Introduction
Liver cancer despite its relatively low incidence rate with a geographically varied global burden is a fatal disease due to its high mortality rate: 5-year survival rate changes from 2% to 33% depending on the stage of the disease at the time of the diagnosis [1, 2]. Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer reaching 80–90% of all primary liver cancers [3]. In 20% of all cases, this is an out of the blue disease developing without any liver cirrhosis [4]. Nevertheless, liver cancer may develop due to a wide variety of reasons, like hepatitis B virus, hepatitis C virus (HCV) related liver cirrhosis, dietary exposure to aflatoxin B1 etc., [5].
Amongst those who are infected with chronic HCV, 15–30% would progress to cirrhosis and 1–5% are expected to die due to decompensated cirrhosis and liver cancer [6]. Anyway, despite the relative low incidence rate of malignancy of an HCV-related cirrhosis and the era of direct antivirals, there is an accumulated pool of patients with HCV-related liver cirrhosis who are at risk group of liver cancer development. Surely, in these cases liver cancer is not that much an out of the blue disease but anyway it may be clinically silent during early stages of the disease, thus, methods for identifying molecular changes developing in pre-clinical stages of the disease are essential.
A valuable source of such molecular information is microvesicles including exosomes, serving as an intracellular whatsapp with huge buffer-exchange volumes. Exosomes are indentified as microvesicles with a size range 30–150 nm [7] or 40–160 nm [8] or 100–200 nm [9], in this paper we assume that exosomes are 30–200 nm microvesicles. The microvesicles may contain different proteins, DNA and various RNA including miRNA, a small non-coding nucleic acid of 18-24 nucleotides involved in epigenetic regulation of the gene expression [10]. Exosomes along with other microvesicles may serve as a target delivery system and mechanism for preserving the structure of the molecules being their defense from different enzymes [11]. Not packed into exosomes miRNAs (hereinafter non-exosomal miRNAs) some years ago were supposed to be non-stable molecules vulnerable to blood RNAses, last years, they are shown to be detectable in the biological fluids in protein and lipoprotein complexes, that prevents their degradation.
The first question to answer in this study was to determine which fraction of extracellular miRNAs in plasma – packed into exosomes (hereinafter, exosomal) or not packed into exosomes (hereinafter, non-exosomal) – is stronger associated with primary liver cancer in plasma. The second question was whether saliva miRNAs – both exosomal and non-exosomal – are associated with primary liver cancer.
Results
Exosome determination in the samples
Exosomes in the samples were identified with indirect method of photon cross-correlation spectroscopy: peaks were identified in a range from 50 nm to 250 nm in all 8 exosomal samples (4 from blood plasma and 4 from saliva) in the control and every study group indicating that in these samples there are microvesicles of exosomal size range, which is 50–200 nm. In all 8 exosome-depleted samples (4 from blood plasma and 4 from saliva) in the control and every study group there were peaks in a range from 50 nm to 70 nm also indicating the presence of the microvesicles of exosomal size range. This may be due to incomplete elimination of microvesicles from plasma and saliva samples. Approximate number of CD63 positive microvesicles in a sample was determined with CD63 based ELISA analysis (shown in Table 1). This data allows to assume the amount of exosomes in exosome-depleted samples is below the meaning of the least amount of the exosomes, provided by manufacturer in calibration curve, and this value can be neglected in further work.
Table 1: Approximate numbers of CD63 positive microvesicles detected with CD63 based ELISA
| Study group 11 | Study group 22 | Control group3 | ||||
|---|---|---|---|---|---|---|
| plasma | saliva | plasma | saliva | plasma | saliva | |
| Exosome depleted samples | not detected4 | not detected | not detected | not detected | not detected | not detected |
| Exosomal samples | ~2,6 E+10 | ~0,9 E+10 | 2,5 E+10 | ~1,1 E+10 | 2,5 E+10 | ~0,8 E+10 |
miRNA concentration in samples
MiRNA concentration in all samples was determined, in exosomal samples concentrations varied from 0,89 μg/mcl to 4,8 μg/mcl; in exosome depleted samples from 2,77 μg/mcl to 7,5 μg/mcl.
Expression level of exosomal and non-exosomal miRNAs
From the set of examined miRNAs two miRNAs – miRNA-145-5p and miRNA-224-5p – were eliminated because there were not detected (Ct value of > 35) both in plasma exosomal and exosome-depleted samples in control and every study group. Expression levels of the following miRNAs – let-7a-5p, miR-16-5p, miR-18a-5p, miR-21-5p, miR-22-3p, miR-34a-5p, miR-103a-3p, miR-122-5p, miR-221-3p, miR-222-3p – were determined both in plasma exosomal and exosome-depleted samples in control and every study group (Table 2) and three miRNAs (miRNA-122-5p, -21-5p, -221-3p) were identified in saliva (Table 3). Expression level of each of these miRNAs was calculated as following: exosomal miRNA-X/exosomal miRNA-16-5p (for plasma miRNAs Figure 1, for saliva miRNAs Figure 3), non-exosomal miRNA-X/non-exosomal miRNA-16-5p (for plasma miRNAs Figure 1, for saliva miRNAs Figure 3), exosomal miRNA-X/non-exosomal miRNA-X (for plasma miRNAs Figure 2, for saliva miRNAs Figure 3), where miRNA-X is a target/studied miRNA.
Table 2: Significance of exosomal and non-exosomal miRNA expression level difference in plasma in healthy volunteers, hepatocellular carcinoma and liver cirrhosis
| Study groups | N | let-7a | mir-18 | mir-21 | mir-22 | mir-34a | mir-103a | mir-122 | mir-221 | mir-222 |
|---|---|---|---|---|---|---|---|---|---|---|
| HCC/LC | e/n-e | 0.00001 | 0.00011 | 0.00181 | 0.00103 | 0.49113 | 0.15647 | 5.34649E-7 | 0.00077 | 0.09031 |
| HCC/HV | e/n-e | 1.41859E-12 | 9.78214E-10 | 0.00028 | 0.00005 | 0.12003 | 0.00117 | 0.00053 | 0.00042 | 0.10107 |
| LC/HV | e/n-e | 0.00016 | 0.00002 | 0.42080 | 0.17264 | 0.21668 | 0.00688 | 0.24411 | 0.42716 | 0.46360 |
| HCC/LC | e/miR16 | 1.75056E-9 | 0.00336 | 1.05143E-6 | 2.32496E-9 | 0.12160 | 2.52250E-10 | 4.3856e-14 | 0.00002 | 0.02189 |
| HCC/HV | e/miR16 | 5.28507E-9 | 3.93833E-8 | 1.19869E-8 | 1.34904E-8 | 0.08330 | 0.00005 | 5.44032E-11 | 8.88518E-7 | 0.00031 |
| LC/HV | e/miR16 | 0.07108 | 0.00010 | 0.00035 | 0.05313 | 0.34435 | 0.22388 | 9.72938E-8 | 0.09019 | 0.00536 |
| HCC/LC | n-e/miR16 | 6.61017E-11 | 0.22048 | 5.58919E-8 | 4.50614E-9 | 0.12806 | 8.2614e-13 | 1.8268e-17 | 5.67705E-8 | 0.32600 |
| HCC/HV | n-e/miR16 | 2.10138E-6 | 0.08421 | 3.230255e-14 | 0.00003 | 0.29906 | 0.00002 | 3.67392E-7 | 2.04322E-7 | 0.00078 |
| LC/HV | n-e/miR16 | 0.28479 | 0.22776 | 7.54786E-7 | 0.01288 | 0.25809 | 0.00002 | 0.00123 | 0.26207 | 0.00209 |
Table 3: Significance of exosomal and non-exosomal miRNA expression level difference in saliva in healthy volunteers, hepatocellular carcinoma and liver cirrhosis
| Study groups | N | mir-122 | mir-21 | mir-221 |
|---|---|---|---|---|
| HCC/LC | e/n-e | 0.00712 | 0.13659 | 0.31759 |
| HCC/HV | e/n-e | 0.20546 | 0.19050 | 1.3845e-14 |
| LC/HV | e/n-e | 0.02305 | 0.47498 | 0.00008 |
| HCC/LC | e/miR16 | 8.52313E-9 | 0.00073 | 0.00814 |
| HCC/HV | e/miR16 | 5.18581E-7 | 1.15359E-12 | 3.37936E-8 |
| LC/HV | e/miR16 | 0.00754 | 0.00007 | 0.00040 |
| HCC/LC | n-e/miR16 | 8.522020274789183e-12 | 4.10744E-6 | 2.3056e-13 |
| HCC/HV | n-e/miR16 | 2.04210E-8 | 0.04090 | 0.04874 |
| LC/HV | n-e/miR16 | 0.41124 | 0.00916 | 5.20617E-9 |
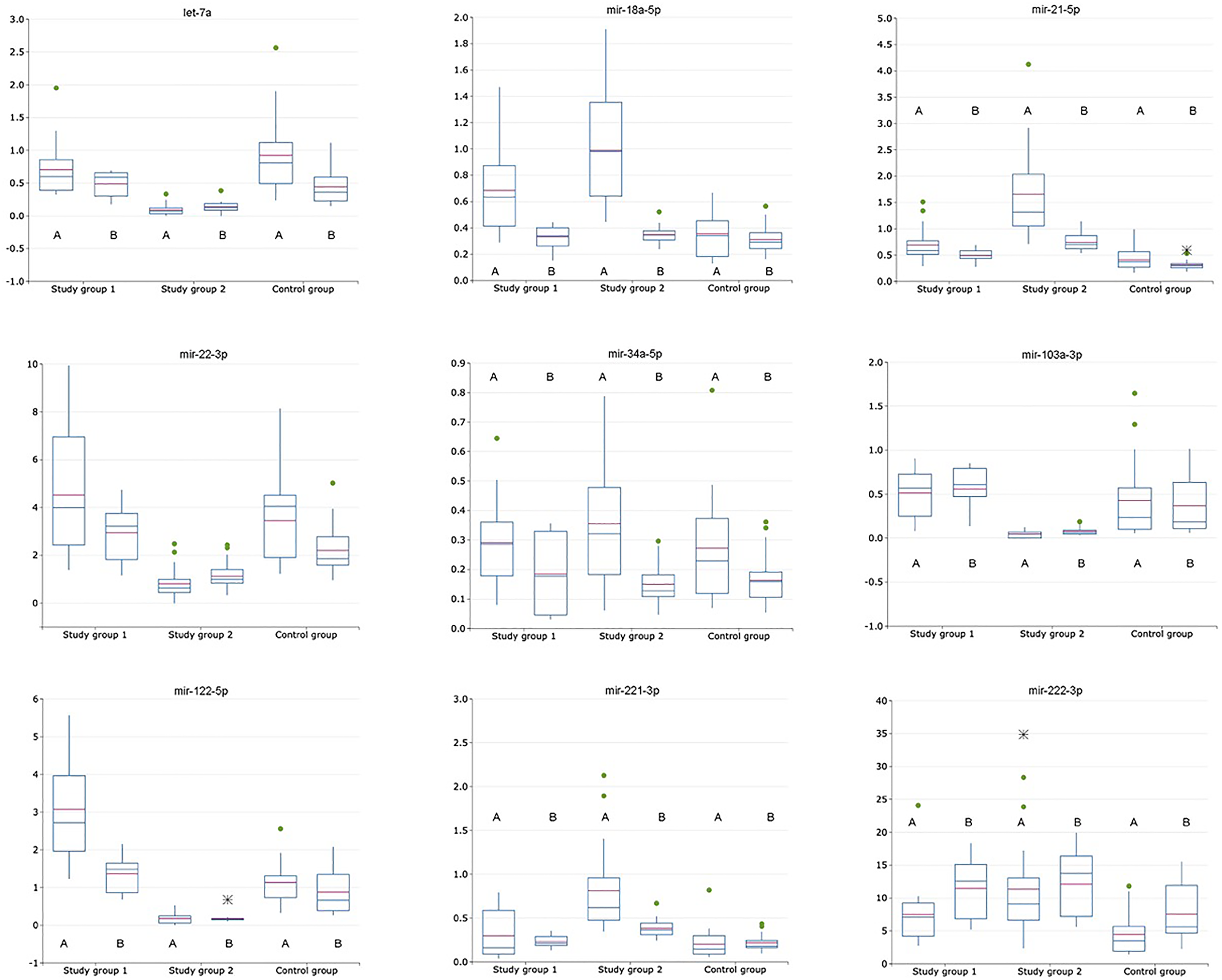
Figure 1: Relative expression of exosomal and non-exosomal miRNAs (normalized to the corresponding miRNA-16-5p) in plasma of patients with HCV-related cirrhosis (study group 1, n = 24), liver cancer (study group 2, n = 24) and healthy volunteers (control group, n = 21); A is for exosomal miRNA; B is for non-exosomal miRNA.
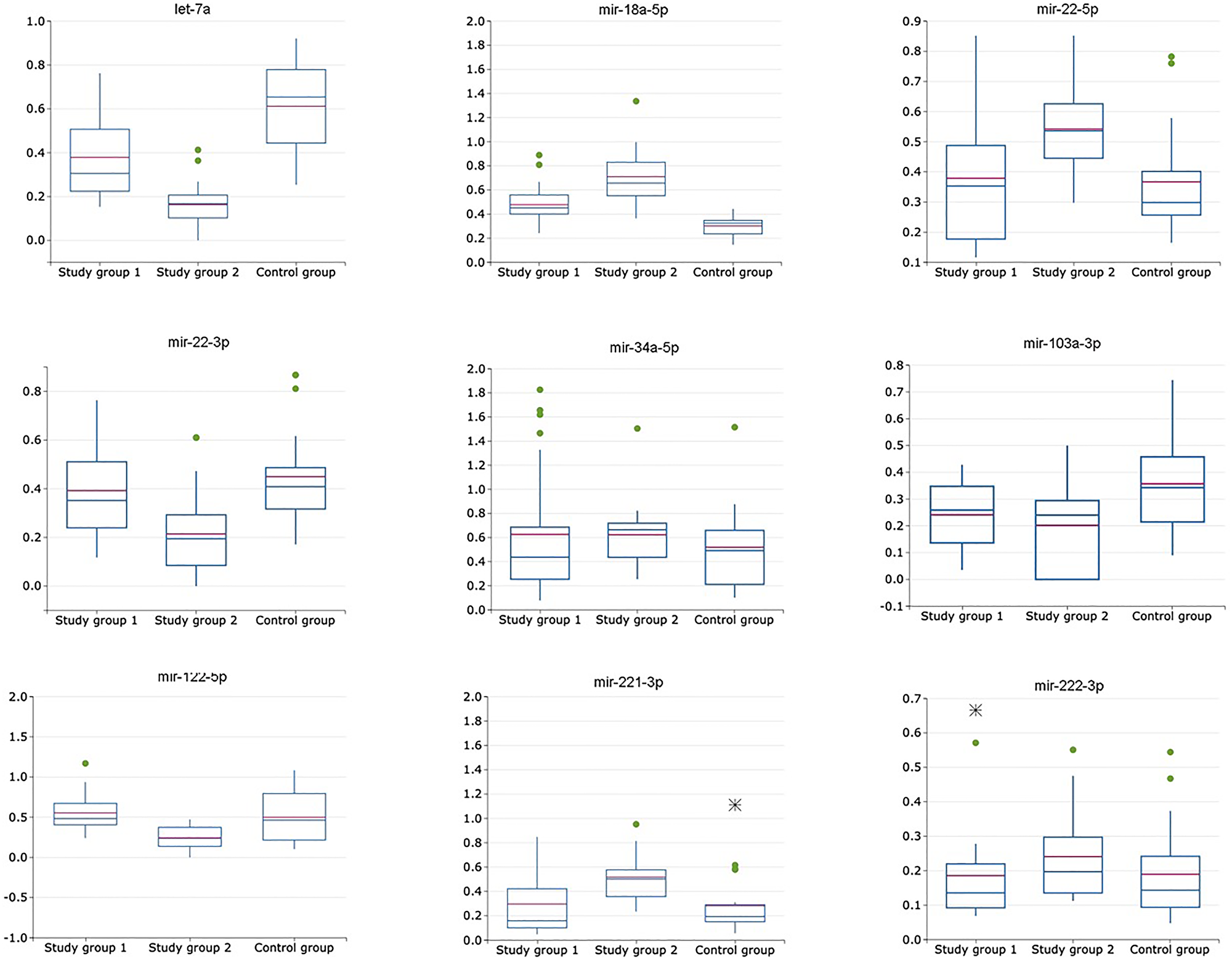
Figure 2: Relative expression of miRNAs (exosomal to non-exosomal miRNAs ratio) in plasma of patients with HCV-related cirrhosis (study group 1, n = 24), liver cancer (study group 2, n = 24) and healthy volunteers (control group, n = 21).
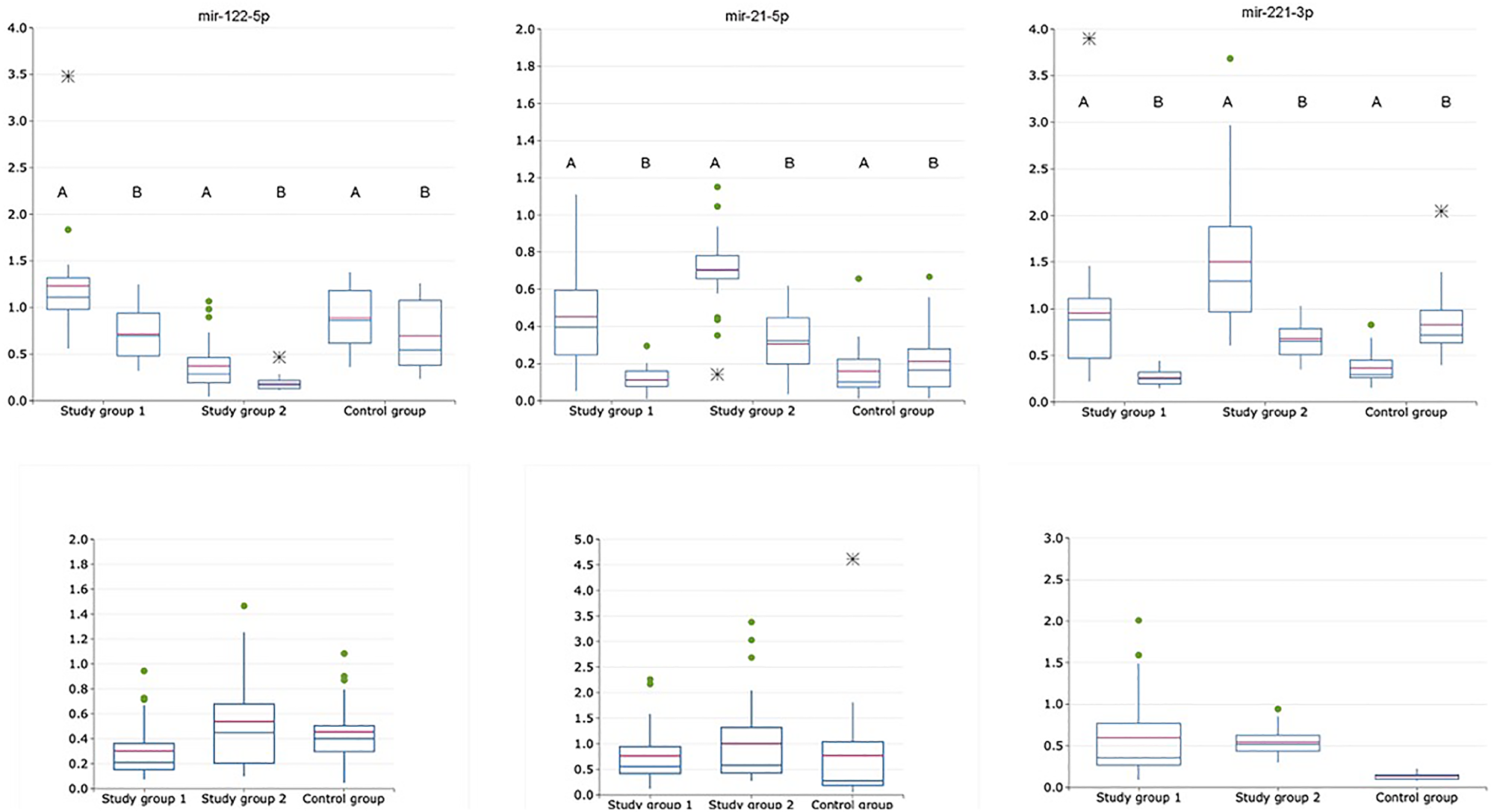
Figure 3: Relative expression of exosomal and non-exosomal miRNAs normalized to the corresponding miRNA-16-5p (upper raw) and normalized as exosomal to non-exosomal miRNAs ratio (lower raw) in saliva of patients with HCV-related cirrhosis (study group 1, n = 24), liver cancer (study group 2, n = 24) and healthy volunteers (control group, n = 21); A is for exosomal samples; B is for non-exosomal samples.
MiRNAs with the most changed relative expression levels (calculated as normalized to the corresponding miRNA-16-5p) compare to control in plasma samples from the cancer study group were following miRNAs: miRNA-21-5p (exosomal increased in 4,04 ± 0,54; non-exosomal increased in 2,35 ± 0,32), miRNA-122-5p (exosomal decreased in 0,15 ± 0,013; non-exosomal decreased in 0,13 ± 0,02), miRNA-221-3p (exosomal increased in 4,01 ± 0,34; non-exosomal increased in 1,77 ± 0,21).
Non-exosomal miRNA relative expression levels calculated as a ratio of target non-exosomal miRNA to non-exosomal miRNA-16-5p ratio
Expression levels of non-exosomal miRNA-34a-5p and miRNA-18a-5p normalized to non-exosomal miRNA-16-5p did not significantly differ (p > 0,05) amongst both study and control groups and were eliminated from the following analysis (Table 2). Expression levels of miRNA-let7a and miRNA-221-3p did not significantly differ (p > 0,05) between the cirrhotic study group and the control group, allowing to suppose there is no significance influence of cirrhosis development on plasma expression levels of these miRNAs. MiRNA-222-3p expression level did not significantly differ (p > 0,05) between the cirrhotic and the cancer study group. ROC analysis for miRNA set consisting of let-7a, miRNA-21-5p, -22-3p, -103a, -122-5p, -221-3p and 222-3p was performed with the values, obtained from the cancer study group and the control group, following AUC values were obtained – 0,98 (sensitivity – 0,9; specificity – 0,98) (Figure 1). Also, ROC analysis for this miRNA set was performed with the values, obtained from the cancer study group and the cirrhotic study group, following AUC values were obtained – 0,97 (sensitivity – 0,9; specificity – 0,99). Non-exosomal miRNA relative expression levels calculated as a ratio of target non-exosomal miRNA to non-exosomal miRNA-16-5p ratio are shown in Figure 1.
Exosomal miRNA relative expression levels calculated as a ratio of target exosomal miRNA to exosomal miRNA-16-5p ratio
Expression level of exosomal miRNA-34a-5p did not significantly differ (p > 0,05) amongst both study and control group and was eliminated from following analysis. Expression levels of exosomal let-7a, miRNA-22-3p, -103a-3p and -221-3p did not significantly differ (p > 0,05) between the liver cirrhosis group and control group. (Table 2) ROC analysis for miRNA set consisting of let-7a, miRNA-18a-5p, miRNA-21-5p, -22-3p, -103a, -122-5p, -221-3p and 222-3p was performed with the values, obtained from the cancer study group and the control group, following AUC values were obtained – 0,96 (sensitivity – 0,82; specificity – 0,95) (Figure 1). Also, ROC analysis for this miRNA set was performed with the values, obtained from the cancer study group and the cirrhotic group, following AUC values were obtained – 0,86 (sensitivity – 0,8; specificity – 0,78). Generally, expression levels of exosomal miRNAs were significantly (p > 0,05) decreased in comparison with non-exosomal miRNA expression levels in all study groups. Exosomal miRNA relative expression levels calculated as a ratio of target exosomal miRNA to exosomal miRNA-16-5p ratio are shown in Figure 1.
Relative miRNA expression levels calculated as exosomal miRNA to non-exosomal miRNA expression levels ratio
This approach to relative miRNA expression level calculation resulted in elimination of miRNA-34a-5p and miRNA-222-3p from the following analysis due to no statistically significant difference in obtained numbers (p > 0,05) (Table 2). Change in expression levels of miRNA-21-5p, -22-3p, -122-5p and -221-3p calculated with this method were not significantly different (p > 0,05) between the liver cirrhosis group and the control group. ROC analysis for miRNA set consisting of let-7a, miRNA-18a-5p, miRNA-21-5p, -22-3p, -103a, -122-5p and -221-3p was performed with the values, obtained from the cancer study group and the control group, following AUC values were obtained – 0,78 (sensitivity – 0,61; specificity – 1,0). Also, ROC analysis for this miRNA set was performed with the values, obtained from the cancer study group and the cirrhotic group, following AUC values were obtained – 0,69 (sensitivity – 0,47; specificity – 0,91). Relative miRNA expression levels calculated as exosomal miRNA to non-exosomal miRNA expression levels ratio are shown in Figure 2.
Exosomal and non-exosomal miRNA expression levels in saliva
Three miRNAs with the most pronounced changes in expression levels in plasma for both exosomal and exosome-depleted samples (normalized to the corresponding miRNA-16-5p) were miRNA-21-5p, miRNA-122-5p and miRNA-221-3p (Table 3). These miRNAs were determined in exosomal and exosome-depleted samples obtained from saliva. Calculating exosomal miRNA to non-exosomal miRNA ratio resulted in no significant difference (p > 0,05) for these miRNAs except for miRNA-221-3p between the cancer study group and control group. Thus, the following analysis was performed with exosomal and non-exosomal miRNA expression levels normalized to the corresponding miRNA-16-5p. ROC analysis for exosomal miRNA set consisting of these three miRNAs was performed with the values, obtained from the cancer study group and the control group, following AUC values were obtained – 0,88 (sensitivity – 0,73; specificity – 1,0). Also, ROC analysis for exosomal miRNA set was performed with the values, obtained from the cancer study group and the cirrhotic group, following AUC values were obtained – 0,54 (sensitivity – 0,78; specificity – 0,5). For non-exosomal miRNAs there were the following values: 0,77 (sensitivity – 0,66; specificity – 0,78) for cancer study group vs control; 0,56 (sensitivity – 0,62; specificity – 0,6) for the cirrhotic study group vs cancer study group. Thus, either exosomal and non-exosomal miRNAs-21-5p, -122-5p and -221-3p determined in saliva do not allow to differ cancer and cirrhotic cases. Exosomal and non-exosomal miRNA expression levels in saliva are shown in Figure 3.
DISCUSSION
MiRNAs are putative molecular harbingers of liver cancer development, which are quite stable molecules detectable in both blood plasma and saliva. Whereas mRNA, rRNA, and tRNA are degraded within several seconds after being placed in nuclease rich extracellular environment [12], most circulating in biological fluids miRNAs are resistant to RNAse activity, pH change and in some researches are supposed to be resistant to multiple numbers of freeze-thaw cycles [13] This resistance is explained by association of miRNAs with protein or lipoprotein complexes or packaging into exosomes what indicates that miRNAs are not just byproducts of cellular activity but are the means of intentional intercellular connections.
In this research, we investigated non-exosomal and exosomal miRNAs expression levels normalized to the corresponding miRNA-16-5p and exosomal miRNA and non-exosomal miRNA ratio in the groups of patients with cirrhosis and liver cancer and in healthy volunteers in blood plasma and the most promising miRNAs in saliva. Expression levels of exosomal miRNAs were significantly (p > 0,05) decreased in comparison with non-exosomal miRNA expression levels in our study for all detected miRNAs in all groups. These data are supported by other studies: in 2011 Turchinovich A, et al. showed, that the amount of plasma supernatant miRNA was ~80 times higher compared to the plasma pellet, thus indicating the overwhelming majority of circulated miRNAs are supernatant (non-exosomal) and is bound to Ago2 protein, but it is worth to admit that exosomes in this paper were defined as vesicles of 50–100 nm [11] Arryo J. D. et al. provided convincing data that about 90% of circulating miRNAs molecules are non-exosomal and associated with different proteins protecting them from external damaging factors [14].
Taking into account a huge role of the exosomes in carcinogenesis and a problem of miRNA normalization it seems to us reasonably despite a minority of exosomal miRNA fraction not to ignore this part of investigated miRNAs. Thus, in this research we used exosomal miRNA X to non-exosomal miRNA X ratio as one of the possible ways of miRNA normalization, where X is the target miRNA. We hypothesized that evaluation of exosomal miRNAs would be more precise compare to non-exosomal miRNAs, but in our research this assumption was not proved: sensitivity and specificity of exosomal miRNAs based panel are significantly inferior to non-exosomal miRNA based panel both for the control group and the study group with liver cancer patients (0, 82 and 0,95 vs 0,9 and 0,98) and for the study groups with patients with HCV-related cirrhosis and liver cancer (0,8 and 0,78 vs 0,9 and 0,98). These results may allow to assume non-exosomal miRNAs have more significant prognostic value compare to exosomal miRNAs. However, it should be admitted, that the sensitivity and specificity were calculated for the whole set of miRNAs with statistically significant difference of relative expression levels between the two groups (cancer and control; cancer and cirrhosis). Such a high values of sensitivity and specificity of non-exosomal miRNAs panel may be explained by the particularities of the sample: all the patients with primary liver cancer involved in this study were at severe stages of the disease (II and more according to AJCC system, 2018), and cancer cells are supposed to be one of the main players in the miRNA expression dysregulation [15]. Nevertheless, there should be admitted that the prevailing part of exosomes circulated in blood plasma are produced by platelets, and their role in primary liver cancer are not that evident and clear, so there is a risk for identification the changes in miRNAs expression levels that are not the consequences of the target process [16].
Generally, identification of non-exosomal miRNAs seems to us more inconvenient than identification of exosomal miRNAs, as far as for the latter case there is a wide variety of commercial kits, some of them are precipitation based and are easy to carry out, allowing relatively standardized exosome isolation method. As for non-exosomal miRNAs there should be performed elimination of microvesicles from the biological fluid, which should allow following investigation of biological fluid for non-exosomal miRNAs. One of the most obvious option is multistep ultracentrifugation which is not a convenient technique for the implementation into clinical routine practice.
The miRNAs for the investigation were chosen according to their target genes, as one of the possible reasons of mRNA profile changes taking place in cancer and adjacent tissues may be the miRNA circulating in blood plasma both in exosomes and in the protein or lipoprotein complexes, which are released by cancer cells and are even able to act like hormones [17]. The second condition was sharing common target genes for several miRNAs. The third condition was identification of the miRNAs by other research groups in the biological fluids and cases of interest for us. Thus, according to MirAnalyze database, let-7a-5p, miRNA-16-5p, -21-5p share common target genes as follows: B-cell translocation gene 2 (BTG 2), and fibroblast growth factor receptor substrate 2 (FGFR2), which change their expression in cancer [18–20]. MiRNA-21-5p was one of the most significantly changed in cancer, what may allow to make a rough assumption that the expression level of its target genes should be decreased. This may find a support in published papers: expression level of BTG2 was significantly reduced in HCC tissues (p = 0.05, n = 44) [28]. The other research demonstrates similar results: BTG2 expression was significantly suppressed in human HCC compared to adjacent non-cancerous tissues. (n = 77) [19]. Nevertheless, the expression level of FGFR2, which is also a target gene for miRNA-21-5p, is increased in HCC samples (n = 54) [20] Surely, the role of miRNA is multivariate and it may induce gene expression in some cases, but for today these cases are insufficiently studied [21]. Common target genes for two others mostly changed miRNAs – decreased miRNA-122-5p and increased miRNA-221-3p - are estrogen receptor 1 (ER-1) and forkhead box P1 (FOXP1). The expression level of both of these target genes - ER-1 and FOXP1 - may be increased in HCV-related HCC samples compare to normal tissue [22, 23]. Anyway, gene expression is a very complex process and miRNAs are not the main factor in gene expression, along with the fact, that the expression levels data of the miRNAs and their target genes are controversial make us considering adding other parameters for including a miRNA to the marker panel. However, increasing in expression levels of miRNA-21-5p and miRNA-221-3p were also found in other studies of liver cancer samples. [24–26].
At last, remembering the primary liver cancer has a strong dependance on geography, its incidence rate is relatively low in Russian Federation [3]. This relative low incidence rate along with other factors including unwillingness of patients of the risk group to come systematically to the research center for blood collection without personal interests make the prospective cases database formation an issue in the Russian Federation. Considering the fact, the Liver Research Center of the RUDN Medical Institute has a database of patients with HCV-associated cirrhosis with the contact information, a possible solution to this problem could be the use of a biomaterial that does not require the presence of medical personnel for its collection, for example, such as a saliva. In our research, we proved saliva was a promising approach as a source for exosomal and non-exosomal miRNAs. The exosomal miRNAs normalized to exosomal miRNA-16-5p seem to be the form of miRNA normalization, which give the strongest association with primary liver cancer in saliva samples.
Materials and Methods
This is a case-control study performed in the Liver Research Center of Institute of Medicine of Peoples’ Friendship University of Russia (RUDN). Patients were informed of the purpose of the acquisition of anonymized clinical data, obtainment of blood and saliva samples and future publication of collected data prior to enrolment in the research project. Informed consent was obtained from all patients. The study was approved by ethics committee of Institute of Medicine of RUDN.
The control group consisted of 21 volunteers: age 30–60, median 46,0 ± 9 (12 male: age 32–60, median 47,5 ± 8,4; 9 female: age 30–59, median 44 ± 9,3), meeting the following criteria: no hospitalization during last 12 months, no cancer/autoimmune disease/alcoholism/drug addiction in anamnesis, no pregnancy or lactation. The first study group (hereinafter, cirrhotic group) consisted of 24 patients with HCV-related liver cirrhosis: age 35–55, median 45,5 ± 6,8 (15 male: 35–55, median 44 ± 7,4; 9 female: 35–53, median 46 ± 6,3) and the second study group (hereinafter, cancer group) consisted of 24 patients with liver cancer: age 43–65, median 49,5 (Q1:44, Q3:58,2); (14 male: age 43–62, median 49,0 (Q1:44, Q3:57); 10 female: age 44–65, median 52,0 ± 8,6). Liver cirrhosis was diagnosed by transient elastography method (METAVIR F4, FibroScan), HCV status was positive in anamnesis, all patients with HCV-related cirrhosis were treated with direct acting antivirals and are HCV negative during more than 1 year, liver cancer was diagnosed by contrast-enhanced MRI with liver-specific contrast agents, 15 cases were identified as hepatocellular carcinoma by pathomorphological examination, 9 cases missed the results of pathomorphological examination at the time of this writing.
Peripheral venous blood was collected in EDTA K2 4 ml tubes from volunteers, patients with liver cirrhosis and liver cancer, immediately after collection blood samples were centrifugated at 3000 g for 10 minutes at room temperature (Eppendorf 5810R, rotor A-4-81, Germany), two aliquots for each patient were frozen at –80°C. Before saliva obtainment patients undergo 2 hours fasting, saliva was obtained in 50 ml tube, about 1-2 ml was transferred to 1.5 ml tubes and centrifugated at 3000 g for 10 minutes at room temperature (Eppendorf Minispin®, Germany), 2 aliquots of supernatant of each sample were collected and frozen at –80°C.
Further exosomal and non-exosomal miRNAs were isolated both from plasma and saliva samples. In order to obtain these miRNA fractions, exosomes were isolated from one aliquot of every sample and proceeded for miRNA isolation, another aliquot of the same sample was ultracentrifugated and exosome-depleted supernatant proceeded for miRNA isolation. Hereinafter, miRNAs, that are isolated from the exosomes are named exosomal miRNA; miRNAs, that are isolated from the exosome-depleted supernatant samples are named non-exosomal miRNAs.
In order to obtain exosomal miRNA fraction exosomes from plasma and saliva samples were isolated with miRCURY Exosome Serum/Plasma Kit (Qiagen, Germany). After thawing saliva and plasma samples were centrifugated at 8000 g for 10 minutes and at 10 000 g for 30 minutes, in order to enrich for microvesicles both saliva and plasma samples (400 μL) after centrifugation were passed through a 0.22 μm filter [27]. The resulting filtrates were processed to exosome isolation, in brief, filtrates were supplemented with PBS till 0.5 mL. Further isolation process was performed according to the manufacturer’s protocols finishing with samples incubation at 4°C for 1 h and centrifugation at 500 g for 5 min (protocol plasma) at room temperature. Pellet fraction was resuspended in 270 μL of Resuspension buffer and frozen at −80°C. These enriched for exosomes samples were named exosomal samples. In order to obtain non-exosomal miRNA fraction, aliquots of plasma and saliva samples after thawing were centrifugated at following regimen: 300 g 10 minutes, 2000 g 10 minutes, 10,000 g 30 minutes, 150,000 g 70 minutes, 150,000 g 70 minutes, at every step the supernatant was taken (Optima Max E, MLA-150 rotor, Beckman Coulter, Inc., CA, USA) [28]. These exosome-depleted samples were proceeded to following miRNA isolation.
8 random exosomal samples and 8 random exosome depleted samples of every group were analyzed with photon cross-correlation spectroscopy (NANOPHOX, Sympatech GmbH, Germany) in order to identify the size spectrum of isolated vesicles. All exosomal and exosome depleted samples were examined with CD63 based ELISA (ExoELISA-ULTRA Complete Kit, System Biosciences, CA, USA) (Table 1).
MiRNAs were isolated with similar kit from both exosomal and exosome-depleted plasma and saliva samples according to the manufacturer’s instruction (miRNeasy Serum/Plasma Advanced kit, Qiagen, Germany), miRNA concentration in samples after isolation was estimated by ultra-violet visible spectrophotometry in “microRNA” mode (NanoPhotometer® NP80, Implen GmbH, Germany). Reverse transcription reactions were performed using a SYBR miRNA Reverse Transcription kit (miRCURY LNA™ RT Kit, Qiagen, Germany). qPCR was performed using CFX96™ Real Time System (Bio-Rad, CA, USA). Each reaction mixture for qPCR contained 5 μL of master mix, 1 μL miRNA-specific LNA primer and 4 μL diluted RT product (1:80) in a total volume of 10 μL. For both plasma exosomal and non-exosomal depleted samples LNA primers for following miRNAs were used: let-7a-5p, miR-16-5p, miR-18a-5p, miR-21-5p, miR-22-3p, miR-34a-5p, miR-103a-3p, miR-122-5p, miR-145-5p, miR-221-3p, miR-222-3p, miR-224-5p. For both saliva exosomal and exosome-depleted samples LNA primers for following miRNAs were used: let-7a, miRNA-21-5p, miRNA-122-5p. Reactions were carried out using the following thermal cycling parameters: 95°C for 10 min, followed by 40 cycles of 95°C for 10 s, and 60°C for 1 min, followed by holding at 4°C. Raw data were analyzed using Bio-Rad CFX Manager Software version 3.1 (Bio-Rad, CA, USA), generally using the automatic Ct setting for assigning baseline and threshold values for Ct determination. Those miRNAs that appeared on the profile with Ct value of < 35 were considered as ‘detected’. The expression level of each miRNA was normalized to miRNA-16 expression level, exosomal and non-exosomal miRNA ratio was calculated.
Conclusions
In this study we have realized that non-exosomal miRNAs normalized to non-exosomal miRNA-16p in plasma samples are stronger associated with primary liver cancer, than exosomal miRNAs. Three miRNAs with the mostly pronounced change of expression levels in plasma in liver cancer samples – miRNA-21-5p, 122-5p, 221-3p – were detected in saliva. In contrast to plasma samples, in saliva exosomal miRNA normalized to exosomal miRNA-16-5p showed stronger association with liver cancer, than non-exosomal miRNAs. Thus, saliva may be a promising approach for collection of prospective primary liver cases for the search of prognostic miRNAs.
CONFLICTS OF INTEREST
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflicts of interest.
FUNDING
This work was supported by grant 5–100.
References
1. Dasgupta P, Henshaw C, Youlden DR, Clark PJ, Aitken JF, Baade PD. Global Trends in Incidence Rates of Primary Adult Liver Cancers: A Systematic Review and Meta-Analysis. Front Oncol. 2020; 10:171. https://doi.org/10.3389/fonc.2020.00171. [PubMed].
2. https://www.cancer.net/cancer-types/liver-cancer/statistics.
3. Yang JD, Hainaut P, Gores GJ, Amadou A, Plymoth A, Roberts LR. A global view of hepatocellular carcinoma: trends, risk, prevention and management. Nat Rev Gastroenterol Hepatol. 2019; 16:589–604. https://doi.org/10.1038/s41575-019-0186-y. [PubMed].
4. Desai A, Sandhu S, Lai JP, Sandhu DS. Hepatocellular carcinoma in non-cirrhotic liver: A comprehensive review. World J Hepatol. 2019; 11:1–18. https://doi.org/10.4254/wjh.v11.i1.1. [PubMed].
5. Hamid AS, Tesfamariam IG, Zhang Y, Zhang ZG. Aflatoxin B1-induced hepatocellular carcinoma in developing countries: Geographical distribution, mechanism of action and prevention. Oncol Lett. 2013; 5:1087–92. https://doi.org/10.3892/ol.2013.1169. [PubMed].
6. Khullar V, Firpi RJ. Hepatitis C cirrhosis: New perspectives for diagnosis and treatment. World J Hepatol. 2015; 7:1843–55. https://doi.org/10.4254/wjh.v7.i14.1843. [PubMed].
7. Doyle LM, Wang MZ. Overview of Extracellular Vesicles, Their Origin, Composition, Purpose, and Methods for Exosome Isolation and Analysis. Cells. 2019; 8:727. https://doi.org/10.3390/cells8070727. [PubMed].
8. Kalluri R, LeBleu VS. The biology, function, and biomedical applications of exosomes. Science. 2020; 367:eaau6977. https://doi.org/10.1126/science.aau6977. [PubMed].
9. Yi YW, Lee JH, Kim SY, Pack CG, Ha DH, Park SR, Youn J, Cho BS. Advances in Analysis of Biodistribution of Exosomes by Molecular Imaging. Int J Mol Sci. 2020; 21:665. https://doi.org/10.3390/ijms21020665. [PubMed].
10. Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015; 15:321–33. https://doi.org/10.1038/nrc3932. [PubMed].
11. Turchinovich A, Weiz L, Langheinz A, Burwinkel B. Characterization of extracellular circulating microRNA. Nucleic Acids Res. 2011; 39:7223–33. https://doi.org/10.1093/nar/gkr254. [PubMed].
12. Turchinovich A, Samatov TR, Tonevitsky AG, Burwinkel B. Circulating miRNAs: cell-cell communication function? Front Genet. 2013; 4:119. https://doi.org/10.3389/fgene.2013.00119. [PubMed].
13. Glinge C, Clauss S, Boddum K, Jabbari R, Jabbari J, Risgaard B, Tomsits P, Hildebrand B, Kääb S, Wakili R, Jespersen T, Tfelt-Hansen J. Stability of Circulating Blood-Based MicroRNAs - Pre-Analytic Methodological Considerations. PLoS One. 2017; 12:e0167969. https://doi.org/10.1371/journal.pone.0167969. [PubMed].
14. Arroyo JD, Chevillet JR, Kroh EM, Ruf IK, Pritchard CC, Gibson DF, Mitchell PS, Bennett CF, Pogosova-Agadjanyan EL, Stirewalt DL, Tait JF, Tewari M. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A. 2011; 108:5003–08. https://doi.org/10.1073/pnas.1019055108. [PubMed].
15. Croce CM. Causes and consequences of microRNA dysregulation in cancer. Nat Rev Genet. 2009; 10:704–14. https://doi.org/10.1038/nrg2634. [PubMed].
16. Tao SC, Guo SC, Zhang CQ. Platelet-derived Extracellular Vesicles: An Emerging Therapeutic Approach. Int J Biol Sci. 2017; 13:828–34. https://doi.org/10.7150/ijbs.19776. [PubMed].
17. Pardini B, Calin GA. MicroRNAs and Long Non-Coding RNAs and Their Hormone-Like Activities in Cancer. Cancers (Basel). 2019; 11:378. https://doi.org/10.3390/cancers11030378. [PubMed].
18. Chen Y, Chen C, Zhang Z, Xiao H, Mao B, Huang H, Ding C, Lei L, Zhang H, Li J, Jiang M, Wang G. Expression of B-cell translocation gene 2 is associated with favorable prognosis in hepatocellular carcinoma patients and sensitizes irradiation-induced hepatocellular carcinoma cell apoptosis in vitro and in nude mice. Oncol Lett. 2017; 13:2366–72. https://doi.org/10.3892/ol.2017.5685. [PubMed].
19. Huang CS, Zhai JM, Zhu XX, Cai JP, Chen W, Li JH, Yin XY. BTG2 Is Down-Regulated and Inhibits Cancer Stem Cell-Like Features of Side Population Cells in Hepatocellular Carcinoma. Dig Dis Sci. 2017; 62:3501–10. https://doi.org/10.1007/s10620-017-4829-y. [PubMed].
20. Jun BG, Lee WC, Jang JY, Jeong SW, Chang Y, Lee SH, Kim YD, Kim SG, Cheon GJ, Kim YS, Kim HS, Jin SY. Relation of fibroblast growth factor receptor 2 expression to hepatocellular carcinoma recurrence after liver resection. PLoS One. 2020; 15:e0227440. https://doi.org/10.1371/journal.pone.0227440. [PubMed].
21. Vaschetto LM. miRNA activation is an endogenous gene expression pathway. RNA Biol. 2018; 15:826–28. https://doi.org/10.1080/15476286.2018.1451722. [PubMed].
22. Iyer JK, Kalra M, Kaul A, Payton ME, Kaul R. Estrogen receptor expression in chronic hepatitis C and hepatocellular carcinoma pathogenesis. World J Gastroenterol. 2017; 23:6802–16. https://doi.org/10.3748/wjg.v23.i37.6802. [PubMed].
23. Wang X, Sun J, Cui M, Zhao F, Ge C, Chen T, Yao M, Li J. Downregulation of FOXP1 Inhibits Cell Proliferation in Hepatocellular Carcinoma by Inducing G1/S Phase Cell Cycle Arrest. Int J Mol Sci. 2016; 17:1501. https://doi.org/10.3390/ijms17091501. [PubMed].
24. Zhang T, Yang Z, Kusumanchi P, Han S, Liangpunsakul S. Critical Role of microRNA-21 in the Pathogenesis of Liver Diseases. Front Med (Lausanne). 2020; 7:7. https://doi.org/10.3389/fmed.2020.00007. [PubMed].
25. Wagenaar TR, Zabludoff S, Ahn SM, Allerson C, Arlt H, Baffa R, Cao H, Davis S, Garcia-Echeverria C, Gaur R, Huang SM, Jiang L, Kim D, et al. Anti-miR-21 Suppresses Hepatocellular Carcinoma Growth via Broad Transcriptional Network Deregulation. Mol Cancer Res. 2015; 13:1009–21. https://doi.org/10.1158/1541-7786.MCR-14-0703. [PubMed].
26. Pineau P, Volinia S, McJunkin K, Marchio A, Battiston C, Terris B, Mazzaferro V, Lowe SW, Croce CM, Dejean A. miR-221 overexpression contributes to liver tumorigenesis. Proc Natl Acad Sci U S A. 2010; 107:264–69. https://doi.org/10.1073/pnas.0907904107. [PubMed].
27. Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-Mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007; 9:654–59. https://doi.org/10.1038/ncb1596. [PubMed].
28. Li P, Kaslan M, Lee SH, Yao J, Gao Z. Progress in Exosome Isolation Techniques. Theranostics. 2017; 7:789–804. https://doi.org/10.7150/thno.18133. [PubMed].

