Introduction
The overall survival (OS) for patients with metastatic colorectal cancer (mCRC) has markedly improved within the last 2 decades, reaching approximately 30 months [1–3].
The therapeutic options include fluoropyrimidines, oxaliplatin and irinotecan combinations (either doublets or triplets) with one anti-vascular endothelial growth factor (anti-VEGF) compound (i.e., bevacizumab, aflibercept and ramucirumab), and/or anti-epidermal growth factor receptor (EGFR) monoclonal antibody (MoAb, namely cetuximab or panitumumab) for patients with RAS/BRAF wild-type status, administered across different treatment lines [4]. Many factors contribute to the choice of the optimal treatment sequence, among these the tumor molecular status.
An important advancement in mCRC patients is based on a better understanding of both tumor biology and microenvironment. The RAS/RAF/MAPK pathway is downstream of EGFR, and its mutations are predictive for a lack of benefit from EGFR targeted therapies [5–9]. Thus, it is crucial to perform an extended RAS and BRAF mutation analysis before considering EGFR inhibitors in mCRC patients. In first-line, data support the use of anti-EGFR MoAbs in patients with left-sided, RAS and BRAF wild-type tumors, whilst bevacizumab is preferred in combination with chemo-doublets or -triplet in RAS/BRAF mutated tumors or RAS/BRAF wild-type right-sided primaries [10, 11].
More recently, two intriguing studies showed that cetuximab sensitivity might be restored either in RAS/BRAF wild-type mCRC patients who acquired resistance to cetuximab-based therapy in first-line therapy [12] or in those with primarily RAS mutated tumors treated with bevacizumab-containing regimens in second-line [13]. These studies may lead to the hypothesis that anti-angiogenic agents through action on RAS mutated cells could revert tumors from RAS mutant to RAS wild-type status, which theoretically could lead to the possibility to treat these patients with anti-EGFR MoAbs, otherwise precluded. In both mentioned trials, molecular analyses were performed by using liquid biopsies.
To substantiate these findings at tissue level, here we analyzed molecular changes in RAS mutated mCRC patients treated with chemotherapy alone or in combination with bevacizumab, by examining tumor tissue samples before and after the systemic therapy using two independent methodologies, next-generation sequencing (NGS) and real-time PCR.
Results
Study population
The cohort included 28 patients (14 male, 14 female); median age at diagnosis was 67.6 years (range 39–87). All patients exhibited RAS mutation in primary tumor, located in 25 cases in the colon and in 3 cases in the rectum. The metastases were limited to the liver.
Our cohort is subdivided in three groups as previously described (details are summarized in Table 1).
Table 1: Details of the patients included in the three treatment groups analyzed by IOT
| n | Group | Sex | Age | PT localization | LM0 (sy) | LM1 (me) | LM2 (me) | LM3 (me) |
|---|---|---|---|---|---|---|---|---|
| 1 | 1 | M | 59 | right colon | ✓ | ✓ | ||
| 2 | 1 | M | 70 | left colon | ✓ | ✓ | ||
| 3 | 1 | M | 69 | left colon | ✓ | |||
| 4 | 1 | M | 64 | rectum | ✓ | ✓ | ||
| 5 | 1 + 3 | F | 61 | right colon | ✓ | ✓ | ||
| 6 | 1 | M | 69 | left colon | ✓ | |||
| 7 | 1 | F | 65 | left colon | ✓ | |||
| 8 | 2 | F | 70 | left colon | ✓ | |||
| 9 | 2 + 3 | F | 57 | right colon | ✓ | ✓ | ||
| 10 | 2 | F | 58 | right colon | ✓ | |||
| 11 | 2 | F | 71 | left colon | ✓ | |||
| 12 | 2 | M | 48 | right colon | ✓ | |||
| 13 | 2 | F | 75 | right colon | ✓ | |||
| 14 | 2 | M | 76 | rectum | ✓ | ✓ | ||
| 15 | 2 | M | 72 | rectum | ✓ | |||
| 16 | 2 | M | 47 | left colon | ✓ | ✓ | ||
| 17 | 2 | F | 81 | right colon | ✓ | |||
| 18 | 2 | F | 58 | right colon | ✓ | ✓ | ✓ | |
| 19 | 2 | M | 81 | left colon | ✓ | ✓ | ||
| 20 | 3 | F | 39 | right colon | ✓ | |||
| 21 | 3 | M | 61 | left colon | ✓ | |||
| 22 | 3 | M | 78 | left colon | ✓ | |||
| 23 | 3 | F | 68 | left colon | ✓ | |||
| 24 | 3 | M | 83 | right colon | ✓ | |||
| 25 | 3 | F | 85 | right colon | ✓ | |||
| 26 | 3 | F | 75 | right colon | ✓ | |||
| 27 | 3 | F | 87 | left colon | ✓ | |||
| 28 | 3 | M | 66 | right colon | ✓ |
Group 1 consisted of 7 patients (5 male, 2 female) who received first-line chemotherapy plus bevacizumab before resection of liver metastases; the median age was 65.3 years (range 59–70).
Group 2 included 12 patients (5 male, 7 female) who underwent first-line chemotherapy alone before resection of liver metastases; the median age was 66.2 years (range 47–81). In Groups 1 and 2 liver metastases were resected metachronously with respect to the primary tumor, at completion of systemic therapy.
The Group 3, control arm, included 11 patients (4 male, 7 female) who did not receive systemic therapy because they underwent concomitant resection of the primary tumor and paired liver metastases. The median age in this group was 69.5 years (range 39–87).
Two cases, out of the 28 included in our cohort, were classified in two distinct groups, namely patient 5 in Groups 1 and 3 and patient 9 in Groups 2 and 3, as they underwent concomitant resection of liver metastases with primary tumor but also secondary resection after systemic therapy.
In some patients we had the opportunity to analyze metastases that emerged later during the follow-up after initial multimodality therapy. In 3 cases of Group 1 (patients 1, 2 and 4) two metastases were available. In Group 2, two metastases were available in 3 cases (patients 14, 16 and 19) and 3 metastases in one case (patient 18).
Assessment of the uniformity of tissue samples: preliminary experiment
We initially selected two cases, patients 8 and 9 (Table 1), for whom abundant tumor tissue was available for molecular analyses. For each patient we examined by NGS and real time PCR 4 different formalin-fixed paraffin-embedded (FFPE) tissue block from the primary tumor (PT, namely PTa, PTb, PTc, PTd) and 4 different FFPE blocks from the liver metastasis (LM, namely LM1a, LM1b, LM1c, LM1d). All details are summarized in Table 2.
Table 2: Results of the pilot experiment to assess the uniformity of the tissue samples
| n | Sample | Group | Localization | KRAS-(%) | APC-(%) | TP53-(%) | PIK3CA-(%) | |
|---|---|---|---|---|---|---|---|---|
| Patient 8 | PT Patient 8 (4 different FFPE blocks) | 8PTa | 2 | left colon | G13D (45.7%) | T1556fs (63.6%) | wt | E545K (43%) |
| 8PTb | 2 | left colon | G13D (47.6%) | T1556fs (65.6%) | wt | E545K (41.8%) | ||
| 8PTc | 2 | left colon | G13D (55.6%) | T1556fs (63.8%) | wt | E545K (42.6%) | ||
| 8PTd | 2 | left colon | G13D (52.6%) | T1556fs (62.6%) | wt | E545K (40.3%) | ||
| LM1 Patient 8 (4 different FFPE blocks) | 8LM1a | 2 | liver | G13D (57.6%) | T1556fs (57.9%) | wt | E545K (42%) | |
| 8LM1b | 2 | liver | G13D (48.7%) | T1556fs (60.5%) | wt | E545K (38.7%) | ||
| 8LM1c | 2 | liver | G13D (50.2%) | T1556fs (63%) | wt | E545K (43.2%) | ||
| 8LM1d | 2 | liver | G13D (41.8%) | T1556fs (61.6%) | wt | E545K (42.8%) | ||
| Patient 9 | PT Patient 9 (4 different FFPE blocks) | 9PTa | 2 | right colon | G12V (34.7%) | wt | wt | E542K (51.6%) |
| 9PTb | 2 | right colon | G12V (35.7%) | wt | wt | E542K (49.2%) | ||
| 9PTc | 2 | right colon | G12V (33.3%) | wt | wt | E542K (50.8%) | ||
| 9PTd | 2 | right colon | G12V (32%) | wt | wt | E542K (52.4%) | ||
| LM1 Patient 9 (4 different FFPE blocks) | 9LM1a | 2 | liver | G12V (36.6%) | wt | wt | E542K (50.1%) | |
| 9LM1b | 2 | liver | G12V (32.4%) | wt | wt | E542K (54%) | ||
| 9LM1c | 2 | liver | G12V (34.2%) | wt | wt | E542K (51.8%) | ||
| 9LM1d | 2 | liver | G12V (36%) | wt | wt | E542K (49.5%) | ||
After normalization based on the tumor content evaluated by the pathologists, the mutations detected in different blocks led us to establish that patient 8 exhibits the same mutations at similar percentages in all areas of the PT (samples 8PTa, 8PTb, 8PTc, 8PTd) and in all different regions of paired liver metastasis (samples 8LM1a, 8LM1b, 8LM1c, 8LM1d). Similar findings were observed in patient 9 after analysis of PT (samples 9PTa, 9PTb, 9PTc, 9PTd) and of the metachronous metastasis (samples 9LM1a, 9LM1b, 9LM1c, 9LM1d). The comparison data concerning the analysis of different tumor areas were not obtained for the synchronous metastasis of patient 9 (i.e.,: 9LM0) because only one tissue block was available (Table 3).
Table 3: Ion Torrent patterns in primary tumour and in the different liver metastases
| n | Treatment group | Sample number | RAS-(%) | APC-(%) | TP53-(%) | Other genes-(%) |
|---|---|---|---|---|---|---|
| 1 | 1 | 1PT | KRAS G12C (15.2) | wt | R280T (50.7) | PIK3CA: E542K (23.0) |
| 1LM1 | KRAS G12C (6.8) | wt | R280T (20.0) | PIK3CA: E542K (12.6) | ||
| 1LM2 | KRAS G12C (8.1) | wt | R280T (36.4) | PIK3CA: E542K (6.44) | ||
| 2 | 1 | 2PT | KRAS G12V (24.8) | wt | wt | wt |
| 2LM1 | KRAS G12V (8.1) | wt | wt | wt | ||
| 2LM2 | KRAS G12V (32.9) | wt | wt | wt | ||
| 3 | 1 | 3PT | KRAS G12S (23.8) | R1114* (34.4) | R282W (35.1) | wt |
| 3LM | KRAS G12S (37.9) | R1114* (59.9) | R282W (64.4) | PIK3CA: E542K (23.0) | ||
| 4 | 1 | 4PT | KRAS G12V (25.9) | E1379* (46.9) | F134L (45.9) | SMAD4: D355V (25.7) |
| 4LM1 | KRAS G12V (7.9) | E1379* (15.2) | F134L (11.8) | SMAD4: D355V (7.1) | ||
| 4LM2 | KRAS G12V (12.7) | E1379* (21.2) | F134L (13.9) | SMAD4: D355V (16.8) | ||
| 5 | 1 + 3 | 5PT | KRAS G12C (38.3) | wt | R273H (50.8) | wt |
| 3 | 5LM0 | KRAS G12C (34.1) | wt | R273H (52.2) | wt | |
| 1 | 5LM1 | KRAS G12C (32.6) | wt | R273H (35.1) | wt | |
| 6 | 1 | 6PT | NRAS G12D (21.2) | Q1291* (61.0) | T256P (37.8) | PIK3CA: E542K (10.2) |
| 1 | 6LM1 | NRAS G12D (6.9) | Q1291* (15.5) | T256P (8.8) | wt | |
| 7 | 1 | 7PT | KRAS G12D (22.2) | E1306K (19.3) | wt | wt |
| 7LM1 | KRAS G12D (14.41) | E1306K (8.8) | wt | wt | ||
| 8 | 2 | 8PT | KRAS G13D (45.7) | T1556fs (63.6) | wt | PIK3CA E545K (43.0) |
| 8LM1 | KRAS G13D (57.6) | T1556fs (57.9) | wt | PIK3CA E545K (42.0) | ||
| 9 | 2 + 3 | 9PT | KRAS G12V (34.7) | wt | wt | PIK3CA E542K (51.6) |
| 3 | 9LM0 | KRAS G12V (35.3) | wt | wt | wt | |
| 2 | 9LM1 | KRAS G12V (36.6) | wt | wt | PIK3CA E542K (50.1) | |
| 10 | 2 | 10PT | KRAS G12S (40.1) | wt | wt | CDKN2A: V106M (39.6) |
| 10LM1 | KRAS G12S (38.6) | wt | wt | CDKN2A: V106M (25.5) | ||
| 11 | 2 | 11PT | KRAS G13D (16.8) | G1499* (15.9) | wt | wt |
| 11LM1 | KRAS G13D (8.4) | G1499* (12.5) | wt | wt | ||
| 12 | 2 | 12PT | KRAS G12S (43.7) | S1503* (24.8) | wt | PIK3CA: C420R (18.2) |
| 12LM1 | KRAS G12S (41.1) | S1503* (22.9) | wt | PIK3CA: C420R (20.4) | ||
| 13 | 2 | 13PT | KRAS G12V (16.2) | I1307Kfs*8 (42.6) | C242fs*5 (29.8) | wt |
| 13LM1 | KRAS G12V (20.9) | I1307Kfs*8 (52.4) | C242fs*5 (28.7) | wt | ||
| 14 | 2 | 14PT | KRAS G13D (27.2) | Q1378* (28) | Y220C (42.1) | wt |
| 14LM1 | KRAS G13D (23.9) | Q1378* (18.4) | Y220C (27.0) | SMAD4: R135* (14.5) | ||
| 14LM2 | KRAS G13D (19.9) | Q1378* (19.1) | Y220C (28.1) | wt | ||
| 15 | 2 | 15PT | KRAS G12D (13.3) | wt | wt | wt |
| 15LM1 | KRAS G12D (13.1) | wt | wt | wt | ||
| 16 | 2 | 16PT | KRAS G12D (21.6) | G1357* (23.8) | R175H (23.0) | wt |
| 16LM1 | KRAS G12D (16.8) | G1357* (30.8) | R175H (35.4) | wt | ||
| 16LM2 | KRAS G12D (29.0) | G1357* (39.7) | wt | wt | ||
| 17 | 2 | 17PT | KRAS G12D (12.4) | wt | wt | CDKN2A: D84N (16.2) |
| 17LM1 | KRAS G12D (18.8) | wt | wt | CDKN2A: D84N (14.5) | ||
| 18 | 2 | 18PT | KRAS G12A (21.9) | wt | H214R (33.8) | wt |
| 18LM1 | KRAS G12A (22.1) | wt | H214R (47.2) | wt | ||
| 18LM2 | KRAS G12A (17.6) | wt | H214R (9.4) | wt | ||
| 18LM3 | KRAS G12A (33.2) | wt | H214R (21.7) | wt | ||
| 19 | 2 | 19PT | KRAS G12A (46.4) | wt | P151T (61.6) | wt |
| 19LM1 | KRAS G12A (41.5) | wt | P151T (76.2) | wt | ||
| 19LM2 | KRAS G12A (48.7) | wt | P151T (63.5) | wt | ||
| 20 | 3 | 20PT | KRAS G13D (32.5) | wt | P98S (33.8) | EGFR: V774M (34.2) |
| 20LM0 | KRAS G13D (24.5) | wt | P98S (29.1) | EGFR: V774M (23.2) | ||
| 21 | 3 | 21PT | KRAS G12D (54.7) | F1491Lfs*16 (38.9) | wt | wt |
| 21LM0 | KRAS G12D (44.8) | F1491Lfs*16 (37.1) | wt | wt | ||
| 22 | 3 | 22PT | KRAS G12C (22.9) | wt | wt | wt |
| 22LM0 | KRAS G12C (18.1) | wt | wt | wt | ||
| 23 | 3 | 23PT | KRAS G12D (29.4) | E1309Dfs*4 (45.3) | wt | SMAD4: W398* (49.7) |
| 23LM0 | KRAS G12D (20.6) | E1309Dfs*4 (42.2) | wt | SMAD4: W398* (35.6) | ||
| 24 | 3 | 24PT | KRAS G12D (49.6) | wt | E271* (54.4) | PTEN: Y240* (63.1); GNAS: R844C (3.0) |
| 24LM0 | KRAS G12D (56.2) | wt | E271* (61.2) | PTEN: Y240* (56.2); GNAS: wt | ||
| 25 | 3 | 25PT | KRAS G12C (51.3) | wt | R196* (37.0) | PIK3CA: E453K (40.4) |
| 25LM0 | KRAS G12C (50.5) | wt | R196* (69.8) | PIK3CA: E453K (32.4) | ||
| 26 | 3 | 26PT | KRAS G12D (28.4) | wt | wt | RB1: D340N (6.9) |
| 26LM0 | KRAS G12D (30.9) | wt | wt | RB1: D340N (6.5) | ||
| 27 | 3 | 27PT | KRAS G12D (41.2) | P1439Lfs*34 (31.6) | Q136* (64.6) | wt |
| 27LM0 | KRAS G12D (39.7) | P1439Lfs*34 (46.7) | Q136* (65.8) | wt | ||
| 28 | 3 | 28PT | NRAS Q61K (32.2) | wt | wt | PIK3CA: E453K (40.4) |
| 28LM0 | NRAS Q61K (35.9) | wt | wt | PIK3CA: Q546K (28.0) |
Overall, the findings obtained from patients 8 and 9 suggested that the mutations are uniformly distributed in both PT and paired liver metastases. Based on this we speculated that only one tissue block from the PT and from the paired liver metastases could reflect the molecular patterns of the whole tumor.
IOT analysis of RAS and other relevant genes in primary tumor and paired liver metastases
In all cases, NGS experiments by Ion Torrent (IOT) provided the calculation of the percentage of the mutant allele of 48 genes in addition to KRAS and NRAS. The data obtained after analysis of PT and liver metastases tissue blocks were normalized with the tumor cells (t cells) % content.
Group 1: patients treated with chemotherapy plus bevacizumab before resection of liver metastases
The comparison of RAS mutation percentages between PT and paired liver metastases resected after systemic therapy showed a reduction of at least 50% of RAS mutant alleles in 4 out of 7 cases (57.1%; patients 1, 2, 4 and 6); in 2 cases (patients 4 and 6) the decrease was greater than 3 times. In the remaining cases, a heterogeneous trend was observed, with a decrease of about 30% in 1 case (patient 7), similar rate in one case (patient 5) and an increase of 30% in patient 3 (Table 3).
In 4 cases, two liver metastases were available. In patient 5, a liver metastasis was synchronous to the PT and showed the same RAS mutant allele content as compared to PT. In patients 1 and 4, the two lesions resected after systemic therapy showed similar trends, with a reduction greater than 50% of RAS mutant allele content in both cases, as compared to the RAS mutant allele content in PT. In patient 2, the two liver metastases were not resected at the same time. The first one, removed immediately after systemic therapy, showed a more than 50% reduction of the RAS mutant allele content. The second metastasis, resected 8 months after initial therapy, showed an increase of 400% of RAS mutant alleles compared to the first liver metastasis, and even an increase of about 30% as compared to the PT.
Concerning the other mutations included in the NGS panel, we observed the same trends of RAS alterations with exception of patient 3 (in whom a new alteration in PIK3CA was detected in the metastasis) and patient 6 (the PIK3CA mutation was not seen in the metastasis).
Group 2: patients treated with chemotherapy alone before resection of liver metastases
The comparison of RAS mutation frequencies between PT and liver metastases showed a reduction of at least 50% of RAS mutant allele content in only one patient (8.3%, patient 11, Table 3). All other cases showed RAS mutant allele frequencies nearly comparable between the PT and liver metastasis. Two or more liver metastases were available in 5 patients: in patient 9, one liver metastasis was synchronous to the PT; in patients 14, 16, 18 and 19 the metastases were resected with an interval of about 6–8 months and after second or third-line therapies not including bevacizumab. In all the cases, the RAS mutant allele content in metastases was similar to the PT, with a trend to increase in late resected metastases (Table 3).
As regards the other mutations included in the IOT panel, we observed the same trend of RAS alterations with few exceptions. Patient 11 did not experience a reduction of APC mutation percentage; patient 14 showed both APC and TP53 reduction and occurrence of SMAD4 frame-shift mutation in the first metastasis only; patient 16 had a higher percentage of APC mutant content and became wild-type in TP53 gene in the second metastasis; patient 18 presented a trend of decrease in TP53 mutation rates starting from the second metachronous metastasis (Table 3).
Group 3 (control group): patients with resected liver metastasis without any systemic therapy
As expected, all the samples of the control group (11/11; 100%) showed the same RAS mutant allele content in PT and synchronous liver metastasis (Table 3).
Concerning the other mutations included in the Cancer Hotspot Panel v2 (CHPv2), we confirmed the same trend of RAS alterations with few exceptions. Patient 9, who did not show PIK3CA mutations in the synchronous metastasis compared to the PT; patient 24, in whom GNAS mutation was not detected in the metastasis; patient 25, who showed a TP53 mutation increase; patient 28 who showed a decrease of PIK3CA mutant allele content in the metastasis (Table 3).
Real-time analysis of RAS
The real-time data obtained after analysis of PT and liver metastases tissue blocks were normalized with the t cells % content. Real-time results mirror those obtained by IOT. In particular, when a decrease of RAS mutations level was observed by IOT, real-time showed an increase of ΔCt comparing PT and the associated metastases. On the contrary, the ΔCt by real-time was similar in patients showing no difference in RAS mutation frequency. An example of increased ΔCt is reported in Figure 1 and an example of stability of ΔCt in Figure 2.
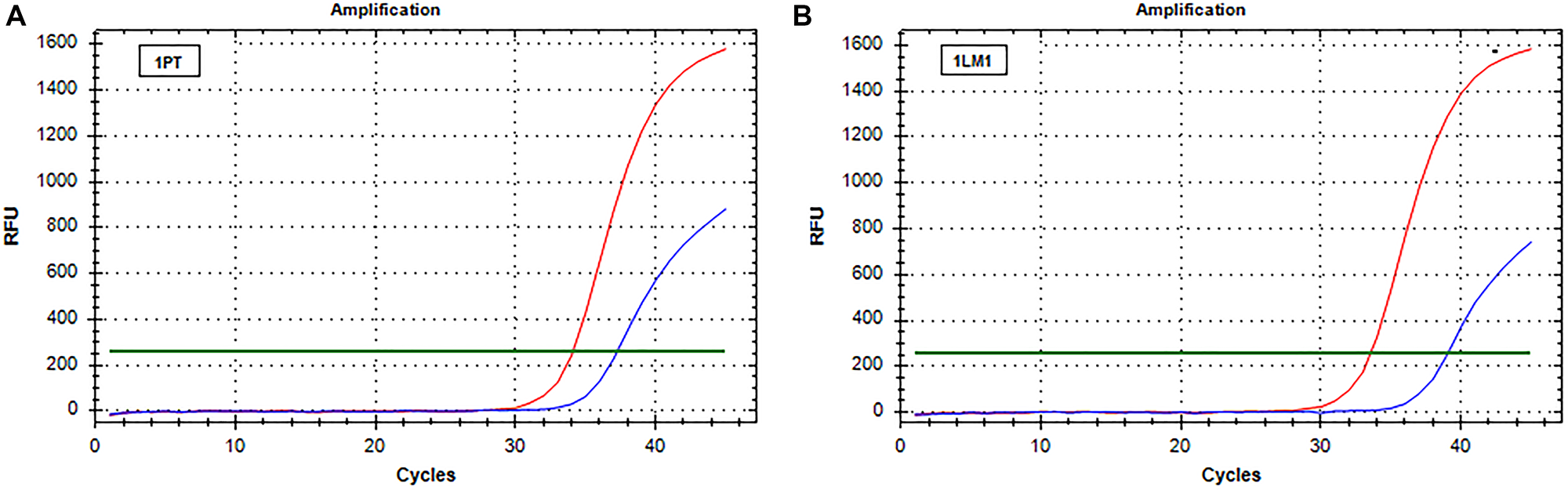
Figure 1: Real-time PCR (SensiScreen™) amplification curves of patient 1 (group 1). X-axis reports real-time PCR cycles and Y-axis reports relative fluorescence unit (RFU). In red is represented the amplification of the reference gene and in blue the amplification of the specific mutation (G12C). (A) Curves obtained from amplification of DNA extracted from sample 1PT; (B) Curves obtained from amplification of DNA extracted from sample 1LM1. Abbreviations: 1LM1, liver metastasis sample (patient 1); 1PT, primary tumor sample (patient 1); RFU, relative fluorescence unit.
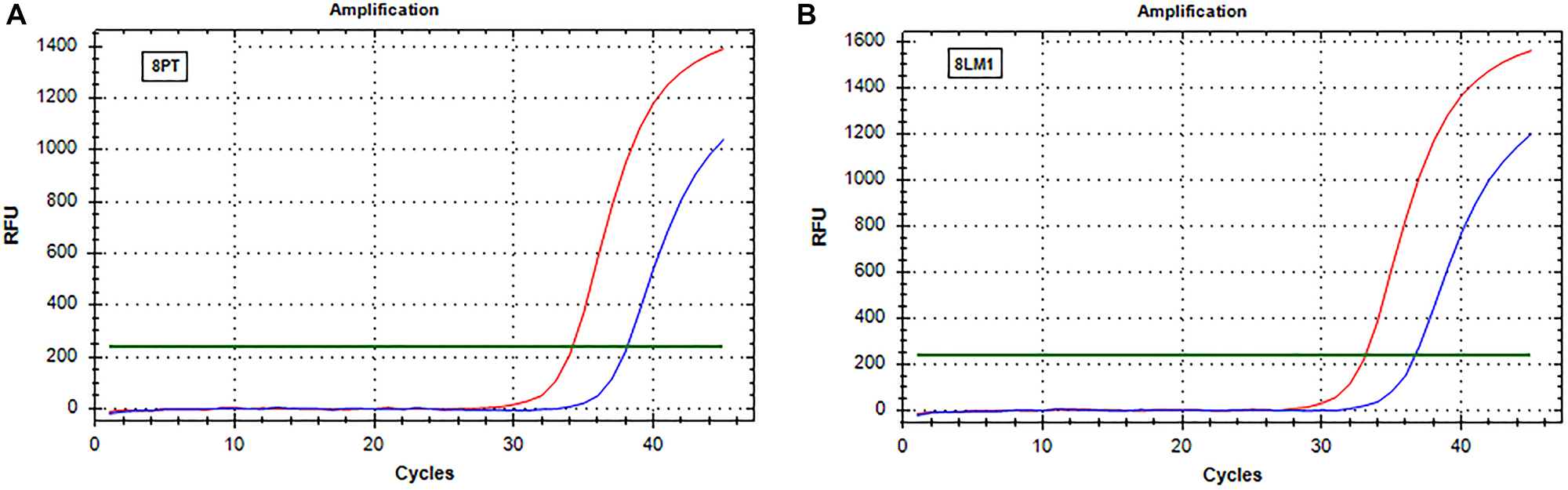
Figure 2: Real-time PCR (SensiScreen™) amplification curves of patient 8 (group 2). X-axis reports real-time PCR cycles and Y-axis reports relative fluorescence unit (RFU). In red is represented the amplification of the reference gene and in blue the amplification of the specific mutation (G13D). (A) Curves obtained from amplification of DNA extracted from sample 8PT; (B) Curves obtained from amplification of DNA extracted from sample 8LM1. Abbreviations: 8LM1, liver metastasis sample (patient 8); 8PT, primary tumor sample (patient 8); RFU, relative fluorescence unit.
Statistical analyses
The evaluation of RAS mutant allele content variations in Groups 1 and 2 showed a significant correlation (p ≤ 0.05) between bevacizumab administration and the decrease of more than 50% of RAS mutations by applying the two-tailed Fisher’s exact test (Table 4).
Table 4: Two-tailed Fisher exact test results
| RAS % (comparison between PT and LM) | ||
|---|---|---|
| Comparable | Decrease of at least 50% | |
| Chemotherapy plus bevacizumab (group 1) | 3 | 4 |
| Chemotherapy (group 2) | 11 | 1 |
| p = 0.038 | ||
DISCUSSION
In patients with mCRC, two preliminary studies reported that bevacizumab could revert RAS mutant status, assessed by liquid biopsies [12, 13].
In the CRICKET trial, twenty-seven RAS wild type mCRC patients were successfully treated for at least 6 months with cetuximab plus chemotherapy. At disease progression, patients underwent second line bevacizumab-containing regimens. In those who experienced a benefit from the second line for at least 4 months, an anti-EGFR rechallenge was proposed at progression. Four cases (14.3%) showed a partial response, with additional 15 cases (53.5%) achieving a disease control. The median progression-free survival (PFS) and median OS were 3.4 months (95% CI: 1.9–3.8 months) and 9.8 months (95% CI: 5.2–13.10 months), respectively [14]. The investigators evaluated RAS status in liquid biopsies collected at progression during bevacizumab and before the EGFR rechallenge, and found that all patients who exhibited a confirmed partial response were RAS wild type.
In another study, eleven RAS mutant mCRC patients were treated with first line bevacizumab-based regimens. Interestingly, the authors reported that at disease progression the RAS status at liquid biopsies reverted to a wild-type pattern in 6 patients (55.5%), and patients were successfully treated with cetuximab (PFS ranging from 6 to 12 months in those treated in second-line, PFS of 4 months in a patient treated in fourth line) [13, 15]. Taken together, these data suggest that bevacizumab is more effective against RAS mutant cells, possibly even leading to a conversion from RAS mutant to RAS wild-type status, at least when evaluated by liquid biopsy.
To date, liquid biopsy represents a promising source of genomic DNA that can be used for diagnostic purposes. The application of liquid biopsies is accepted in the management of patients with non-small cell lung cancer (NSCLC), but its role in CRC is still marginal. If on the one hand liquid biopsy may provide a dynamic characterization of the mutational status over time, the analysis of samples requires techniques characterized by a very low limit of detection (LOD), equal or less than 0.1% because the mutant alleles are extremely diluted in blood vessels, thus could be potentially classified as false wild type. The concordance between tumor tissue and liquid biopsy is only observed in highly vascularized tumors. For example, in NSCLC the concordance is up to 70%, while in CRC concordance reaches values above 90% [16–18].
In our study we aimed to assess whether exposure to bevacizumab could revert a RAS mutant tumor to RAS wild-type status, or at least induce a significant decrease of RAS mutant t cells at tissue level in patients with mCRC. To explore this concept, we identified three cohorts of previously untreated patients with RAS mutated mCRC, treated either with (Group 1) or without (Group 2) bevacizumab-based regimens before resection of liver metastases. The control arm (Group 3) consisted of patients with RAS mutant tumors, who underwent resection of liver metastases without any systemic preoperative therapy. Before the analyses of the whole cohort, in two cases selected on the basis of availability of multiple tissue blocks of both PT and paired liver metastases, we evaluated the RAS mutant allele content in different portions of the two lesions. After normalization with the t cell content provided by the pathologists, we found that RAS mutations were distributed homogeneously in the PT as well as in the liver metastasis. Thus, we can assume that the analysis of a single tissue block is representative of the totality of the malignancy and we can be confident in analysing only one block in the patients of the aforementioned three groups.
In the control arm (group 3) we observed 100% concordance of RAS mutant allele content between PT and liver metastases. This finding suggests that there is no clonal selection for RAS mutant cells in the natural history of mCRC patients on the one hand, and that cells with RAS mutations are not more aggressive than RAS wild-type cells on the other hand, confirming the absence of correlation between RAS mutation and worse prognosis [19]. Then we focused our attention to patients treated with chemotherapy alone before resection of liver metastases. Here we observed a similar pattern with respect to the control group. Indeed, only 1 out of 12 cases shows a drop of more than 50% of RAS mutant cell content, suggesting that chemotherapy alone is effective independently from the RAS mutational status. The situation is completely different in patients treated with a bevacizumab-containing regimen. In the majority of cases (4 out of 7 patients) we observed a dramatic decrease of RAS mutant allele content (at least 50%) in liver metastasis with respect to the PT resected before systemic therapy. Interestingly, results changed in the patient who underwent resection of additional liver metastases after a second line systemic therapy not including bevacizumab. This single case presented an increase of 400% of the RAS mutant allele content compared to RAS mutant allele observed in liver metastases removed at the end of a bevacizumab-containing regimen, even greater than the value observed in the PT. The difference in the decrease of the RAS mutant allele content between PT and liver metastasis after systemic therapy was statistically significant when Group 1 and Group 2 are compared (p = 0.038). In line with previously mentioned studies using liquid biopsies [13–15], our results overall confirm that bevacizumab-based systemic therapy seems to reduce RAS mutant cells burden, even at the tissue level. In contrast with these studies, in our cohort we did not observe a complete regression of RAS mutant cells in liver metastases. The differences may be explained by methodological issues: indeed the assays used for molecular characterization of plasma samples, including also the IOT panels for plasma analyses, are usually not sensitive enough to identify very low copy numbers of the mutant allele. It is therefore not surprising that our results show a significant decrease but not complete disappearance of RAS mutant cells, especially taking into account that bevacizumab is not a RAS-targeted agent. How bevacizumab could exerts its effect on RAS status changes still needs to be elucidated, but inflammation and neo-angiogenesis can be taken in consideration, as suggested in studies on transgenic murine models [20–22].
The precise estimation of the RAS mutant allele content in tumor tissues is challenging. In particular, the t cell content represents a crucial factor for such analysis because the RAS mutant allele content is provided by pathologists, based on a subjective histological estimation. The comparison of tissue samples from the same patient and the evaluation performed by different skilled pathologists should minimize inter-observer variability and interpretation bias. In our study two pathologists with experience in gastrointestinal tract cancer examined cases and provided an estimated t cell content in a blinded manner. As the concordance between pathologists was 100%, the subsequent use of a semi-quantitative method for the molecular characterization (IOT CHPv2 panel) should provide a percentage of the mutant allele truly reflecting the real content. To substantiate our evaluation, we also applied another semi-quantitative method (real-time PCR-based) for the analysis of RAS mutations, confirming the results obtained by the NGS analysis. The overlapping of NGS results and real-time PCR data observed in our work has been also described in literature. Indeed, a relevant number of papers, concerning KRAS evaluation in CRC and NSCLC samples, reports a significant concordance (from 93 to 100%) between these two methodologies [23–25].
The main limitation of our study is represented by the limited number of patients. Thus, our findings deserve to be confirmed in larger and independent cohorts.
In conclusion, this retrospective observational study, strongly suggests the existence of a link between bevacizumab exposure and RAS status changes in mCRC patients. In addition to other reports using liquid biopsies, our findings on tissue samples corroborate the hypothesis that bevacizumab could revert RAS mutant mCRC to a wild-type pattern, conceptually opening to the possibility to treat with anti-EGFR MoAbs mCRC patients otherwise excluded based on initial RAS mutated status.
Materials and Methods
Patients
All patients included in this retrospective study were treated in two institutions, the Oncology Institute of Southern Switzerland (Switzerland) and the Hospital of Legnano (Italy), from 2011 to 2019. Inclusion criteria were: age ≥ 18 years, histology proven mCRC, presence of mutations in RAS genes in primary tumor or paired resected liver metastases, availability of tumor tissue before and after systemic therapy. Exclusion criteria were: insufficient amount of t cells (< 10%), inadequate material for molecular purposes on primary tumor and paired liver metastasis.
Our study population cohort consisted of three groups: 1) patients deemed potentially suitable for secondary liver resection after preoperative chemotherapy plus bevacizumab; 2) patients deemed potentially suitable for secondary liver resection after chemotherapy alone; 3) the control group, namely patients with liver metastases deemed resectable without any systemic therapy.
As most of these patients were deceased before entry in this retrospective study, the permission to perform the molecular analyses was given by the Regional Ethical Committee (No.: 2019-02224). For those who were alive at study entry a signed informed consent was requested. The study was conducted in accordance with the Declaration of Helsinki.
Molecular analyses
All tumor tissue specimens were evaluated by two senior pathologists of the Institute of Pathology in Locarno (Switzerland). The pathologists selected the appropriate tumor area on hematoxylin and eosin (HE) stained slides, and independently established the t cells %. Genomic DNA was extracted from six 7 μm-thick serial sections of each selected FFPE tissue block from the primary tumor and paired liver metastasis using the QIAamp DNA FFPE Tissue kit (Qiagen, USA), according to the manufacturer’s instructions. DNA was analyzed using an NGS approach on S5 IOT platform, by applying the Ion AmpliSeq™ Cancer Hotspot Panel v2 (CHPv2) (ThermoFisher Scientific, USA) [26, 27]. The CHPv2 panel generated semi-quantitative and qualitative data on the mutational status of RAS and 49 additional genes, encompassing the most relevant and frequently mutated genes in CRC (i.e., APC, TP53, PIK3CA, BRAF and PTEN). NGS data give an idea of how many cells of the sample are mutated in each gene, through the evaluation of the Variant Allele Frequency (VAF) of each mutation. The quantitative level of RAS mutations in both primary tissue and liver metastasis was obtained by normalizing VAF of RAS mutation obtained from the NGS experiment with the t cells % calculated by the pathologists. The presence of RAS mutations was also confirmed quantitatively using an independent methodology based on a real-time PCR amplification (SensiScreen, PentaBase ApS, Denmark) [28]. Real-time data were evaluated comparing ΔCt values of PT and paired liver metastasis. ΔCt was calculated as the difference between the threshold cycle (Ct) of reference gene and the Ct of the specific RAS mutation previously detected by IOT.
Statistical analyses
The loss of RAS mutations was considered meaningful in presence of a difference greater than 50% between the primary tumor and paired liver metastases. The two-tailed Fisher’s exact test was used to calculate p-values and the level of significance was set at p < 0.05.
Ethics approval and consent to participate
This work has been conducted in compliance with the current version of the Declaration of Helsinki as well as all national legal and regulatory requirements. Data and samples have been collected and analyzed for the study purpose only after the required authorizations from the competent Ethics Committees were obtained (No.: 2019-02224). As most patients has deceased before study entry, the permission to perform the molecular analyses was given by the Regional Ethics Committee. Conversely, for those who were alive at study entry a signed informed consent was requested. To partecipate in this study a consent for the publication of the anonymized data has also been obtained.
All the data reported in this paper have been anonymized and they have been obtained following the national legal and regulatory requirements.
Abbreviations
EGFR: epidermal growth factor receptor; FFPE: formalin fixed paraffin embedded; HE: hematoxylin and eosin staining; MoAbs: monoclonal antibodies; mCRC: metastatic colorectal cancer; NGS: next-generation sequencing; NSCLC: non-small cell lung cancer; ORR: overall response rate; OS: overall survival; PFS: progression-free survival; PT: primary tumor; t cells: tumor cells; VEGF: vascular endothelial growth factor.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
FUNDING
This study was supported by the Advisory Board of Ente Ospedaliero Cantonale (ABREOC) and by scientific funds of the Institute of Pathology of Locarno (Switzerland). (Project ID: IOSI-ICP-001).
References
1. Heinemann V, von Weikersthal LF, Decker T, Kiani A, Vehling-Kaiser U, Al-Batran SE, Heintges T, Lerchenmüller C, Kahl C, Seipelt G, Kullmann F, Stauch M, Scheithauer W, et al. FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): a randomised, open-label, phase 3 trial. Lancet Oncol. 2014; 15:1065–75. https://doi.org/10.1016/S1470-2045(14)70330-4. [PubMed].
2. Venook AP, Niedzwiecki D, Lenz HJ, Innocenti F, Fruth B, Meyerhardt JA, Schrag D, Greene C, O'Neil BH, Atkins JN, Berry S, Polite BN, O'Reilly EM, et al. Effect of First-Line Chemotherapy Combined With Cetuximab or Bevacizumab on Overall Survival in Patients With KRAS Wild-Type Advanced or Metastatic Colorectal Cancer: A Randomized Clinical Trial. JAMA. 2017; 317:2392–401. https://doi.org/10.1001/jama.2017.7105. [PubMed].
3. Cremolini C, Loupakis F, Antoniotti C, Lupi C, Sensi E, Lonardi S, Mezi S, Tomasello G, Ronzoni M, Zaniboni A, Tonini G, Carlomagno C, Allegrini G, et al. FOLFOXIRI plus bevacizumab versus FOLFIRI plus bevacizumab as first-line treatment of patients with metastatic colorectal cancer: updated overall survival and molecular subgroup analyses of the open-label, phase 3 TRIBE study. Lancet Oncol. 2015; 16:1306–15. https://doi.org/10.1016/S1470-2045(15)00122-9. [PubMed].
4. Xie YH, Chen YX, Fang JY. Comprehensive review of targeted therapy for colorectal cancer. Signal Transduct Target Ther. 2020; 5:22. https://doi.org/10.1038/s41392-020-0116-z. [PubMed].
5. Douillard JY, Oliner KS, Siena S, Tabernero J, Burkes R, Barugel M, Humblet Y, Bodoky G, Cunningham D, Jassem J, Rivera F, Kocákova I, Ruff P, et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med. 2013; 369:1023–34. https://doi.org/10.1056/NEJMoa1305275. [PubMed].
6. Douillard JY, Rong A, Sidhu R. RAS mutations in colorectal cancer. N Engl J Med. 2013; 369:2159–60. https://doi.org/10.1056/NEJMc1312697. [PubMed].
7. Sorich MJ, Wiese MD, Rowland A, Kichenadasse G, McKinnon RA, Karapetis CS. Extended RAS mutations and anti-EGFR monoclonal antibody survival benefit in metastatic colorectal cancer: a meta-analysis of randomized, controlled trials. Ann Oncol. 2015; 26:13–21. https://doi.org/10.1093/annonc/mdu378. [PubMed].
8. Peeters M, Oliner KS, Price TJ, Cervantes A, Sobrero AF, Ducreux M, Hotko Y, André T, Chan E, Lordick F, Punt CJ, Strickland AH, Wilson G, et al. Analysis of KRAS/NRAS Mutations in a Phase III Study of Panitumumab with FOLFIRI Compared with FOLFIRI Alone as Second-line Treatment for Metastatic Colorectal Cancer. Clin Cancer Res. 2015; 21:5469–79. https://doi.org/10.1158/1078-0432.CCR-15-0526. [PubMed].
9. Martini G, Ciardiello D, Vitiello PP, Napolitano S, Cardone C, Cuomo A, Troiani T, Ciardiello F, Martinelli E. Resistance to anti-epidermal growth factor receptor in metastatic colorectal cancer: What does still need to be addressed? Cancer Treat Rev. 2020; 86:102023. https://doi.org/10.1016/j.ctrv.2020.102023. [PubMed].
10. Shen H, Yang J, Huang Q, Jiang MJ, Tan YN, Fu JF, Zhu LZ, Fang XF, Yuan Y. Different treatment strategies and molecular features between right-sided and left-sided colon cancers. World J Gastroenterol. 2015; 21:6470–78. https://doi.org/10.3748/wjg.v21.i21.6470. [PubMed].
11. Ulivi P, Scarpi E, Chiadini E, Marisi G, Valgiusti M, Capelli L, Casadei Gardini A, Monti M, Ruscelli S, Frassineti GL, Calistri D, Amadori D, Passardi A. Right- vs. Left-Sided Metastatic Colorectal Cancer: Differences in Tumor Biology and Bevacizumab Efficacy. Int J Mol Sci. 2017; 18:1240. https://doi.org/10.3390/ijms18061240. [PubMed].
12. Cremolini C, Rossini D, Dell'Aquila E, Lonardi S, Conca E, Del Re M, Busico A, Pietrantonio F, Danesi R, Aprile G, Tamburini E, Barone C, Masi G, et al. Rechallenge for Patients With RAS and BRAF Wild-Type Metastatic Colorectal Cancer With Acquired Resistance to First-line Cetuximab and Irinotecan: A Phase 2 Single-Arm Clinical Trial. JAMA Oncol. 2019; 5:343–50. https://doi.org/10.1001/jamaoncol.2018.5080. [PubMed].
13. Gazzaniga P, Raimondi C, Nicolazzo C, Gradilone A, Cortesi E. ctDNA might expand therapeutic options for second line treatment of KRAS mutant mCRC. Ann Oncol. 2017; 28:V568.
14. Rossini D, Cremolini C, Conca E, Del Re M, Busico A, Pietrantonio F, Bergamo F, Danesi R, Cardellino G, Tamburini E, Dell’Aquila E, Strippoli A, Masi G, et al. O-007Liquid biopsy allows predicting benefit from rechallenge with cetuximab(cet)+irinotecan(iri) in RAS/BRAF wild-type mCRC patients(pts) with resistance to 1st-line cet+iri: final results and translational analyses of the CRICKET study by GONO. Ann Oncol. 2018; 29:v102. https://doi.org/10.1093/annonc/mdy149.006.
15. Raimondi C, Nicolazzo C, Belardinilli F, Loreni F, Gradilone A, Mahdavian Y, Gelibter A, Giannini G, Cortesi E, Gazzaniga P. Transient Disappearance of RAS Mutant Clones in Plasma: A Counterintuitive Clinical Use of EGFR Inhibitors in RAS Mutant Metastatic Colorectal Cancer. Cancers (Basel). 2019; 11:42. https://doi.org/10.3390/cancers11010042. [PubMed].
16. Pi C, Zhang MF, Peng XX, Zhang YC, Xu CR, Zhou Q. Liquid biopsy in non-small cell lung cancer: a key role in the future of personalized medicine? Expert Rev Mol Diagn. 2017; 17:1089–96. https://doi.org/10.1080/14737159.2017.1395701. [PubMed].
17. García-Foncillas J, Tabernero J, Élez E, Aranda E, Benavides M, Camps C, Jantus-Lewintre E, López R, Muinelo-Romay L, Montagut C, Antón A, López G, Díaz-Rubio E, et al. Prospective multicenter real-world RAS mutation comparison between OncoBEAM-based liquid biopsy and tissue analysis in metastatic colorectal cancer. Br J Cancer. 2018; 119:1464–70. https://doi.org/10.1038/s41416-018-0293-5. [PubMed].
18. Wu Z, Yang Z, Dai Y, Zhu Q, Chen LA. Update on liquid biopsy in clinical management of non-small cell lung cancer. Onco Targets Ther. 2019; 12:5097–109. https://doi.org/10.2147/OTT.S203070. [PubMed].
19. Oh HH, Joo YE. Novel biomarkers for the diagnosis and prognosis of colorectal cancer. Intest Res. 2020; 18:168–83. https://doi.org/10.5217/ir.2019.00080. [PubMed].
20. Kranenburg O, Gebbink MF, Voest EE. Stimulation of angiogenesis by Ras proteins. Biochim Biophys Acta. 2004; 1654:23–37. https://doi.org/10.1016/j.bbcan.2003.09.004. [PubMed].
21. Tang Y, Kim M, Carrasco D, Kung AL, Chin L, Weissleder R. In vivo assessment of RAS-dependent maintenance of tumor angiogenesis by real-time magnetic resonance imaging. Cancer Res. 2005; 65:8324–30. https://doi.org/10.1158/0008-5472.CAN-05-0027. [PubMed].
22. Figueras A, Arbos MA, Quiles MT, Viñals F, Germà JR, Capellà G. The impact of KRAS mutations on VEGF-A production and tumour vascular network. BMC Cancer. 2013; 13:125. https://doi.org/10.1186/1471-2407-13-125. [PubMed].
23. Tuononen K, Mäki-Nevala S, Sarhadi VK, Wirtanen A, Rönty M, Salmenkivi K, Andrews JM, Telaranta-Keerie AI, Hannula S, Lagström S, Ellonen P, Knuuttila A, Knuutila S. Comparison of targeted next-generation sequencing (NGS) and real-time PCR in the detection of EGFR, KRAS, and BRAF mutations on formalin-fixed, paraffin-embedded tumor material of non-small cell lung carcinoma-superiority of NGS. Genes Chromosomes Cancer. 2013; 52:503–11. https://doi.org/10.1002/gcc.22047. [PubMed].
24. Xu X, Yang Y, Li H, Chen Z, Jiang G, Fei K. Assessment of the clinical application of detecting EGFR, KRAS, PIK3CA and BRAF mutations in patients with non-small cell lung cancer using next-generation sequencing. Scand J Clin Lab Invest. 2016; 76:386–92. https://doi.org/10.1080/00365513.2016.1183813. [PubMed].
25. Gao J, Wu H, Wang L, Zhang H, Duan H, Lu J, Liang Z. Validation of targeted next-generation sequencing for RAS mutation detection in FFPE colorectal cancer tissues: comparison with Sanger sequencing and ARMS-Scorpion real-time PCR. BMJ Open. 2016; 6:e009532. https://doi.org/10.1136/bmjopen-2015-009532. [PubMed].
26. Zhang L, Chen L, Sah S, Latham GJ, Patel R, Song Q, Koeppen H, Tam R, Schleifman E, Mashhedi H, Chalasani S, Fu L, Sumiyoshi T, et al. Profiling cancer gene mutations in clinical formalin-fixed, paraffin-embedded colorectal tumor specimens using targeted next-generation sequencing. Oncologist. 2014; 19:336–43. https://doi.org/10.1634/theoncologist.2013-0180. [PubMed].
27. Butler KS, Young MY, Li Z, Elespuru RK, Wood SC. Performance characteristics of the AmpliSeq Cancer Hotspot panel v2 in combination with the Ion Torrent Next Generation Sequencing Personal Genome Machine. Regul Toxicol Pharmacol. 2016; 74:178–86. https://doi.org/10.1016/j.yrtph.2015.09.011. [PubMed].
28. Riva A, BØrgesen M, Guldmann-Christensen M, Hauge Kyneb M, Voogd K, Andersen C, Epistolio S, Merlo E, Yding Wolff T, Hamilton-Dutoit S, Lorenzen J, Christensen UB, Frattini M. SensiScreen®KRAS exon 2-sensitive simplex and multiplex real-time PCR-based assays for detection of KRAS exon 2 mutations. PLoS One. 2017; 12:e0178027. https://doi.org/10.1371/journal.pone.0178027. [PubMed].

