Introduction
Peripheral nervous system plays an important role in neoplastic invasion. The glial cells present in this system, specifically Schwann cells, have been associated with the process known as perineural invasion. Perineural invasion occurs mainly in pancreas, prostate, breast, lung and head and neck cancers; and causes morbidity of patient through pain and paralysis, associated with a high risk of local recurrence and decreasing survival patients rates [1–3]. Understanding how cancer invades nerves is an essential step in developing more effective treatment strategies. The main questions are how the neoplastic cells interact with the cells of the nervous system and how they acquire mobile and invasive characteristics from these interactions.
Schwann cells performs several roles, depends on their ability to reversibly dedifferentiate and re-differentiate into subtypes with different phenotypes. After the nerve damage, Schwann cells become dedifferentiated, lose their capacity to myelinate and, promote neuronal orientation during repair [3]. This is accompanied by the reexpression of proteins lost during the myelinizing differentiation program, such as the glial fibrillary acidic protein (GFAP) and the neural cell adhesion molecule 1 (NCAM1) [1, 4]. Paracrine signaling has been implicated in perineural invasion, with factors secreted by nerves, including glial cell line derived neurotrophic factor (GDNF), increasing the invasion of tumor cells along the nerves [5, 6].
Schwann cells were identified in the tumor surrounding area prior to initiate the invasion process underlying connective tissue. These cells promote cancer invasion through direct contact, while paracrine signaling and matrix remodeling are not sufficient to proceed [4]. These cells stimulate tumor cells at the cell-cell contact sites and promote detachment and spread of individual cancer cells from neighboring tissue. Physical contact between Schwann cells and tumor cells is necessary to potentiate the process of tumor invasion, by induction of specific chemokines, such as CX3CL1 and its receptor CX3CL1 [2, 7, 8]. Schwann cells are also able to degrade the extracellular matrix, forming tunnels or bands coated with laminin. Tumor cells use these structures to migrate during tissue invasion. These activities strongly promote tumor spread and are dependent on NCAM1 expression by Schwann cells [2, 3, 7, 8].
Considering the intertwined structure of signaling, regulatory, and metabolic processes within a cell, we employed a genome-scale biomolecular network. Accordingly, a meta-analysis of Schwann cells associated transcriptomic datasets was performed, and the core information on differentially expressed genes (DEGs) was obtained by statistical analyses. Gene set over-representation analyses was performed on core DEGs to identify significantly functional and pathway enrichment analysis. DEGs were further integrated with genome-scale human biomolecular networks. miRNAs were proposed by the reconstruction of a transcriptional and post-transcriptional regulatory network. Moreover, microarray-based transcriptome profiling was performed, and the prognostic power of selected dedifferentiated Schwann cell biomolecules was predicted. Consequently, this systematic study reports candidate biomolecules that can be considered as biomarkers or potential targets for further experimental and clinical trials for perineural invasion risk in lung cancer patients.
Results
Identification of differentially expressed genes, gene ontology enrichment, and functional classification
We obtained gene expression data belonging to specimens across two cancer types (LUAD and LUSC) from TCGA; these data were preprocessed using standard methods. After the identification of DEGs, a DAVID analysis was performed using them. Downregulated genes in both lung cancers belong to “complement and coagulation cascades” and “cell adhesion molecules (CAMs)” pathways. While “cell cycle” and “DNA replication” were the most active pathways in both lung cancers (Tables 1–4).
Table 1: Functional annotation analysis of downregulated differentially expressed genes (DEGs) in lung adenocarcinoma (LUAD) datasets using the DAVID tool
| Term | Count | P Value | Genes | Pop Hits | Fold Enrichment |
|---|---|---|---|---|---|
| hsa04610:Complement and coagulation cascades | 17 | 2,02E-05 | F11, C7, A2M, F10, C5AR1, MASP1, C6, F8, SERPING1, C1QA, C8B, VWF, C1QB, THBD, CFD, PROS1, CPB2 | 69 | 3,423022096 |
| hsa04270:Vascular smooth muscle contraction | 21 | 1,10E-04 | RAMP3, ADCY4, RAMP2, NPR1, PRKCE, PRKG1, MYL9, AGTR1, ACTG2, PRKCQ, PTGIR, GNAQ, ADCY9, GUCY1A2, PLA2G1B, MYH11, CALCRL, PLA2G3, PLA2G5, MYLK, PPP1R14A | 112 | 2,605020492 |
| hsa04514:Cell adhesion molecules (CAMs) | 21 | 0,00103288 | SELP, ITGAL, CLDN18, PTPRM, CADM1, ICAM2, CLDN5, NFASC, CLDN11, HLA-E, CDH5, SIGLEC1, CD34, ITGA8, ESAM, JAM2, JAM3, NEGR1, SELE, SELPLG, SPN | 132 | 2,210320417 |
| hsa04080:Neuroactive ligand-receptor interaction | 32 | 0,00245857 | F2RL3, THRB, LEPR, PTH1R, FPR1, GRIN3B, FPR2, VIPR1, APLNR, AGTR1, EDNRB, AGTR2, PTGIR, S1PR1, NMUR1, S1PR4, CNR1, P2RY1, S1PR5, CALCRL, GHR, GRID1, C5AR1, PTGER4, PTGFR, ADRB2, ADRB1, P2RX1, SSTR1, GRIA1, P2RY14, CTSG | 256 | 1,736680328 |
| hsa04614:Renin-angiotensin system | 6 | 0,0055758 | AGTR1, ACE, AGTR2, MME, CPA3, CTSG | 17 | 4,903567985 |
| hsa02010:ABC transporters | 9 | 0,0117617 | ABCA8, ABCC9, ABCA9, ABCB1, CFTR, ABCA3, ABCG1, ABCA6, ABCG2 | 44 | 2,841840537 |
| hsa04670:Leukocyte transendothelial migration | 16 | 0,02073563 | ITGAL, CLDN18, NCF2, NCF1, CLDN5, CLDN11, CXCL12, CDH5, MYL9, CYBB, ESAM, PIK3R5, RAPGEF4, JAM2, JAM3, PIK3R1 | 118 | 1,883856627 |
| hsa03320:PPAR signaling pathway | 11 | 0,02420575 | LPL, CD36, CYP27A1, SORBS1, OLR1, PPARG, RXRG, FABP4, SCD5, ACADL, FABP5 | 69 | 2,21489665 |
| hsa04062:Chemokine signaling pathway | 22 | 0,02578677 | ADCY4, CCL2, FGR, NCF1, PREX1, HCK, CXCL3, CXCL2, CXCR1, GNG11, CXCR2, CXCL12, CCL14, CCL23, PPBP, ADCY9, ARRB1, RASGRP2, CX3CR1, PIK3R5, GRK5, PIK3R1 | 187 | 1,634522662 |
| hsa04060:Cytokine-cytokine receptor interaction | 28 | 0,03410983 | CSF3, ACVRL1, CCL2, PDGFB, CXCL3, LEPR, CXCL2, BMPR2, CXCR1, CXCR2, TNFSF13, TNFSF12, IL7R, CXCL12, CCL23, PLEKHO2, FIGF, GHR, IL18R1, IL6, BMP2, FLT4, TGFBR2, LIFR, CCL14, PPBP, IL20RA, CX3CR1, IL3RA | 262 | 1,484795395 |
| hsa05020:Prion diseases | 7 | 0,0361229 | C1QA, EGR1, C1QB, C8B, C7, IL6, C6 | 35 | 2,778688525 |
| hsa00590:Arachidonic acid metabolism | 9 | 0,04474724 | ALOX15, PTGIS, PTGDS, GPX3, PLA2G1B, LTC4S, ALOX5, PLA2G3, HPGDS, PLA2G5 | 56 | 2,232874707 |
| hsa04350:TGF-beta signaling pathway | 12 | 0,04546484 | BMP2, SMAD9, ACVRL1, CDKN2B, ZFYVE9, SMAD6, LEFTY2, TGFBR2, BMPR2, ID4, DCN, ID3 | 87 | 1,916336914 |
Table 2: Functional annotation analysis of upregulated differentially expressed genes (DEGs) in lung adenocarcinoma (LUAD) datasets using the DAVID tool
| Term | Count | P Value | Genes | Pop Hits | Fold Enrichment |
|---|---|---|---|---|---|
| hsa04110:Cell cycle | 43 | 2,82E-09 | E2F1, E2F2, E2F3, E2F5, DBF4, PRKDC, TTK, PKMYT1, CHEK1, CHEK2, SFN, PTTG1, CCNE2, CCNE1, CDC45, MCM7, CDKN2A, ORC6L, BUB1, CCNA2, CDC7, CDC6, CDK1, RBL1, SKP2, ESPL1, CDC20, MCM2, CDC25C, MCM3, CDK4, MCM4, ORC1L, CDC25A, MCM6, CCNB1, MAD2L1, CCNB2, PLK1, PCNA, BUB1B, MAD2L2, SMC1B | 125 | 2,591466667 |
| hsa03030:DNA replication | 17 | 4,23E-06 | LIG1, POLE, MCM2, MCM3, RNASEH2A, MCM4, MCM6, RFC5, PRIM1, DNA2, RFC3, RFC4, MCM7, POLE2, PRIM2, PCNA, FEN1 | 36 | 3,557407407 |
| hsa04512:ECM-receptor interaction | 26 | 5,15E-05 | IBSP, TNC, COL3A1, ITGA11, ITGB4, COL2A1, VTN, ITGB3, CHAD, HMMR, LAMB3, ITGB8, COMP, COL6A3, SV2A, COL11A2, COL11A1, THBS2, SPP1, THBS4, ITGA2, COL5A2, COL5A1, LAMA1, COL1A2, COL1A1 | 84 | 2,331746032 |
| hsa00270:Cysteine and methionine metabolism | 13 | 9,00E-04 | DNMT3A, LDHB, LDHA, AHCY, SRM, IL4I1, CTH, GOT1, SDS, MAT1A, DNMT1, DNMT3B, CBS | 34 | 2,880392157 |
| hsa00330:Arginine and proline metabolism | 17 | 9,64E-04 | PYCRL, ODC1, ALDH18A1, CKMT1B, SRM, AGMAT, CPS1, PYCR1, ABP1, CKMT1A, GOT1, CKM, ALDH1B1, P4HA1, P4HA3, GAMT, OAT, ADC | 53 | 2,416352201 |
| hsa00250:Alanine, aspartate and glutamate metabolism | 12 | 0,00139536 | ADSSL1, GOT1, ACY3, GFPT1, IL4I1, GPT, CAD, ASNS, CPS1, GAD1, GPT2, PPAT | 31 | 2,916129032 |
| hsa00010:Glycolysis / Gluconeogenesis | 18 | 0,00148413 | ALDOA, LDHB, LDHA, ALDOB, PFKP, ADH1C, ALDH3B2, PGAM2, PCK1, ALDH3A1, GPI, TPI1, G6PC, ALDH1B1, PKM2, ENO3, GAPDH, ENO1 | 60 | 2,26 |
| hsa00601:Glycosphingolipid biosynthesis | 10 | 0,0033785 | B4GALT3, B4GALT2, FUT9, B3GALT5, B3GNT4, FUT6, FUT3, B3GNT3, FUT2, B4GALT4 | 25 | 3,013333333 |
| hsa04115:p53 signaling pathway | 18 | 0,00619724 | CDK1, LRDD, CHEK1, CHEK2, PMAIP1, SFN, CDK4, GTSE1, CCNB1, CCNE2, CCNE1, CCNB2, CDKN2A, SERPINB5, RRM2, BAI1, PERP, IGFBP3 | 68 | 1,994117647 |
| hsa00051:Fructose and mannose metabolism | 11 | 0,01033738 | ALDOA, TPI1, AKR1B15, SORD, GMDS, AKR1B10, GMPPA, ALDOB, PFKP, TSTA3, MTMR7, PMM2 | 34 | 2,437254902 |
| hsa04950:Maturity onset diabetes of the young | 9 | 0,0124429 | HNF1A, HNF4A, BHLHA15, FOXA3, MNX1, NEUROD1, IGF2, HNF4G, PDX1 | 25 | 2,712 |
| hsa00260:Glycine, serine and threonine metabolism | 10 | 0,01604608 | CTH, SHMT2, SDS, GCAT, DMGDH, GAMT, PSAT1, PSPH, CBS, GLDC | 31 | 2,430107527 |
| hsa03440:Homologous recombination | 9 | 0,02497133 | XRCC3, XRCC2, BLM, EME1, SHFM1, BRCA2, RAD54B, RAD54L, RAD51 | 28 | 2,421428571 |
| hsa00533:Keratan sulfate biosynthesis | 6 | 0,02885381 | B4GALT3, B4GALT2, FUT8, CHST6, CHST4, B4GALT4 | 14 | 3,228571429 |
| hsa00140:Steroid hormone biosynthesis | 12 | 0,0343766 | AKR1C3, AKR1C2, UGT1A6, UGT1A10, UGT1A9, AKR1C4, HSD17B2, HSD17B1, CYP11B1, SRD5A3, UGT2B4, SRD5A1, UGT2B15, AKR1C1 | 46 | 1,965217391 |
| hsa00512:O-Glycan biosynthesis | 9 | 0,03707018 | GALNT3, GCNT3, GALNT7, GALNT6, GALNT4, B3GNT6, C1GALT1, GALNT14, ST6GALNAC1 | 30 | 2,26 |
| hsa05219:Bladder cancer | 11 | 0,04377571 | E2F1, E2F2, E2F3, CDKN2A, PGF, MMP9, ERBB2, CDH1, EGF, CDK4, MMP1 | 42 | 1,973015873 |
| hsa00830:Retinol metabolism | 13 | 0,04676524 | BCMO1, CYP2C18, CYP2B6, ADH1C, RDH5, UGT1A10, UGT1A6, UGT1A9, RDH10, LRAT, DGAT1, DGAT2, CYP2A6, UGT2B4, UGT2B15 | 54 | 1,813580247 |
| hsa00980:Metabolism of xenobiotics by cytochrome P450 | 14 | 0,04737766 | CYP2C18, CYP2B6, ADH1C, ALDH3B2, CYP2E1, ALDH3A1, AKR1C3, UGT1A10, UGT1A6, AKR1C2, UGT1A9, AKR1C4, UGT2B4, UGT2B15, AKR1C1, MGST1 | 60 | 1,757777778 |
Table 3: Functional annotation analysis of downregulated differentially expressed genes (DEGs) in Lung squamous cell carcinoma (LUSC) datasets using the DAVID tool
| Term | Count | P Value | Genes | Pop Hits | Fold Enrichment |
|---|---|---|---|---|---|
| hsa04610:Complement and coagulation cascades | 31 | 1,15E-10 | C3AR1, C7, A2M, MASP1, C3, F13A1, C6, C5, C1QC, FGG, SERPINA1, C2, CFI, CFD, F11, CR1, F10, C5AR1, C4A, F8, SERPING1, C4BPA, C1QA, C1QB, VWF, C8B, CD55, TFPI, SERPIND1, CPB2, PROS1 | 69 | 3,586444611 |
| hsa04514:Cell adhesion molecules (CAMs) | 44 | 8,51E-10 | HLA-DQB1, ITGAL, CADM3, CLDN18, CADM1, HLA-DRB1, CLDN5, ITGB2, HLA-DMB, SDC4, HLA-DMA, CDH5, ITGAM, CD22, HLA-DRB5, ESAM, CD4, HLA-DPB1, HLA-DOA, SELPLG, NEGR1, SPN, ICAM1, SELP, PTPRC, PTPRM, ICAM2, HLA-E, HLA-DQA2, CLDN23, HLA-DQA1, SIGLEC1, NCAM2, ITGA9, CD86, CD34, ITGA8, CLDN2, HLA-DPA1, JAM2, JAM3, SELE, CD226, HLA-DRA | 132 | 2,660910518 |
| hsa04640:Hematopoietic cell lineage | 30 | 2,22E-07 | CSF3, IL1R1, HLA-DRB1, CSF1, MME, ANPEP, IL7R, ITGAM, FCGR1C, FCGR1A, CSF3R, CD22, HLA-DRB5, CD4, CSF2RA, CSF1R, IL6, CR1, CD1C, ITGA1, IL6R, IL11RA, CD1E, CD55, CD37, CD36, CD34, CD33, EPOR, IL3RA, HLA-DRA | 86 | 2,784673798 |
| hsa05416:Viral myocarditis | 25 | 2,37E-06 | HLA-DQB1, PRF1, ITGAL, CAV1, HLA-DRB1, ITGB2, HLA-DMB, HLA-DMA, HLA-DRB5, HLA-DPB1, HLA-DOA, ICAM1, HLA-E, HLA-DQA2, HLA-DQA1, LAMA2, CD86, CD55, SGCG, MYH11, SGCD, HLA-DPA1, SGCA, HLA-DRA, MYH10 | 71 | 2,81082097 |
| hsa04060:Cytokine-cytokine receptor interaction | 58 | 1,18E-05 | ACVRL1, PDGFB, IL6ST, LEPR, TNFSF15, CXCR1, CXCR2, TNFSF13, TNFSF12, CXCL12, TGFB2, CSF3R, CSF2RB, CSF2RA, IL18RAP, LIFR, IL6R, IL11RA, TNFRSF10C, PPBP, TNFRSF10D, CCR2, CX3CR1, TNFSF12-TNFSF13, CSF3, CCL3, IL1R1, CCL2, CXCL5, CSF1, CCR1, CXCL3, CXCL2, BMPR2, IL7R, CCL4, LIF, TNFRSF1B, IL12RB1, CCL21, IL10RA, PLEKHO2, FIGF, CSF1R, IL18R1, IL6, BMP2, FLT1, FLT4, TGFBR2, EDA2R, HGF, CCL18, KDR, CCL17, CCL13, CCL14, CXCL16, EPOR, IL3RA | 262 | 1,767169581 |
| hsa04062:Chemokine signaling pathway | 45 | 1,61E-05 | ADCY4, PRKCZ, CCL3, CCL2, GNAI2, CXCL5, FGR, PREX1, CXCL3, CCR1, CXCL2, NCF1C, NFKBIA, CXCR1, GNG11, CXCR2, CCL4, CXCL12, DOCK2, CCL21, RASGRP2, GNG2, PIK3R5, SHC3, PLCB2, GNG7, PIK3CG, ITK, NCF1, HCK, WAS, CCL18, ELMO1, PRKCB, CCL17, GNGT2, CCL13, CCL14, PPBP, ADCY9, ARRB2, ARRB1, CXCL16, CCR2, CX3CR1, GRK5 | 187 | 1,920978181 |
| hsa05310:Asthma | 14 | 1,62E-05 | FCER1A, HLA-DQB1, HLA-DRB1, HLA-DMB, HLA-DMA, HLA-DQA2, HLA-DQA1, HLA-DRB5, MS4A2, FCER1G, HLA-DPA1, HLA-DPB1, HLA-DOA, HLA-DRA | 29 | 3,853732474 |
| hsa04270:Vascular smooth muscle contraction | 31 | 2,82E-05 | ADCY4, ADORA2A, PPP1R12B, MRVI1, PRKG1, MYL9, EDNRA, AGTR1, ACTG2, PTGIR, GUCY1A2, PLA2G1B, GUCY1A3, CALCRL, PLCB2, PPP1R14A, RAMP3, RAMP2, PLA2G10, NPR1, PRKCE, ITPR1, PRKCB, PRKCQ, ADCY9, GNAQ, MYH11, CACNA1C, CACNA1D, PLA2G5, MYLK | 112 | 2,209506055 |
| hsa05332:Graft-versus-host disease | 16 | 3,38E-05 | HLA-DQB1, PRF1, IL6, HLA-DRB1, HLA-DMB, HLA-E, HLA-DMA, HLA-DQA2, HLA-DQA1, CD86, HLA-DRB5, HLA-DPA1, HLA-DPB1, HLA-DOA, KLRD1, HLA-DRA | 39 | 3,274966791 |
| hsa05322:Systemic lupus erythematosus | 27 | 1,37E-04 | HLA-DQB1, C7, HLA-DRB1, C3, C6, C5, HLA-DMB, C1QC, HLA-DMA, FCGR1C, FCGR1A, HLA-DRB5, HLA-DPB1, C2, FCGR3A, HLA-DOA, FCGR3B, C4A, HLA-DQA2, HLA-DQA1, C1QA, C8B, C1QB, CD86, HLA-DPA1, FCGR2A, CTSG, HLA-DRA | 99 | 2,177108606 |
| hsa04672:Intestinal immune network for IgA production | 17 | 1,84E-04 | HLA-DQB1, IL6, HLA-DRB1, TNFSF13, TNFSF12, PIGR, HLA-DMB, HLA-DMA, CXCL12, HLA-DQA2, HLA-DQA1, TGFB2, CD86, HLA-DRB5, HLA-DPA1, HLA-DPB1, TNFSF12-TNFSF13, HLA-DOA, HLA-DRA | 49 | 2,769519111 |
| hsa05330:Allograft rejection | 14 | 2,39E-04 | HLA-DQB1, PRF1, HLA-DRB1, HLA-DMB, HLA-E, HLA-DMA, HLA-DQA2, HLA-DQA1, CD86, HLA-DRB5, HLA-DPA1, HLA-DPB1, HLA-DOA, HLA-DRA | 36 | 3,104395604 |
| hsa04940:Type I diabetes mellitus | 15 | 3,63E-04 | HLA-DQB1, PRF1, HLA-DRB1, PTPRN2, HLA-DMB, HLA-E, HLA-DMA, HLA-DQA2, HLA-DQA1, CD86, HLA-DRB5, HLA-DPA1, HLA-DPB1, HLA-DOA, HLA-DRA | 42 | 2,850975555 |
| hsa05414:Dilated cardiomyopathy | 24 | 7,06E-04 | ADCY4, SLC8A1, TNNC1, ITGA1, CACNG4, ITGA10, CACNB4, TTN, CACNA2D2, TGFB2, LAMA2, ITGA9, DES, ADRB1, SGCG, ADCY9, PLN, ITGA8, ITGA7, RYR2, SGCD, CACNA1C, CACNA1D, SGCA | 92 | 2,08245171 |
| hsa05410:Hypertrophic cardiomyopathy (HCM) | 22 | 0,00139878 | SLC8A1, IL6, TNNC1, ITGA1, CACNG4, ITGA10, CACNB4, TTN, CACNA2D2, TGFB2, LAMA2, ITGA9, ACE, DES, SGCG, ITGA8, ITGA7, RYR2, SGCD, CACNA1C, CACNA1D, SGCA | 85 | 2,066118755 |
| hsa04614:Renin-angiotensin system | 8 | 0,00290177 | AGTR1, ACE, AGTR2, MME, CPA3, ANPEP, ENPEP, CTSG | 17 | 3,756579555 |
| hsa05320:Autoimmune thyroid disease | 14 | 0,0083188 | HLA-DQB1, PRF1, HLA-DRB1, HLA-DMB, HLA-E, HLA-DMA, HLA-DQA2, HLA-DQA1, CD86, HLA-DRB5, HLA-DPA1, HLA-DPB1, HLA-DOA, HLA-DRA | 51 | 2,191338074 |
| hsa04670:Leukocyte transendothelial migration | 25 | 0,00994356 | ITGAL, CLDN18, GNAI2, CLDN5, NCF1C, ITGB2, CXCL12, CDH5, ITGAM, MYL9, ESAM, PIK3R5, RAPGEF4, RAPGEF3, PIK3CG, ICAM1, ITK, NCF2, NCF1, NCF4, CLDN23, PRKCB, CYBB, CLDN2, JAM2, JAM3 | 118 | 1,691256685 |
| hsa04612:Antigen processing and presentation | 19 | 0,01258302 | HLA-DQB1, CIITA, HLA-DRB1, IFI30, CTSS, HLA-DMB, HLA-E, HLA-DMA, HLA-DQA2, HLA-DQA1, CD74, B2M, HLA-DRB5, CD4, HLA-DPA1, HLA-DPB1, HLA-DOA, KLRD1, HLA-DRA | 83 | 1,827372283 |
| hsa05412:Arrhythmogenic right ventricular cardiomyopathy (ARVC) | 17 | 0,02357362 | SLC8A1, ITGA1, CACNG4, ITGA10, CACNB4, CACNA2D2, LAMA2, ITGA9, DES, SGCG, ITGA8, ITGA7, RYR2, SGCD, CACNA1C, CACNA1D, SGCA | 76 | 1,785611006 |
| hsa04530:Tight junction | 26 | 0,02458398 | PRKCZ, CLDN18, GNAI2, MRAS, CLDN5, MYL9, RRAS, PARD6B, MAGI3, MAGI1, MPDZ, PRKCE, CLDN23, PRKCB, EPB41L2, EPB41L3, PRKCQ, TJP1, CGN, MYH11, CLDN2, TJP3, JAM2, JAM3, MYH10, SPTAN1 | 134 | 1,548888212 |
| hsa05020:Prion diseases | 10 | 0,02477695 | C1QA, EGR1, C1QB, NCAM2, C8B, C7, IL6, C6, C5, C1QC | 35 | 2,280780444 |
| hsa04960:Aldosterone-regulated sodium reabsorption | 11 | 0,0263487 | PIK3CG, ATP1B2, HSD11B1, NR3C2, PIK3R5, NEDD4L, ATP1A2, SCNN1G, SCNN1B, SLC9A3R2, PRKCB | 41 | 2,141708466 |
| hsa03320:PPAR signaling pathway | 15 | 0,04364393 | ACOX2, LPL, OLR1, PPARG, RXRG, ACADL, ACSL1, CD36, SORBS1, CYP27A1, HMGCS2, FABP3, FABP4, ACSL4, ACSL5 | 69 | 1,735376425 |
| hsa04666:Fc gamma R-mediated phagocytosis | 19 | 0,04492025 | PIK3CG, DNM3, PTPRC, WASF3, NCF1, HCK, NCF1C, PIP5K1B, PRKCE, WAS, PRKCB, DOCK2, GAB2, FCGR1C, CFL2, FCGR1A, PLA2G4F, PIK3R5, FCGR2A, FCGR3A, PPAP2B | 95 | 1,596546311 |
| hsa04020:Calcium signaling pathway | 31 | 0,04546262 | GNA14, ADCY4, ERBB4, CYSLTR1, ADORA2A, TNNC1, EDNRA, AGTR1, EDNRB, PDE1B, PLCB2, SLC8A1, BST1, PTGFR, ITPR1, PRKCB, ADRB2, P2RX7, PLCE1, ADRB1, P2RX1, ADCY9, GNAQ, PLN, TBXA2R, CACNA1H, RYR2, CACNA1C, CACNA1D, MYLK, PTAFR | 176 | 1,406049308 |
Table 4: Functional annotation analysis of upregulated differentially expressed genes (DEGs) in Lung squamous cell carcinoma (LUSC) datasets using the DAVID tool
| Term | Count | P Value | Genes | Pop Hits | Fold Enrichment |
|---|---|---|---|---|---|
| hsa04110:Cell cycle | 54 | 7,45E-13 | E2F1, E2F2, E2F3, DBF4, PKMYT1, TTK, PTTG1, CCNE2, CCNE1, CDC45, MCM7, CDKN2A, CCNA1, CCNA2, CDC7, CDK1, CDC6, RBL1, SKP2, CDK6, ESPL1, MCM2, MCM3, CDK4, MCM4, MCM5, ORC1L, CDK2, MCM6, MAD2L1, BUB1B, ORC5L, ANAPC7, MAD2L2, YWHAZ, PRKDC, CHEK1, CHEK2, SFN, ORC6L, BUB1, TFDP2, TFDP1, CDC20, ATR, CDC25C, CDC25A, CCNB1, HDAC2, CCNB2, HDAC1, PLK1, PCNA, SMC1B | 125 | 2,715352287 |
| hsa03030:DNA replication | 25 | 9,11E-12 | RNASEH1, POLA2, RPA3, PRIM1, MCM7, POLE2, POLE3, PRIM2, FEN1, LIG1, POLE, MCM2, RNASEH2A, MCM3, MCM4, MCM5, MCM6, RFC5, DNA2, RFC3, RFC4, RFC2, POLD1, POLD2, PCNA | 36 | 4,364956737 |
| hsa04115:p53 signaling pathway | 27 | 6,14E-06 | BID, LRDD, RPRM, CHEK1, SFN, CHEK2, PMAIP1, GTSE1, SESN3, CCNE2, CCNE1, CDKN2A, BAI1, TP53AIP1, CDK1, CYCS, CDK6, ATR, CDK4, CDK2, TP73, CCNB1, CCNB2, SERPINB5, RRM2, PERP, IGFBP3 | 68 | 2,495728205 |
| hsa00480:Glutathione metabolism | 21 | 3,38E-05 | GSTA1, ODC1, GCLC, GGCT, PGD, GCLM, GSS, GSTM1, GPX2, GSTM2, GSR, GSTM3, GSTM4, G6PD, OPLAH, RRM2, RRM1, IDH2, GSTO2, GPX7, SMS | 50 | 2,639925834 |
| hsa03410:Base excision repair | 16 | 1,29E-04 | APEX2, UNG, NEIL3, LIG1, POLE, LIG3, POLB, SMUG1, POLE2, POLE3, POLD1, POLD2, PCNA, TDG, PARP1, FEN1 | 35 | 2,873388663 |
| hsa05217:Basal cell carcinoma | 21 | 1,64E-04 | FZD9, WNT5A, DVL3, WNT10A, WNT16, WNT10B, WNT5B, LEF1, FZD3, GLI2, FZD7, FZD6, GLI1, WNT2B, SMO, FZD10, WNT7B, WNT4, WNT3, WNT11, PTCH2 | 55 | 2,399932577 |
| hsa03430:Mismatch repair | 12 | 3,26E-04 | RFC5, EXO1, MSH6, RFC3, RFC4, RFC2, MSH2, POLD1, LIG1, POLD2, PCNA, RPA3 | 23 | 3,279410974 |
| hsa00010:Glycolysis / Gluconeogenesis | 21 | 6,17E-04 | ALDOA, LDHA, ALDOC, PGAM1, HK2, PFKP, ADH1C, ALDH3B2, PGAM2, ADH7, PCK1, ALDH3A1, PGM2, GPI, TPI1, PKM2, ENO2, ENO3, PGK1, GAPDH, ENO1 | 60 | 2,199938195 |
| hsa05200:Pathways in cancer | 75 | 6,70E-04 | FGF19, E2F1, E2F2, E2F3, HRAS, PGF, MMP9, FGF11, FGF12, GLI2, MMP1, ACVR1C, GLI1, CCNE2, CCNE1, WNT4, WNT3, CDKN2A, SLC2A1, TGFA, NOS2, CCNA1, FGF3, EGFR, WNT10A, RET, PLD1, WNT10B, CYCS, SKP2, LEF1, FADD, CDK6, CDK4, CDK2, RAD51, JUP, SMO, PIAS3, LAMC2, WNT11, WNT5A, BID, CKS1B, WNT16, WNT5B, FGFR3, EGLN3, TFG, CDH1, LAMB3, RAC3, EGF, TRAF4, PIK3R2, FZD9, MSH6, DVL3, MSH2, BRCA2, ITGA2, FZD3, BIRC5, COL4A6, FZD7, FZD6, WNT2B, LAMA1, CBLC, FZD10, WNT7B, HDAC2, ITGA6, HDAC1, PTCH2 | 328 | 1,437241852 |
| hsa04114:Oocyte meiosis | 32 | 6,81E-04 | YWHAZ, ADCY2, PKMYT1, AURKA, PTTG1, PRKX, CCNE2, CCNE1, CALML3, FBXO43, BUB1, STAG3, FBXO5, CAMK2B, CALML5, CDK1, SGOL1, CHP2, IGF2, ESPL1, CDC20, CDC25C, CDK2, CCNB1, RPS6KA6, MAD2L1, CCNB2, MAPK12, PLK1, ANAPC7, MAD2L2, SMC1B | 110 | 1,828520058 |
| hsa00240:Pyrimidine metabolism | 28 | 0,00129063 | POLR2H, CTPS, PNPT1, DTYMK, CAD, POLA2, POLR2D, TK1, PRIM1, TYMS, POLE2, NT5M, POLE3, PRIM2, ENTPD3, UCK2, POLR3G, POLE, POLR1C, POLR2J3, NME4, UMPS, NME2, NME1-NME2, NME1, RRM2, POLD1, POLD2, RRM1, TXNRD1 | 95 | 1,852579533 |
| hsa00250:Alanine, aspartate and glutamate metabolism | 13 | 0,00179493 | GLS2, GOT2, ADSSL1, ALDH4A1, ADSL, IL4I1, GPT, CAD, ASNS, CPS1, GAD1, GPT2, PPAT | 31 | 2,635870649 |
| hsa04340:Hedgehog signaling pathway | 19 | 0,00182644 | WNT5A, WNT10A, WNT16, WNT10B, WNT5B, GLI2, PRKX, ZIC2, GLI1, WNT2B, SMO, WNT7B, WNT4, WNT3, WNT11, PTCH2, BMP7, BMP8B, BMP8A | 56 | 2,132593149 |
| hsa04512:ECM-receptor interaction | 25 | 0,0021767 | TNC, COL3A1, ITGB4, ITGA11, ITGA2, COL2A1, COL5A2, COL5A1, COL4A6, HMMR, LAMA1, LAMB3, SDC1, ITGA6, ITGB8, COMP, COL1A2, LAMC2, SV2B, SV2A, COL1A1, COL11A2, THBS2, COL11A1, SPP1 | 84 | 1,870695744 |
| hsa03440:Homologous recombination | 12 | 0,00241416 | XRCC3, XRCC2, BLM, POLD1, POLD2, EME1, SHFM1, BRCA2, RAD54B, RAD54L, RAD51, RPA3 | 28 | 2,693801872 |
| hsa00360:Phenylalanine metabolism | 10 | 0,00433744 | GOT2, DDC, NAA20, LCLAT1, ALDH3B2, IL4I1, PAH, PNPLA3, MIF, ALDH3A1 | 22 | 2,857062591 |
| hsa00230:Purine metabolism | 37 | 0,00801809 | XDH, POLR2H, GDA, ADCY2, PNPT1, POLA2, POLR2D, HPRT1, AK3L1, ADA, PPAT, PRIM1, POLE2, ATIC, POLE3, NT5M, PRIM2, ENTPD3, ADCY10, ENTPD2, POLR3G, ADSSL1, POLE, POLR1C, GMPS, POLR2J3, GART, NME4, NME2, NME1-NME2, NME1, RRM2, POLD1, PKM2, POLD2, RRM1, ADSL, PAICS, PRPS2 | 153 | 1,520031993 |
| hsa00980:Metabolism of xenobiotics by cytochrome P450 | 18 | 0,01003269 | GSTA1, CYP2C18, CYP2S1, ADH1C, ALDH3B2, ADH7, CYP2E1, UGT1A1, ALDH3A1, AKR1C3, GSTM1, UGT1A7, GSTM2, AKR1C2, UGT1A10, UGT1A6, UGT1A9, GSTM3, GSTM4, UGT1A8, UGT2A1, GSTO2, AKR1C1 | 60 | 1,88566131 |
| hsa00030:Pentose phosphate pathway | 10 | 0,01140598 | PGM2, ALDOA, GPI, TALDO1, G6PD, ALDOC, PGD, PFKP, TKTL1, PRPS2 | 25 | 2,51421508 |
| hsa00330:Arginine and proline metabolism | 16 | 0,01525979 | PYCRL, ODC1, ALDH18A1, CKMT1B, AGMAT, CPS1, GLS2, GOT2, ABP1, PYCR1, CKMT1A, P4HA1, P4HA3, ALDH4A1, GAMT, NOS2, SMS | 53 | 1,897520815 |
| hsa04310:Wnt signaling pathway | 35 | 0,0189717 | WNT5A, WNT16, WNT5B, MMP7, PRKX, WNT4, CSNK2A1, WNT3, RAC3, CACYBP, CAMK2B, FZD9, WNT10A, TBL1XR1, DVL3, WNT10B, VANGL1, VANGL2, CHP2, LEF1, FZD3, CSNK2A1P, PORCN, FZD7, WNT2B, FZD6, DKK4, SENP2, WNT7B, FZD10, DKK1, SFRP1, SFRP2, WNT11, RUVBL1, TBL1X | 151 | 1,456912712 |
| hsa04916: Melanogenesis | 25 | 0,01931079 | WNT5A, HRAS, WNT16, ADCY2, WNT5B, POMC, PRKX, WNT4, WNT3, MC1R, CALML3, CAMK2B, CALML5, TUBB3, FZD9, DVL3, WNT10A, WNT10B, LEF1, FZD3, FZD7, FZD6, WNT2B, WNT7B, FZD10, WNT11 | 99 | 1,587256995 |
| hsa05219:Bladder cancer | 13 | 0,02646666 | E2F1, EGFR, E2F2, E2F3, HRAS, FGFR3, PGF, MMP9, CDH1, CDK4, MMP1, CDKN2A, EGF | 42 | 1,945523574 |
| hsa00670:One carbon pool by folate | 7 | 0,0303469 | TYMS, MTHFD2, SHMT2, ALDH1L1, DHFR, ATIC, GART | 16 | 2,749922744 |
| hsa05222:Small cell lung cancer | 21 | 0,03694956 | E2F1, E2F2, CKS1B, E2F3, CYCS, SKP2, ITGA2, CDK6, CDK4, CDK2, COL4A6, CCNE2, LAMA1, CCNE1, LAMB3, ITGA6, PIAS3, LAMC2, NOS2, TRAF4, PIK3R2 | 84 | 1,571384425 |
As Schwann cells act on both of the aforementioned upregulated pathways, we selected datasets from the GEO database containing gene expression profiles of both early and late passage human Schwann cells exposed to cancer growth factors heregulin and forskolin (GSE4030). These expression profiles were used to identify DEGs with the aid of the online tool GEO2R. We found 638 upregulated and 1250 downregulated genes. Then, we performed a GO term enrichment and functional classification by KEGG analysis, using DAVID platform, to investigate the biological and functional roles of these DEGs. Upregulated DEG enrichment included “neuroactive ligand-receptor interaction,” while “pathways in cancer” and “focal adhesion” were enriched for downregulated DEGs (Supplementary Tables 1-2).
To facilitate the analysis of the large throughput of DEGs, a protein classification analysis was performed using the PANTHER classification system. According to the study, upregulated genes categories were identified, in which hydrolase (PC00121), enzyme modulator (PC00095), nucleic acid binding (PC00171), receptor (PC00197), transcription factor (PC00218), transporter (PC00227), and signaling molecule (PC00207) are the top seven most abundant protein classes. However, protease (PC00190) is the top one most abundant protein class among downregulated DEGs (Figure 1A-B).
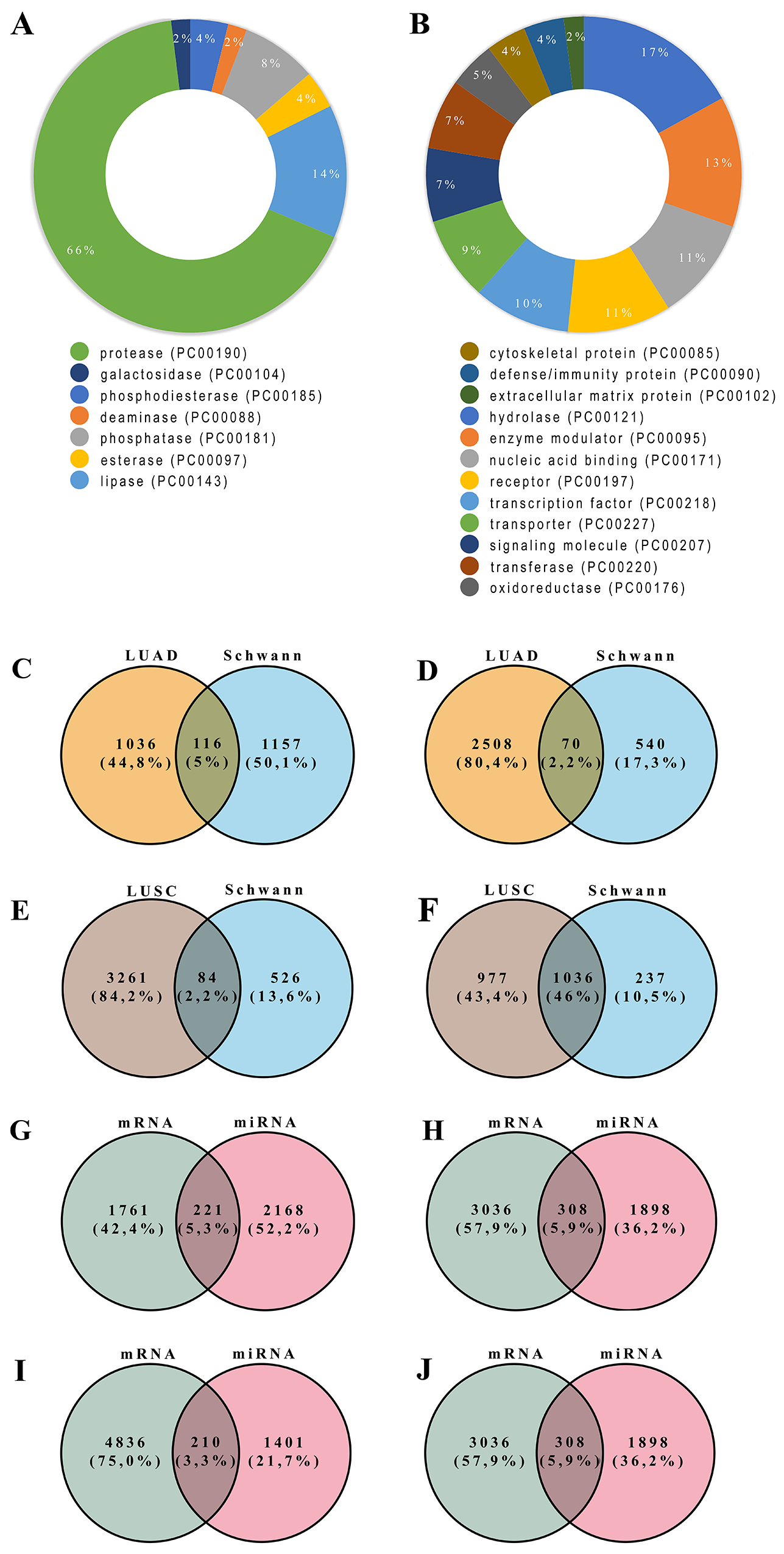
Figure 1: Molecular function and protein class terms of downregulated. (A) and upregulated (B) among the differentially regulated genes in Schwann cells. Figure 1C-D - Venn Diagrams of combined overrepresented differentially expressed genes in Schwann cells and lung adenocarcinoma (C and D), and Lung squamous cell carcinoma (E and F) cancer samples. The results are showing overlapping downregulated (Figure 1C) and upregulated (Figure 1D) genes between lung adenocarcinoma and Schwan cells. Overlapped downregulated and upregulated genes between Lung squamous cell carcinoma and Schwan cells are showed in Figure 1E and Figure 1F, respectively. Figure 1G-H - Venn Diagrams of downregulated miRNA target genes combined with upregulated genes in lung adenocarcinoma (G) and Lung squamous cell carcinoma (H). Figure 1I-J - Venn Diagrams of upregulated miRNA target genes combined with downregulated genes in lung adenocarcinoma (I) and Lung squamous cell carcinoma (J). The number in each intersecting region represents the number of overlapping genes.
Overview of the cancer transcriptomic analysis
We conducted a systematic and integrative analysis to explore cancer type-specific and Schwann cell specific DEGs, to construct a cancer network. We first determined DEGs by comparing gene expression levels between tumor and normal samples. A Venn diagram was then constructed to visualize the overlap between DEG genes, both upregulated and downregulated, from both cancer types and from Schwann cells. Results are shown in Figure 1C-F. Among upregulated genes from LUSC and LUAD, we identified 84 and 70 co-expressed DEGs with Schwann cells, respectively.
We also ran a KEGG analysis to investigate pathways with major expression changes in studied cell lines and lung cancer, based on previously overlapping identified DEGs. The analysis revealed that “neuroactive ligand-receptor interaction” and “p53 signaling pathway” were among the most active pathways according to overlapped Schwann cells and LUSC DEGs. In contrast, overlapped Schwann cells and LUAD DEGs samples show “neuroactive ligand-receptor interaction” pathway as mainly downregulated, while “p53 signaling pathway” is upregulated (Supplementary Tables 3-6). Grm1, Chrm3, Gabra5, Gpr50, and Pth2r were the common upregulated genes associated with the “neuroactive ligand-receptor interaction” pathway; Ccne1, Cdkn2a, and Perp were commonly associated to “p53 signaling pathway”.
To confirm our results, we performed a differentially expressed miRNA analysis over cancer data. Data from identified miRNAs target genes were crossed with DEGs data from Schwann cells (Figure 1G-J). We found that downregulated miRNAs had target genes associated with “neuroactive ligand-receptor interaction” in LUAD and “pathways in cancer” in both cancers (Supplementary Tables 7-8). In contrast, upregulated miRNAs had target genes associated with the “axon guidance” pathway in both lung cancers (Supplementary Tables 7-10). Genes from the “axon guidance” pathway, common to all our analyses, were Robo2 and Slit2 genes. ROBO2 and SLIT2 proteins were highly expressed in normal tissue compared to cancer samples (data not shown).
Hallmarks of cancer analysis
In order to understand the mechanism by which Schwann cells aid in neoplastic development, we analyzed the behavior of genes associated with cell differentiation processes (Gata3, Cdh1, and Cdh2), apoptosis (Casp3, Casp9, Bax, and Bcl2), motility (Cxcr2, Cxcl5, Mmp9, and Ccl12), and cell proliferation (Mki67). While Cdh1 and Cdh2 showed an increased mRNA expression in LUSC samples, only CDH2 protein expression was decreased in LUAD when compared with LUSC (data not shown). Also, Mmp9 and Mki67 were overexpressed in both lung cancers (Supplementary Figures 1-2).
Analysis of genes involved in dedifferentiation of Schwann cells
Schwann cells produce cell differentiation maintenance proteins (Sox10, S100, Egr2, Mbp, and Mpz) that, after nerve damage, diminish their expression, provoking cellular dedifferentiation. Consequently, the expression of a new set of proteins that form non-myelinated cells, Sox10, Gap43, S100, Ncam1, Ngfr1, and Gfap, is augmented. We observed increased expression of Gap43 and Gfap genes in both lung cancers, whereas Ngfr1 was upregulated only in LUSC cells (Supplementary Figures 1-2).
Methylation and protein analyses
Methylation analysis of the Cdh1, Cdh2, Mmp9, Mki67, Gfap, Gap43, Robo2, and Slit2 genes from tumor samples demonstrated that all of them were methylated in their promoter regions unlike those from normal tissues. However, there was no significant correlation between methylation and Gfap gene expression in lung cancers. Whereas there was a positive correlation between methylation and Gap43 gene expression in LUSC samples (Supplementary Figures 3-24).
Copy number alteration
Copy number alteration data demonstrated that Cdh1, Cdh2, Gfap, Perp, and Robo2 had a higher mRNA expression than normal tissues; increased expression was associated with gain or amplification alterations in LUAD samples (Supplementary Figure 25). Similarly, in LUSC samples, Ccne1, Cdh1, Cdkn2a, Gap43, and Perp had a higher mRNA expression associated with gain or amplification alterations (Figure 2).
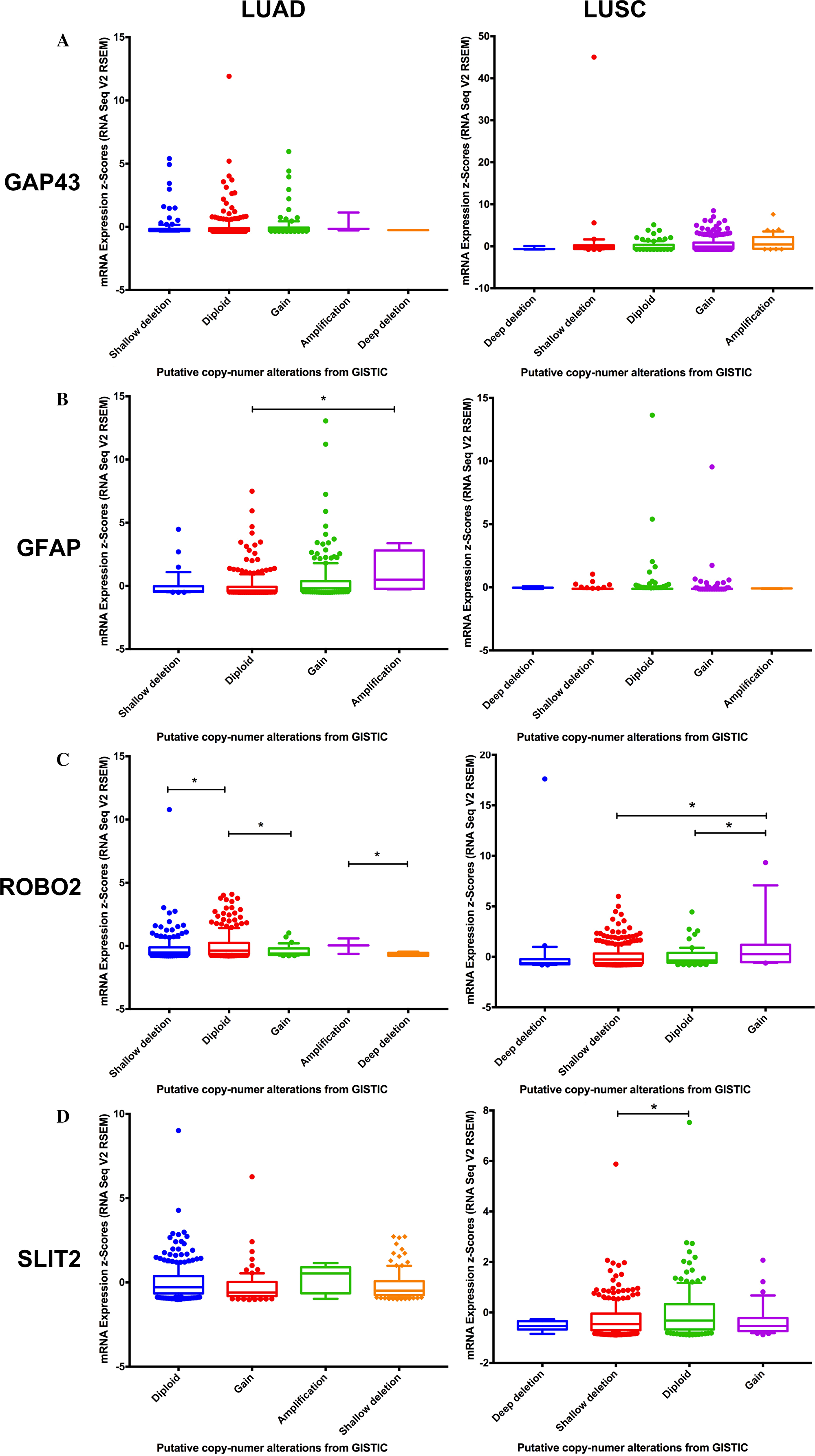
Figure 2: Correlation of copy number variation and expression of. Gap43 (A), Gfap (B), Robo2 (C) and Slit2 (D) genes in lung adenocarcinom a (LUAD) and lung squamous cell carcinoma (LUSC). Figures were generated using Cbioportal data.
Schwann cell differentiation protein expression in lung cancer samples
For analysis of proteins expressed in dedifferentiated Schwann cells, we initially identified the gene list in Pubmed and GeneCards databases. We only analyzed genes with relevance score higher than seven. The relevance scores of elements related to genes are based on the analysis of co-occurrences of two elements in Medline documents. The observed are compared to an expected value based on a hypergeometric distribution. We identified 325 Schwann cell dedifferentiated related genes in both databases. Data were then cross-checked with previously published protein data expression analysis [9], which the expression of normal lung tissue and lung cancer were compared. 10 proteins (GFAP, STAT3, SRC, CD36, CAV1, PCNA, HDAC9, AQP1, APOA1, RALA) associated with dedifferentiation of Schwann cells were increased in lung cancer, including GFAP. No downregulated protein expression was associated with Schwann cell dedifferentiation.
Cancer protein expression patterns correspond to pathway activation levels
We also performed an RPPA protein analysis. Only CDH1 and CDH2 protein expression data were available for analysis. We observed that CDH2 was overexpressed in LUSC compared to LUAD. Additionally, no significant difference was found in CDH1 analysis (Supplementary Figure 26).
In order to analyze the pathway by which Schwann cells induce neoplastic and their own cell proliferation and migration, we evaluated the expression of MEK1 (MEK1 and MEK1_pS217S221), ERK2, AKT (PRAS40_pT246, AKT_pT308 and AKT_pS473), RAF (CRAF and CRAF_pS338), and GSK3Β (GSK3_pS9 and GSK3ALPHABETA_pS21S9) proteins. We observed higher expression of ERK2, AKT, CRAF and GSK3Β proteins in LUSC when compared with LUAD. Only MEK1_pS217S221 presented higher expression in LUAD (Supplementary Figure 26).
Prognostic analyses
TIMER survival analyses showed that Ccne1, Mki67, and Gap43 mRNA higher expression are associated with poor LUAD prognostic (Supplementary Figure 27). Similarly, PRECOG analysis showed that Grm1, Gap43 and Mki67 higher mRNA expression is associated with poor prognostic in LUAD. However, PRECOG analysis demonstrated Slit2 and Robo2 downregulation are associated with poor survival in LUAD samples (Figure 3). Figure 4 illustrates the possible association between Schwann cells and lung cancer.
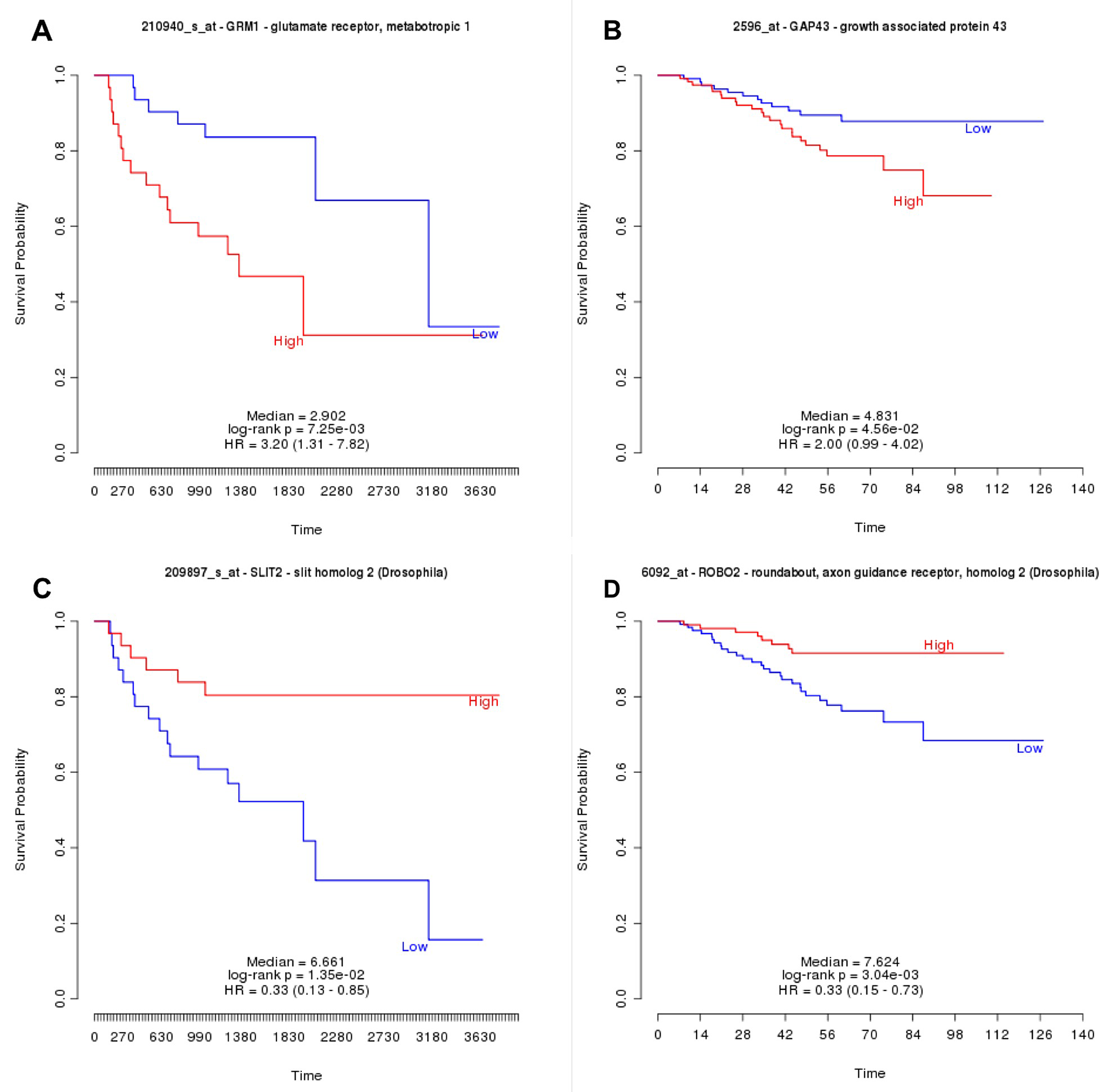
Figure 3: PRECOG Survival analyses in lung adenocarcinoma for Grm1. (A), Gap43 (B), Slit2 (C) and Robo2 (D) mRNA expression. Note that Grm1 and Gap43 upregulation and, Slit2 and Robo2 downregulation are associated with poor survival.
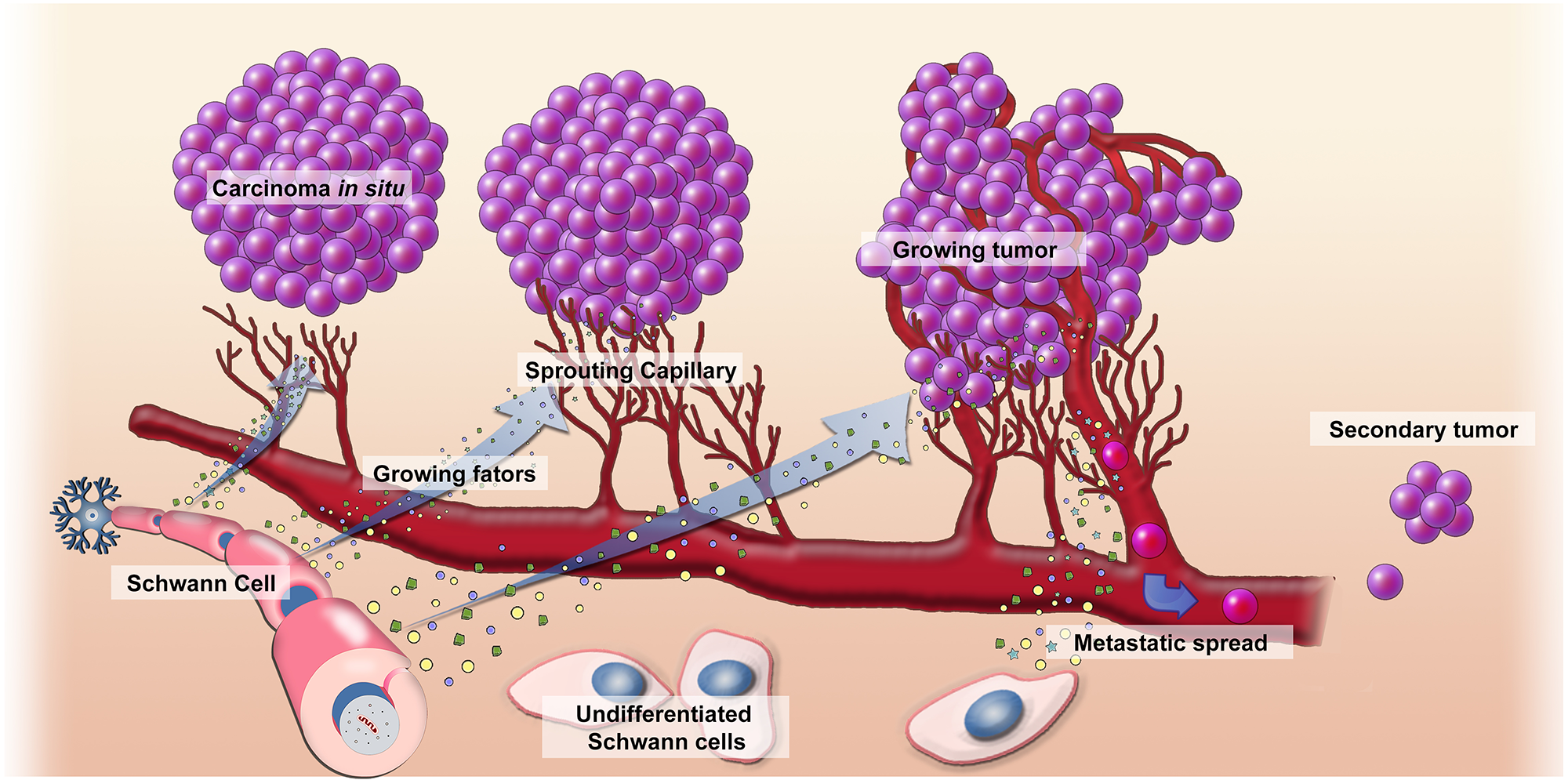
Figure 4: Mechanism of action of Schwann cells in the development of cancer. We suppose that Schwann cells assist the neoplastic lung cells to invade surrounding tissue during the early stages of carcinogenesis. This mechanism would be possible through the dedifferentiation of the Schwann cells and blocking axon guidance signaling pathways. Migration of Schwann cells to the peritumoral region would be associated with perineural invasion and, subsequently, central nervous system metastases.
DISCUSSION
Cells surrounding a tumor form a molecular microenvironment known as stroma. Stroma formation is influenced by tumor cells and, in turn, it influences tumor growth, migration, and invasion [17]. Depending on the characteristics of the primary tumor, the stroma, and the intrinsic ability of metastatic tumor cells to adapt to a new location, malignant cells use distinct mechanisms for proliferation, survival, and dissemination. These cells often reactivate the expression of genes employed during embryogenesis [17, 18]. In order to leave the primary tumor and disseminate to distant organs, metastatic cells lose the ability to adhere to adjacent cells, enhancing their migratory and invasive capacity. This mechanism is accompanied by several modifications in the expression of genes, such as loss of epithelial receptor expression and increased expression of mesenchymal markers, a phenomenon also known as epithelial-mesenchymal transition [18, 19].
We investigated the association between Schwann cells and those lung cancer types often associated with perineural invasion. Initially, we used the GEO DataSets platform from the GEO repository to identify a database reporting gene expression in Schwann cells in a neoplastic context. Briefly, the database contains the expression results from experiments in which two factors produced by tumor cells were added into cell cultures. Comparisons were made between samples from the first and third passages. We then used these data to perform differential gene expression analysis and crossed data from upregulated genes with differential expression data from LUAD and LUSC. After identifying the genes in common, we ran several analysis tools to identify molecular pathways associated to these genes. Interestingly, we noted that the “neuroactive ligand-receptor interaction” pathway was active in LUSC. Studies have demonstrated the association between neuroactive ligand-receptors in the control of Schwann cell differentiation as well as in neoplastic cell proliferation. For example, GRM1 can activate the RAS/MEK1/ERK/AKT/RAF/GSK3Β phosphorylation cascade [20, 21], leading to an increase in cell proliferation and invasion. Grm1 was upregulated in LUSC, suggesting that this gene participates in lung cancer development.
In these context, it has also been demonstrated that neuroactive ligand-receptors are active during Wallerian degeneration of peripheral nerves. Together with macrophages, Schwann cells remove axon and myelin debris, and clear a path for subsequent axonal regrowth and nerve regeneration [22]. Tumor cells benefit from nerve regeneration machinery to promote cell proliferation, migration, and invasion. It has been reported that Schwann cells are induced to migrate to the region close to the tumor at the beginning of the carcinogenic process. It was also suggested that Schwann cells promote neoplastic invasion by direct contact with cancer cells, since paracrine signaling and matrix remodeling are not yet sufficient to induce the migration process [4, 23]. Cell-cell contact between Schwann cells and tumor tissue is necessary to potentiate the ability of neoplastic cells to penetrate into the underlying tissue [1, 3]. After contact, the degradation of the extracellular matrix by Schwann cells provokes the formation of tunnels or bands coated with laminin, due to Schwann cells’ capacity to express matrix metalloproteins, especially MMP2 and MMP9. The mechanism of extracellular matrix degradation that promotes neoplastic migration also depends on the production of NCAM1 and N-cadherin (CDH2) by Schwann cells [1, 7].
In the present study, we analyzed the expression of genes associated with the hallmark of cancer. The importance of analyzing these genes derives from the hypothesis that there may be a correlation between the presence of Schwann cells and the aggressiveness of the neoplastic cell. Increased expression of genes related to cell differentiation (Cdh1 and Cdh2), motility (Mmp9), and proliferation (Mki67), mainly in LUSC, suggests this correlation. We observed that these genes are mutated and had higher expression in lung cancer samples when compared with normal tissue.
Decreased E-cadherin protein expression after contact of Schwann cells with the tumor resembles the mechanism followed by cells during axonal repair process [8]. The loss of E-cadherin during the epithelial-mesenchymal transition in cancer is associated with a positive regulation of NCAM1 and CDH2 [1, 3, 4, 8]. When E-cadherin is suppressed, NCAM1 and CDH2 are upregulated; they associate with the p59fyn protein, whose subsequent activation leads to inhibition of focal adhesion and an increase in cell migration. A study using oncogenic K-ras pancreatic cancer cell lines identified increased levels of polysialylated NCAM1 expression, which interacts with E-cadherin to create steric hindrance of homophilic binding and decrease cell adhesion [1]. In our study, we observed upregulation of E-cadherin and CDH2 in both lung cancers. A CDH2 protein RPPA analysis showed that it is overexpressed in LUSC. Copy number alteration analysis also showed that amplification and gain are associated with E-cadherin and N-cadherin mRNA levels in both cancers. However, we showed that Cdh2 mRNA expression is higher in LUSC when compared to LUAD. No difference was observed neither in Ncam1 mRNA nor in E-cadherin protein analyses.
Differentiation of myelinating Schwann cells may undergo interference from inhibitory pathways that negatively control the expression of genes responsible for myelin sheath formation. NOTCH1 and JUN stand out among the negative regulators of the myelination program, in the same way as SOX-2 and PAX-3 [2, 24, 25]. The myelinating phenotype also involves the inactivation of a number of genes linked to the production of immature Schwann cell markers. Some transcription factors are responsible for ensuring proper maturation in Schwann cells. SOX-10 acts synergistically with a second factor, OCT-6, resulting in the expression of Krox-20. In turn, KROX-20 is a key inducer of expression of myelin genes, such as Mbp, Mpz and Prx. The maintenance of the myelinating phenotype therefore requires the continuous expression of KROX-20 and SOX-10, considering that inactivation of both proteins results in dedifferentiation of Schwann cells [24].
Previous studies have shown that Schwann cells induce cellular aggressiveness in lung cancer [26]. During dedifferentiation, Schwann cells express proteins initially lost during the myelination process. Among these proteins are GAP43, NCAM1, P75NTR (Ngfr), GFAP and SOX-2 [2]. Data from our study showed that Gfap, Ngfr and Gap43 gene expression is increased in lung cancers, especially in LUSC. To confirm our data, we performed an analysis by using protein expression data published before [9]. We selected only genes associated with Schwann cells dedifferentiation. The results showed that proteins associated with dedifferentiated Schwann cells are overexpressed in lung cancer, when compared with normal lung samples. Besides, Gfap and Gap43 copy number alterations are associated with higher mRNA levels in LUAD and LUSC, respectively. It is likely that Schwann cells in these tissues are dedifferentiated, aiding tumors in their mechanisms of cell proliferation, migration, and tissue invasion.
During the process of carcinogenesis, there is an inactivation of the Slit2 and Robo2 genes, being therefore considered as tumor suppressor genes [27]. Both genes are extremely important during the process of nerve formation and repair. Both SLIT2 and ROBO2 have been reported to inhibit migration of Schwann cells [28]. In the present study, we observed that Slit2 and Robo2 mRNAs are decreased in lung cancer samples when compared to normal tissue. Also, there is a participation of miRNAs and of methylation of their promoter regions in the regulation of these genes. Similarly, both genes are targets for upregulated miRNAs. Immunohistochemical expression analyses revealed low expression of both genes in the lung. Thus, we suggest that decreased expression of Slit2 and Robo2 genes in lung cancer may favor the migration of Schwann cells; consequently, favoring invasion by neoplastic tissue.
We next analyzed the RAS/ MEK1/ ERK/ AKT/ RAF/ GSK3Β phosphorylation cascade. This pathway is activated during the proliferation process of Schwann cells and lung cancer. The authors observed that SC-treated lung cancer cells exhibit an increased phosphorylation of Akt and GSK-3β. Besides, they blocked CXCL5 expression in SC significantly decreases the phosphorylation of AKT and GSK-3β in SC-treated lung cancer cells; PI3K inhibitor LY294002 blocks the activation of AKT/GSK-3β signaling in SC-conditioned lung cancer cells. In our study, we observed that the RAS/ MEK1/ ERK/ AKT/ RAF/ GSK3Β phosphorylation is increased in LUSC when compared to LUAD, suggesting that, in LUSC samples, this pathway could be more active.
In summary, we observed that the “neuroactive ligand-receptor interaction” pathway was upregulated in lung squamous cell carcinoma and downregulated in lung adenocarcinoma. The “p53 signaling pathway” was active in both lung cancers, since Ccne1, Cdkn2a, and Perp were upregulated. Meanwhile, upregulated miRNAs inactivate the “axon guidance” pathway, targeting Robo2 and Slit2 genes. Both genes are also associated with Schwann cells migration inhibition. Also, Gfap and Gap43 are overexpressed, leading to Schwann cells dedifferentiation. We believe that Schwann cells’ dedifferentiation and proliferation are induced by neoplastic tissue; consequently, Schwann cells produce different factors that will participate in several processes of tumor progression. These processes may also be involved in tumor invasion into the perineural tissue in lung cancer. Our data stimulate efforts in news studies to achieve the experimental and clinical validation about these biomolecules.
Materials and Methods
Collection and inclusion criteria of studies
We searched the GEO database (https://www.ncbi.nlm.nih.gov/geo/) for publicly available studies. After a systematic review, only one GSE studies were retrieved. The inclusion criteria for studies were as follows: (1) experiments involving both cancer and Schwann cells and, (2) gene expression profiling of mRNA. Then, one gene expression profile (GSE4030) was collected for our analysis.
Microarray data and Data processing
One gene expression profiles (GSE4030) was downloaded from the GEO database. GEO2R was applied to compare gene expression profiles in early and late passage human Schwann cells exposed to the cancer growth factors heregulin and forskolin. Because both groups in this comparison have been exposed to mitogens, differences in gene expression profiles will be interpreted as indicative of changes caused by prolonged versus short term exposure to mitogens.
GEO2R (http://www.ncbi.nlm.nih.gov/geo/geo2r/) is an interactive web tool for comparing two groups of data that can analyze any GEO series. The adjusted p-values using Benjamini and Hochberg false discovery rate method by default were applied to correct the occurrence of false positive results. An adj. P < 0.05 and a logFC ≥ 1 were set as the cut-off criteria.
Gene list collection
We used Entrez Gene from NCBI (www.ncbi.nlm.nih.gov/gene/) and GeneCards (https://www.genecards.org/), which were used as the identifiers for Schwann cell dedifferentiation-related genes. The gene list and downregulated/ upregulated proteins of a previous published data [9] were combined and identified with a Venn Diagram 2.1.0 (http://bioinfogp.cnb.csic.es/tools/venny/index.html).
Functional and pathway enrichment analysis
Gene ontology (GO) analysis of the relevant biological processes, cellular components, and molecular functions was performed using the protein analysis through evolutionary relationships program (PANTHER, www.pantherdb.org), a curated database of protein families, functions, and pathways. GO terms assigned into identified molecules were classified according to their functions [10].
The Kyoto Encyclopedia of Genes and Genomes (KEGG) is an integrated database resource for biological interpretation of genome sequences and other high-throughput data [11]. KEGG analyses were available at the DAVID database (https://david.ncifcrf.gov/), a data resource composed of an integrated biology knowledge base and analysis tools to extract meaningful biological information from large quantities of genes and protein collections. A p-value < 0.05 was set as the cut-off criterion [12].
RNA-seq and clinical information data from The Cancer Genome Atlas (TCGA)
We used TCGAbiolinks, an R/Bioconductor software (http://bioconductor.org/packages/release/bioc/ html/TCGAbiolinks.html) [13] and the interphase TCGAbiolinksGUI [14] to download genomic and clinical data of both normal and solid tumor tissues for two different types of cancer from TCGA. Selected cancer types were lung squamous cell carcinoma (LUSC) and lung adenocarcinoma (LUAD). We retrieved level data for raw count mRNA and miRNA expression (Illumina HiSeq 2000). Co-expressed upregulated and downregulated DEGs from the gene expression profiles were combined and identified with a Venn Diagram 2.1.0 (http://bioinfogp.cnb.csic.es/tools/venny/index.html). An adj. P < 0.05 and a logFC ≥ 1 were set as the cut-off criteria.
DNA methylation analyses were performed using MEXPRESS dataset (https://mexpress.be/?ref=labworm). The cancer dataset, consisting of DNA methylation data (Illumina Infinium Human Methylation 450 K Bead array, Illumina, USA) and the β value, was considered as significantly hypermethylated only if the value was found in more than 5% of the tumors [15]. Copy number alteration analysis was performed using the cBio Cancer Genomics Portal (http://cbioportal.org).
Prediction of miRNA targets
The target miRNAs of the DEGs from TCGA data were predicted with miRDB (http://mirdb.org/miRDB/), which is an online database for predicting microRNA targets [16].
Lung cancer expression analyses
To obtain individual gene-protein data, relevant information from The Cancer Proteome Atlas (TCPA) website (https://tcpaportal.org/tcpa/analysis.html) was used as the primary source of information for reverse phase protein array (RPPA) analysis. Several other web resources were used as source of information while some more were used as analysis tools for lung cancer expression studies: Immunohistochemistry image-based protein data for both normal and cancer samples are available at the Human Protein Atlas (https://www.proteinatlas.org/). LOCATE database for protein subcellular location was included on the analysis (http://locate.imb.uq.edu.au/cgi-bin/sort_search.cgi). Survival analysis of the TCGA data was performed using the Survival module of the Tumor Immune Estimation Resource (TIMER) and Prediction of clinical outcomes from genomics (PRECOG). Kaplan–Meier plots were drawn using TIMER to explore the association between clinical outcome and gene expression, and to visualize survival differences.
Statistical analysis
Statistical analyses involving copy number variation were performed using GraphPad Prism 7 software. A two-way ANOVA and a two-tailed unpaired t test were used to compare the means between groups.
Note
LOCATE database for protein subcellular location was included in the analysis (http://locate.imb.uq.edu.au/cgi-bin/sort_search.cgi), that was used in the study design has been removed by the owner and is no longer available.
ACKNOWLEDGMENTS AND FUNDING
This study was supported by grants from the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Estado de Alagoas (FAPEAL), Fundação de Amparo à Ciência e Tecnologia de Pernambuco (FACEPE), Programa de Pós-graduação Multicêntrico de Bioquímica e Biologia Molecular (PMBqBM) and, Programa de Pós-graduação em Ciências Médicas (PPGCM).
Novelty and Significance: What Is New? This is the first study showing that Schwann cells is associated with lung cancer, by alteration in mRNA, miRNA, methylation and protein expressions, deregulating several pathways such as axon guidance and neuroactive ligand receptor-interaction. What Is Relevant? Pathway in cancer were enriched for the upregulated differentially expressed genes (DEGs) in Schwann cells after treatment with tumor growth factors. Axon guidance pathway and correspondent genes Slit2 and Robo2 was inactive in cancer samples. Neuroactive ligand receptor-interaction was enriched for upregulated DEGs in lung cancer. Gfap (protein and mRNA) was upregulated in lung cancer samples. Gap43 had a negative impact on survival in lung adenocarcinoma samples.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
Ethics approval and consent to participate
Not applicable.
References
1. Deborde S, Omelchenko T, Lyubchik A, Zhou Y, He S, McNamara WF, Chernichenko N, Lee SY, Barajas F, Chen CH, Bakst RL, Vakiani E, He S, et al. Schwann cells induce cancer cell dispersion and invasion. J Clin Invest. 2016; 126:1538–54. https://doi.org/10.1172/JCI82658. [PubMed].
2. Gokey NG, Srinivasan R, Lopez-Anido C, Krueger C, Svaren J. Developmental regulation of MicroRNA expression in Schwann cells. Mol Cell Biol. 2012; 32:558–68. https://doi.org/10.1128/MCB.06270-11. [PubMed].
3. Napoli I, Noon LA, Ribeiro S, Kerai AP, Parrinello S, Rosenberg LH, Collins MJ, Harrisingh MC, White IJ, Woodhoo A, Lloyd AC. A central role for the ERK-signaling pathway in controlling Schwann cell plasticity and peripheral nerve regeneration in vivo. Neuron. 2012; 73:729–42. https://doi.org/10.1016/j.neuron.2011.11.031. [PubMed].
4. Azam SH, Pecot CV. Cancer’s got nerve: Schwann cells drive perineural invasion. J Clin Invest. 2016; 126:1242–4. https://doi.org/10.1172/JCI86801. [PubMed].
5. Demir IE, Boldis A, Pfitzinger PL, Teller S, Brunner E, Klose N, Kehl T, Maak M, Lesina M, Laschinger M, Janssen KP, Algül H, Friess H, et al. Investigation of Schwann cells at neoplastic cell sites before the onset of cancer invasion. J Natl Cancer Inst. 2014; 106. https://doi.org/10.1093/jnci/dju184. [PubMed].
6. Wang C, Slikker W Jr, Zhang X, Patterson TA, Paule MG, Wang C, Slikker W. Schwann cells. Neural Cell Biology. 2017; 14–18. https://doi.org/10.1201/9781315370491.
7. Deborde S, Wong RJ. How Schwann cells facilitate cancer progression in nerves. Cellular and Molecular Life Sciences. 2017; 24:4405–4420. https://doi.org/10.1007/s00018-017-2578-x. [PubMed].
8. Webber CA, Christie KJ, Cheng C, Martinez JA, Singh B, Singh V, Thomas D, Zochodne DW. Schwann cells direct peripheral nerve regeneration through the Netrin-1 receptors, DCC and Unc5H2. Glia. 2011; 10:1503–17. https://doi.org/10.1002/glia.21194. [PubMed].
9. Tenzer S, Leidinger P, Backes C, Huwer H, Hildebrandt A, Lenhof HP, Wesse T, Franke A, Meese E, Keller A. Integrated quantitative proteomic and transcriptomic analysis of lung tumor and control tissue: a lung cancer showcase. Oncotarget. 2016; 12:14857–70. https://doi.org/10.18632/oncotarget.7562. [PubMed].
10. Bastos HP, Tavares B, Pesquita C, Faria D, Couto FM. Application of gene ontology to gene identification. Methods Mol Biol. 2011; 760: 141–57. https://doi.org/10.1007/978-1-61779-176-5_9. [PubMed].
11. Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016; 44: D457–62. https://doi.org/10.1093/nar/gkv1070. [PubMed].
12. Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009; 4: 44–57. https://doi.org/10.1038/nprot.2008.211. [PubMed].
13. Colaprico A, Silva TC, Olsen C, Garofano L, Cava C, Garolini D, Sabedot TS, Malta TM, Pagnotta SM, Castiglioni I, Ceccarelli M, Bontempi G, Noushmehr H. TCGAbiolinks: An R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Res. 2016; 44: e71. https://doi.org/10.1093/nar/gkv1507. [PubMed].
14. Silva TC, Colaprico A, Olsen C, Malta TM, Bontempi G, Ceccarelli M, Berman BP, Noushmehr H. TCGAbiolinksGUI: A graphical user interface to analyze cancer molecular and clinical data. F1000Research. 2018; 7: 439. https://doi.org/10.12688/f1000research.14197.1.
15. Guo S, Yan F, Xu J, Bao Y, Zhu J, Wang X, Wu J, Li Y, Pu W, Liu Y, Jiang Z, Ma Y, Chen X, et al. Identification and validation of the methylation biomarkers of non-small cell lung cancer (NSCLC). Clin Epigenetics. 2015; 7:3. https://doi.org/10.1186/s13148-014-0035-3. [PubMed].
16. Wong N, Wang X. miRDB: An online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015; 43: D146–52. https://doi.org/10.1093/nar/gku1104. [PubMed].
17. Zhou H, Huang S. Role of mTOR signaling in tumor cell motility, invasion and metastasis. Curr Protein Pept Sci. 2011; 1:30–42. https://doi.org/10.2174/138920311795659407. [PubMed].
18. Lo HC, Zhang XH. EMT in Metastasis: Finding the Right Balance. Developmental Cell. 2018; 6:663–665. https://doi.org/10.1016/j.devcel.2018.05.033. [PubMed].
19. Shaw R. The epigenetics of oral cancer. International Journal of Oral and Maxillofacial Surgery. 2006; 35:101–108. https://doi.org/10.1016/j.ijom.2005.06.014. [PubMed].
20. Namkoong J, Shin SS, Hwa JL, Marín YE, Wall BA, Goydos JS, Chen S. Metabotropic glutamate receptor 1 and glutamate signaling in human melanoma. Cancer Res. 2007; 5:2298–305. https://doi.org/10.1158/0008-5472.CAN-06-3665. [PubMed].
21. Wen Y, Li J, Koo J, Shin SS, Lin Y, Jeong BS, Mehnert JM, Chen S, Cohen-Sola KA, Goydos JS. Activation of the glutamate receptor GRM1 enhances angiogenic signaling to drive melanoma progression. Cancer Res. 2014; 9:2499–509. https://doi.org/10.1158/0008-5472.CAN-13-1531. [PubMed].
22. Cheng Q, Wang YX, Yu J, Yi S. Critical signaling pathways during wallerian degeneration of peripheral nerve. Neural Regen Res. 2017; 6: 995–1002. https://doi.org/10.4103/1673-5374.208596. [PubMed].
23. Kim HS, Lee J, Lee DY, Kim YD, Kim JY, Lim HJ, Lim S, Cho YS. Schwann Cell Precursors from Human Pluripotent Stem Cells as a Potential Therapeutic Target for Myelin Repair. Stem Cell Reports. 2017; 6:1714–1726. https://doi.org/10.1016/j.stemcr.2017.04.011. [PubMed].
24. Beirowski B, Wong KM, Babetto E, Milbrandt J. mTORC1 promotes proliferation of immature Schwann cells and myelin growth of differentiated Schwann cells. Proc Natl Acad Sci U S A. 2017; 21:E4261–E4270. https://doi.org/10.1073/pnas.1620761114. [PubMed].
25. Clements MP, Byrne E, Camarillo Guerrero LF, Cattin AL, Zakka L, Ashraf A, Burden JJ, Khadayate S, Lloyd AC, Marguerat S, Parrinello S. The Wound Microenvironment Reprograms Schwann Cells to Invasive Mesenchymal-like Cells to Drive Peripheral Nerve Regeneration. Neuron. 2017; 1:98–114.e7. https://doi.org/10.1016/j.neuron.2017.09.008. [PubMed].
26. Zhou Y, Shurin GV, Zhong H, Bunimovich YL, Han B, Shurin MR. Schwann cells augment cell spreading and metastasis of lung cancer. Cancer Res. 2018; 20:5927–5939. https://doi.org/10.1158/0008-5472.CAN-18-1702. [PubMed].
27. Gara RK, Kumari S, Ganju A, Yallapu MM, Jaggi M, Chauhan SC. Slit/Robo pathway: A promising therapeutic target for cancer. Drug Discovery Today. 2015; 1:156–64. https://doi.org/10.1016/j.drudis.2014.09.008. [PubMed].
28. Carr L, Parkinson DB, Dun XP. Expression patterns of Slit and Robo family members in adult mouse spinal cord and peripheral nervous system. PLoS One. 2017; 2:e0172736. https://doi.org/10.1371/journal.pone.0172736. [PubMed].

