EDITORIAL COMMENT
Lung cancer is the leading cause of cancer death worldwide [1]. Systemic treatment for lung cancer is a continuously evolving landscape with several options, such as cytotoxic chemotherapy, molecularly targeted therapy and immunotherapy [2]. Imaging is critical for the assessment of tumor response [3]. However, standard imaging metrics are limited when assessing response and progression with emerging targeted and immune therapies. A change in the size of a subset of target lesions, which remains the gold standard for assessing response under treatment [3, 4], does not or cannot fully capture the complexity of lung cancer behavior, including the primary tumor and metastases, driven by its high cellular heterogeneity [5]. In the era of precision medicine, the challenge entails choosing the right treatment for the right patient at the right time. The core concept is to personalize medical care and optimize cost-effectiveness. Recent updates of the Response Evaluation Criteria in Solid Tumors (RECIST) to iRECIST [6], irRECIST [7], and many others [8–10] have proposed novel patterns of tumor response and progression to immunotherapy. These updates begin to address part of the challenge, however, the paradigm of response assessment is shifting toward new imaging methods.
Radiomics -the use of imaging features from radiographic medical images transform images into quantitative data- has emerged a decade ago [11]. This high-throughput feature extraction procedure significantly increases the radiologist's analysis capability. However, despite the growing availability of dedicated software, numerous methodological challenges have to be faced with validating the procedure, which explains the slow implementation of such a revolutionary approach in clinical practice: the difficulty to collect massive structured imaging data from suitable target populations; the implementation and harmonization of operational multilevel imaging pipelines including imaging acquisition, tumor segmentation, feature extraction, and finally the feature selection process, adapted to a particular output task which requires AI-based validation procedures [12].
The recent study entitled "Identification of Non-Small Cell Lung Cancer Sensitive to Systemic Cancer Therapies Using Radiomics" offers a relevant clinical illustration of the current predictive capabilities of radiomics in NSCLC tumors to several systemic treatments [13]. Using standard-of-care CT-scan images (baseline and first-treatment assessment) of NSCLC patients collected from clinical trials, machine-learning algorithms were trained to predict NSCLC sensitivities to the following treatments: nivolumab, docetaxel, and gefitinib. To this end, this study combined Radiomic features derived from the largest measurable lung lesion of each patient. These radiomics signatures achieved areas under the receiver operating characteristic curve (AUC) of 0.77, 0.67 and 0.82, for nivolumab, docetaxel, and gefitinib, respectively. The radiomic features used in these signatures characterized 1) tumor burden, 2) tumor spatial heterogeneity, and 3) density change around tumor-parenchyma boundaries. Interestingly, the radiomic signatures in nivolumab (immunotherapy) and gefitinib (EGFR-targeted) arms were dominated by intra-tumor spatial heterogeneity and tumor-parenchyma density transition descriptors. In contrast, volume descriptors were more relevant in the docetaxel (chemotherapy) cohort. This proof-of-concept study paves the way for future use of radiomics and AI in tumor response assessment. Of note, the ultimate goal is to find a signature that could generalize to all treatment types.
In perspective, a fundamental breakthrough would be to integrate multiparametric imaging data into the radiomic framework (Figure 1). Multiparametric imaging offers unique opportunities to extend the image-based tumor analyses to more holistic characteristics [14], including both non-parametric (qualitative) and parametric (quantitative) imaging data [15, 16]. A multiparametric radiomics signature could be a suitable combination of features extracted from non-fused or fused multimodal data [17, 18] or higher-order composite vector representation of the tumor temporal and spatial changes between multidimensional features [19, 20]. This is a new era to investigate the combination of imaging features extracted through various methods to guide clinical care. AI techniques would bring a remarkable transformation and redefine medical image interpretation from an inherently subjective human-based interpretation to an objective computer-based pattern-recognition algorithm. Successful implementation of these technologies would allow for faster and more accurate treatment efficacy predictions, which will probably significantly impact clinical outcomes and decision-making.
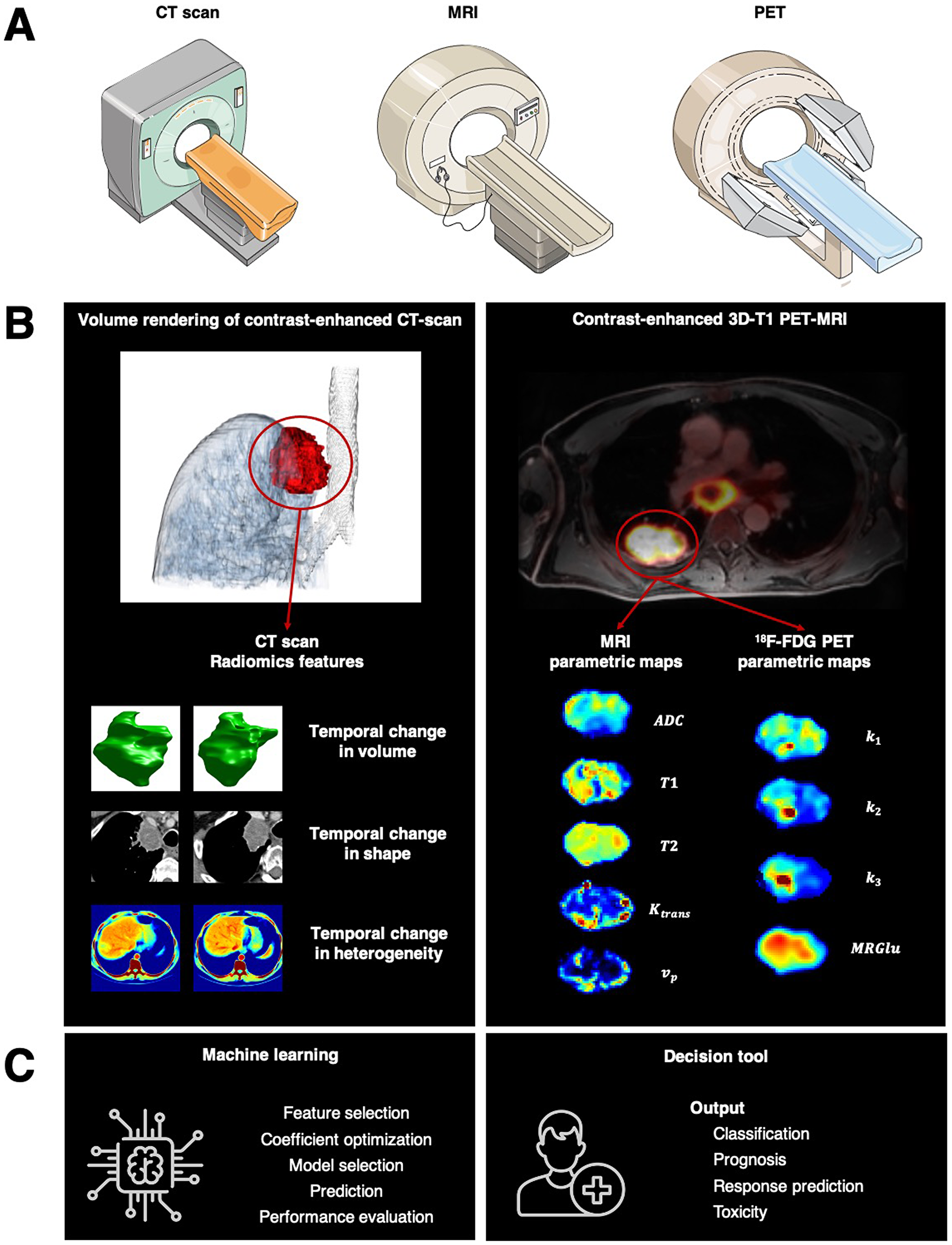
Figure 1: Toward radiomics for assessment of response to systemic therapies in lung cancer. (A) Multiple imaging modalities can characterize tumor imaging phenotypes such as CT scan, MRI, and PET. Multiparametric imaging offers unique opportunities to extend the image-based tumor analyses to more holistic characteristics. (B) left. In “Identification of Non-Small Cell Lung Cancer Sensitive to Systemic Cancer Therapies Using Radiomics,” the authors demonstrated that change over serial radiographic measurements in radiomics features deciphering tumor volume, invasion of tumor boundaries, or spatial tumor heterogeneity predicted tumor sensitivity to treatment, offering an approach that could enhance clinical decision-making to continue systemic therapies and forecast overall survival. (B) right. A fundamental breakthrough would be integrating multiparametric imaging data into the radiomic framework, such as imaging features extracted from functional MRI and 18F-FDG PET. (C) Machine-learning approaches can unravel among these imaging features, new imaging biomarkers predicting tumor sensitivity to treatment.
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
References
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424. https://doi.org/10.3322/caac.21492. [PubMed].
2. Yuan M, Huang LL, Chen JH, Wu J, Xu Q. The emerging treatment landscape of targeted therapy in non-small-cell lung cancer. Signal Transduct Target Ther. 2019; 4:61. https://doi.org/10.1038/s41392-019-0099-9. [PubMed].
3. Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, Dancey J, Arbuck S, Gwyther S, Mooney M, Rubinstein L, Shankar L, Dodd L, et al. New response evaluation criteria in solid tumors: revised RECIST guideline (version 1.1). Eur J Cancer. 2009; 45:228–247. https://doi.org/10.1016/j.ejca.2008.10.026. [PubMed].
4. Therasse P, Arbuck SG, Eisenhauer EA, Wanders J, Kaplan RS, Rubinstein L, Verweij J, Van Glabbeke M, van Oosterom AT, Christian MC, Gwyther SG. New guidelines to evaluate the response to treatment in solid tumors. European Organization for Research and Treatment of Cancer, National Cancer Institute of the United States, National Cancer Institute of Canada. J Natl Cancer Inst. 2000; 92:205–216. https://doi.org/10.1093/jnci/92.3.205. [PubMed].
5. Chen Z, Fillmore CM, Hammerman PS, Kim CF, Wong KK. Non-small-cell lung cancers: a heterogeneous set of diseases. Nat Rev Cancer. 2015; 15:247. https://doi.org/10.1038/nrc3775. [PubMed].
6. Seymour L, Bogaerts J, Perrone A, Ford R, Schwartz LH, Mandrekar S, Lin NU, Litière S, Dancey J, Chen A, Hodi FS, Therasse P, Hoekstra OS, et al. iRECIST: guidelines for response criteria for use in trials testing immunotherapeutics. Lancet Oncol. 2017; 18:e143–e152. https://doi.org/10.1016/S1470-2045(17)30074-8. [PubMed].
7. Le Lay J, Jarraya H, Lebellec L, Penel N. irRECIST and iRECIST: the devil is in the details. Ann Oncol. 2017; 28:1676–1678. https://doi.org/10.1093/annonc/mdx168. [PubMed].
8. Cheson BD, Ansell S, Schwartz L, Gordon LI, Advani R, Jacene HA, Hoos A, Barrington SF, Armand P. Refinement of the Lugano Classification lymphoma response criteria in the era of immunomodulatory therapy. Blood. 2016; 128:2489–2496. https://doi.org/10.1182/blood-2016-05-718528. [PubMed].
9. Okada H, Weller M, Huang R, Finocchiaro G, Gilbert MR, Wick W, Ellingson BM, Hashimoto N, Pollack IF, Brandes AA, Franceschi E, Herold-Mende C, Nayak L, et al. Immunotherapy response assessment in neuro-oncology: a report of the RANO working group. Lancet Oncol. 2015; 16:e534–e542. https://doi.org/10.1016/S1470-2045(15)00088-1. [PubMed].
10. Goldmacher GV, Khilnani AD, Andtbacka RHI, Luke JJ, Hodi FS, Marabelle A, Harrington K, Perrone A, Tse A, Madoff DC, Schwartz LH. Response Criteria for Intratumoral Immunotherapy in Solid Tumors: itRECIST. J Clin Oncol. 2020; 38:2667–2676. https://doi.org/10.1200/JCO.19.02985. [PubMed].
11. Lambin P, Leijenaar RTH, Deist TM, Peerlings J, de Jong EEC, van Timmeren J, Sanduleanu S, Larue RTHM, Even AJG, Jochems A, van Wijk Y, Woodruff H, van Soest J, et al. Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017; 14:749–762. https://doi.org/10.1038/nrclinonc.2017.141. [PubMed].
12. Vallière M, Zwanenburg A, Badic B, Cheze Le Rest C, Visvikis D, Hatt M. Responsible Radiomics Research for Faster Clincial Translation. J Nucl Med. 2018; 59:189–193. https://doi.org/10.2967/jnumed.117.200501. [PubMed].
13. Dercle L, Fronheiser M, Lu L, Du S, Hayes W, Leung DK, Roy A, Wilkerson J, Guo P, Fojo AT, Schwartz LH, Zhao B. Identification of Non-Small Cell Lung Cancer Sensitive to Systemic Cancer Therapies Using Radiomics. Clin Cancer Res. 2020; 26:2151–2162. https://doi.org/10.1158/1078-0432.CCR-19-2942. [PubMed].
14. Padhani AR, Miles KA. Multiparametric Imaging of Tumor Response to Therapy. Radiology. 2010; 256:348–64. https://doi.org/10.1148/radiol.10091760. [PubMed].
15. Peerlings J, Woodruff HC, Winfield JM, Ibrahim A, Van Beers BE, Heerschap A, Jackson A, Wildberger JE, Mottaghy FM, DeSouza NM, Lambin P. Stability of radiomics features in apparent diffusion coefficient maps from a multi-centre test-retest trial. Sci Rep. 2019; 9:4800. https://doi.org/10.1038/s41598-019-41344-5. [PubMed].
16. Elshafeey N, Kotrotsou A, Hassan A, Elshafei N, Hassan I, Ahmed S, Abrol S, Agarwal A, El Salek K, Bergamaschi S, Acharya J. Multicenter study demonstrates radiomic features derived from magnetic resonance perfusion images identify pseudoprogression in glioblastoma. Nat Commun. 2019; 10:3170. https://doi.org/10.1038/s41467-019-11007-0. [PubMed].
17. Vallière M, Freeman CR, Skamene SR, El Naqa I. A radiomic model from joint FDG-PET and MRI texture features for the prediction of lung metastases in soft-tissue sarcomas of the extremities. Phys Med Biol. 2015; 60:5471–96. https://doi.org/10.1088/0031-9155/60/14/5471. [PubMed].
18. Lucia F, Visvikis D, Desseroit MC, Miranda O, Malhaire JP, Robin P, Pradier O, Hatt M, Schick U. Prediction of outcome using pretreatment 18F-FDG PET/CT and MRI radiomics in locally advanced cervical cancer treated with chemoradiotherapy. Eur J Nucl Med Mol Imaging. 2018; 45:768–86. https://doi.org/10.1007/s00259-017-3898-7. [PubMed].
19. Parekh VS, Jacobs MA. Deep learning and radiomics in precision medicine. Expert Rev Precis Med Drug Dev. 2019; 4:59–72. https://doi.org/10.1080/23808993.2019.1585805. [PubMed].
20. Parekh VS, Jacobs MA. Multiparametric radiomics methods for breast cancer tissue characterization using radiological imaging. Breast Cancer Res Treat. 2020; 180:407–421. https://doi.org/10.1007/s10549-020-05533-5. [PubMed].

