Introduction
The von Hippel-Lindau (VHL) tumor suppressor gene located on chromosome 3p25.3 encodes a multifunctional protein. One important function for tumor biology is serving as part of an E3 ubiquitin ligase that recognizes and ubiquitylates the α-subunits of hypoxia-induced factors (HIF-α) 1, 2 and 3, which play a crucial role for regulation of vascular formation. Inactivation of VHL mimics the consequences of hypoxia as it abrogates proteosomal HIF-α degradation, resulting in HIF-α accumulation and activation of antiapoptotic signals [1–3]. Germline mutations of VHL cause a rare familial tumor syndrome that is characterized by highly vascularized tumors. Sporadic biallelic genetic or epigenetic VHL alterations, including VHL gene mutations, chromosome 3p deletions, or VHL gene methylations, are found in the vast majority of cell renal cell carcinomas and have widely been investigated over the past years [4, 5]. Most of these studies have analyzed either mutations or deletions but rarely both. Overall the data suggest that the vast majority (> 90%) of clear cell carcinomas harbor VHL aberrations [6, 7].
Although VHL alterations are established markers of clear cell renal cell carcinomas, their role is much less clear in other kidney tumor entities. Anecdotal reports exist on VHL deletions in papillary carcinoma and clear cell (tubulo) papillary carcinoma [8–10] as well as VHL mutations in individual cases of oncocytoma as well as chromophobe and papillary cancer [11]. Regarding other less frequent tumor entities – including tumors introduced or modified by the latest WHO classification of tumors of the Urological Tract – data on VHL mutation and deletion status are also rare.
In this study, we took advantage of a cohort of more than 1,800 tumors recently reviewed according to the latest WHO classification criteria to study the combined impact of VHL mutations and deletions in kidney tumors of all types and to study the role of these alterations on clinical outcome.
Results
Technical issues
A total of 705/1,805 (39.1%) samples of renal tumors were successfully sequenced for VHL. The successful sequencing of 3 exons was considered a prerequisite for the identification of tumors without mutations, whereas the successful sequencing of at least one altered exon was sufficient to consider tumors as positive for VHL mutations.
There were 256 (36.3%) tumors without and 449 (63.7%) tumors with VHL mutations. In 1,100 samples, an unambiguous result could not be obtained due to insufficient amount or quality of DNA (n = 166) or inadequate quality of the sequencing result (n = 934).
VHL mutations
The frequency of VHL mutations is summarized in Table 1 for all kidney tumors and the different main histologic subtypes. Mutation rates are shown separately for tumors with sequencing data of 1–2 exons and of all 3 exons.
Table 1: Frequency of VHL mutations and VHL depletion of 3p in all renal cancers and in different histologic subtypes of renal cell carcinomas (RCC)
| VHL mutation analysis* | 3p deletion analysis | |||||
| 1–2 exons analyzed | 3 exons analyzed | |||||
| n | mutated (%) | n | mutated (%) | n | deletion (%) | |
| all cancers | 705 | 63.7 | 431 | 40.6 | 1403 | 43.3 |
| clear cell RCC | 515 | 78.3 | 275 | 59.3 | 918 | 57.2 |
| papillary RCC | 68 | 19.1 | 58 | 5.2 | 210 | 17.6 |
| oncocytoma | 49 | 22.4 | 41 | 7.3 | 118 | 11.9 |
| chromophobic RCC | 38 | 18.4 | 32 | 3.1 | 79 | 17.7 |
| clear cell (tubulo) papillary RCC | 13 | 84.6 | 5 | 60.0 | 24 | 29.2 |
| nephroblastoma | 4 | 0.0 | 4 | 0.0 | 12 | 16.7 |
| Xp11.2 translocated carcinoma | 4 | 25.0 | 4 | 25.0 | 10 | 10.0 |
| collecting duct carcinoma | 0 | 0 | 3 | 0.0 | ||
| metanephrogenic adenoma | 0 | 0 | 3 | 33.3 | ||
| tubular cystic RCC | 1 | 0.0 | 1 | 0.0 | 2 | 0.0 |
| multilocular cystic renal cell neoplasia of low malignant potential | 2 | 50.0 | 2 | 50.0 | 2 | 50.0 |
| acquired renal cystic disease associated RCC | 1 | 0.0 | 1 | 0.0 | 1 | 0.0 |
| cystic nephroma/mixed epithelial stromal tumor | 1 | 0.0 | 1 | 0.0 | 1 | 0.0 |
| medullary carcinoma | 0 | 0 | 1 | 0.0 | ||
| reninoma | 0 | 0 | 1 | 100.0 | ||
| n. o. s. | 9 | 22.2 | 7 | 0.0 | 18 | 22.2 |
The lower percentage of VHL mutations in the different tumor types with complete data sets for all 3 exons was regarded as the more “objective” real mutation rate as compared to the results of only 1–2 interpretable exons. For example, the overall VHL mutation frequency decreased from 63.7% (3 exons) to 40.6% (1–2 exons) in all cancers and dropped from 78.3% to 59.3% in clear cell carcinomas. Both chromophobe and papillary renal cell carcinomas showed comparable mutation rates of 19.1% and 18.4%, respectively, with a “corrected” value of 5.2% and 3.1%, respectively, for 3 exons. The majority of kidney tumor subtypes had – at least in some cases – VHL mutations. The number of examined tumors was very low for these four subtypes without detectable mutations: nephroblastoma (n = 4), tubulocystic renal cell carcinoma (n = 1), acquired cystic disease–associated renal cell carcinoma (n = 1), and cystic nephroma/mixed epithelial stromal tumor (n = 1). There were no associations between the mutation rate and the tumor phenotype in the two major histologic subtypes of clear cell (Table 2) and papillary renal cell carcinomas (Table 3).
Table 2: Association of histopathological parameters versus VHL mutations and 3p deletions in the subgroup of clear cell renal cell carcinoma
| VHL mutation analysis | 3p deletion analysis | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1–2 exons analyzed | 3 exons analyzed | n | del. (%) | P | ||||||
| n | mut. (%) | P | n | mut. (%) | P | |||||
| ISUP | 1 | 123 | 82.9 | 0.3106 | 57 | 63.2 | 0.9009 | 246 | 61.0 | 0.4581 |
| 2 | 185 | 79.5 | 93 | 59.1 | 302 | 56.6 | ||||
| 3 | 166 | 74.7 | 98 | 57.1 | 298 | 54.4 | ||||
| 4 | 33 | 72.7 | 21 | 57.1 | 64 | 54.7 | ||||
| Fuhrmann | 1 | 25 | 72.0 | 0.1269 | 15 | 53.3 | 0.6446 | 47 | 63.8 | 0.4702 |
| 2 | 285 | 82.1 | 137 | 62.8 | 494 | 58.7 | ||||
| 3 | 171 | 73.7 | 100 | 55.0 | 305 | 54.1 | ||||
| 4 | 33 | 72.7 | 22 | 59.1 | 71 | 56.3 | ||||
| Thoenes | 1 | 162 | 79.6 | 0.1238 | 83 | 60.2 | 0.6302 | 310 | 60.3 | 0.3801 |
| 2 | 290 | 79.7 | 149 | 60.4 | 489 | 56.0 | ||||
| 3 | 62 | 67.7 | 42 | 52.4 | 118 | 54.2 | ||||
| UICC | 1 | 221 | 77.8 | 0.2904 | 120 | 59.2 | 0.3478 | 394 | 55.1 | 0.5170 |
| 2 | 36 | 69.4 | 22 | 50.0 | 66 | 56.1 | ||||
| 3 | 57 | 86.0 | 30 | 73.3 | 112 | 58.9 | ||||
| 4 | 45 | 77.8 | 24 | 58.3 | 100 | 63.0 | ||||
| pT | 1 | 303 | 78.5 | 0.5410 | 154 | 57.8 | 0.1851 | 534 | 55.8 | 0.414 |
| 2 | 72 | 73.6 | 37 | 48.6 | 113 | 55.8 | ||||
| 3 | 130 | 81.5 | 77 | 68.8 | 252 | 61.9 | ||||
| 4 | 6 | 66.7 | 5 | 60.0 | 13 | 53.8 | ||||
| pN | 0 | 66 | 86.4 | 0.4757 | 35 | 74.3 | 0.6521 | 128 | 55.5 | 0.2800 |
| ≥1 | 14 | 78.6 | 9 | 66.7 | 27 | 66.7 | ||||
| pM | 0 | 67 | 83.6 | 0.4432 | 40 | 72.5 | 0.1983 | 121 | 58.7 | 0.4890 |
| ≥1 | 45 | 77.8 | 23 | 56.5 | 98 | 63.3 | ||||
Table 3: Association of histopathological parameters versus VHL mutations and 3p deletions in the subgroup of papillary renal cell carcinoma
| VHL mutation analysis | 3p deletion analysis | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1-2 exons analyzed | 3 exons analyzed | n | del. (%) | P | ||||||
| n | mut. (%) | P | n | mut. (%) | P | |||||
| ISUP | 1 | 15 | 20.0 | 0.8595 | 13 | 7.7 | 0.396 | 38 | 21.1 | 0.7056 |
| 2 | 25 | 16.0 | 21 | 0.0 | 97 | 15.5 | ||||
| 3 | 27 | 22.2 | 23 | 8.7 | 71 | 18.3 | ||||
| 4 | 1 | 0.0 | 1 | 0.0 | 2 | 0.0 | ||||
| Fuhrmann | 1 | 1 | 0.0 | 0.7131 | 1 | 0.0 | 0.7989 | 2 | 0.0 | 0.823 |
| 2 | 37 | 18.9 | 31 | 3.2 | 133 | 17.3 | ||||
| 3 | 28 | 21.4 | 24 | 8.3 | 69 | 17.4 | ||||
| 4 | 2 | 0.0 | 2 | 0.0 | 4 | 25.0 | ||||
| Thoenes | 1 | 13 | 23.1 | 0.4968 | 11 | 9.1 | 0.7264 | 49 | 18.4 | 0.8205 |
| 2 | 52 | 19.2 | 44 | 4.5 | 146 | 16.4 | ||||
| 3 | 3 | 0.0 | 3 | 0.0 | 13 | 23.1 | ||||
| UICC | 1 | 25 | 12.0 | 0.518 | 22 | 0.0 | 0.1528 | 105 | 17.1 | 0.6210 |
| 2 | 10 | 30.0 | 8 | 12.5 | 22 | 22.7 | ||||
| 3 | 5 | 20.0 | 5 | 20.0 | 10 | 10.0 | ||||
| 4 | 6 | 33.3 | 5 | 20.0 | 14 | 28.6 | ||||
| pT | 1 | 37 | 16.2 | 0.5774 | 31 | 0.0 | 0.1206 | 138 | 16.7 | 0.2838 |
| 2 | 19 | 26.3 | 16 | 12.5 | 43 | 16.3 | ||||
| 3 | 7 | 28.6 | 6 | 16.7 | 19 | 21.1 | ||||
| 4 | 2 | 0.0 | 2 | 0.0 | 3 | 66.7 | ||||
| pN | 0 | 8 | 37.5 | 0.2014 | 7 | 28.6 | 0.2817 | 23 | 0.0 | 0.0036 |
| ≥1 | 2 | 0.0 | 2 | 0.0 | 9 | 33.3 | ||||
| pM | 0 | 7 | 14.3 | 0.6623 | 6 | 0.0 | - | 29 | 0.0 | 0.0028 |
| ≥1 | 45 | 77.8 | 23 | 56.5 | 98 | 63.3 | ||||
VHL deletions (3p25)
The frequency of 3p deletions is summarized in Table 1 for all renal tumors and for the individual histological subtypes. In addition to mutations, clear cell carcinomas also showed the highest percentage of 3p deletions with 57.2% of 918 analyzable cancers, while papillary renal cell carcinoma (17.6%), chromophobe renal cell carcinoma (17.7%) and oncocytoma (11.9%) had fewer deletions. While there was no association between 3p deletion and any unfavorable pathological parameter in clear cell carcinomas (Table 2), 3p deletions were significantly linked to the presence of lymph node and distant metastasis (<0.005 each) in papillary renal cell carcinomas (Table 3).
Types of VHL alterations in the different renal cancer subtypes
Highly variable ratios of VHL mutations and 3p deletions were seen in the different histologic subtypes of kidney tumors, as shown in Figure 1. Clear cell RCC (n = 452) not only showed highest rates of VHL alterations exceeding 80% of all cases but also high overlap of both mutation and deletion (around 50%). In contrast, VHL alterations did not exceed 30% in the other renal subtypes showing either small or no mutation/deletion overlap in oncocytoma (n = 43) or papillary renal cell carcinoma (n = 58), respectively, and a strikingly low mutation rate of approx. 10% in chromophobe renal cell carcinomas. Since sequencing analysis was not repeated for validation in cases with mutations, the number of insertions and deletions, representing the more reliable fraction of the detected mutations, were separately compared to all mutations (Figure 2, upper panel). Since their relationship was quite consistent for all 3 analyzed exons, an overall homogenous mutational pattern without hot spot regions was supported (Figure 2, lower panel).
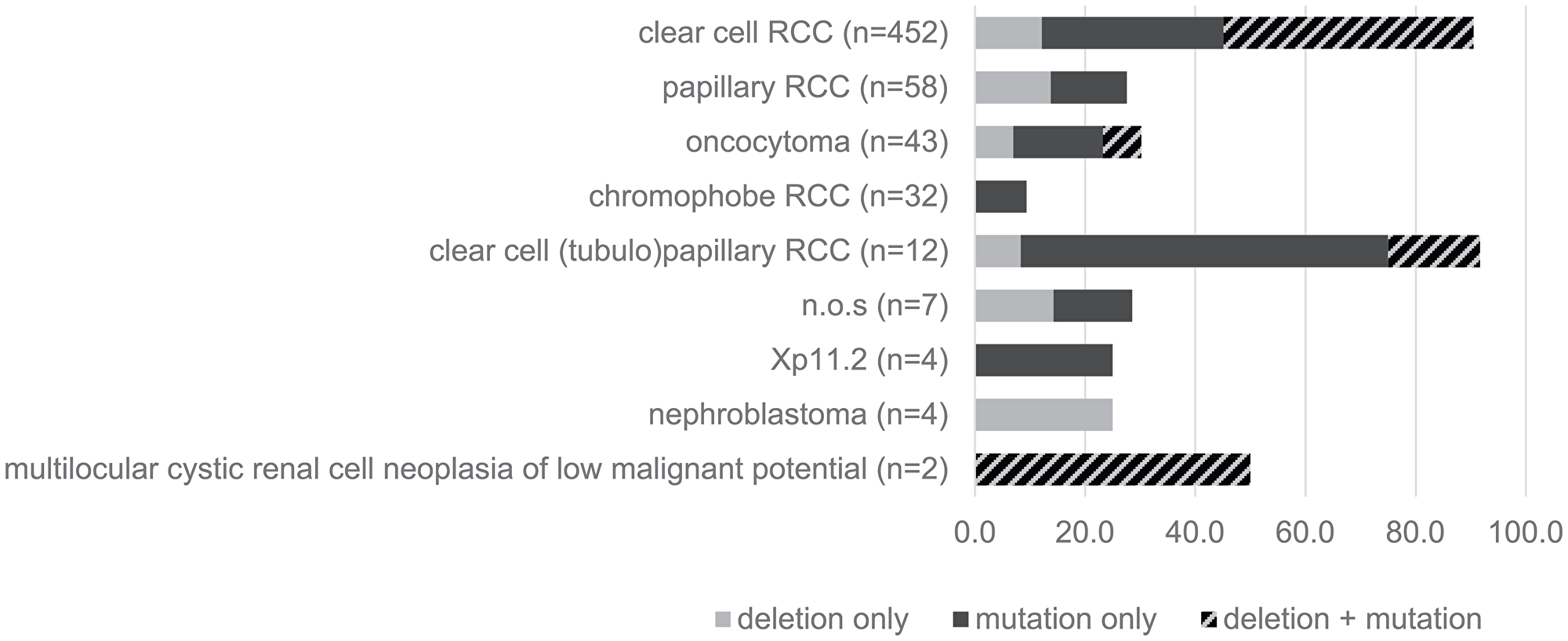
Figure 1: Type of VHL-alterations in different histologic subtypes of renal cell carcinomas (RCC).
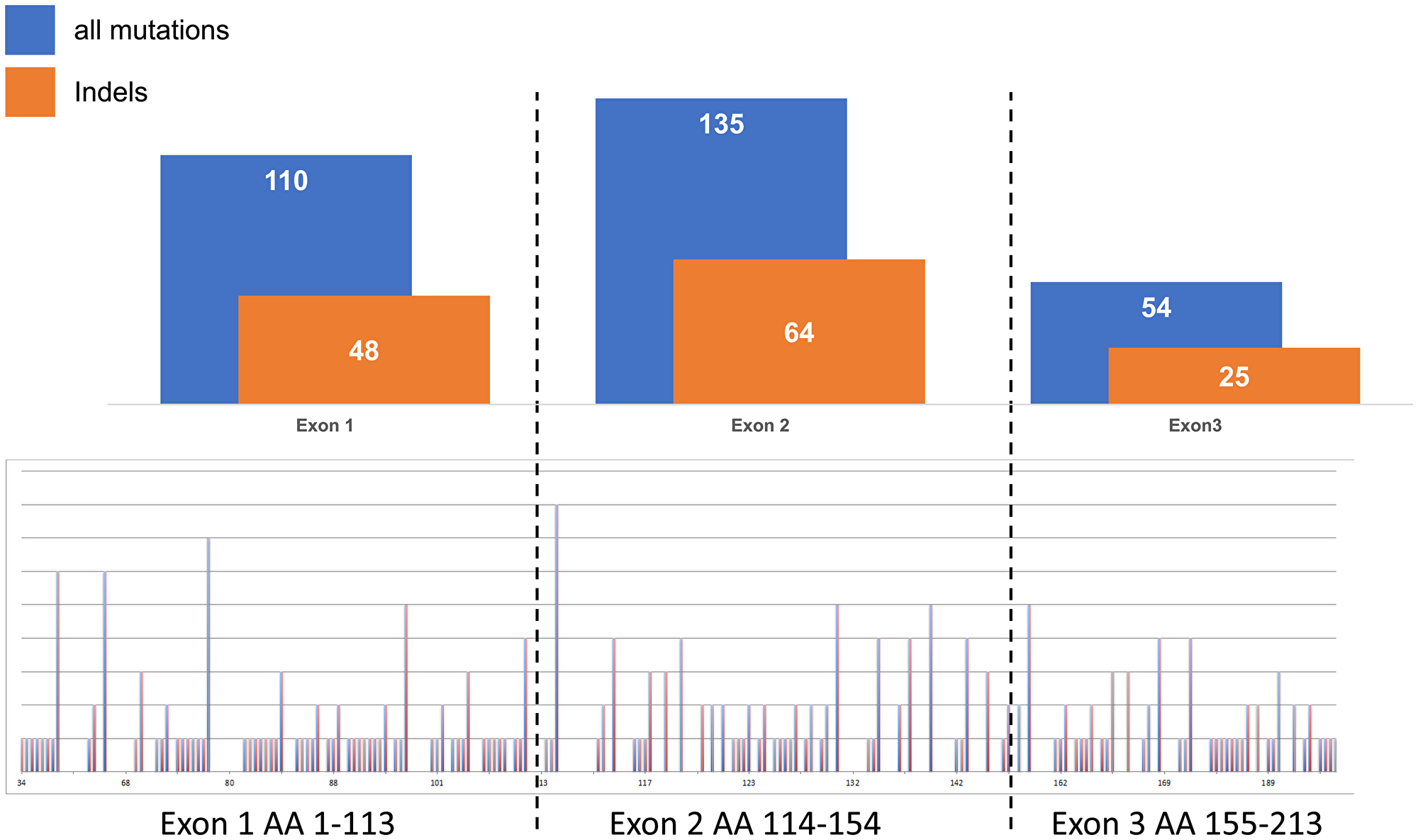
Figure 2: Relationship of all mutations (point and indels) versus insertions and/or deletions (indels) in exons 1–3 in clear cell renal cell carcinomas.
Association of VHL alterations with prognosis
In clear cell carcinoma, VHL alterations were not associated with patient prognosis regarding overall survival, relapse-free survival or cancer specific death (Figure 3A–3F). In addition, VHL mutations and 3p deletions (Figure 5B) did not show prognostic relevance for relapse-free survival in chromophobe renal cell carcinoma, based on the low number of cases. In papillary renal cell carcinoma, 3p deletions (Figure 4D–4F) but not VHL mutations (Figure 4A–4C) were significantly associated with both relapse-free survival and cancer specific death (p < 0.05, each).
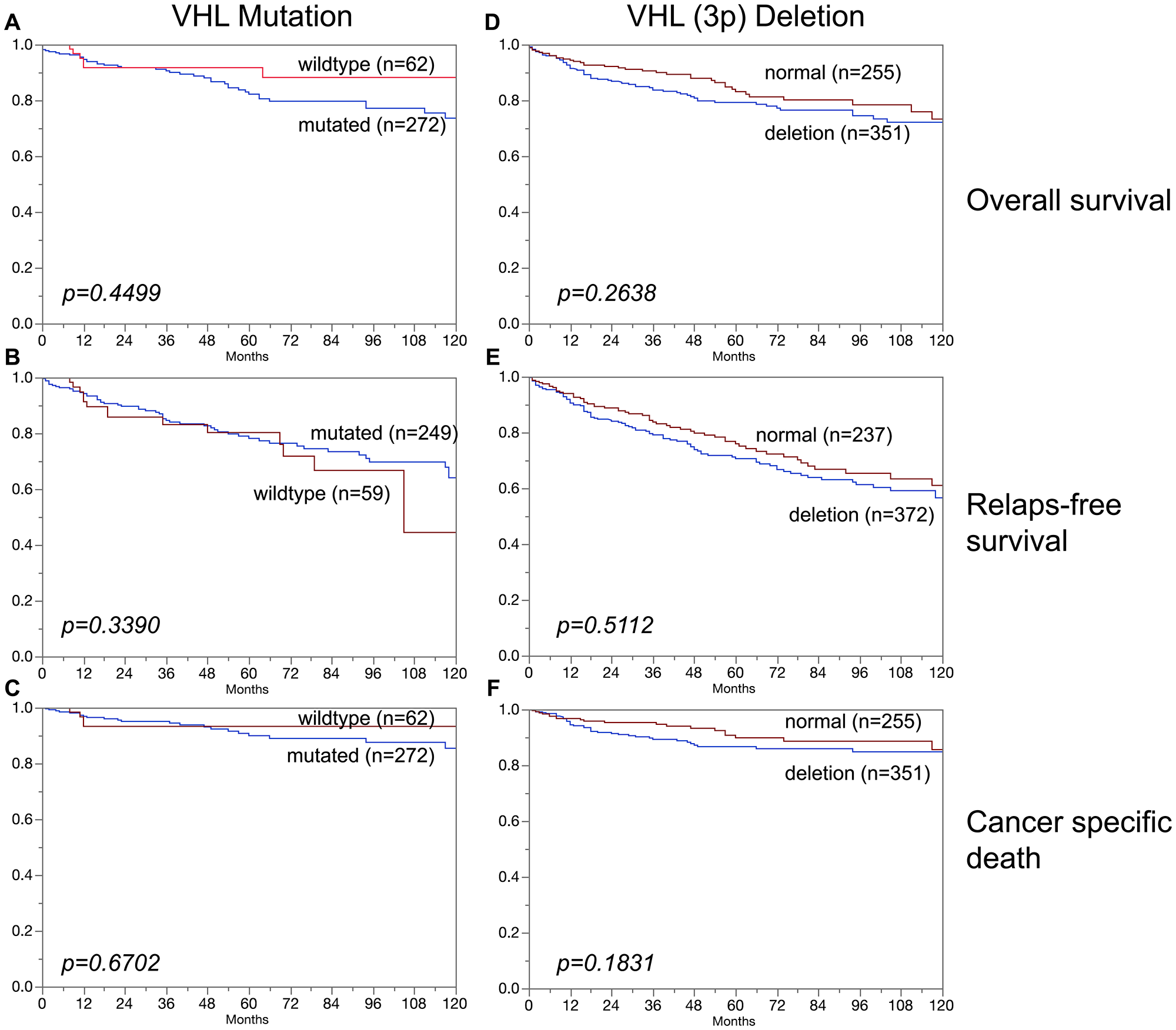
Figure 3: Association of (A–C) VHL mutation and (D–F) VHL deletions (3p) in clear cell carcinoma and overall survival (A, D), relapse-free survival (B, E), and cancer specific death (C, F).
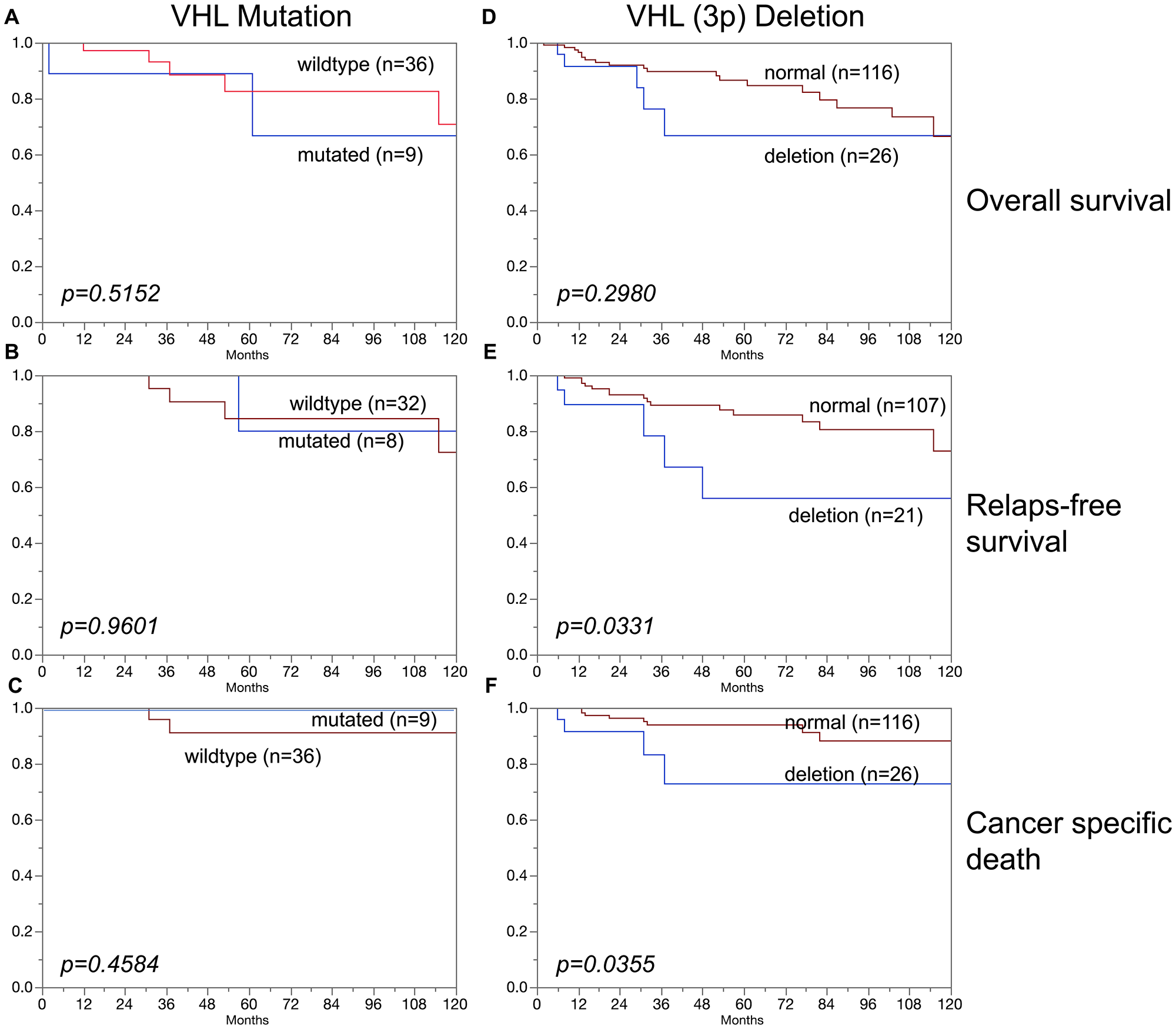
Figure 4: Association of (A–C) VHL mutations and (D–F) VHL deletions (3p) in papillary carcinoma and overall survival (A, D), relapse-free survival (B, E), and cancer specific death (C, F).
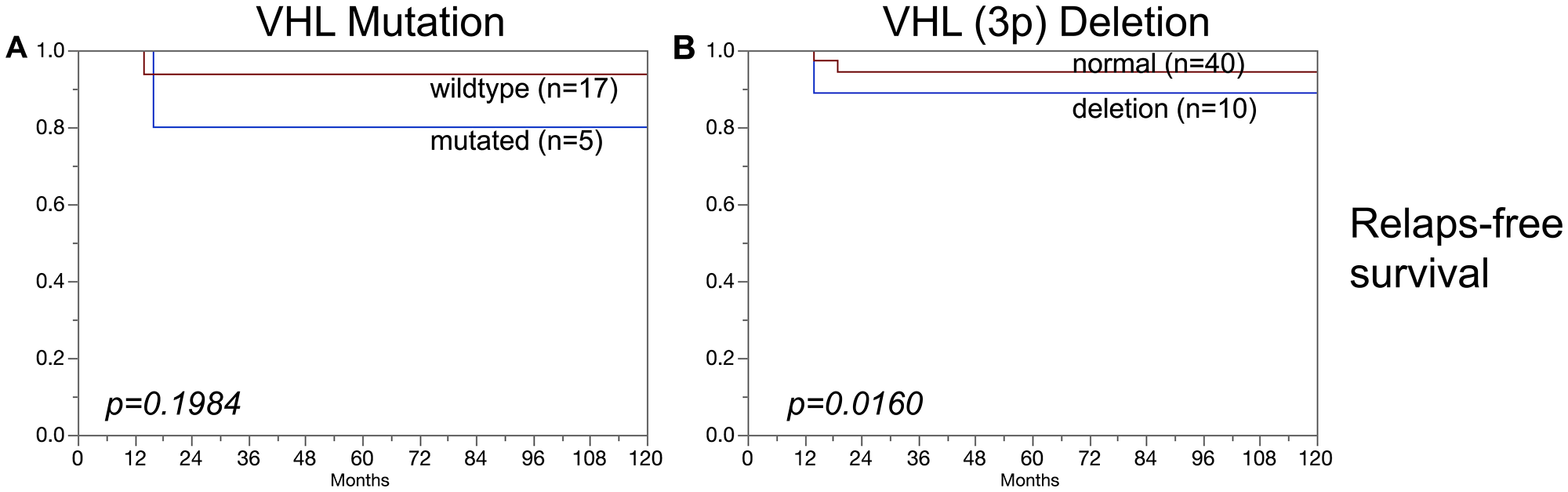
Figure 5: Association of (A) VHL mutations and (B) VHL deletions (3p) in chromophobe renal cell carcinomas and relapse-free survival.
DISCUSSION
A total of 1,805 formalin fixed paraffin embedded tumors collected between 1993 and 2015 were sequenced for all three VHL exons in this study. The quality of tissue (and DNA) preservation depends on the quality and duration of fixation. For example, it is necessary that the quantity of formalin exceeds the quantity of submitted tissue by a factor of 10 in transportation vessels. Because such parameters can hardly be standardized in clinical practice, it is not surprising that sequencing was not always successful. Since a mutation can be diagnosed if only one exon is successfully analyzed and this exon contains a mutation, “mutation detection” is easier than “mutation exclusion”. To exclude VHL mutation, successful analysis of all three exons is required. The mutation frequency of 59.3% found in these 275 clear cell carcinomas with complete analysis of all 3 VHL exons is likely to represent the “real” prevalence of VHL mutations in our patient cohort. This frequency ranks in the middle range of frequencies described in numerous earlier publications (Figure 6) [12–21, 4, 22–25]. Deviations between these studies might be due to cohort size or technical issues. The two studies reporting mutation rates lower than 30% studied particularly small patient cohorts (n = 12 and n = 56) [26, 12]. As in other studies, single-base point mutations were more frequent than indels, affecting several bases, and were found in all exons [4, 11].
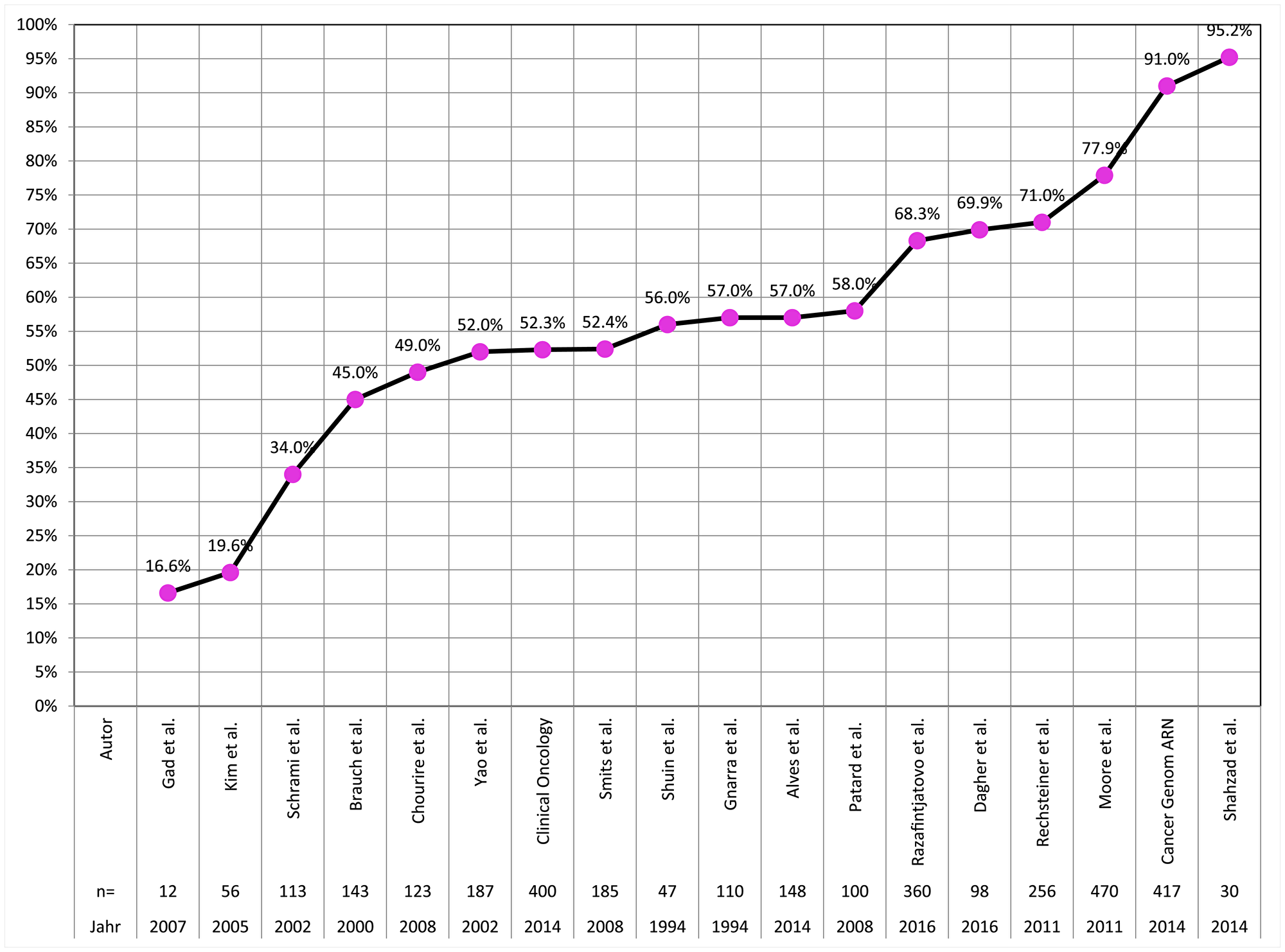
Figure 6: Frequency of VHL mutations in clear cell carcinoma in the literature.
The large number of cancers in combination with a thorough review of all tumors according to the recently published WHO criteria enabled us to collect information on VHL mutations in classical kidney tumor types and also in newly defined entities. These data demonstrate that VHL mutations predominantly occur in clear cell and clear cell (tubulo) papillary carcinoma but can – perhaps at lower frequencies – also be found in most other kidney tumor entities. In this study, we detected VHL mutations in all entities for which at least 10 cases could be successfully analyzed including 11 of 49 oncocytomas and in 7 of 38 chromophobe carcinomas (Table 1). It is of note that most others have not found VHL mutations in chromophobe carcinoma and oncocytoma [14, 27, 28], which may be explained by the low numbers of analyzed cases (n = 2 – 17). However, individual cases of reported VHL mutations can be found also for these entities in the literature (1 of 6 oncocytomas, 2 of 7 chromophobe carcinomas) [11]. We have validated our diagnoses of oncocytoma and/or chromophobe carcinoma in all mutated cases by histological slide review and – where necessary – by immunohistochemical analyses. Our finding of VHL mutations in papillary carcinoma matches quite well with previous reports of smaller cohorts (3/28 and 2/30 cases) [14, 11]. With respect of newly defined entities, we found VHL mutations in 1 of 2 multilocular cystic renal cell neoplasias of low malignant potential and in 1 of 4 Xp11.2 translocation carcinomas. Obviously – due to the low number of analyzed cases – we cannot exclude mutations in tubulocystic carcinoma, cystic nephroma, acquired cystic disease–associated renal cell carcinoma or nephroblastoma where we failed to find mutations in 1 to 4 analyzed specimens.
Deletion of the short arm of chromosome 3 (3p) is another common and well-known mechanism for VHL inactivation. It is not surprising that the distribution of 3p deletion among kidney cancer entities was comparable to the frequencies of VHL deletion. The complete lack of an association between 3p deletions and VHL mutations in our 452 clear cell carcinomas with FISH and sequencing data is unexpected, however (Figure 7). Other authors, employing LOH or CGH analysis, have earlier suggested that mutation and deletion may be statistically linked [27, 18]. The complete independency of VHL deletion and VHL mutation is consistent with a model suggesting that VHL is (almost) always completely inactivated in clear cell kidney cancer by a combination of multiple mechanisms including deletion, mutation, methylation and possibly other pathways. In such a scenario, our data suggest that there is no specific advantage for tumor cells to use the specific combination of deletion and mutation for knocking out the VHL gene.
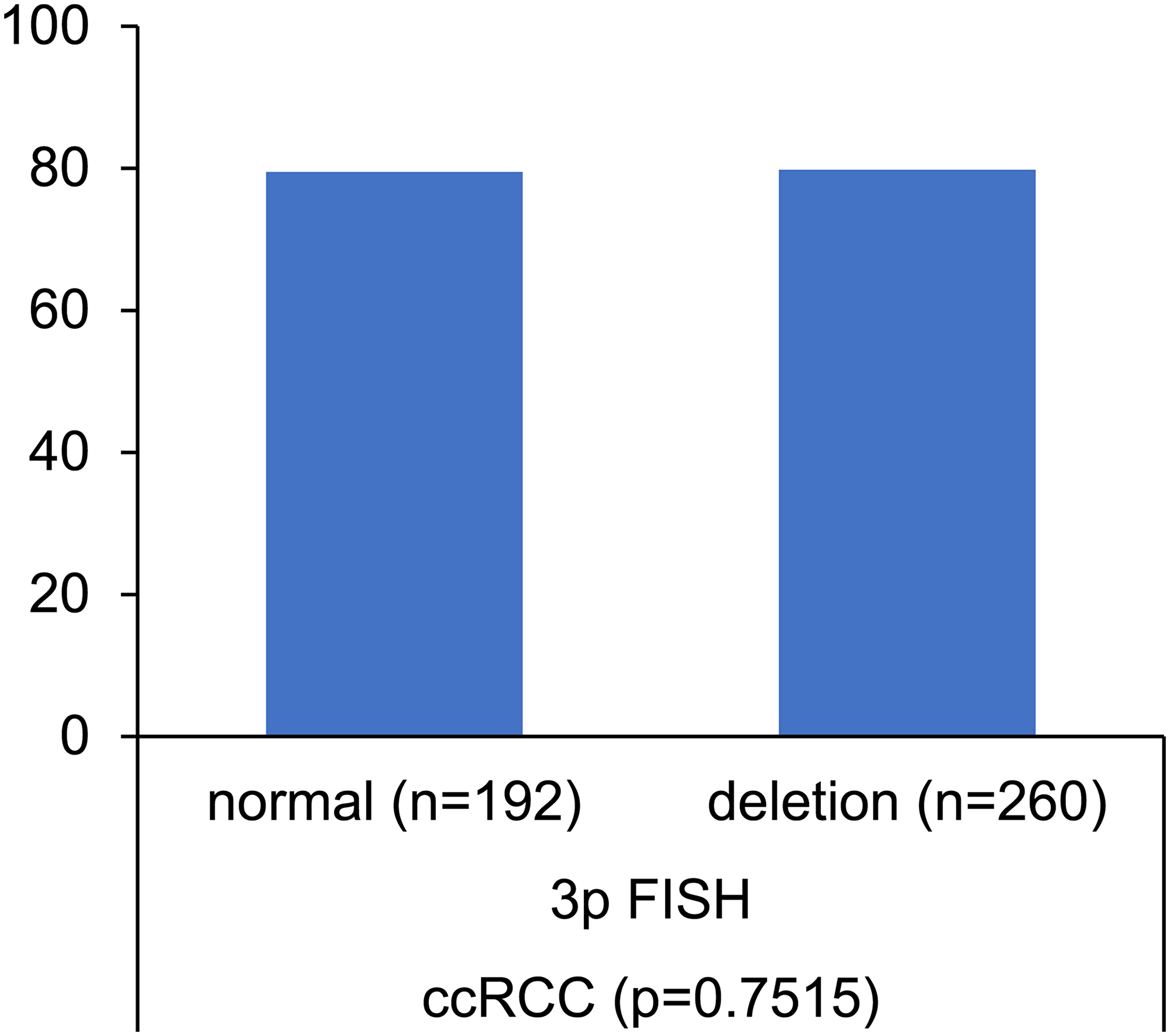
Figure 7: Association between VHL mutation and VHL (3p) deletion. Norm = no 3p deletion.
No association was found in this study between VHL aberrations (mutation/deletion) and patient prognosis. This also hold true for the subgroups of clear cell and chromophobe carcinomas. An overwhelming majority of earlier studies had come to similar conclusions for clear cell carcinomas [29, 30]. However, one other study had suggested a prognostic role of 3p deletions in chromophobe carcinomas [31]. This is not confirmed by our data, neither for mutations nor for deletions (Figures 3, 5). It is of note that the cohorts of both Velickovic et al. (n = 16) and of us (n = 101) are too small to strongly suggest a prognostic role of VHL aberrations in chromophobe carcinomas. The cohort of analyzed papillary carcinomas was also small in our study, but evidence for a possible prognostic role of 3p deletions but not of VHL mutations for both relapse-free survival and cancer specific death arises from this analysis (Figure 4). This is supported by Velickovic et al., reporting 3p deletions in 59% of 29 papillary renal cell carcinomas significantly associated with tumor histology and patient outcome [31].
Other studies analyzing 22–37 papillary renal cell carcinoma described 3p deletions as absent or representing rare events [32–34]. Of note, the WHO/ISUP kidney cancer grading was also unrelated to presence of VHL mutations and deletions.
In summary, our data show that VHL deletions and mutations occur frequently in kidney tumors and are largely unrelated to clinical or pathological parameters. They preferentially occur in clear cell cancers but they can also be found in virtually every other kidney cancer entity. This is important because it suggests that therapies targeting the VHL pathway may not be limited to clear cell kidney cancer. It also precludes the diagnostic use of VHL aberrations for classification of kidney tumors.
Materials and Methods
Patients and TMA construction
A total of 1,805 tissue specimens were available from patients with renal cell tumors, undergoing surgery between 1994–2015 at the Department of Urology, University Medical Center Hamburg-Eppendorf. Research using pseudonymized human left-over tissue samples from routine diagnosis was performed in compliance with the Helsinki Declaration, and is covered by §12a of the Hamburgisches Krankenhausgesetz (HmbKHG). Manufacturing and usage of tissue microarrays (TMAs) for research purposes has been approved by the Review Board of the Ärztekammer Hamburg (WF-049/09). The cohort mainly comprised of 1,176 (65%) patients with clear cell renal carcinoma, 270 (15%) with papillary renal cell carcinoma, 101 (6%) with chromophobe renal cell carcinoma and 149 (8%) with oncocytoma. A detailed overview of the patient cohort is given in Table 4. The mean age of the patients at date of surgery was 62 years, ranging between 1 and 94 years. Follow-up data were available from 1,171 patients with a median follow-up of 61.8 months (range: 0.3–105.4 months). Routinely formalin-fixed and paraffin embedded tissue specimens were analyzed. A tissue microarray (TMA) was constructed from all samples. The TMA manufacturing procedure was described earlier in detail [35, 36]. In brief, one 0.6 mm core per patient was taken from a representative tissue block. All 1,805 tissue cores were distributed among 4 TMA blocks, each containing 385 to 510 tumor samples.
Table 4: Pathological and clinical data of the arrayed renal cancers
| Study cohort on TMA | |
| (n = 1,805) | |
| Follow-up (months) | |
| Available (n) | 1,171 |
| Mean (months) | 47 |
| Median (months) | 61.8 |
| Age (y) | |
| <50 | 267 |
| 50–70 | 1,011 |
| >70 | 527 |
| Histology | |
| Clear cell | 1,176 |
| Chromophobic | 101 |
| Papillary | 270 |
| Oncocytoma | 149 |
| UICC stage | |
| I | 732 |
| II | 131 |
| III | 175 |
| IV | 158 |
| pT category | |
| pT1 | 995 |
| pT2 | 223 |
| pT3-pT4 | 408 |
| ISUP grade | |
| 1 | 398 |
| 2 | 537 |
| 3 | 469 |
| 4 | 100 |
| Fuhrman grade | |
| 1 | 72 |
| 2 | 851 |
| 3 | 480 |
| 4 | 110 |
| Thoenes grade | |
| 1 | 497 |
| 2 | 839 |
| 3 | 177 |
| pN category | |
| pN0 | 232 |
| pN+ | 59 |
| pM category | |
| pM0 | 220 |
| pM+ | 148 |
Von Hippel-Lindau (VHL) polymerase chain reaction and sequencing
Following deparaffinization of the TMA sections for 30 min, DNA was isolated using the QIAmp DNA Micro kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. Polymerase-chain-reaction (PCR) was performed in a 96-well plate format using 100 ng DNA in a total reaction volume of 25 µl. Three VHL-exons were amplified using the specific primers specified in Supplementary Table 1. Subsequent to PCR reaction, the service laboratory of the Eurofins-Genomics institute (Eversberg, Germany) performed sequence analysis of all PCR products using the didesoxy-method [37]. Graphical depiction and analysis of the obtained sequences were conducted using Geneious software 9.1.6 (Biomatters Ltd., New Zealand).
3p deletion analysis
Chromosome 3p deletion was assessed by fluorescence in-situ hybridization analysis (FISH) using a commercial VHL-locus specific FISH probe (Zytolight SPEC VHL/CEN 3 Dual Color Probe, ZytoVision). Prior to hybridization, 4 µm TMA sections were deparaffinized, air-dried and dehydrated in an ethanol dilution series followed by denaturation for 5 min at 74°C in 70% formamide-2xSSC solution. After overnight hybridization at 37°C in a humidified chamber, slides were washed and counterstained with 0.2 µmol/L of 4′,6-diamidino-2-phenylindole in an antifade solution. Each spot was evaluated, and the predominant signal numbers were recorded for each FISH probe. Deletion of VHL was defined as the presence of fewer VHL signals than centromere 3 probe signals in at least 60% of tumor cell nuclei.
Statistics
Statistical calculations were performed using JMP 11 software (SAS Institute Inc., NC, USA). Contingency tables were calculated with the Chi2 test to search for associations between molecular parameters and tumor phenotype. Survival curves were calculated according to Kaplan-Meier. The log-Rank test was applied to detect significant differences between groups. Cox proportional hazards regression analysis was conducted to assess statistical independence and significance between clinico-pathological and molecular parameters.
ACKNOWLEDGMENTS
We thank Janett Lüttgens, Sünje Seekamp and Inge Brandt for excellent technical assistance.
Ethics statement
The ethics committee of the Hamburger Ärtzekammer approved the study (WF-049/09). In compliance with the local law (HmbKHG, 12a) informed consent was not obtained.
CONFLICTS OF INTEREST
All authors declare that no competing interests exist.
References
1. Maxwell PH, Wiesener MS, Chang GW, Clifford SC, Vaux EC, Cockman ME, Wykoff CC, Pugh CW, Maher ER, Ratcliffe PJ. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature. 1999; 399:271–275. https://doi.org/10.1038/20459. [PubMed].
2. Iliopoulos O, Kibel A, Gray S, Kaelin WG Jr. Tumour suppression by the human von Hippel-Lindau gene product. Nat Med. 1995; 1:822–826. https://doi.org/10.1038/nm0895-822. [PubMed].
3. Ohh M, Park CW, Ivan M, Hoffman MA, Kim TY, Huang LE, Pavletich N, Chau V, Kaelin WG. Ubiquitination of hypoxia-inducible factor requires direct binding to the beta-domain of the von Hippel-Lindau protein. Nat Cell Biol. 2000; 2:423–427. https://doi.org/10.1038/35017054. [PubMed].
4. Razafinjatovo C, Bihr S, Mischo A, Vogl U, Schmidinger M, Moch H, Schraml P. Characterization of VHL missense mutations in sporadic clear cell renal cell carcinoma: hotspots, affected binding domains, functional impact on pVHL and therapeutic relevance. BMC Cancer. 2016; 16:638. https://doi.org/10.1186/s12885-016-2688-0. [PubMed].
5. Yang L, Zhao Z, Zhao S, Chen C, Cong X, Li Z, Ren M. The Clinicopathological Significance of Epigenetic Silencing of VHL Promoter and Renal Cell Carcinoma: A Meta-Analysis. Cell Physiol Biochem. 2016; 40:1465–1472. https://doi.org/10.1159/000453198. [PubMed].
6. Favazza L, Chitale DA, Barod R, Rogers CG, Kalyana-Sundaram S, Palanisamy N, Gupta NS, Williamson SR. Renal cell tumors with clear cell histology and intact VHL and chromosome 3p: a histological review of tumors from the Cancer Genome Atlas database. Mod Pathol. 2017; 30:1603–1612. https://doi.org/10.1038/modpathol.2017.72. [PubMed].
7. Frew IJ, Moch H. A clearer view of the molecular complexity of clear cell renal cell carcinoma. Annu Rev Pathol. 2015; 10:263–289. https://doi.org/10.1146/annurev-pathol-012414-040306. [PubMed].
8. Klatte T, Pantuck AJ, Said JW, Seligson DB, Rao NP, LaRochelle JC, Shuch B, Zisman A, Kabbinavar FF, Belldegrun AS. Cytogenetic and molecular tumor profiling for type 1 and type 2 papillary renal cell carcinoma. Clin Cancer Res. 2009; 15:1162–1169. https://doi.org/10.1158/1078-0432.CCR-08-1229. [PubMed].
9. Marsaud A, Dadone B, Ambrosetti D, Baudoin C, Chamorey E, Rouleau E, Lefol C, Roussel JF, Fabas T, Cristofari G, Carpentier X, Michiels JF, Amiel J, et al. Dismantling papillary renal cell carcinoma classification: The heterogeneity of genetic profiles suggests several independent diseases. Genes Chromosomes Cancer. 2015; 54:369–382. https://doi.org/10.1002/gcc.22248. [PubMed].
10. Shi SS, Shen Q, Xia QY, Tu P, Shi QL, Zhou XJ, Rao Q. Clear cell papillary renal cell carcinoma: a clinicopathological study emphasizing ultrastructural features and cytogenetic heterogeneity. Int J Clin Exp Pathol. 2013; 6:2936–2942. [PubMed].
11. van Houwelingen KP, van Dijk BA, Hulsbergen-van de Kaa CA, Schouten LJ, Gorissen HJ, Schalken JA, van den Brandt PA, Oosterwijk E. Prevalence of von Hippel-Lindau gene mutations in sporadic renal cell carcinoma: results from The Netherlands cohort study. BMC Cancer. 2005; 5:57. https://doi.org/10.1186/1471-2407-5-57. [PubMed].
12. Kim JH, Jung CW, Cho YH, Lee J, Lee SH, Kim HY, Park J, Park JO, Kim K, Kim WS, Park YS, Im YH, Kang WK, et al. Somatic VHL alteration and its impact on prognosis in patients with clear cell renal cell carcinoma. Oncol Rep. 2005; 13:859–864. https://doi.org/10.3892/or.13.5.859. [PubMed].
13. Schraml P, Struckmann K, Hatz F, Sonnet S, Kully C, Gasser T, Sauter G, Mihatsch MJ, Moch H. VHL mutations and their correlation with tumour cell proliferation, microvessel density, and patient prognosis in clear cell renal cell carcinoma. J Pathol. 2002; 196:186–193. https://doi.org/10.1002/path.1034. [PubMed].
14. Brauch H, Weirich G, Brieger J, Glavac D, Rodl H, Eichinger M, Feurer M, Weidt E, Puranakanitstha C, Neuhaus C, Pomer S, Brenner W, Schirmacher P, et al. VHL alterations in human clear cell renal cell carcinoma: association with advanced tumor stage and a novel hot spot mutation. Cancer Res. 2000; 60:1942–1948. [PubMed].
15. Choueiri TK, Vaziri SA, Jaeger E, Elson P, Wood L, Bhalla IP, Small EJ, Weinberg V, Sein N, Simko J, Golshayan AR, Sercia L, Zhou M, et al. von Hippel-Lindau gene status and response to vascular endothelial growth factor targeted therapy for metastatic clear cell renal cell carcinoma. J Urol. 2008; 180:860–865, discussion 865–866. https://doi.org/10.1016/j.juro.2008.05.015. [PubMed].
16. Yao M, Yoshida M, Kishida T, Nakaigawa N, Baba M, Kobayashi K, Miura T, Moriyama M, Nagashima Y, Nakatani Y, Kubota Y, Kondo K. VHL tumor suppressor gene alterations associated with good prognosis in sporadic clear-cell renal carcinoma. J Natl Cancer Inst. 2002; 94:1569–1575. https://doi.org/10.1093/jnci/94.20.1569. [PubMed].
17. Smits KM, Schouten LJ, van Dijk BA, Hulsbergen-van de Kaa CA, Wouters KA, Oosterwijk E, van Engeland M, van den Brandt PA. Genetic and epigenetic alterations in the von hippel-lindau gene: the influence on renal cancer prognosis. Clin Cancer Res. 2008; 14:782–787. https://doi.org/10.1158/1078-0432.CCR-07-1753. [PubMed].
18. Shuin T, Kondo K, Torigoe S, Kishida T, Kubota Y, Hosaka M, Nagashima Y, Kitamura H, Latif F, Zbar B, Lerman MI, Yao M. Frequent somatic mutations and loss of heterozygosity of the von Hippel-Lindau tumor suppressor gene in primary human renal cell carcinomas. Cancer Res. 1994; 54:2852–2855. [PubMed].
19. Gnarra JR, Tory K, Weng Y, Schmidt L, Wei MH, Li H, Latif F, Liu S, Chen F, Duh FM, Lubensky I, Duan DR, Florence C, et al. Mutations of the VHL tumour suppressor gene in renal carcinoma. Nat Genet. 1994; 7:85–90. https://doi.org/10.1038/ng0594-85. [PubMed].
20. Alves MR, Carneiro FC, Lavorato-Rocha AM, da Costa WH, da Cunha IW, de Cassio Zequi S, Guimaraes GC, Soares FA, Carraro DM, Rocha RM. Mutational status of VHL gene and its clinical importance in renal clear cell carcinoma. Virchows Arch. 2014; 465:321–330. https://doi.org/10.1007/s00428-014-1629-z. [PubMed].
21. Patard JJ, Fergelot P, Karakiewicz PI, Klatte T, Trinh QD, Rioux-Leclercq N, Said JW, Belldegrun AS, Pantuck AJ. Low CAIX expression and absence of VHL gene mutation are associated with tumor aggressiveness and poor survival of clear cell renal cell carcinoma. Int J Cancer. 2008; 123:395–400. https://doi.org/10.1002/ijc.23496. [PubMed].
22. Dagher J, Kammerer-Jacquet SF, Brunot A, Pladys A, Patard JJ, Bensalah K, Perrin C, Verhoest G, Mosser J, Lespagnol A, Vigneau C, Dugay F, Belaud-Rotureau MA, et al. Wild-type VHL Clear Cell Renal Cell Carcinomas Are a Distinct Clinical and Histologic Entity: A 10-Year Follow-up. Eur Urol Focus. 2016; 1:284–290. https://doi.org/10.1016/j.euf.2015.06.001. [PubMed].
23. Rechsteiner MP, von Teichman A, Nowicka A, Sulser T, Schraml P, Moch H. VHL gene mutations and their effects on hypoxia inducible factor HIFalpha: identification of potential driver and passenger mutations. Cancer Res. 2011; 71:5500–5511. https://doi.org/10.1158/0008-5472.CAN-11-0757. [PubMed].
24. Moore LE, Nickerson ML, Brennan P, Toro JR, Jaeger E, Rinsky J, Han SS, Zaridze D, Matveev V, Janout V, Kollarova H, Bencko V, Navratilova M, et al. Von Hippel-Lindau (VHL) inactivation in sporadic clear cell renal cancer: associations with germline VHL polymorphisms and etiologic risk factors. PLoS Genet. 2011; 7:e1002312. https://doi.org/10.1371/journal.pgen.1002312. [PubMed].
25. Shahzad H, Kehar SI, Ali S, Tariq N. Expression of Von Hippel - Lindau (VHL) gene mutation in diagnosed cases of renal cell carcinoma. Pak J Med Sci. 2014; 30:880–885. https://doi.org/10.12669/pjms.304.4733. [PubMed].
26. Gad S, Sultan-Amar V, Meric JB, Izzedine H, Khayat D, Richard S, Rixe O. Somatic von Hippel-Lindau (VHL) gene analysis and clinical outcome under antiangiogenic treatment in metastatic renal carcinoma: preliminary results. Target Oncol. 2007; 2:3–6. https://doi.org/10.1007/s11523-006-0039-9.
27. Foster K, Prowse A, van den Berg A, Fleming S, Hulsbeek MM, Crossey PA, Richards FM, Cairns P, Affara NA, Ferguson-Smith MA, Buys CH, Maher ER. Somatic mutations of the von Hippel-Lindau disease tumour suppressor gene in non-familial clear cell renal carcinoma. Hum Mol Genet. 1994; 3:2169–2173. https://doi.org/10.1093/hmg/3.12.2169. [PubMed].
28. Zhuang Z, Gnarra JR, Dudley CF, Zbar B, Linehan WM, Lubensky IA. Detection of von Hippel-Lindau disease gene mutations in paraffin-embedded sporadic renal cell carcinoma specimens. Mod Pathol. 1996; 9:838–842. [PubMed].
29. Banks RE, Tirukonda P, Taylor C, Hornigold N, Astuti D, Cohen D, Maher ER, Stanley AJ, Harnden P, Joyce A, Knowles M, Selby PJ. Genetic and epigenetic analysis of von Hippel-Lindau (VHL) gene alterations and relationship with clinical variables in sporadic renal cancer. Cancer Res. 2006; 66:2000–2011. https://doi.org/10.1158/0008-5472.CAN-05-3074. [PubMed].
30. Kondo K, Yao M, Yoshida M, Kishida T, Shuin T, Miura T, Moriyama M, Kobayashi K, Sakai N, Kaneko S, Kawakami S, Baba M, Nakaigawa N, et al. Comprehensive mutational analysis of the VHL gene in sporadic renal cell carcinoma: relationship to clinicopathological parameters. Genes Chromosomes Cancer. 2002; 34:58–68. https://doi.org/10.1002/gcc.10054. [PubMed].
31. Velickovic M, Delahunt B, Storkel S, Grebem SK. VHL and FHIT locus loss of heterozygosity is common in all renal cancer morphotypes but differs in pattern and prognostic significance. Cancer Res. 2001; 61:4815–4819. [PubMed].
32. Kovacs G, Fuzesi L, Emanual A, Kung HF. Cytogenetics of papillary renal cell tumors. Genes Chromosomes Cancer. 1991; 3:249–255. https://doi.org/10.1002/gcc.2870030403. [PubMed].
33. Alimov A, Kost-Alimova M, Liu J, Li C, Bergerheim U, Imreh S, Klein G, Zabarovsky ER. Combined LOH/CGH analysis proves the existence of interstitial 3p deletions in renal cell carcinoma. Oncogene. 2000; 19:1392–1399. https://doi.org/10.1038/sj.onc.1203449. [PubMed].
34. Rohan SM, Xiao Y, Liang Y, Dudas ME, Al-Ahmadie HA, Fine SW, Gopalan A, Reuter VE, Rosenblum MK, Russo P, Tickoo SK. Clear-cell papillary renal cell carcinoma: molecular and immunohistochemical analysis with emphasis on the von Hippel-Lindau gene and hypoxia-inducible factor pathway-related proteins. Mod Pathol. 2011; 24:1207–1220. https://doi.org/10.1038/modpathol.2011.80. [PubMed].
35. Kononen J, Bubendorf L, Kallioniemi A, Barlund M, Schraml P, Leighton S, Torhorst J, Mihatsch MJ, Sauter G, Kallioniemi OP. Tissue microarrays for high-throughput molecular profiling of tumor specimens. Nat Med. 1998; 4:844–847. https://doi.org/10.1038/nm0798-844. [PubMed].
36. Mirlacher M, Simon R. Recipient block TMA technique. Methods Mol Biol. 2010; 664:37–44. https://doi.org/10.1007/978-1-60761-806-5_4. [PubMed].
37. Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977; 74:5463–5467. https://doi.org/10.1073/pnas.74.12.5463. [PubMed].

