Introduction
Small cell lung cancer (SCLC) accounts for 10–15% of lung cancer, and its prognosis has remained relatively dismal for years [1]. Most patients have metastatic spread at the time of diagnosis [2]. Currently, conventional platinum-based chemotherapy regimens with or without radiation remain the standard first-line treatment for SCLC. Although atezolizumab was approved for use in combination with carboplatin and etoposide as a first-line treatment for adult patients with extensive-stage SCLC, the median overall survival period, compared with that for chemotherapy alone, was only prolonged for a few months [3]. On the other hand, the role of surgery has been limited to rare (less than 5% of patients) for early-stage disease [4]. Although SCLC is more responsive to initial cytotoxic chemotherapy than non-small cell lung cancer, most patients relapse with a relatively resistant disease.
Genome-wide sequencing studies of SCLC have failed to identify targetable driver mutations such as EGFR, ALK, ROS1, and BRAF that are frequently observed in lung adenocarcinoma. Recurrent mutations of SCLC include the loss of the tumor suppressors TP53 and RB1, inactivating mutations in NOTCH family genes, and the amplification of MYC family genes, all of which are difficult to target [5]. The loss of PTEN, activating PI3K mutations, and aurora kinase activation have been reported as potential therapeutic targets [6]. There are ongoing trials for small molecule inhibitors of poly-ADP-ribose polymerase (PARP) [7–9] and an enhancer of zeste homolog 2 (EZH2), which regulate the DNA damage response and chromatin modifications, respectively [10]. A recent study proposed a new model of SCLC subtypes defined by the differential expressions of four key transcription regulators, ASCL1, NeuroD1, YAP1, and POU2F3, which would help to accelerate therapeutic research leading to targeted approaches [11]. Novel therapeutic modalities for SCLC are long awaited.
Antibody-drug conjugates (ADCs) are an emerging technology that has already been implemented in clinical practice for some malignancies. An ADC is a monoclonal antibody conjugated with a cytotoxic drug via a chemical linker, enabling selective drug delivery by binding to specific cell surface proteins [12, 13]. Considering the high sensitivity of SCLC to chemotherapy, the selective delivery of a cytotoxic agent using ADC could be a novel treatment strategy for SCLC [14].
Five ADCs have been approved by the Food and Drug Administration: brentuximab vedotin for Hodgkin lymphoma [15], ado-trastuzumab emtansine for HER2-positive metastatic breast cancer [16, 17], inotuzumab ozogamicin for acute lymphoblastic leukemia [18], gemtuzumab ozogamicin for CD33-positive acute myeloid leukemia [19], and trastuzumab deruxtecan for unresectable or metastatic HER2-positive breast cancer patients who have received two or more prior anti-HER2-based regimens in a metastatic setting [20]. To date, ADCs targeting solid tumors other than metastatic breast cancer have not exhibited distinct clinical benefits [21–29]. In SCLC, DLL3, a cell surface Notch ligand that appear to be a direct downstream target of ASCL1 [30, 31], has been identified as a novel target for ADCs [32]. However, a phase III trial comparing rovalpituzumab tesirine with topotecan as a second-line therapy had to be halted because of a shorter overall survival period in the ADC arm [33]. Trop-2, a glycoprotein overexpressed in many epithelial cancers, has also been reported to be a candidate target of ADCs [34, 35]. Sacituzumab govitecan, a Trop-2-targeting ADC, showed a potential efficacy and was deemed safe in a phase I/II trial in SCLC patients [36]. A phase I/II study of a CD56-targeting ADC in combination with carboplatin and etoposide showed no improvement in efficacy over standard carboplatin and etoposide therapy in SCLC [37]. Recently, promiximab-duocarmycin, a new CD56-targeting ADC, was shown to demonstrate promising activity in a preclinical study of SCLC.
In this study, we aimed to identify novel molecular targets for ADCs in SCLC. The candidates were cell surface proteins overexpressed specifically in tumors with little or no expression in normal tissues. We searched for transmembrane proteins of SCLC using a computational-biological approach. We herein report that NRXN1-mediated ADC exhibited anti-tumor activity in vitro, and thus NRXN1 could be a novel target of ADCs for SCLC.
Results
mRNA expression profile of cell surface proteins in SCLC
We analyzed the expressions of cell surface proteins using microarray data available in the Cancer Cell Line Encyclopedia using an unsupervised clustering analysis. Among the 565 genes coding membrane proteins, 31 genes showed an increased compensated fluorescence signal by three times or more, on average. The National Center for Biotechnology Information (NCBI) RNA sequencing data was used to select genes with little or no expression in normal tissues. Interestingly, SCLC cell lines were divided into two subgroups with and without overexpression of the 31 genes. Since Rudin et al. reported a new model of SCLC subtypes based on the expressions of four key transcription regulators (ASCL1, NeuroD1, YAP1, and POU2F3) [11], we compared the results of our clustering analysis with their subtypes (Supplementary Figure 1). The NRXN1-positive SCLC cell lines generally overlapped with ASCL1-high or NEUROD1-high subtypes.
NRXN1 expression in cell lines, surgical specimens, and human normal tissues
Among the 31 membrane proteins, we focused on NRXN1. NRXN1 expression was analyzed using two SCLC cell lines (SHP77 and NCI-H526) and HEK293 cells. SHP77 had the highest mRNA expression, NCI-H526 displayed moderate expression, and HEK293 showed little expression (Figure 1A). Patient-derived cells (PDC) showed a moderate NRXN1 expression level that was slightly lower than that in the NCI-H526 cell line. Cell-surface NRXN1 protein expression was verified using flow cytometry (Figure 1B). The percentage of NRXN1-positive cells determined using flow cytometry is generally correlated with mRNA expression (Figure 1C). An analysis of the surgical specimens revealed a high NRXN1 expression in a subset of primary SCLCs (Figure 1D).
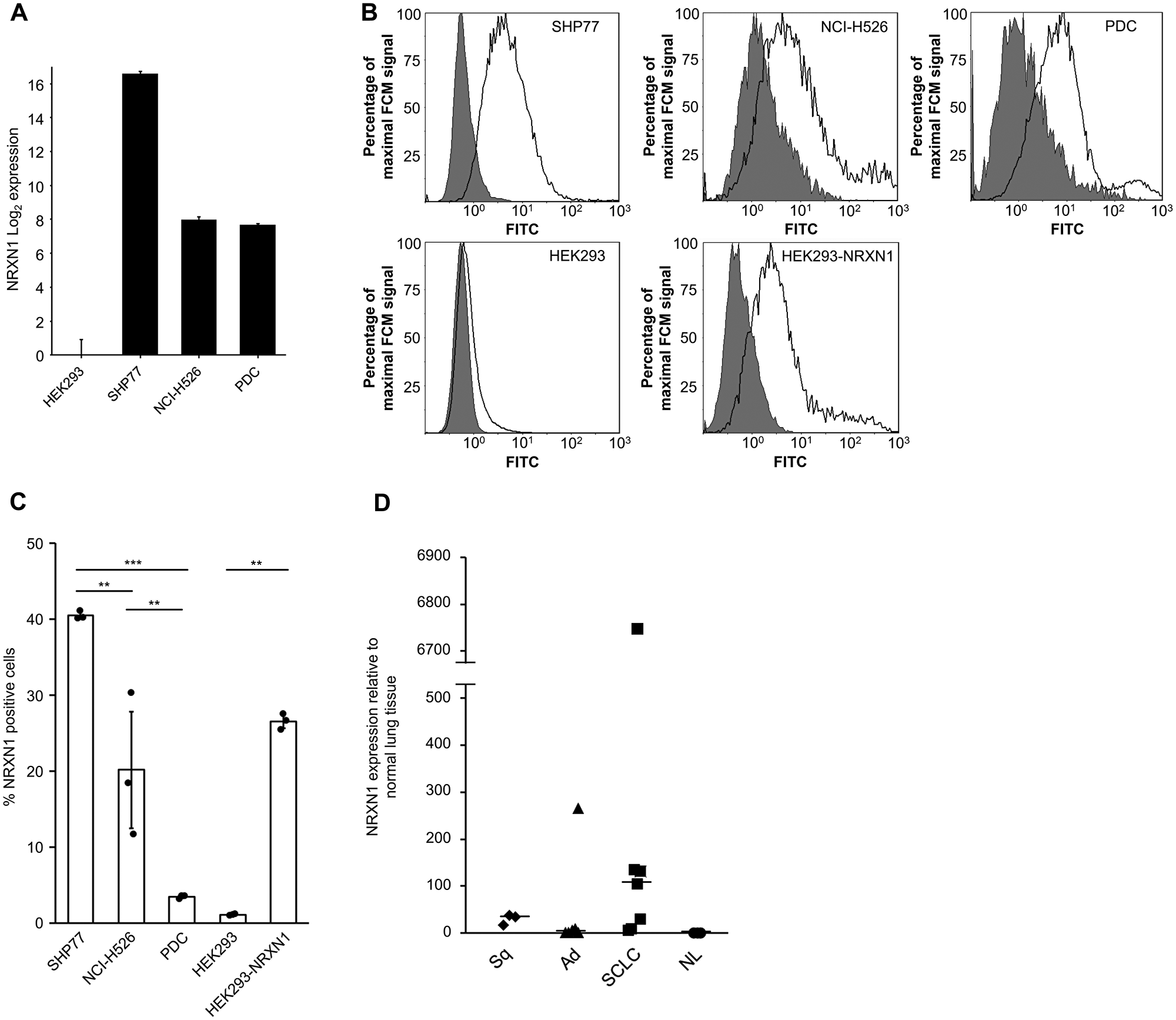
Figure 1: NRXN1 expression in cell lines and surgical specimens of lung tissue including non-SCLC and normal tissues. (A) The relative expressions in cell lines were examined using qRT-PCR with the SYBR green dye assay; data for NRXN1 is shown. The log2-scale relative gene expression is indicated on the y-axis. Error bars, SD. (B) Flow cytometry of NRXN1 on cell lines. Cell surface NRXN1 was assessed using FITC-conjugated anti-rabbit polyclonal antibody following rabbit anti-NRXN1 (Black trace) or IgG isotype control (gray-filled) antibodies. FCM, flow cytometry. (C) Percentage of NRXN1-positive cells examined for each cell line using flow cytometry. Cells were stained with rabbit anti-NRXN1 polyclonal antibody followed by FITC conjugated goat anti-rabbit IgG. A one-way analysis of variance (ANOVA) followed by the Tukey test was performed. (*P < 0.05; **P < 0.01; ***P < 0.0001; Tukey test). Error bars, SD. (D) Relative expression of NRXN1 in surgical specimens including normal lung by qRT-PCR using the SYBR green dye assay. The y-axis shows the NRXN1 expression levels relative to normal lung tissue. The horizontal bars indicate the median gene expression levels for each group. Error bars, SD. Ad, adenocarcinoma. NL, normal lung. SCLC, small cell lung cancer. Sq, squamous cell carcinoma.
NRXN1 expression in a commercially available human RNA panel was also analyzed to confirm the minimal or absence of NRXN1 expression in normal tissues. Consistent with the NCBI RNA sequencing data, NRXN1 expression was relatively limited to the brain. In other normal tissues, the expression of NRXN1 was less than one-third of the level observed in the brain (Figure 2).
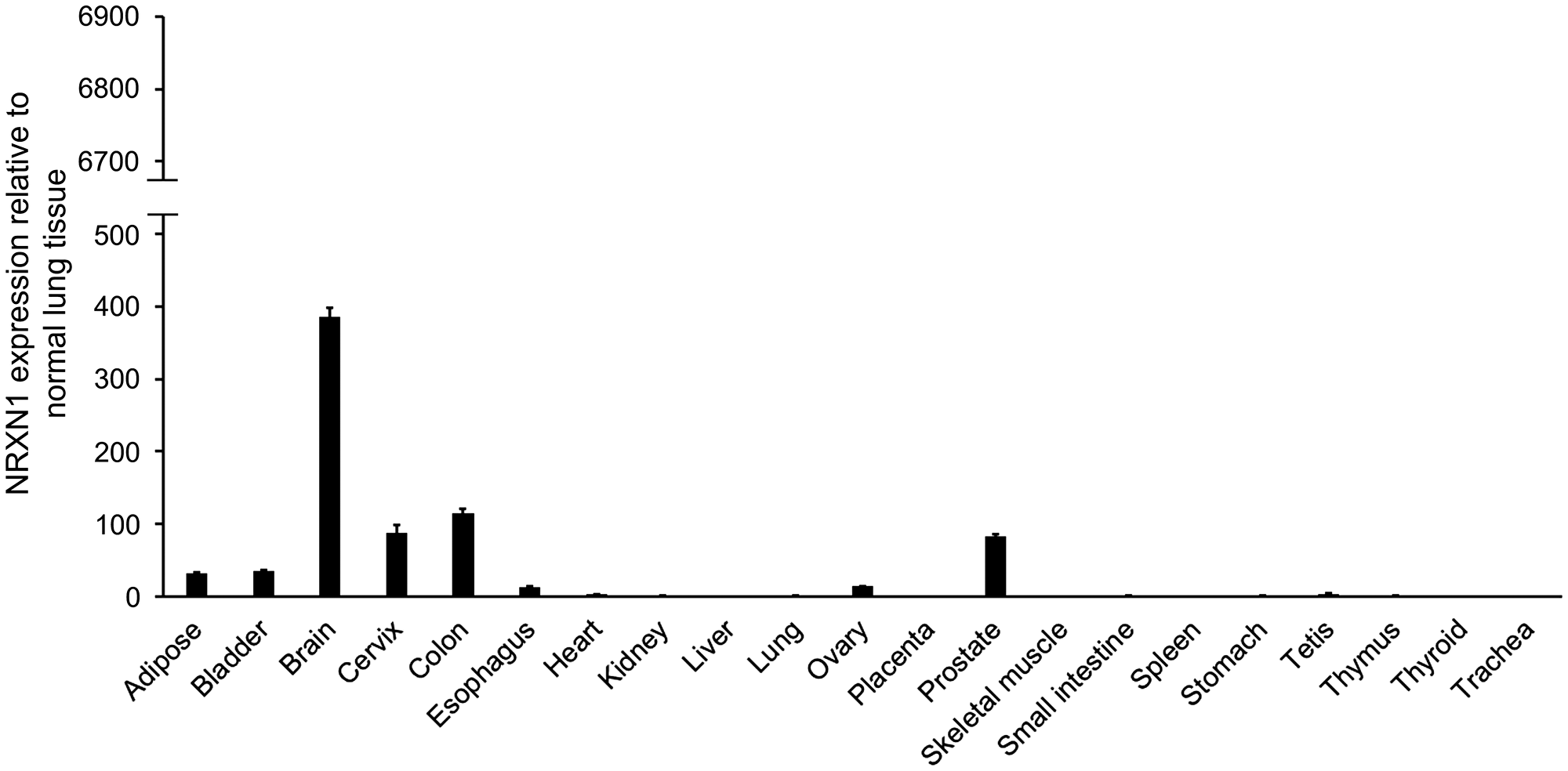
Figure 2: NRXN1 expression in normal multiple organs in humans. The relative expressions of NRXN1 in normal multiple organs of human tissue were examined using qRT-PCR with the SYBR green dye assay. The y-axis shows the NRXN1 expression levels relative to normal lung tissue. The y-axis is set to the same scale as that of Figure 1D so that the expression intensities of both surgical specimens and normal organs can be visually compared. Error bars, SD.
Growth inhibition using a primary anti-NRXN1 monoclonal antibody and a secondary ADC
To screen the ADC activity, we used a monoclonal antibody against NRXN1 and a common secondary ADC capable of binding the primary antibody. In SHP77 cells, the secondary ADC caused a dose-dependent cell growth inhibition in the presence of anti-NRXN1 monoclonal antibody, whereas anti-NRXN1 monoclonal antibody alone, secondary ADC alone, or an IgG isotype control plus the secondary ADC did not inhibit cell growth (Figure 3A). The inhibitory concentration 50 (IC50) value of the secondary ADC in the presence of anti-NRXN1 monoclonal antibody was 3.8 nM, and the potency was more than 3-fold higher than that in the presence of the IgG isotype control antibody. On the other hand, NRXN1-defficient SHP77 cells showed little susceptibility to the secondary ADC with anti-NRXN1 monoclonal antibody (Figure 3B). NRXN1 overexpression sensitized HEK293 cells to the primary anti-NRXN1 monoclonal antibody and the secondary ADC (Figure 3C and 3D). In NCI-H526, growth inhibition by the secondary ADC in the presence of anti-NRXN1 monoclonal antibody was still observed, but to a lesser extent compared to that in SHP77 (Figure 3E). The PDC displayed reduced cell growth by secondary ADC, similar to that in the SHP77 cells, even in the presence of the IgG isotype control (Figure 3F).
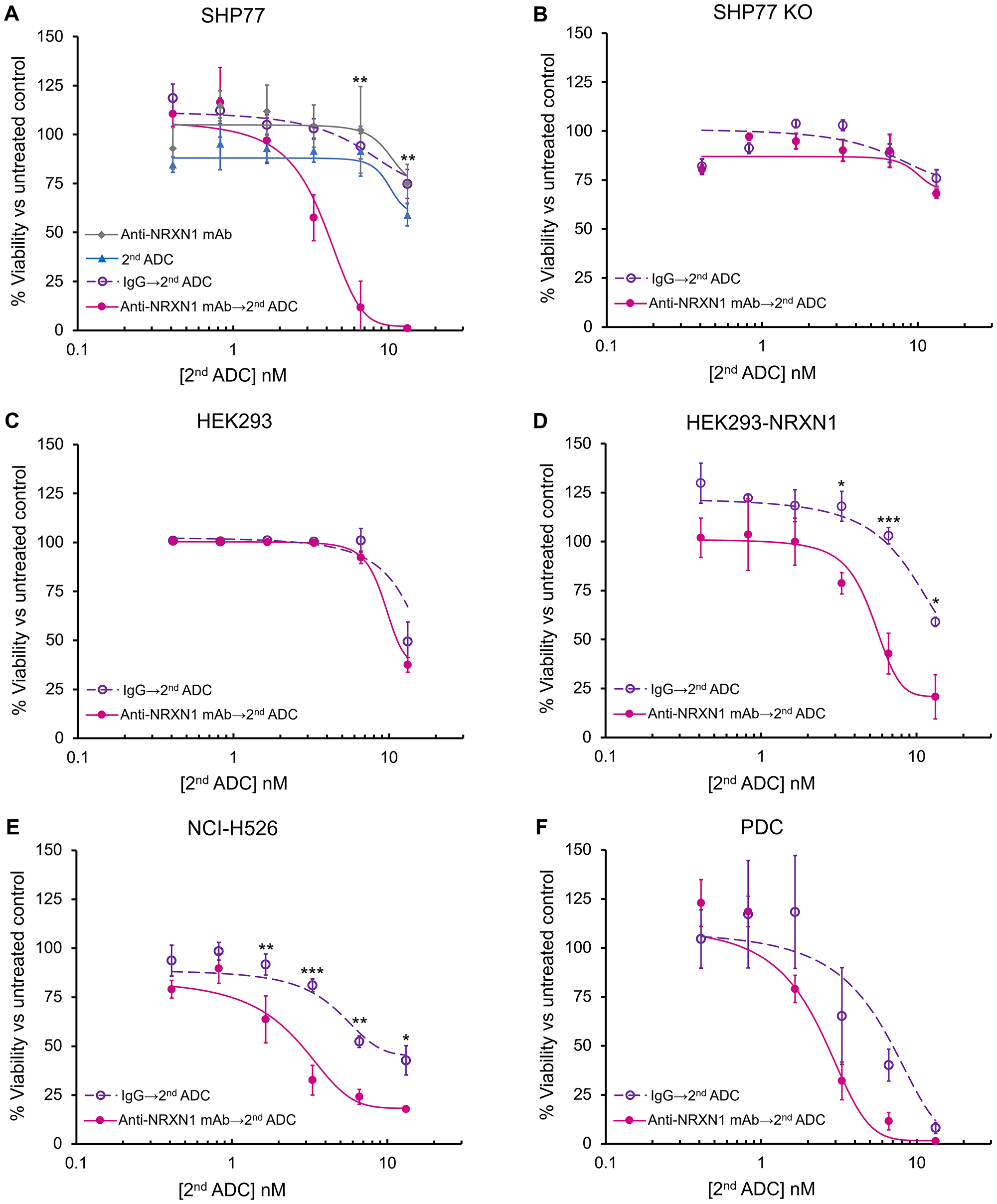
Figure 3: In vitro growth inhibition of NRXN1-targeted ADC. (A–F) In vitro growth inhibition of anti-NRXN1 monoclonal antibody only, secondary ADC only, isotype control antibody (IgG) followed by secondary ADC, and anti-NRXN1 monoclonal antibody followed by secondary ADC on incubation with (A) SHP77, (B) SHP77 KO, (C) HEK293, (D) HEK293-NRXN1, (E) NCI-H526, and (F) PDC. All the assays were performed in triplicate. A two-way ANOVA followed by the Tukey test was performed to assess the difference between IgG with the second ADC group and anti-NRXN1 mAb with the second ADC group. A P value < 0.05 was considered significant (*P < 0.05; **P < 0.01; ***P < 0.0001). Error bars represent the SD of the mean. mAb, Monoclonal antibody. ns, Not significant.
Collectively, these results demonstrate that secondary ADC with anti-NRXN1 monoclonal antibody mediated cytotoxicity in a NRXN1 expression-dependent manner.
Induction of apoptosis by the primary anti-NRXN1 monoclonal antibody and the secondary ADC
To verify actual cell death induced by secondary ADC with anti-NRXN1 monoclonal antibody, cells were treated with secondary ADC plus anti-NRXN1 monoclonal antibody, anti-NRXN1 monoclonal antibody alone, secondary ADC alone, and secondary ADC plus an IgG isotype control. Consistent with the growth inhibition assay shown in Figure 3A, apoptosis was significantly induced in SHP77 cells by the combination of the anti-NRXN1 monoclonal antibody and the secondary ADC (Figure 4). The induction of the apoptosis of PDC was also observed by secondary ADC alone or secondary ADC plus the IgG isotype control, suggesting that PDC had a high sensitivity to a low level of cytotoxic drug (PNU-159682) degraded from the secondary ADC.
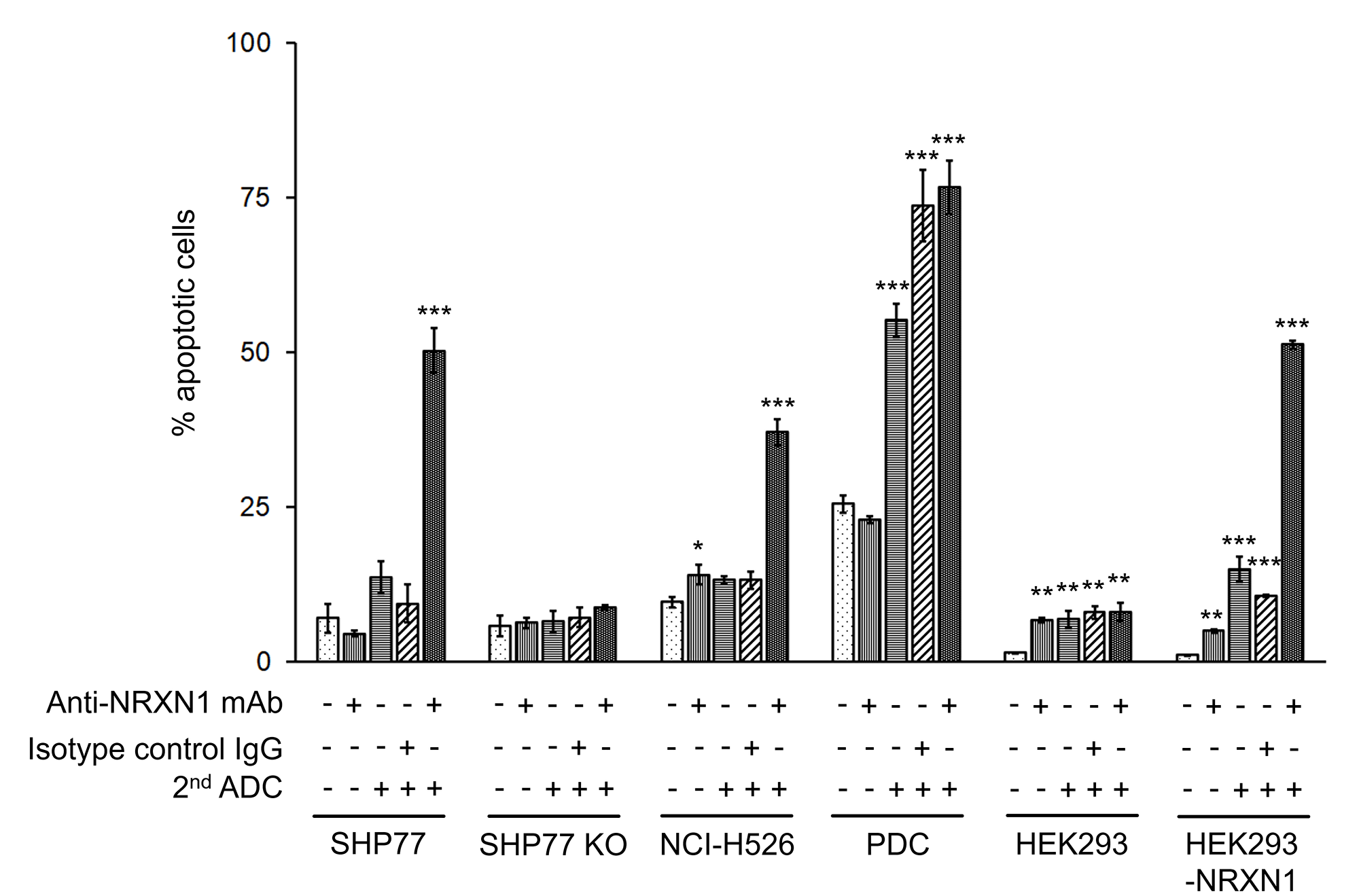
Figure 4: Apoptosis assay of NRXN1-targeted ADC at IC50 dose calculated by growth inhibition curves. Late apoptotic cells were quantified by Cy7-conjugated annexin-V and PI using flow cytometry. Results were analyzed using a one-way ANOVA followed by the Dunnett multiple comparisons test (*P < 0.05; **P < 0.01; ***P < 0.0001 versus no-treatment control group; Dunnett test). Error bars represent the SD of the mean. mAb, monoclonal antibody.
DISCUSSION
We identified 31 membrane proteins as candidates for novel ADC targets and demonstrated that NRXN1 is a promising target for ADCs in SCLC.
Neurexins are single-pass transmembrane proteins encoded by three genes (NRXN1, 2, and 3), and they function as cell adhesion molecules in synaptic transmission. Neurexins expressed in the presynaptic terminal bind postsynaptic neuroligins in a Ca2+-dependent manner, and the neurexin/neuroligin complex plays an important role in synaptic transmission [38]. The relationship between genomic alterations in NRXN genes and a wide variety of neuropsychiatric disorders, including autism spectrum disorders and schizophrenia, has been described in medical literature [39].
Each NRXN gene has at least two alternative promoters. An upstream promotor generates a longer α-isoform and a downstream promotor generates a shorter β-isoform. Moreover, multiple alternative splicing in both isoforms generates thousands of variants [40, 41], and it is difficult to strictly prove the selectivity of the used antibodies. As for the ADC experiments, the loss of cytotoxicity by specific NRXN1 knockout supported the selectivity of the primary antibody. In the flow cytometry, the loss of NRXN1 expression by specific NRXN1 knockout also provided evidence for the selectivity of the antibody.
To reduce the toxicity of ADC therapy, the expressions of the ADC target should be low or absent in normal tissues. NRXN1 is expressed in a limited manner in the central nervous system (CNS) and is a favorable target, since ADCs are unable to pass through the blood-brain barrier because of their large molecular size. A CD56-mediated ADC did not induce obvious damage to the CNS in xenograft models [42]. However, given that antineuronal autoantibodies are detected in paraneoplastic neurological syndrome, possible CNS side effects should be further validated using in vivo experiments.
Given the fact that NRXN1-mediated ADC exhibited favorable cytotoxic activity, monoclonal antibodies directly bound to cytotoxic agents should be generated and optimized for further study. Specifically, the use of different cytotoxic agents, cleavable or non-cleavable linkers, the optimal drug-antibody ratio, and epitopes should be explored and integrated comprehensively using many candidate antibodies. A smaller molecular size using a single antibody instead of two would contribute to more efficient drug delivery. The optimized ADCs should achieve their maximum effect while exhibiting a lower IC50 than that reported in this study through improvements in specific binding, efficient internalization, degradation and the potency of the cytotoxic agent.
Recently, a new nomenclature for SCLC subtypes based on the expressions of four key transcription regulators was reported [11]. NRXN1-positive cell lines generally overlapped with ASCL1-high or NeuroD1-high subtypes, suggesting that NRXN1-positive cell lines belong to subtypes with enhanced neuroendocrine characteristics (Supplementary Figure 1). This result is convincing, considering that NRXN1 is expressed at synapses in the CNS. As to NRXN1 expression in SCLC, not all cases of SCLC are NRXN1-positive, and the proportion of NRXN1-positive cases remains unclear. We tested commercially available anti-NRXN1 polyclonal antibodies (ANR-031 from Alomone Labs and ab214191 from Abcam) for immunohistochemistry using SCLC tissue microarrays, but they lacked specificity (data not shown). Still, the results of our study suggest that an NRXN1-enriched SCLC group could benefit from NRXN1-mediated ADCs.
In the study, we did not directly verify the internalization and trafficking of anti-NRXN1 monoclonal antibody with secondary ADC. We could not conjugate sensor or fluorescent dyes directly to the secondary ADC, because the chemical information of the secondary ADC was a proprietary information of the manufacturer. Moreover, using the third antibody conjugated to sensor or fluorescent dyes to detect the secondary ADC would alter its behavior due to the enlarged size of the complex. In addition, the host species of the secondary ADC was also undisclosed by the manufacturer. Although the results of cell growth inhibition and the apoptosis assay in the present study imply the internalization of the anti-NRXN1 antibody with the secondary ADC because of the significant anti-tumor effects that were observed, the internalization, efficiency of the internalization, and intracellular trafficking of NRXN1-mediated ADCs should be addressed in future research using optimized monoclonal antibodies directly conjugated to payloads.
In conclusion, we identified NRXN1 as a new target for ADCs by screening membrane proteins using a computational-biological approach. The combination of the primary anti-NRXN1 monoclonal antibody and the secondary ADC exhibited anti-tumor activity in an NRXN1-expression dependent manner. NRXN1 could be a novel potential target of ADCs for SCLC that is worth further research.
Materials and Methods
In silico selection of new potential targets for ADCs
We exported the gene expression profiles of 51 SCLC cell lines available at the Cancer Cell Line Encyclopedia and those of 30 normal lung tissue samples examined using the Human Genome U133 Plus 2.0 Array (Thermo Fisher Scientific, Waltham, MA) at the Gene Expression Omnibus (GEO) of the NCBI (Supplementary Table 1). Genes coding membrane proteins were identified using Gene Ontology (GO) and its subontology, known as the Cellular Component Ontology, with the GO terms “plasma membrane” (GO: 0005886) and “anchored component of membrane” (GO: 0031225) or “integral component of membrane” (GO: 0016021).
Cell lines and clinical samples
SCLC cell lines were obtained from the American Type Culture Collection and were cultured according to the manufacturer’s instructions. SHP77 and H526 cells were cultured in Roswell Park Memorial Institute (RPMI 1640) medium with L-glutamine and phenol red (Wako Pure Chemical Industries, Osaka, Japan) containing 10% fetal bovine serum (FBS, Biowest, Nuaillé, France) and 5% Antibiotic-Antimycotic Mixed Stock Solution (Nacalai Tesque, Kyoto, Japan). HEK293 and HEK293-NRXN1 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) with L-glutamine and phenol red (Wako Pure Chemical Industries) containing 10% FBS and 5% Antibiotic-Antimycotic Mixed Stock Solution (Nacalai Tesque). All the cells were cultured at 37°C in a humidified incubator with 5% CO2 and passaged every 3 to 4 days. As to patient-derived cells, the circulating tumor cells of an 84-year-old male with SCLC were derived from his peripheral blood after the approval of the Institutional Review Board at the University of Tokyo Hospital. The circulating tumor cells were purified using RosetteSep™ CTC Enrichment Cocktail Containing Anti-CD56 (STEMCELL Technologies, Vancouver, Canada) according to the manufacturer’s protocol. The purified cells were cultured in RPMI-1640 supplemented with bFGF (20 μg/L, #13256029, Thermo Fisher Scientific), EGF (20 μg/L, #PHG0314, Thermo Fisher Scientific), and B-27™ supplement (#17504044, Thermo Fisher Scientific) using Coster 24 Well Clear Flat Bottom Ultra Low Attachment Multiple Well Plates (Corning, Corning, NY) [43].
Generation of NRXN1 KO cells
We knocked out the NRXN1 gene using the lentiviral CRISPR/Cas9 sgRNA mediated expression knockout protocol. Candidates for the sgRNA were configured using the CRISPR design tool CHOPCHOP [44] and CRISPRdirect [45]. Selected candidates for the sgRNA (Merck, Darmstadt, Germany) were cloned into Cas9 SmartNuclease All-in-one Vector (System Biosciences, Palo Alto, CA). The insert coding Cas9 and the sgRNA was ligated into the lentivirus vector CSII-CMV-MCS-IRES2-Bsd (RIKEN BioResource Research Center, Tsukuba, Japan) after being removed from the Cas9 SmartNuclease All-in-one Vector. The SHP77 cells underwent lentiviral transduction using polybrene. After incubation for 14 days with medium containing 10 μg/mL blasticidin, the cell populations were screened using flow cytometry and qRT-PCR to confirm the efficiency of the CRISPR knockout of NRXN1 (Supplementary Figure 2).
mRNA expression analysis
RNA was extracted using RNAiso plus (TAKARA BIO, Shiga, Japan). Complementary DNA (cDNA) was generated from 1 μg of RNA using SuperScript III Reverse Transcriptase (Thermo Fisher Scientific). Each cDNA and primers specific for NRXN1 (forward primer 5′- GAT TCT TAC CAC AAC GGG CTA CA-3′, and reverse primer 5′-GGG TTT CAA AGG TGA TTG GGT C-3′) and GAPDH (forward primer 5′-CAC CAC CAA CTG CTT AGC AC-3′, and reverse primer 5′-TGG CAG GTT TTT CTA GAC GG-3′) were mixed with THUNDERBIRD SYBR qPCR Mix (TOYOBO, Osaka, Japan). The reaction mixes were run on the 7500 Fast Real-Time PCR System (Thermo Fisher Scientific). Relative gene expression was calculated using the ddCt method. NRXN1 expression in human normal tissues was examined using he FirstChoice Human Total RNA Survey Panel (Thermo Fisher Scientific).
Antibodies
Mouse anti-NRXN1α monoclonal antibody (sc-136001, Santa Cruz Biotechnology, Dallas, TX) recognizing amino acids 1063-1184 of rabbit NRXN1α was applied to the cell viability assay. Anti-NRXN1α polyclonal antibody raised in rabbits (ANR-031; Alomone Labs, Jerusalem, Israel) against amino acid residues 546-560 of rat NRXN1α was applied for NRXN1 measurements using flow cytometry. The secondary ADC used in the study was αMFc-CL-PNU (AM-102-PN; Moradec, San Diego, CA), an anti-mouse IgG Fc-specific antibody conjugated to PNU-159682 with a cleavable linker. PNU-159682 is a derivative of nemorubicin, which induces cell death by intercalating DNA and topoisomerase inhibition. The antibody portion is a polyclonal antibody specific to the Fc region of mouse IgGs. The cleavable linker is stable in extracellular fluid, but is cleaved by cathepsin in endosomes once the conjugate is internalized into cells.
NRXN1 overexpression
cDNA was generated from RNA extracted from SHP77 cells. The DNA sequence coding NRXN1 was reconstructed by polymerase reaction with the first half and the latter half of the entire coding sequence [46]. The first half of the sequence was amplified with forward primer 5′-TCC CGC CTT TCC CCT TAC-3′ and reverse primer 5′-GCT GGA ATT ACA GTT AAT CCT GAT AC-3′. The latter half of the sequence was amplified with forward primer 5′-GGA GCA TGT TTA TGA AAA TTC AG-3′ and reverse primer 5′-CAT TCC CTG TCT TCT TTT GTA TG-3′. The full-length sequence was subcloned to the pGEM-T Easy Vector plasmid (Promega, Madison, WI). The plasmid was transformed into E. coli DH-5α Competent Cells (TaKaRa Bio) according to the manufacture’s protocol. The sequences of plasmids from colonies of the competent cells were verified after being extracted from transformed DH-5α using the PureYield™ Plasmid Miniprep System (Promega, Madison, WI). The collected plasmids were extracted from competent cells incubated overnight using the PureYield™ Plasmid Midiprep System (Promega). Fragments including the NRXN1 sequence were subcloned into pcDNA™3.1/Zeo(+)(Thermo Fisher Scientific). HEK293 cells were transfected with HilyMax Reagent (Dojindo, Kumamoto, Japan) and incubated for 2 days, at which time the cells were examined to confirm the overexpression of the target genes using qRT-PCR and flow cytometry.
Flow cytometry
Cells were incubated with 2.5 μL of rabbit anti-NRXN1 polyclonal antibody (ANR-031; Alomone Labs) or an isotype control at 1 × 105 cells/100 μL in PBS with 2% FBS for 30 min in the dark. After two washes, the cells were incubated with 2 μL of goat anti-rabbit IgG FITC conjugate (ab97050; Abcam, Cambridge, United Kingdom) for 30 min.
After two washes, 2 μL of PI (Biolegend, San Diego, CA) were added to the cells for 15 min before the FC500 flow cytometer (Beckman Coulter, Brea, CA) run. Data were analyzed using FC500 analysis software (Beckman Coulter).
In vitro cytotoxic assay
On day 1, the cancer cells, HEK293, and HEK293-NRXN1 cells were seeded at a density of 1000 cells per well in 50 μL of medium on 96-well culture plates. PDCs were plated at a density of 2500 cells per well. On day 2, the primary antibodies (anti-NRXN1 monoclonal antibody or IgG isotype control; 50 μL per well) were added to each well. After incubation in the presence of the primary antibodies for 10 min, 2 μL of the secondary ADC was added as a serial dilution. The primary antibody (anti-NRXN1 monoclonal antibody or IgG isotype control) and the secondary ADC were added at a constant volume ratio of 2:1 in each well. The cell lines and PDCs were incubated for 3 and 7 days, respectively, and were subjected to a cell growth assay using the Cell Counting Kit-8 (Dojindo).
Apoptosis assay
Apoptosis was analyzed using double staining with Annexin-V Cy7 conjugate (BioLegend) and PI (BioLegend) according to the manufacturer’s instructions. Briefly, cells were incubated in Annexin-V binding buffer. After incubation with 5 μL of Annexin-V Cy7 conjugate and 2 μL of PI for 15 min, the cells were examined using FC500. Data were analyzed using FC500 software.
Statistical methods
Data are presented as the mean ± SD, as stated in the figure legends. The statistical analysis of the gene expression profiles obtained from CCLE and NCBI GEO was performed using R (version 3.1.1, R Foundation for Statistical Computing) and Bioconductor. The statistical analysis of the NRXN1-positive cells detected using flow cytometry, the in vitro growth inhibition assay, and the apoptosis assay was performed using JMP Pro 14.2 software (SAS Institute Inc., Cary, NC). The results of the NRXN1-positive cells detected using flow cytometry were analyzed using a one-way ANOVA followed by the Tukey test (*P < 0.05; **P < 0.01; ***P < 0.0001; Tukey test). Regarding the in vitro growth inhibition assay, a two-way ANOVA followed by the Tukey test was performed for multiple comparison analyses of the in vitro cytotoxic activity (*P < 0.05; **P < 0.01; ***P < 0.0001; Tukey test). The percentage of late apoptotic cells (Annexin V and PI double-positive cells) were compared with those of untreated cells using a one-way ANOVA followed by the Dunnett test (*P < 0.05; **P < 0.01; ***P < 0.0001 versus no-treatment control group; Dunnett test). Differences were considered significant when the P value was less than 0.05.
Abbreviations
ADC: antibody-drug conjugate; ALK: anaplastic lymphoma kinase; ANOVA: analysis of variance; ASCL1: achaete-scute homolog 1; bFGF: basic fibroblast growth factor; Cas9: CRISPR-associated proteins 9; cDNA: complementary deoxyribonucleic acid; CRISPR: clustered regularly interspaced short palindromic repeats; CSC: cancer stem cell; Cy: cyanine; DLL3: delta-like ligand 3; EGF: epidermal growth factor; EGFR: epidermal growth factor receptor; EZH2: enhancer of zeste homolog 2; FITC: fluorescein isothiocyanate; GABA: gamma-aminobutyric acid; GAPDH: Glyceraldehyde 3-phosphate dehydrogenase; HER2: erb-b2 receptor tyrosine kinase 2; KRAS: kirsten rat sarcoma viral oncogene homologue; mAb: monoclonal antibody; MYC: cellular myelocytomatosis oncogene; NEUROD1: Neurogenic differentiation 1; NRXN: neurexin; PARP: poly-ADP- ribose polymerase; PBS: phosphate buffered saline; PI: propidium iodide; PI3K: phosphoinositide 3 kinase; POU2F3: POU class 2 homeobox 3; PTEN: phosphatase and tensin homologue deleted on chromosome 10; RB1: retinoblastoma; SCLC: small cell lung cancer; TP53: tumor suppressor protein p53; TROP2: Trophoblast cell surface antigen 2; YAP1: Yes-associated protein 1.
CONFLICTS OF INTEREST
Authors have no conflicts of interest to declare.
FUNDING
This study was supported by a Grant in Aid for Young Scientists (JSPS KAKENHI No. 18K15266, No.19K17666), a Grant in Aid for Scientific Research (C) (JSPS KAKENHI No.16K09574, No.19K08622), and a Grant in Aid for Scientific Research (A) (JSPS KAKENHI No.16H02653) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
References
1. Travis WD. Pathology and diagnosis of neuroendocrine tumors: Lung neuroendocrine. Thorac Surg Clin. 2014; 24:257–266. https://doi.org/10.1016/j.thorsurg.2014.04.001. [PubMed].
2. Parsons HM, Harlan LC, Stevens JL, Ullmann CD. Treatment of small cell lung cancer in academic and community settings: Factors associated with receiving standard therapy and survival. Cancer J. 2014; 20:97–104. https://doi.org/10.1097/PPO.0000000000000039. [PubMed].
3. Horn L, Mansfield AS, Szczęsna A, Havel L, Krzakowski M, Hochmair MJ, Huemer F, Losonczy G, Johnson ML, Nishio M, Reck M, Mok T, Lam S, et al. First-Line Atezolizumab plus Chemotherapy in Extensive-Stage Small-Cell Lung Cancer. N Engl J Med. 2018; 379:2220–2229. https://doi.org/10.1056/NEJMoa1809064. [PubMed].
4. Kalemkerian GP, Loo BW, Akerley W, Attia A, Bassetti M, Boumber Y, Decker R, Dobelbower MC, Dowlati A, Downey RJ, Florsheim C, Ganti AKP, Grecula JC, et al. NCCN guidelines insights: Small cell lung cancer. J Natl Compr Canc Netw. 2018; 16:1171–1182. https://doi.org/10.6004/jnccn.2018.0079. [PubMed].
5. George J, Lim JS, Jang SJ, Cun Y, Ozreti’c L, Kong G, Leenders F, Lu X, Fernández-Cuesta L, Bosco G, Müller C, Dahmen I, Jahchan NS, et al. Comprehensive genomic profiles of small cell lung cancer. Nature. 2015; 524:47–53. https://doi.org/10.1038/nature14664. [PubMed].
6. Tsoukalas N, Aravantinou-Fatorou E, Baxevanos P, Tolia M, Tsapakidis K, Galanopoulos M, Baxevanos P, Tolia M, Tsapakidis K, Galanopoulos M, Liontos M, Kyrgias G. Advanced small cell lung cancer (SCLC): new challenges and new expectations. Ann Transl Med. 2018; 6:145. https://doi.org/10.21037/atm.2018.03.31. [PubMed].
7. Atrafi F, Groen HJM, Byers LA, Garralda E, Lolkema MP, Sangha RS, Viteri S, Chae YK, Camidge DR, Gabrail NY, Hu B, Tian T, Nuthalapati S, et al. A phase I dose-escalation study of veliparib combined with carboplatin and etoposide in patients with extensive-stage small cell lung cancer and other solid tumors. Clin Cancer Res. 2019; 25:496–505. https://doi.org/10.1158/1078-0432.CCR-18-2014. [PubMed].
8. Owonikoko TK, Dahlberg SE, Sica GL, Wagner LI, Wade JL, Srkalovic G, Lash BW, Leach JW, Leal TB, Aggarwal C, Ramalingam SS. Randomized phase II trial of cisplatin and etoposide in combination with veliparib or placebo for extensive-stage small-cell lung cancer: ECOG-ACRIN 2511 study. J Clin Oncol. 2019; 37:222–229. https://doi.org/10.1200/JCO.18.00264. [PubMed].
9. Pietanza MC, Waqar SN, Krug LM, Dowlati A, Hann CL, Chiappori A, Owonikoko TK, Woo KM, Cardnell RJ, Fujimoto J, Long L, Diao L, Wang J, et al. Randomized, double-blind, phase II study of temozolomide in combination with either veliparib or placebo in patients with relapsed-sensitive or refractory small-cell lung cancer. J Clin Oncol. 2018; 36:2386–2394. https://doi.org/10.1200/JCO.2018.77.7672. [PubMed].
10. Gardner EE, Lok BH, Schneeberger VE, Desmeules P, Miles LA, Arnold PK, Ni A, Khodos I, de Stanchina E, Nguyen T, Sage J, Campbell JE, Ribich S, et al. Chemosensitive relapse in small cell lung cancer proceeds through an EZH2-SLFN11 axis. Cancer Cell. 2017; 31:286–299. https://doi.org/10.1016/j.ccell.2017.01.006. [PubMed].
11. Rudin CM, Poirier JT, Byers LA, Dive C, Dowlati A, George J, Heymach JV, Johnson JE, Lehman JM, MacPherson D, Massion PP, Minna JD, Oliver TG, et al. Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat Rev Cancer. 2019; 19:289–297. https://doi.org/10.1038/s41568-019-0133-9. [PubMed].
12. Polakis P. Antibody Drug Conjugates for Cancer Therapy. Pharmacol Rev. 2016; 68:3–19. https://doi.org/10.1124/pr.114.009373. [PubMed].
13. Chalouni C, Doll S. Fate of Antibody-Drug Conjugates in Cancer Cells. J Exp Clin Cancer Res. 2018; 37:20. https://doi.org/10.1186/s13046-017-0667-1. [PubMed].
14. Deneka AY, Boumber Y, Beck T, Golemis EA. Tumor-Targeted Drug Conjugates as an Emerging Novel Therapeutic Approach in Small Cell Lung Cancer (SCLC). Cancers (Basel). 2019; 11:1297. https://doi.org/10.3390/cancers11091297. [PubMed].
15. Senter PD, Sievers EL. The discovery and development of brentuximab vedotin for use in relapsed Hodgkin lymphoma and systemic anaplastic large cell lymphoma. Nat Biotechnol. 2012; 30:631–637. https://doi.org/10.1038/nbt.2289. [PubMed].
16. Lambert JM, Chari RV. Ado-trastuzumab Emtansine (T-DM1): an antibody-drug conjugate (ADC) for HER2-positive breast cancer. J Med Chem. 2014; 57:6949–6964. https://doi.org/10.1021/jm500766w. [PubMed].
17. Lewis Phillips GD, Li G, Dugger DL, Crocker LM, Parsons KL, Mai E, Blättler WA, Lambert JM, Chari RV, Lutz RJ, Wong WL, Jacobson FS, Koeppen H, et al. Targeting HER2-Positive Breast Cancer with Trastuzumab-DM1, an Antibody–Cytotoxic Drug Conjugate. Cancer Res. 2008; 68:9280–9290. https://doi.org/10.1158/0008-5472.CAN-08-1776. [PubMed].
18. Kantarjian HM, DeAngelo DJ, Stelljes M, Martinelli G, Liedtke M, Stock W, Gökbuget N, O’Brien S, Wang K, Wang T, Paccagnella ML, Sleight B, Vandendries E, et al. Inotuzumab Ozogamicin versus Standard Therapy for Acute Lymphoblastic Leukemia. N Engl J Med. 2016; 375:740–753. https://doi.org/10.1056/NEJMoa1509277. [PubMed].
19. Appelbaum FR, Bernstein ID. Gemtuzumab ozogamicin for acute myeloid leukemia. Blood. 2017; 130:2373–2376. https://doi.org/10.1182/blood-2017-09-797712. [PubMed].
20. Modi S, Saura C, Yamashita T, Park YH, Kim SB, Tamura K, Andre F, Iwata H, Ito Y, Tsurutani J, Sohn J, Denduluri N, Perrin C, et al. Trastuzumab Deruxtecan in Previously Treated HER2-Positive Breast Cancer. N Engl J Med. 2020; 382:610–621. https://doi.org/10.1056/NEJMoa1914510. [PubMed].
21. Wong OK, Tran TT, Ho WH, Casas MG, Au M, Bateman M, Lindquist KC, Rajpal A, Shelton DL, Strop P, Liu SH. RN765C, a low affinity EGFR antibody drug conjugate with potent anti-tumor activity in preclinical solid tumor models. Oncotarget. 2018; 9:33446–33458. https://doi.org/10.18632/oncotarget.26002. [PubMed].
22. Purcell JW, Tanlimco SG, Hickson J, Fox M, Sho M, Durkin L, Uziel T, Powers R, Foster K, McGonigal T, Kumar S, Samayoa J, Zhang D, et al. LRRC15 Is a Novel Mesenchymal Protein and Stromal Target for Antibody–Drug Conjugates. Cancer Res. 2018; 78:4059–4072. https://doi.org/10.1158/0008-5472.CAN-18-0327. [PubMed].
23. Damelin M, Bankovich A, Bernstein J, Lucas J, Chen L, Williams S, Park A, Aguilar J, Ernstoff E, Charati M, Dushin R, Aujay M, Lee C, et al. A PTK7-targeted antibody-drug conjugate reduces tumor-initiating cells and induces sustained tumor regressions. Sci Transl Med. 2017; 9:eaag2611. https://doi.org/10.1126/scitranslmed.aag2611. [PubMed].
24. Puca L, Gavyert K, Sailer V, Conteduca V, Dardenne E, Sigouros M, Isse K, Kearney M, Vosoughi A, Fernandez L, Pan H, Motanagh S, Hess J, et al. Delta-like protein 3 expression and therapeutic targeting in neuroendocrine prostate cancer. Sci Transl Med. 2019; 11:eaav089. https://doi.org/10.1126/scitranslmed.aav0891. [PubMed].
25. Merlino G, Fiascarelli A, Bigioni M, Bressan A, Carrisi C, Bellarosa D, Salerno M, Bugianesi R, Manno R, Bernadó Morales C, Arribas J, Dusek RL, Ackroyd JE, et al. MEN1309/OBT076, a First-In-Class Antibody-Drug Conjugate Targeting CD205 in Solid Tumors. Mol Cancer Ther. 2019; 18:1533–1543. https://doi.org/10.1158/1535-7163.MCT-18-0624. [PubMed].
26. Sau S, Petrovici A, Alsaab HO, Bhise K, Iyer AK. PDL-1 Antibody Drug Conjugate for Selective Chemo-Guided Immune Modulation of Cancer. Cancers (Basel). 2019; 11:232. https://doi.org/10.3390/cancers11020232. [PubMed].
27. Sano R, Krytska K, Larmour CE, Raman P, Martinez D, Ligon GF, Lillquist JS, Cucchi U, Orsini P, Rizzi S, Pawel BR, Alvarado D, Mossé YP. An antibody-drug conjugate directed to the ALK receptor demonstrates efficacy in preclinical models of neuroblastoma. Sci Transl Med. 2019; 11:eaau9732. https://doi.org/10.1126/scitranslmed.aau9732. [PubMed].
28. Gan HK, van den Bent M, Lassman AB, Reardon DA, Scott AM. Antibody-drug conjugates in glioblastoma therapy: the right drugs to the right cells. Nat Rev Clin Oncol. 2017; 14:695–707. https://doi.org/10.1038/nrclinonc.2017.95. [PubMed].
29. Abrams T, Connor A, Fanton C, Cohen SB, Huber T, Miller K, Hong EE, Niu X, Kline J, Ison-Dugenny M, Harris S, Walker D, Krauser K, et al. Preclinical Antitumor Activity of a Novel Anti-c-KIT Antibody-Drug Conjugate against Mutant and Wild-type c-KIT -Positive Solid Tumors. Clin Cancer Res. 2018; 24:4297–4308. https://doi.org/10.1158/1078-0432.CCR-17-3795. [PubMed].
30. Henke RM, Meredith DM, Borromeo MD, Savage TK, Johnson JE. Ascl1 and Neurog2 form novel complexes and regulate Delta-like3 (Dll3) expression in the neural tube. Dev Biol. 2009; 328:529–540. https://doi.org/10.1016/j.ydbio.2009.01.007. [PubMed].
31. Augustyn A, Borromeo M, Wang T, Fujimoto J, Shao C, Dospoy PD, Lee V, Tan C, Sullivan JP, Larsen JE, Girard L, Behrens C, Wistuba II, et al. ASCL1 is a lineage oncogene providing therapeutic targets for high-grade neuroendocrine lung cancers. Proc Natl Acad Sci U S A. 2014; 111:14788–14793. https://doi.org/10.1073/pnas.1410419111. [PubMed].
32. Saunders LR, Bankovich AJ, Anderson WC, Aujay MA, Bheddah S, Black K, Desai R, Escarpe PA, Hampl J, Laysang A, Liu D, Lopez-Molina J, Milton M, et al. A DLL3-targeted antibody-drug conjugate eradicates high-grade pulmonary neuroendocrine tumor-initiating cells in vivo. Sci Transl Med. 2015; 7:302ra136. https://doi.org/10.1126/scitranslmed.aac9459. [PubMed].
33. Phase 3 Trial of Rova-T as Second-line Therapy for Advanced Small-Cell Lung Cancer (TAHOE Study) Halted. https://news.abbvie.com/news/phase-3-trial-rova-t-as-second-line-therapy-for-advanced-small-cell-lung-cancer-tahoe-study-halted.htm.
34. Goldenberg DM, Cardillo TM, Govindan SV, Rossi EA, Sharkey RM. Trop-2 is a novel target for solid cancer therapy with sacituzumab govitecan (IMMU-132), an antibody-drug conjugate (ADC). Oncotarget. 2015; 6:22496–22512. https://doi.org/10.18632/oncotarget.4318. [PubMed].
35. Inamura K, Yokouchi Y, Kobayashi M, Ninomiya H, Sakakibara R, Subat S, Nagano H, Nomura K, Okumura S, Shibutani T, Ishikawa Y. Association of tumor TROP2 expression with prognosis varies among lung cancer subtypes. Oncotarget. 2017; 8:28725–28735. https://doi.org/10.18632/oncotarget.15647. [PubMed].
36. Gray JE, Heist RS, Starodub AN, Camidge DR, Kio EA, Masters GA, Purcell WT, Guarino MJ, Misleh J, Schneider CJ, Schneider BJ, Ocean A, Johnson T, et al. Therapy of Small Cell Lung Cancer (SCLC) with a Topoisomerase-I–inhibiting Antibody–Drug Conjugate (ADC) Targeting Trop-2, Sacituzumab Govitecan. Clin Cancer Res. 2017; 23:5711–5719. https://doi.org/10.1158/1078-0432.CCR-17-0933. [PubMed].
37. Socinski MA, Kaye FJ, Spigel DR, Kudrik FJ, Ponce S, Ellis PM, Majem M, Lorigan P, Gandhi L, Gutierrez ME, Nepert D, Corral J, Ares LP. Phase 1/2 Study of the CD56-Targeting Antibody-Drug Conjugate Lorvotuzumab Mertansine (IMGN901) in Combination With Carboplatin/Etoposide in Small-Cell Lung Cancer Patients With Extensive-Stage Disease. Clin Lung Cancer. 2017; 18:68–76.e2. https://doi.org/10.1016/j.cllc.2016.09.002. [PubMed].
38. Reissner C, Klose M, Fairless R, Missler M. Mutational analysis of the neurexin/neuroligin complex reveals essential and regulatory components. Proc Natl Acad Sci U S A. 2008; 105:15124–15129. https://doi.org/10.1073/pnas.0801639105. [PubMed].
39. Kasem E, Kurihara T, Tabuchi K. Neurexins and neuropsychiatric disorders. Neurosci Res. 2018; 127:53–60. https://doi.org/10.1016/j.neures.2017.10.012. [PubMed].
40. Schreiner D, Nguyen TM, Russo G, Heber S, Patrignani A, Ahrné E, Scheiffele P. Targeted combinatorial alternative splicing generates brain region-specific repertoires of neurexins. Neuron. 2014; 84:386–398. https://doi.org/10.1016/j.neuron.2014.09.011. [PubMed].
41. Harkin LF, Lindsay SJ, Xu Y, Alzu’bi A, Ferrara A, Gullon EA, James OG, Clowry GJ. Neurexins 1-3 Each Have a Distinct Pattern of Expression in the Early Developing Human Cerebral Cortex. Cereb Cortex. 2017; 27:216–232. https://doi.org/10.1093/cercor/bhw394. [PubMed].
42. Yu L, Lu Y, Yao Y, Liu Y, Wang Y, Lai Q, Zhang R, Li W, Wang R, Fu Y, Tao Y, Yi S, Gou L, et al. Promiximab-duocarmycin, a new CD56 antibody-drug conjugates, is highly efficacious in small cell lung cancer xenograft models. Oncotarget. 2017; 9:5197–5207. https://doi.org/10.18632/oncotarget.23708. [PubMed].
43. Yu M, Bardia A, Aceto N, Bersani F, Madden MW, Donaldson MC, Desai R, Zhu H, Comaills V, Zheng Z, Wittner BS, Stojanov P, Brachtel E, et al. Cancer therapy. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science. 2014; 345:216–220. https://doi.org/10.1126/science.1253533. [PubMed].
44. Labun K, Montague TG, Gagnon JA, Thyme SB, Valen E. CHOPCHOP v2: a web tool for the next generation of CRISPR genome engineering. Nucleic Acids Res. 2016; 44:272–276. https://doi.org/10.1093/nar/gkw398. [PubMed].
45. Naito Y, Hino K, Bono H, Ui-Tei K. CRISPRdirect: software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics. 2015; 31:1120–1123. https://doi.org/10.1093/bioinformatics/btu743. [PubMed].
46. Sunohara M, Kawakami M, Kage H, Watanabe K, Emoto N, Nagase T, Ohishi N, Takai D. Polymerase reaction without primers throughout for the reconstruction of full-length cDNA from products of rapid amplification of cDNA ends (RACE). Biotechnol Lett. 2011; 33:1301–1307. https://doi.org/10.1007/s10529-011-0580-1. [PubMed].

