| 14 | c.990del p.Gln331Argfs*14 | – | – | MET (Pro1364Ser), NF1 (Ala188Glu) |
| 15 | c.742C>T p.Arg248Trp | – | – | ROS1 (Arg167Gln), NF1 (Asp2465Glu), ERBB2 (Arg487Gln), AR (Glu494Ala) |
| 16 | c.722C>T p.Ser241Phe | – | PIK3CA (Glu545Lys), NRAS (Gly60Arg) | BRCA2 (Arg1160Gly), BRCA2 (Cys1159Tyr), CDKN2A (Arg10Trp), RET (Glu768Lys), STK11(Val66Met), STK11 (Gly408Ser), NF1 (Ala2485Gly) |
| 17 | c.743G>A p.Arg248Gln | – | – | KMT2C (Glu1625Lys) |
| 18 | c.638G>T p.Arg213Leu | – | – | KDR (Leu625Phe), CDK4 (Glu265Lys), CDK4 (Met264Ile) |
| 19 | c.817C>T pArg273Cys | – | – | ROS1 (Arg1942Trp) KMT2C (Gly4411Arg) |
| 20 | c.455C>T p.Pro152Leu | – | – | ROS1 (Thr145Pro), MET (Pro1382Ser), KMT2C (Asp2692Ala) MYC (Asn26Ser) |
| 21 | c.743G>A: p.Arg248Gln | – | – | ROS1 (Arg167Gln) KMT2C (Arg1292Gln) |
| 22 | c.681_682insT p.Asp228Ter | – | – | MTOR (Arg1896Gln) ROS1 (Gly2245Ser) KMT2C (Ile439Val) |
| 23 | c.626_627del: p.Arg209LysfsTer6 | – | – | PIK3CA (Gly439Ala) CDK6 (Arg90Thr) JAK2 (Glu1024Lys) |
Survival analysis
Disease-free survival (DFS) was evaluated in cases and controls. DFS was 16.3 months in cases harboring TP53 somatic mutations (95% CI; 11.38–21.25) versus 62.9 months in TP53 wild-type controls (95% CI; 40.8–85). Somatic TP53 mutations were associated with shorter DFS in our study. Figure 1 presents Kaplan Meier DFS curves for cases and controls. TP53 mutation was associated with a 8-fold risk of recurrence in the Cox regression analysis (OR = 8.530, 95% CI: 1.81–40.117; p = 0.007).
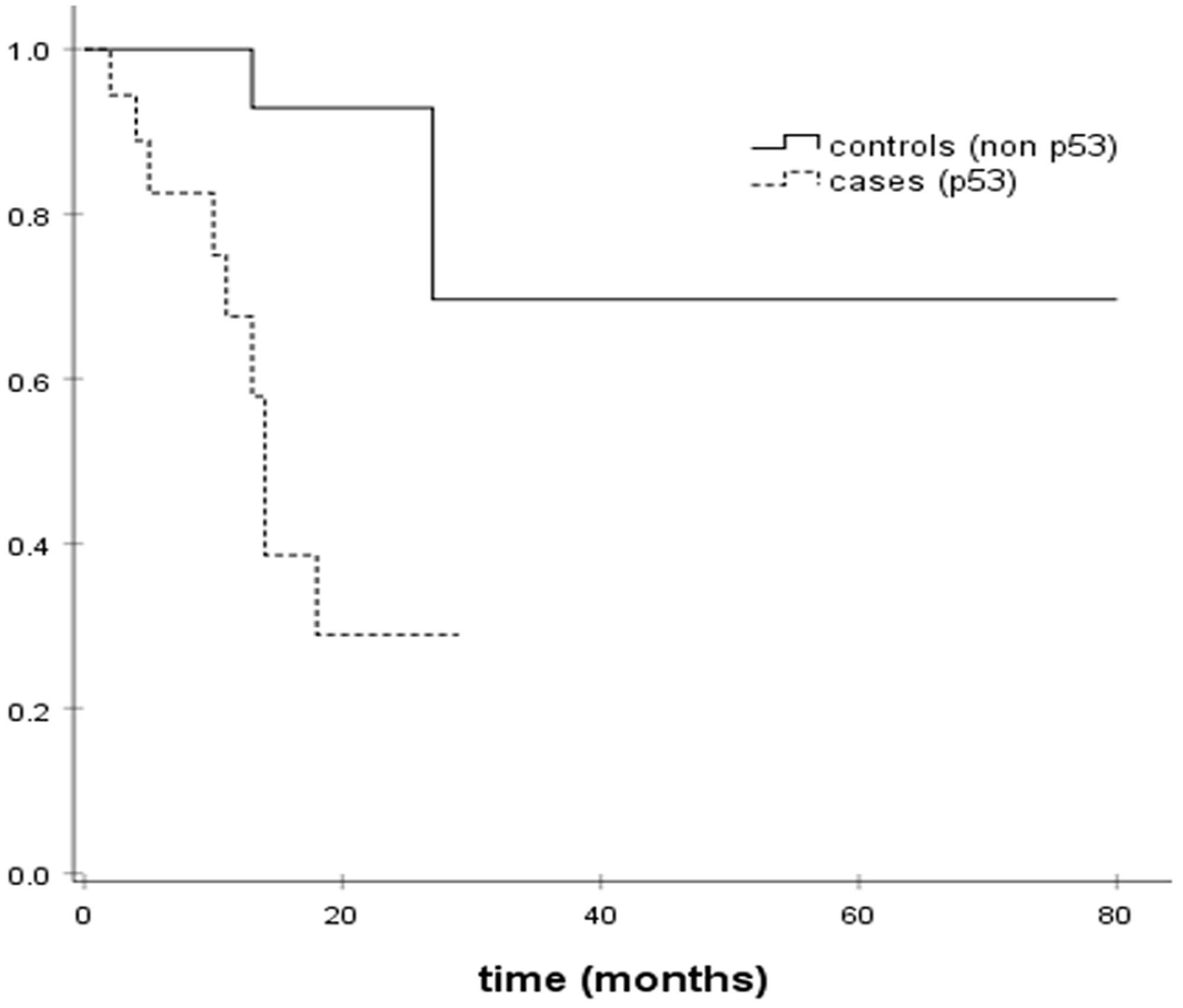
Figure 1: Kaplan–Meier DFS estimates.
DISCUSSION
We here demonstrate that pathogenic somatic TP53 mutations are associated with a decreased disease-free survival in patients with early-stage breast cancer. We retrospectively identified 23 breast cancer patients harboring TP53 somatic mutations identified via next-generation sequencing and 23 TP53 wild-type controls matched on age and clinicopathological characteristics. DFS was significantly reduced in TP53-mutated cases (16 vs. 63 months).
In our study TP53 somatic mutations were associated with high grade (G2/3: 22; 95.7%) and high proliferating (Ki67 > 20%: 87%) breast tumors. This is in accordance with previous results that TP53 mutations are more frequently identified in HER2-positive and triple negative breast tumors (TNBC), while the incidence is low in luminal A tumors. TP53 mutation is indicative of a more aggressive entity and affects tumor response to treatment [15]. P53 regulates epithelial to mesenchymal (EMT) process and stem cell characteristics through upregulation of miR200c [15]. Mutant TP53 tumors suppress transcriptional factors involved in the TGF-β signaling pathway (e.g. ZEB2/SIP1, Snail, Twist) and induce epithelial-mesenchymal transition (EMT). Moreover, genes involved in cell migration like the matrix metalloproteinase family member MMP1 were overexpressed in TP53 mutated tumors [15]. Overall, TP53 mutational status was related to reduced cell differentiation and increased metastatic ability.
An interesting finding of our study was the co-existence of TP53 mutations with pathogenic somatic PIK3CA mutations in the 17.4% of the cases. PIK3CA mutations are detected in more than one third of HR-positive breast tumors (34.5%) and less frequently in HER2-overexpressing tumors (22.7%), whereas the incidence drops to 8.3% in triple-negative and basal-like breast cancer [16]. It has been previously shown that co-mutation of TP53 and PIK3CA account for more than 6% of breast cancers and for approximately 30% of TP53-mutated tumors [17]. Co-mutated tumors represent an aggressive entity and were associated with a worse progression-free survival [17]. In our study, co-existence of these mutations was not a rare event. The effect of the simultaneous presence of TP53 and PIK3CA mutations on response to chemotherapy and prognosis needs to be further addressed.
Our case-control study demonstrated that TP53 pathogenic somatic mutations are associated with a reduced disease-free survival in early-stage breast cancer. In agreement with our findings, previous studies report an adverse prognosis in patients harboring TP53 mutations [6]. Some other studies suggest that TP53 mutations have a distinct role in different breast cancer subtypes. TP53 somatic mutations were associated with a worse prognosis in patients with luminal B and HER2-positive breast tumors, but not in patients with luminal A and basal-like tumors [18]. Moreover, TP53 mutations are linked to different response rates to different treatment regimens. One study suggests that TP53 wild-type tumors respond better to hormone therapy, while the opposite effect is identified in patients that receive chemotherapy only [19]. TP53 status may have a distinct clinical role according to the tumor subtype and the type of treatment administered.
One strength of our study was the detection method applied for the identification of TP53 mutations. DNA sequencing is currently the gold standard for identification of TP53 mutations [20]. Recently, next generation sequencing has emerged as a highly accurate alternative since it offers the ability of detecting mutations outside exons 5–8. Accuracy of immunohistochemistry (IHC) is limited by the presence of null mutations (nonsense mutations, deletions, insertions etc) in TP53 gene that result in a detectable but unstable protein. In these cases that account for up to ~40% of TP53 mutations in breast cancer IHC will fail to detect TP53 mutation. Consequently, IHC detection of TP53 status should be evaluated with caution. Our study overcomes this limitation by applying next generation sequencing in patient samples further increasing the validity of the data presented. Despite the originality, limitations of this case-control study should be acknowledged. Our study is confined to a single institution and thus the sample size is limited. More studies with a larger sample size should be performed to confirm our results. A multicenter study with a similar design could generate more robust scientific data.
We here show that TP53 pathogenic somatic mutations are associated with a shorter DFS in early-stage breast cancer patients. In addition, TP53 mutations often coexist with PIK3CA mutations in breast tumors (17.4%). Future well designed studies should be performed to address the clinical role of the co-existence of these mutations in breast cancer.
Materials and Methods
Subjects
Incident cases of 82 patients with histologically confirmed Stage I-III breast cancer that underwent NGS in paraffin blocks and blood samples during the period 25/09/2019 to 25/05/2021 were retrospectively collected. Among them 23 cases of somatic TP53 mutations were detected and were matched on age at diagnosis (±5 years), histological subtype (luminal A, luminal B, HER2-enriched, TNBC), histological grade (1 vs. 2/3), menopausal status, ki67 expression and disease stage as classified by TNM classification system with controls; controls included 23 women with TP53 wild-type breast cancer. All women were treated in a single Institute at the Oncology Department of “Alexandra” Hospital, Medical School, University of Athens, Greece. Immunohistochemical (IHC) analysis was performed to quantify expression of human epidermal growth factor receptor 2 (HER2), hormone receptors (HR) and Ki67. Estrogen receptor (ER) and progesterone receptor (PR) were considered positive if tumors had more than 1% nuclear-stained cells. HER2 status was considered positive when graded as 3+, while 0 to 1+ were negative and 2+ was an inconclusive result and in situ hybridization (ISH) was performed in those cases to confirm positivity. Hormone receptor positive tumors characterized by ki67 expression of over 20% were considered as luminal B. Information on histological characteristics (tumor subtype, grade, ER/PR expression, HER2 expression, expression levels of ki67), TNM stage (tumor size, lymph node infiltration, metastasis), type of surgery performed, type of chemotherapy administered, disease-free survival and overall survival were collected from patient files and were registered on an electronic database. Both cases and controls were Caucasian and reside in the same geographical region. This case–control study is in accordance with the Helsinki Declaration and has been approved by the Review Board of Alexandra General Hospital of Athens. An informed consent form was obtained from each of the eligible patients.
DNA extraction
For breast cancer patients, paraffin-embedded breast tissues derived from mastectomy or breast conserving operation before adjuvant treatment and blood samples were analyzed. Paraffin-embedded breast tissues were cut at slices of 10 μm diameter. Tumor DNA was isolated from paraffin-embedded breast tissues using the QIAamp DNA FFPE Tissue or the kit Ion Ampliseq Custom Next Generation Sequencing (NGS) DNA panel (Amplicon Sequencing following the manufacturer’s instructions. Plasma blood samples were collected in Vacutainer tubes. Within 4 hours after collection, plasma was separated from whole blood samples through centrifugation for 10 min at 3000 rpm at room temperature and stored at −80°C until further use. Isolation of plasma DNA was performed using QIAsymphony DSP DNA Mini Kit and the genomic library was constructed using Trusight™ Comprehensive Hereditary Cancer Panel – Nextera™ DNA Flex Pre-enrichment Library Prep according to the manufacturer’s instructions.
Targeted sequencing
The NGS study on paraffin-embedded breast tissues was performed using Ion Torrent platform (Ion S5Prime) with a median amplicon cover 2000x (whenever DNA extraction was performed with kit Ion Ampliseq). For the data annotation and analysis IonReporter (v5.12) (Thermo Scientific) was used. The sequences were aligned to the human genome reference sequence GRCh37-hg19. An additional manual data curation was performed using data from OncomineReporter (v4.4) and relevant databases (CinVar, dbSNP, Ensemble, COSMIC, CIVIC, PharmGKB, OMIM, My Cancer Genome, Vasome etc.). For the tissues that underwent DNA extraction via the QIAamp DNA FFPE Tissue, libraries were constructed using AmpliSeq for Illumina Comprehensive Panel v3. The NGS study was performed using the Illumina platform (MiSeq, NextSeq500 or NovaSeq) in these cases with a median amplicon cover 500x for the 88.8% of the targeted regions.
Plasma DNA sequencing
Genomic libraries were constructed using Trusight™ Comprehensive Hereditary Cancer Panel – Nextera™ DNA Flex Pre-enrichment Library Prep according to the manufacturer’s instructions. Plasma sequencing was performed using Illumina platform (NextSeq500/NovaSeq). The validation of results was performed according to criteria of American College of medical Genetics – ACMG [21] and NCCN guidelines.
Statistical analysis
The statistical analysis was performed with SPSS 24.0 statistical software. Differences between cases and controls were examined by Student’s t-test for continuous variables or the chi-square test for categorical variables. Pearson’s correlation and Fisher’s exact test (for categorical variables) were used. The threshold for statistical significance was set at p < 0.05. Univariate Cox regression analysis was performed to evaluate the association of TP53 mutation with disease-free survival in breast cancer patients. Kaplan–Meier survival curves were estimated to graphically represent the results.
Abbreviations
TP53: Tumor protein 53; NGS: next-generation sequencing; PIK3CA: phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit alpha; DFS: disease-free survival; ER: estrogen receptor; PR: progesterone receptor; HER2: human epidermal growth factor receptor 2; IHC: immunohistochemistry; IDC: invasive ductal carcinoma; VUS: variants of unknown significance; TNBC: triple negative breast cancer; EMT: epithelial to mesenchymal transition; FFPE: Formalin-fixed paraffin-embedded.
CONFLICTS OF INTEREST
M.L. has received honoraria from Roche, Astra Zeneca, Astellas, MSD, Janssen, Bristol-Myers-Squibb and IPSEN. M.-A.D. has received honoraria from participation in advisory boards from Amgen, Bristol-Myers-Squibb, Celgene, Janssen, Takeda. F.Z. has received honoraria for lectures and has served in an advisory role for Astra-Zeneca, Daiichi, Eli-Lilly, Merck, Novartis, Pfizer, and Roche. The remaining authors declare no conflicts of interest.
FUNDING
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
References
1. Surget S, Khoury MP, Bourdon JC. Uncovering the role of p53 splice variants in human malignancy: a clinical perspective. Onco Targets Ther. 2013; 7:57–68. https://doi.org/10.2147/OTT.S53876. [PubMed].
2. Lane DP. Cancer. p53, guardian of the genome. Nature. 1992; 358:15–16. https://doi.org/10.1038/358015a0. [PubMed].
3. Lane D, Levine A. p53 Research: the past thirty years and the next thirty years. Cold Spring Harb Perspect Biol. 2010; 2:a000893. https://doi.org/10.1101/cshperspect.a000893. [PubMed].
4. Schon K, Tischkowitz M. Clinical implications of germline mutations in breast cancer: TP53. Breast Cancer Res Treat. 2018; 167:417–23. https://doi.org/10.1007/s10549-017-4531-y. [PubMed].
5. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012; 490:61–70. https://doi.org/10.1038/nature11412. [PubMed].
6. Shahbandi A, Nguyen HD, Jackson JG. TP53 Mutations and Outcomes in Breast Cancer: Reading beyond the Headlines. Trends Cancer. 2020; 6:98–110. https://doi.org/10.1016/j.trecan.2020.01.007. [PubMed].
7. Yamashita H, Toyama T, Nishio M, Ando Y, Hamaguchi M, Zhang Z, Kobayashi S, Fujii Y, Iwase H. p53 protein accumulation predicts resistance to endocrine therapy and decreased post-relapse survival in metastatic breast cancer. Breast Cancer Res. 2006; 8:R48. https://doi.org/10.1186/bcr1536. [PubMed].
8. Iwaya K, Tsuda H, Hiraide H, Tamaki K, Tamakuma S, Fukutomi T, Mukai K, Hirohashi S. Nuclear p53 immunoreaction associated with poor prognosis of breast cancer. Jpn J Cancer Res. 1991; 82:835–40. https://doi.org/10.1111/j.1349-7006.1991.tb02710.x. [PubMed].
9. Ostrowski JL, Sawan A, Henry L, Wright C, Henry JA, Hennessy C, Lennard TJ, Angus B, Horne CH. p53 expression in human breast cancer related to survival and prognostic factors: an immunohistochemical study. J Pathol. 1991; 164:75–81. https://doi.org/10.1002/path.1711640113. [PubMed].
10. Rossner P Jr, Gammon MD, Zhang YJ, Terry MB, Hibshoosh H, Memeo L, Mansukhani M, Long CM, Garbowski G, Agrawal M, Kalra TS, Gaudet MM, Teitelbaum SL, et al. Mutations in p53, p53 protein overexpression and breast cancer survival. J Cell Mol Med. 2009; 13:3847–57. https://doi.org/10.1111/j.1582-4934.2008.00553.x. [PubMed].
11. Olivier M, Langerød A, Carrieri P, Bergh J, Klaar S, Eyfjord J, Theillet C, Rodriguez C, Lidereau R, Bièche I, Varley J, Bignon Y, Uhrhammer N, et al. The clinical value of somatic TP53 gene mutations in 1,794 patients with breast cancer. Clin Cancer Res. 2006; 12:1157–67. https://doi.org/10.1158/1078-0432.CCR-05-1029. [PubMed].
12. Blaszyk H, Hartmann A, Cunningham JM, Schaid D, Wold LE, Kovach JS, Sommer SS. A prospective trial of midwest breast cancer patients: a p53 gene mutation is the most important predictor of adverse outcome. Int J Cancer. 2000; 89:32–38. https://doi.org/10.1002/(sici)1097-0215(20000120)89:1%3C32::aid-ijc6%3E3.0.co;2-g. [PubMed].
13. Rong G, Yi Z, Ma F, Guan Y, Xu Y, Li L, Xu B. DNA damage response as a prognostic indicator in metastatic breast cancer via mutational analysis. Ann Transl Med. 2021; 9:220. https://doi.org/10.21037/atm-20-2137. [PubMed].
14. Chen MB, Zhu YQ, Xu JY, Wang LQ, Liu CY, Ji ZY, Lu PH. Value of TP53 status for predicting response to neoadjuvant chemotherapy in breast cancer: a meta-analysis. PLoS One. 2012; 7:e39655. https://doi.org/10.1371/journal.pone.0039655. [PubMed].
15. Bertheau P, Lehmann-Che J, Varna M, Dumay A, Poirot B, Porcher R, Turpin E, Plassa LF, de Roquancourt A, Bourstyn E, de Cremoux P, Janin A, Giacchetti S, et al. p53 in breast cancer subtypes and new insights into response to chemotherapy. Breast. 2013 (Suppl 2); 22:S27–29. https://doi.org/10.1016/j.breast.2013.07.005. [PubMed].
16. Stemke-Hale K, Gonzalez-Angulo AM, Lluch A, Neve RM, Kuo WL, Davies M, Carey M, Hu Z, Guan Y, Sahin A, Symmans WF, Pusztai L, Nolden LK, et al. An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res. 2008; 68:6084–91. https://doi.org/10.1158/0008-5472.CAN-07-6854. [PubMed].
17. Kotoula V, Karavasilis V, Zagouri F, Kouvatseas G, Giannoulatou E, Gogas H, Lakis S, Pentheroudakis G, Bobos M, Papadopoulou K, Tsolaki E, Pectasides D, Lazaridis G, et al. Effects of TP53 and PIK3CA mutations in early breast cancer: a matter of co-mutation and tumor-infiltrating lymphocytes. Breast Cancer Res Treat. 2016; 158:307–21. https://doi.org/10.1007/s10549-016-3883-z. [PubMed].
18. Silwal-Pandit L, Vollan HK, Chin SF, Rueda OM, McKinney S, Osako T, Quigley DA, Kristensen VN, Aparicio S, Børresen-Dale AL, Caldas C, Langerød A. TP53 mutation spectrum in breast cancer is subtype specific and has distinct prognostic relevance. Clin Cancer Res. 2014; 20:3569–80. https://doi.org/10.1158/1078-0432.CCR-13-2943. [PubMed].
19. Ungerleider NA, Rao SG, Shahbandi A, Yee D, Niu T, Frey WD, Jackson JG. Breast cancer survival predicted by TP53 mutation status differs markedly depending on treatment. Breast Cancer Res. 2018; 20:115. https://doi.org/10.1186/s13058-018-1044-5. [PubMed].
20. Robles AI, Harris CC. Clinical outcomes and correlates of TP53 mutations and cancer. Cold Spring Harb Perspect Biol. 2010; 2:a001016. https://doi.org/10.1101/cshperspect.a001016. [PubMed].
21. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, Grody WW, Hegde M, Lyon E, Spector E, Voelkerding K, Rehm HL, and ACMG Laboratory Quality Assurance Committee. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015; 17:405–24. https://doi.org/10.1038/gim.2015.30. [PubMed].


