INTRODUCTION
The circadian clock system in mammals is hierarchically organized comprising a central master clock and peripheral clocks [1, 2]. The central clock localizes to the suprachiasmatic nuclei (SCN) of the anterior hypothalamus and is mainly synchronized by environmental light [3, 4]. In contrast, synchronization of peripheral clocks, which are present in nearly all tissues and organs, occurs by hormonal and neuronal cues and/or food intake [5–7]. In central and peripheral clocks, the molecular clockwork is composed of intertwined positive and negative feedback loops at the transcriptional and (post)translational level [4, 8–10]. In particular, core molecular clockwork components encoded by so-called clock genes regulate positively (Clock, Bmal1) or negatively (Cry1/Cry2 and Per1/Per2) rhythmic expression of their target genes (i.e., clock output genes) through transcriptional control via so-called E-box elements [11]. Important clock output genes include members of the family of PAR-domain basic leucine zipper (PAR bZip) transcription factors, i.e. hepatic leukemia factor (HLF), thyrotrophic embryonic factor (TEF), and albumin D-box binding protein (DBP) [12]. The latter activates gene expression of Per1/2 via DBP binding sites (so-called D-box elements) therefore modulating the levels of core clock gene products [13, 14]. Moreover, PAR bZip proteins control the circadian expression of hepatic enzymes and regulators involved in endobiotic and xenobiotic biotransformation and drug metabolism [15], including isoenzymes of the cytochrome P450 family of monooxygenases.
Several findings indicate an important role of the Gs/Gi-protein-controlled cAMP-CREB signaling pathway in the regulation of clock gene expression. Treatment of Rat-1 fibroblasts with forskolin, which activates adenylyl cyclase and therefore increases intracellular cAMP concentrations, resulted in elevated pSer133-CREB levels concomitant with an enhanced circadian accumulation of DBP, Per1, and Per2 [16]. Moreover, the cAMP-CREB signaling pathway positively regulates Per1 expression in human hepatoma cells [17] as well as the light-induced Per1 expression in the SCN [18, 19]. In the SCN, CREB is rapidly phosphorylated at Ser133 after light at night [20–22], but circadian oscillation of intracellular cAMP is also important for intercellular coupling of SCN neurons [23]. In liver, intracellular cAMP concentrations and CREB activation are modulated by Cry1 [24].
Signaling by GPCRs via heterotrimeric Gs (i.e., Gαsβγ) and Gi (i.e., Gαiβγ) proteins activates or inhibits, respectively, cAMP production by adenylyl cyclases, thereby regulating downstream cAMP-PKA (protein kinase A)-CREB signaling [25]. The Gαi subunits comprise three highly homologous isoforms, Gαi1, Gαi2, and Gαi3 [26, 27], which display a high amino acid sequence identity together with overlapping as well as cell- and tissue-type specific expression profiles. Overall, Gαi2 represents the predominantly and ubiquitously expressed isoform, whereas Gαi3 is found in all peripheral tissues with only low detectable expression in neuronal cells [28–30]. Of note, both Gαi2 and Gαi3 are expressed in the liver where Gαi3 exhibits an isoform-specific crucial function in the anti-autophagic action of insulin in hepatocytes [29]. However, the function of pertussis toxin-sensitive Gi proteins in circadian signaling and gene expression is largely unclear.
Here, we analyzed the role of Gαi3 in rhythmic pSer133-CREB activation and clock gene expression in the liver in mice. The phase of clock genes Per1, Per2, Bmal1, Rev-Erba, Cry1, and Cry2 as well as clock output genes Hlf and Tef was not different between Gαi3−/− and wild-type littermates of both genders. In contrast, in Gαi3−/− females the phase of the clock output gene DBP was advanced. This was associated with a slight increase in pCREB and DBP protein levels in females. Moreover, the expression level of the DBP target gene and cytochrome P450 family member Cyp3a11 were significantly higher in females at mid-day and mid-night, and in males at early day. Thus, our data indicate that the Gαi3-pCREB signaling pathway functions as a novel direct (via DNA binding) or indirect regulator of sexual-dimorphic hepatic expression of DBP and its xenobiotic targets.
RESULTS
Rhythmic clock gene expression is not affected by Gαi3 deficiency
The cAMP-CREB signaling pathway, that is under the control of heterotrimeric Gs and Gi proteins, regulates the expression of core clock and clock-regulated genes, including Per1 and DBP [16, 17]. To characterize the role of the Gαi isoform Gαi3 in this pathway, Gαi3 deficient mice [29] and wild-type control animals were kept in a 12 h light/12 h dark cycle setting and liver tissue was collected every 6 h for up to 24 h followed by gene expression analysis via quantitative real-time polymerase chain reaction (qRT-PCR). As depicted in Figure 1, the phases of rhythms in transcript levels of the core molecular clockwork components (Per1, Per2, Bmal1, Rev-erba, Cry1, and Cry2) and clock-regulated genes (Hlf and Tef) and their expression levels were comparable between Gαi3−/− mice and wild-type controls of both genders. In male Gαi3−/− mice, the peak in Cry1 expression at ZT18 was significantly higher as compared to wild-type controls (Figure 1B). To address these molecular findings at a functional level, we next isolated primary tail fibroblasts from wild-type and Gαi3−/− female mice followed by lentiviral transduction of a Bmal1 promoter-driven luciferase reporter construct and subsequent real-time bioluminescence monitoring. As indicated in Supplementary Figure 1A, and consistent with the expression profiling results in Figure 1, recording of luciferase activity revealed a circadian period length that was nearly identical between Gαi3 deficient fibroblasts and wild-type cells (25.1 vs. 25.0 h, respectively; Supplementary Figure 1B).
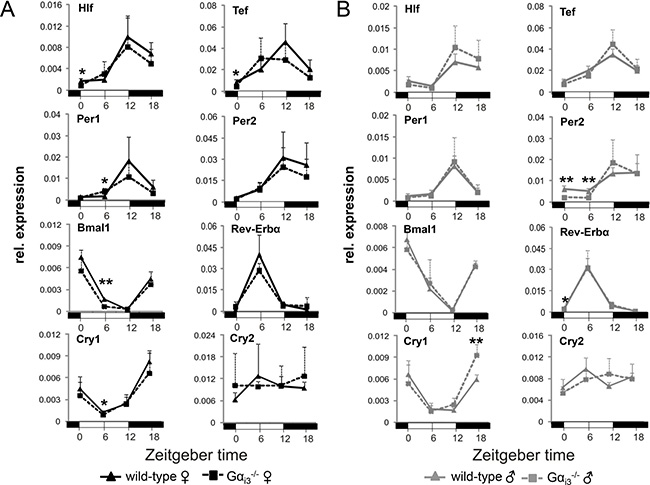
Figure 1: Rhythmic expression of core clock genes and clock-regulated genes in the liver of Gαi3-/- mice. Quantitative real-time PCR analysis of rhythmic expression of core clock genes and clock output genes in the liver of female (A) and male (B) Gαi3 deficient mice vs. wild-type control animals. Mice were sacrificed and analyzed every six hours at the indicated time points (ZT 0 to ZT 18). Transcript levels of the indicated genes were normalized to the endogenous control Gapdh. Shown are 2-ΔCt values. Results are expressed as mean ± s.d. of six animals analyzed per genotype and time point (*p < 0.05; **p < 0.01 as compared to corresponding wild-type control animals).
Gαi3 deficient female mice exhibit a phase-shifted rhythm of hepatic DBP expression
Interestingly, in contrast to the clock output genes and PAR bZip members Tef and Hlf, whose transcript levels were apparently not influenced by Gαi3 deficiency (Figure 1A), rhythmic expression of the PAR bZip member DBP was phase advanced by six hours in Gαi3−/− females as compared to wild-type females (Figure 2A and Supplementary Figure 2A). Here, comparable results were obtained using Gapdh, β-Actin, or EF1a as controls to normalize gene expression analysis. This phase-shifted expression was associated with increased DBP protein levels in Gαi3−/− female vs. wild-type livers during the dark phase (ZT12 and ZT18) (Figure 2C). In contrast, the rhythm in hepatic expression of DBP in male mice was comparable in both genotypes at the mRNA (Figure 2B and Supplementary Figure 2B) and protein level (Figure 2D). DBP mRNA levels in male mice showed the same peak as wild-type females peaking at ZT12 similarly as previously described for C57Bl/6 wild-type mice [15]. The sexual dimorphic, phase-shifted expression of DBP became particularly evident when the data from Figure 2A, B were compared between the wild-type (Supplementary Figure 3A) and Gαi3 knockout (Supplementary Figure 3B) of both sexes. In contrast, an analogous re-analysis of the data from Figure 1 revealed rather comparable circadian rhythms in the transcript levels of core molecular clockwork components (with the exception of Per1 and Per2 which displayed slightly to significantly higher transcript levels in females as compared to males of both genotypes) and other clock-regulated genes (like Hlf and Tef) (Supplementary Figure 4). Of note, Gαi3 in the liver per se showed no rhythmic expression at the mRNA level (data now shown) or the protein level (Supplementary Figure 5). The latter is consistent with a lack of temporal changes of Gαi amounts in the murine SCN [23]. Taken together, hepatic expression of the PAR bZip member DBP in female mice seems to underlie a sexual-dimorphic regulation that is dependent on Gαi3.
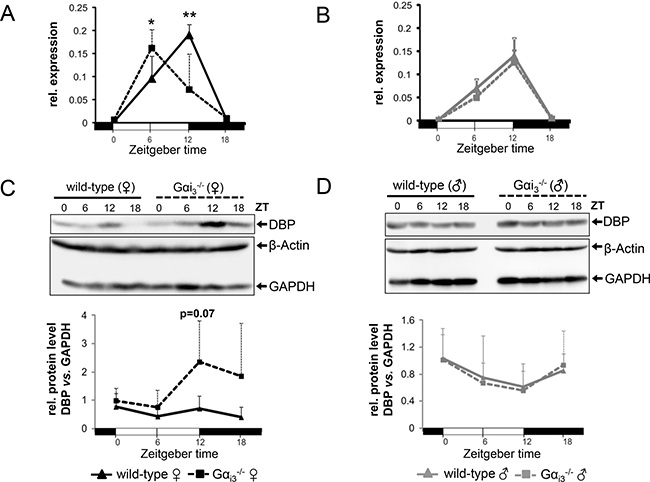
Figure 2: Rhythmic expression of DBP in the liver of female Gαi3-/- mice is phase advanced (A, B) Quantitative real-time PCR analysis of rhythmic expression of DBP mRNA in the liver of female (A) and male (B) Gαi3 deficient mice as compared to wild-type control animals. Dbp transcript levels were normalized to the endogenous control Gapdh. Shown are 2-ΔCt values. (C, D) Representative immunoblots of rhythmic expression of the DBP protein in the liver of female (C) and male (D) Gαi3-deficient mice and wild-type controls. GAPDH and β-Actin were employed as loading controls. Relative protein levels (DBP/GAPDH) were determined by densitometric analysis using ImageJ software (lower panels in C and D). Mice were sacrificed and analyzed every six hours at the indicated time points (ZT 0 to ZT 18). Results are expressed as mean ± s.d. of six animals (mRNA) or four animals (immunoblot) analyzed per genotype and time point (*p < 0.05; **p < 0.01 as compared to corresponding wild-type control animals).
Evidence for increased circadian cAMP-CREB signaling in the liver of Gαi3 deficient female mice
We next addressed the role of Gαi3 in the GPCR/Gαi controlled cAMP-CREB signaling pathway as an important regulator of hepatic expression of core clock and clock-regulated genes [16, 17, 19]. Initial analysis of total cell lysates of liver tissue indicated that the cellular levels of pCREB (pSer133-CREB) were higher in Gαi3−/− female mice as compared to corresponding wild-type female controls (Supplementary Figure 6A). In contrast, total pCREB levels were rather comparable between male Gαi3−/− and wild-type mice (as judged by pCREB/CREB signal ratios) thereby following a similar circadian rhythm (Supplementary Figure 6B). Given that pSer133-CREB localizes in the nucleus thereby functioning as transcription factor [31], we next analyzed nuclear extracts isolated from liver tissue. Consistent with the findings above, Gαi3−/− female mice displayed slightly higher nuclear levels of pCREB as compared to wild-type females (Figure 3A, 3B), whereas there was no obvious difference in nuclear pCREB levels between male Gαi3−/− and wild-type mice (Supplementary Figure 7). These findings indicate a sexual-dimorphic inhibitory impact of Gαi3 on the cAMP-CREB pathway and suggest that the Gαi3-pCREB axis may function as a candidate determinant of female specific expression of DBP. The important role of CREB is emphasized by the observation that selective inhibition of CREB led to a clear decrease in the expression of DBP (Supplementary Figures 9 and 10).
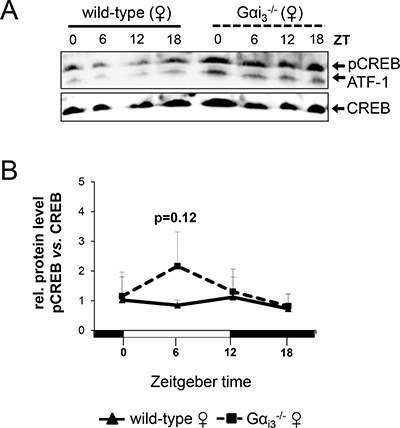
Figure 3: Gαi3-/- female mice display increased nuclear pSer133-CREB levels in the liver. (A) Immunoblot analysis of pSer133-CREB and CREB protein levels in nuclear extracts from livers of female wild-type vs. Gαi3-/- mice. (B) Quantitative analysis of relative protein levels (pSer133-CREB/CREB) was performed by ImageJ software. Results are expressed as mean ± s.d. of three independent experiments. Mice were sacrificed and analyzed every six hours at the indicated time points (ZT 0 to ZT 18). ATF1, activating transcription factor 1.
Elevated transcript levels of the DBP cytochrome P450 target gene Cyp3a11 in Gαi3 deficient female mice
DBP drives transcriptionally the rhythmic expression of the xeno- and endobiotic target genes CYP2A4 (Steroid 15α-hydroxylase), CYP2A5 (Coumarin 7-hydroxylase), and CYP3A4 (Cyp3a11 in mouse) [32], which are involved in cholesterol/steroid metabolism or biotransformation of lipophilic xenobiotics. In Gαi3 deficient females, transcript levels of Cyp3a11 at ZT06 and ZT18 were significantly higher as compared to levels in the corresponding wild-type females (Figure 4A). In comparison, Gαi3 deficient males displayed levels of Cyp3a11 that were significantly higher at ZT0 as compared to corresponding wild-type males (Figure 4B). In contrast, levels of CYP2A4/CYP2A5 were not different between Gαi3 deficient and wild-type mice of both genders (Figure 4A, 4B). Taken together, upon Gαi3 deficiency, increased signaling by the cAMP-pCREB axis seems to translate into higher hepatic Cyp3a11 gene expression that was more pronounced in female mice and potentially dependent on transcriptional regulation via DBP. The important role of CREB signaling is emphasized by a clear decrease in the expression of DBP and its target gene CYP3A4 in human HepG2 cells upon selective inhibition of CREB (Supplementary Figures 9 and 10).
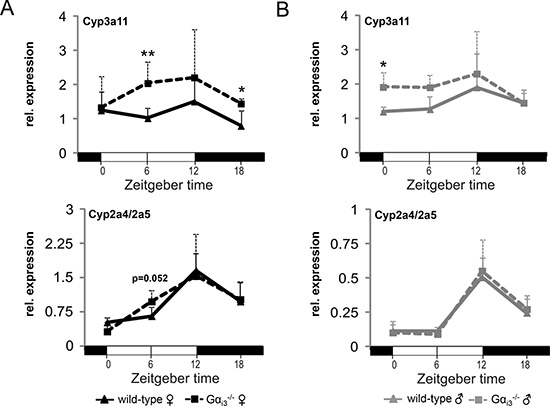
Figure 4: Transcript levels of the DBP target gene and cytochrome P450 family member Cyp3a11 are increased in the liver of Gαi3-/- female mice. Quantitative real-time PCR analysis of rhythmic expression of Cyp3a11 and Cyp2a4/a5 genes in the liver of female (A) and male (B) Gαi3 deficient mice vs. wild-type control animals. Mice were sacrificed and analyzed every six hours at the indicated time points (ZT 0 to ZT 18). Transcript levels of Cyp3a11 and Cyp2a4/a5 were normalized to the endogenous control Gapdh. Shown are 2-ΔCt values. Results are expressed as mean ± s.d. of five animals analyzed per genotype and time point (*p < 0.05; **p < 0.01 as compared to corresponding wild-type control animals).
DISCUSSION
This study provides novel insight into the role of the heterotrimeric G protein Gαi3 as upstream regulator of the cAMP-pCREB signalling pathway in rhythmic gene expression in the liver. Our data indicate (i) that gene ablation of Gαi3 in mice has only modest effects on overall core clock and clock output gene expression and does not affect the period length of clock gene expression; (ii) interestingly, slightly increased levels of nuclear activated CREB (pSer133-CREB) which are linked to a phase-shifted and increased expression of its potential target gene and PAR bZip transcription factor DBP is detectable in livers from Gαi3 deficient female mice; (iii) accordingly, the DBP target gene and cytochrome P450 family member Cyp3a11 is found at higher transcript levels in the liver of Gαi3−/− female mice. Thus, Gαi3 may function as novel regulator of sexual-dimorphic, cAMP-PKA-pCREB driven expression of DBP together with its xenobiotic target genes in the liver (Figure 5). Of note, selective inhibition of CREB led to a clearly decreased expression of DBP and CYP3A4 (human Cyp3a11 homologue) in HepG2 hepatoma cells (Supplementary Figures 9 and 10) further stressing the downstream role of CREB in the regulation of DBP and its target genes.
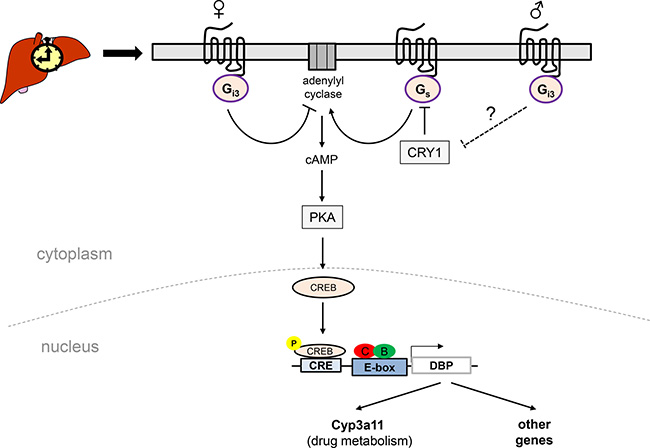
Figure 5: Schematic model suggesting a putative role of the Gas/Gαi3-CREB signalling pathways in sexually-dimorphic regulation of DBP expression in female vs. male mouse liver. In females, loss of Gαi3 leads presumably via desinhibition of adenylyl cyclase and hence increased pSer133-CREB levels to a phase-advanced and increased expression of DBP in the liver, concomitant with an increase in transcript levels of Cyp3a11, a putative DBP target gene. In Gαi3-/- males, this signalling pathway may be less active provided that an increased CRY1 expression (as indicated in Figure 1) results in a stronger inhibitory role of CRY1 on cAMP levels (via Gs inhibition) [24]. B: BMAL1; C: CLOCK; CRE: cAMP responsive element; CREB, cAMP response element binding protein; CRY1, cryptochrome circadian clock 1; Cyp3a11: cytochrome P450, family 3, subfamily a, polypeptide 11; DBP: albumin D-box binding protein; PKA: protein kinase A; Gi3 and Gs: heterotrimeric guanine nucleotide binding proteins which upon activation cause a decrease or increase in intracellular cAMP levels, respectively.
The phenotypic characterization of Gαi deficient mouse lines has been expanded in the recent years revealing that Gαi2 and Gαi3 take over gene-specific as well as shared physiological functions in various organs and cell types in-vivo [28, 29, 33–41]. The liver expresses both Gαi2 and Gαi3 with the latter exhibiting an isoform-specific and crucial function in the anti-autophagic action of insulin in hepatocytes [29]. Although a comparative characterization of the hepatic expression of core clock and clock-regulated genes in Gαi2 knockout mice is not yet available, we provide evidence that Gαi3 deficiency per se alters expression of DBP and its target genes in a Gαi isform-specific manner apparently via a desinhibited cAMP-pCREB pathway.
Several observations emphasize a critical role of GPCR/Gs (cAMP↑) and GPCR/Gi (cAMP↓) pathways in the regulation of rhythmic gene expression. Interestingly, Cry1 interacts directly with Gas at the GPCR level and thereby inhibits adenylyl cyclase activity and accumulation of cAMP and activation of CREB [24]. These findings provided an explanation how the rhythmic expression of Cry1 translates directly into a circadian regulation of cAMP signaling in hepatic glucose metabolism, in particular gluconeogenesis [24]. Moreover, functional inactivation of Gi-proteins by pertussis toxin [42] or a reduction of cAMP levels by MDL-12330A mediated inhibition of adenylyl cyclase led to a desynchronized circadian gene expression in the SCN and an altered circadian rhythmicity [43, 44]. Here, although the identity of the involved GPCRs in the regulation of circadian signaling is currently only speculative, the Gi/Gq-coupled melatonin receptors MT1 and MT2 [45] represent prime candidates given that they play an important role in central clock synchronization. MT1 and MT2 are both expressed in the liver [46] where they display circadian rhythmic changes in gene expression [47]. Interestingly, and consistent with our findings in Figure 2, DBP mRNA levels are phase-advanced in mice lacking either MT1, MT2, or both receptors [46, 48], indicating that these GPCRs are involved in the regulation of DBP expression. However, since the mouse lines analyzed in this study were on the melatonin-deficient C57Bl/6 genetic background [49], backcrossing with mice on the melatonin proficient C3H background is a prerequisite to ultimately determine changes in melatonin receptor signaling and rhythmic hepatic gene expression upon Gαi3 deficiency.
In terms of the regulation of DBP expression, previous work showed that maximal and minimal levels of DBP mRNA occur approximately four hours later in the liver as compared to the SCN [50], suggesting that additional regulatory mechanisms (besides transcriptional control via E-box elements) must be involved in the control of rhythmic DBP expression in peripheral tissues when compared to the SCN [13, 51–54]. Indeed, treatment of Rat-1 fibroblasts with the adenylyl cyclase activator forskolin was sufficient to increase cAMP-pCREB signalling and trigger circadian expression of DBP [16, 55]. These findings indicate that candidate cAMP inducible promoter elements (CRE) and pCREB binding sites [56], which are regulated by Gαi3-mediated signaling, are present in the Dbp gene (“direct model”). Indeed, besides the two E-box binding sites (5’-CACGTC-3’), two CRE half sites (5’-CGTCA-3’) can be detected in the promoter region of the murine Dbp gene (Singh and Piekorz, unpublished). However, it remains to be determined whether these sites function as pCREB binding elements. Alternatively, one could envision that the subcellular (nuclear vs. cytoplasmic) distribution of BMAL-1/CLOCK, that regulates Dbp gene expression through E-box binding [57], has become disturbed by Gαi3 deficiency in hepatocytes (“indirect model”). So far, however, we failed to detect any obvious differences in nuclear vs. cytoplasmic CLOCK protein levels between wild-type and Gαi3 deficient hepatocytes at ZT6 and ZT12 (Supplementary Figure 11).
Increasing evidence argues for a role of sexual dimorphism in life span regulation, disease sensitivity, and particularly drug metabolism in females vs. males [3, 58–61]. The liver represents a bona fide sexual-dimorphic organ with a rhythmic physiology especially in terms of its detoxification function [62, 63]. In these regards, indications for a sexual dimorphic signalling and expression of core clock and clock regulated genes in the liver include based on our study (i) slightly to significantly higher transcript levels of Per1 and Per2 in females vs. males (Supplementary Figure 4), (ii) potential differences in pCREB signalling given the levels and rhythms of pSer133-CREB in female vs. male animals (Supplementary Figure 6; Figure 3 and Supplementary Figure 7), (iii) a phase-shifted expression of DBP that became particularly evident when comparing wild-type (Supplementary Figure 3A) and Gαi3 knockout (Supplementary Figure 3B) of both sexes, and (iv) a significantly higher circadian expression of CYP2A4/2A5 in females vs. males independent of the genotype (Supplementary Figure 8). As summarized in the model in Figure 5, in Gαi3−/− males, the cAMP-PKA-pCREB signalling pathway may be less active provided that the increased CRY1 expression (Figure 1B) results in a stronger inhibitory role of CRY1 on cAMP levels (via Gs inhibition) essentially as suggested by Zhang et al. [24]. Of note, the increased CRY1 expression in Gαi3−/− males could be due to a phase shift/earlier onset of Cry1 expression between ZT12 and ZT18, given that ZT intervals of six hours (instead of four hours) have been analyzed in this study.
CYP2A4/2A5 belongs to the cytochrome P450 family of monooxygenases, whose expression is under diurnal control by the circadian clock via PAR bZip proteins, including DBP. These transcription factors also control the expression of additional enzymes and regulators involved in endobiotic and xenobiotic biotransformation and drug metabolism [15]. In particular, DBP per se is able to transcriptionally regulate the circadian accumulation of the target genes CYP2A4 (Steroid 15α-hydroxylase), CYP2A5 (Coumarin 7-hydroxylase), and CYP3A4 (Cyp3a11 in mouse [32]), given that these genes display D-box elements in their promotor regions [64, 65]. CYP2A4/CYP2A5 is involved in cholesterol and bile acid metabolism [64], whereas CYP3A4/Cyp3a11 plays an important role in the metabolism of endogenous steroids and is responsible for biotransformation of approximately 50% of all prescription drugs [65, 66]. The profound role of PAR bZip family members especially in the regulation of drug metabolism and biotransformation [15] becomes particularly obvious in triple knockout mice lacking Dbp, Hlf, and Tef. These animals display a deregulated expression of numerous cytochrome P450 gene family members, an increased sensitivity to xenobiotics, and phenotypic alterations reminiscent of premature aging [12].
In summary, taking our findings in account it remains to be particularly tested whether an increased signaling by the cAMP-pCREB-DBP axis in the liver of Gαi3−/− female mice translates into an increased Cyp3a11 activity and therefore improved biotransformation.
MATERIALS AND METHODS
Mice
All experiments were performed with Gαi3-deficient mice on a C57Bl/6 background with corresponding C57BL/6 wild-type animals used as controls. For breeding and maintenance, mice were kept under specific pathogen-free conditions (SPF) with 12 h light / 12 h dark cycles and free access to food and water at the local animal house of the Heinrich-Heine-University Düsseldorf. The study was performed in accordance with the national and local guidelines on animal care.
Isolation of liver tissue
Experimental animals (male and female mice between twelve to eighteen weeks of age) were maintained under standard conditions with 12 h light / 12 h dark cycles. Animals were killed by cervical dislocation and livers were obtained at the Zeitgeber times (ZT) 0, 6, 12, and 18 (ZT 0: Light ON; ZT 12: Light OFF) and immediately frozen in aliquots in liquid nitrogen for further analysis.
RNA isolation and quantitative real-time PCR (qRT-PCR)
Total RNA from liver tissue was extracted using RNeasy mini kit spin columns (QIAGEN). RNA purity and concentration was determined using a NanoDrop spectrophotometer. One μg of liver RNA was reverse-transcribed using oligo(dT)15 primers and the ImProm™ II Reverse Transcription System (Promega) according to the manufacturer’s specifications. Relative quantification of mRNA was carried out using quantitative real-time PCR (qRT-PCR; 7500 Real-Time PCR System; Applied Biosystems) and specific TaqMan probes (Applied Biosystems) for core clock genes and clock regulated genes [67]. Expression of target sequences was normalized to an endogenous control, glyceraldehyde-3-phosphate dehydrogenase (GAPDH: Part No.: 4331182, Assay-ID: Mm99999915_g1). For further normalization to other endogenous controls, β-Actin and EF1a were used (Schneider et al., 2014). qRT-PCR for cytochrome P450 isoforms was performed using TaqMan probes for Cyp3a11 (Part No.: 4331182, Assay-ID: Mm00731567_m1) and Cyp2a4/Cyp2a5 (Part No.: 4331182, Assay-ID: Mm00487248_g1).
Total cell lysates and immunoblot analysis
Total cell lysates from mouse liver were prepared as described [68] with minor modifications. Briefly, snap-frozen liver tissue was homogenized with an Ultra-Turrax for 30–60 s at 4° C in 800 μl of lysis buffer (50 mM Tris-HCl, pH 8.0; 150 mM NaCl; 1 mM EDTA; 1% Triton-X100; 0.5% deoxycholic acid; 1% SDS) supplemented with cocktails of protease inhibitors (Roche) and phosphatase inhibitors (Sigma-Aldrich)]. After sonication for 45 seconds, tissue homogenates were cleared by a centrifugation step (30.000 × g, 15 min). Lysates were subjected to protein electrophoresis employing 8.5–10% SDS-acrylamide gels or 6 M urea/SDS-PAGE gels and blotted onto nitrocellulose membrane [29]. Membranes were blocked in 5% non-fat dry milk and incubated with primary antibodies at 4° C overnight (anti-Gαi3, Santa Cruz sc-262, 1:1000; anti-DBP, abcam ab22824, 1:1000; anti-phospho-CREB [pSer133-CREB], Cell Signaling 87G3 #9198, 1:1000; anti-CREB, Cell Signaling 48H2 #9197, 1:1000; anti-GAPDH, abcam ab8245, and anti-Actin C4, Millipore, both 1:5000). The anti-phospho-CREB antibody recognizes the phosphorylated form of CREB (pSer133-CREB) and the activating transcription factor-1 (ATF-1). Nuclear and cytoplasmatic CLOCK levels were detected using a primary antibody from Cell Signaling (D45B10, rabbit mAb #5157; 1:1000). Antibodies against total Histone (Cell Signaling #9715, 1:1000) or Actin (C4, Millipore, 1:5000) was employed to control protein loading of nuclear and cytoplasmatic fractions, respectively. Following three washing steps membranes were incubated with a horseradish peroxidase (HRP)-conjugated anti-rabbit IgG antibody (Cell Signaling #7074) or a HRP-conjugated polyclonal rabbit-anti-mouse IgG (Dako P0161) both at a dilution of 1:5000 for 60 min at room temperature. Protein signals were visualized by the ECL detection system (GE Healthcare) and images were collected using an INTAS chemostar imager.
Isolation of nuclear proteins from mouse liver
Nuclear proteins from 12–18 weeks old Gαi3−/− and wild-type livers (ZT 0, 6, 12, and 18) were prepared essentially as described [69]. In brief, 0.5–0.6 g of liver tissue was washed in phosphate buffered saline (PBS) buffer (pH 7.4) and disintegrated in 4 ml of ice-cold buffer A (250 mM sucrose, 5 mM MgCl2, 10 mM Tris-HCl [pH 7.4]) using 5 ml syringe plungers and 40 μm pore size cell strainers. Samples were centrifuged at 800 × g for 10 min at 4° C. The supernatant was collected as cytoplasmatic fraction and stored at −80° C for further analysis. The pellet was gently resuspended in 14 ml of ice-cold buffer A and centrifuged at 1000 × g for 10 min. The resulting pellet was resuspended in 1 ml of ice-cold buffer B (2.0 M sucrose, 1 mM MgCl2, 10 mM Tris-HCl [pH 7.4]) followed by a centrifugation step at 16.000 × g and 4° C for 30 min. The final pellet was resuspended in Laemmli SDS sample buffer and kept at −80° C for further analysis. Alternatively, subcellular fractionation of liver tissue was performed using the Nuclear Extract Kit (Version D3) from Active Motif, Inc. (La Hulpe, Belgium) essentially as decribed by the manufacturer.
Densitometric analysis
Quantitative immunoblot analysis was performed using ImageJ software (http://imagej.nih.gov/ij/).
Statistical analysis
Results are given as mean ± s.d. To evaluate statistical significance, Student’s t-tests or ANOVA tests were performed. P-values ≤ 0.05 were considered statistically significant.
Abbreviations
ATF1: activating transcription factor 1; Cyp3a11: cytochrome P450: family 3: subfamily a: polypeptide 11; CREB: cAMP responsive element binding protein; DBP: albumin D-box binding protein; Gαi: inhibitory G protein alpha subunit; Gi: heterotrimeric Gαiβγ protein; PAR bZip: PAR-domain basic leucine zipper; PKA: protein kinase A; qRT-PCR: quantitative real-time PCR; ZT: Zeitgeber time.
ACKNOWLEDGMENTS AND FUNDING
We thank Jutta Schröder for introduction into the ImageJ software and all current and former members of the institute for discussion and support. This work was supported by the research commission of the medical faculty of the Heinrich-Heine-University (grant 9772517), the Deutsche Forschungsgemeinschaft (SFB 612, project A8), and the Deutsche Jose Carreras Leukämie-Stiftung e.V. (grant number DJCLS R15/10 to H.H.).
CONFLICTS OF INTEREST
The authors state no conflicts of interest.
REFERENCES
1. Dibner C, Schibler U, Albrecht U. The mammalian circadian timing system: organization and coordination of central and peripheral clocks. Annu Rev Physiol. 2010; 72:517–49.
2. Dibner C, Schibler U. Circadian timing of metabolism in animal models and humans. J Intern Med. 2015; 277:513–27.
3. Levi F, Schibler U. Circadian rhythms: mechanisms and therapeutic implications. Annu Rev Pharmacol Toxicol. 2007; 47:593–628.
4. Reppert SM, Weaver DR. Molecular analysis of mammalian circadian rhythms. Annu Rev Physiol. 2001; 63:647–76.
5. Reinke H, Asher G. Circadian Clock Control of Liver Metabolic Functions. Gastroenterology. 2016; 150:574 –80. https://doi.org/10.1053/j.gastro.2015.11.043.
6. Richards J, Gumz ML. Advances in understanding the peripheral circadian clocks. FASEB J. 2012; 26:3602–13.
7. Stokkan KA, Yamazaki S, Tei H, Sakaki Y, Menaker M. Entrainment of the circadian clock in the liver by feeding. Science. 2001; 291:490–3.
8. Crane BR, Young MW. Interactive features of proteins composing eukaryotic circadian clocks. Annu Rev Biochem. 2014; 83:191–219.
9. Robinson I, Reddy AB. Molecular mechanisms of the circadian clockwork in mammals. FEBS Lett. 2014; 588:2477–83.
10. Schibler U, Sassone-Corsi P. A web of circadian pacemakers. Cell. 2002; 111:919–22.
11. Stratmann M, Suter DM, Molina N, Naef F, Schibler U. Circadian Dbp transcription relies on highly dynamic BMAL1-CLOCK interaction with E boxes and requires the proteasome. Mol Cell. 2012; 48:277–87.
12. Gachon F, Olela FF, Schaad O, Descombes P, Schibler U. The circadian PAR-domain basic leucine zipper transcription factors DBP, TEF, and HLF modulate basal and inducible xenobiotic detoxification. Cell Metab. 2006; 4:25–36.
13. Yamaguchi S, Mitsui S, Yan L, Yagita K, Miyake S, Okamura H. Role of DBP in the circadian oscillatory mechanism. Mol Cell Biol. 2000; 20:4773–81.
14. Yamajuku D, Shibata Y, Kitazawa M, Katakura T, Urata H, Kojima T, Takayasu S, Nakata O, Hashimoto S. Cellular DBP and E4BP4 proteins are critical for determining the period length of the circadian oscillator. FEBS Lett. 2011; 585:2217–22.
15. Zhang YK, Yeager RL, Klaassen CD. Circadian expression profiles of drug-processing genes and transcription factors in mouse liver. Drug Metab Dispos. 2009; 37:106–15.
16. Yagita K, Okamura H. Forskolin induces circadian gene expression of rPer1, rPer2 and dbp in mammalian rat-1 fibroblasts. FEBS Lett. 2000; 465:79–82.
17. Motzkus D, Loumi S, Cadenas C, Vinson C, Forssmann WG, Maronde E. Activation of human period-1 by PKA or CLOCK/BMAL1 is conferred by separate signal transduction pathways. Chronobiol Int. 2007; 24:783–92.
18. Gau D, Lemberger T, von Gall C, Kretz O, Le Minh N, Gass P, Schmid W, Schibler U, Korf HW, Schutz G. Phosphorylation of CREB Ser142 regulates light-induced phase shifts of the circadian clock. Neuron. 2002; 34:245–53.
19. Tischkau SA, Mitchell JW, Tyan SH, Buchanan GF, Gillette MU. Ca2+/cAMP response element-binding protein (CREB)-dependent activation of Per1 is required for light-induced signaling in the suprachiasmatic nucleus circadian clock. J Biol Chem. 2003; 278:718–23.
20. Ding JM, Faiman LE, Hurst WJ, Kuriashkina LR, Gillette MU. Resetting the biological clock: mediation of nocturnal CREB phosphorylation via light, glutamate, and nitric oxide. J Neurosci. 1997; 17:667–75.
21. Ginty DD, Kornhauser JM, Thompson MA, Bading H, Mayo KE, Takahashi JS, Greenberg ME. Regulation of CREB phosphorylation in the suprachiasmatic nucleus by light and a circadian clock. Science. 1993; 260:238–41.
22. Mayr B, Montminy M. Transcriptional regulation by the phosphorylation-dependent factor CREB. Nat Rev Mol Cell Biol. 2001; 2:599–609.
23. Doi M, Ishida A, Miyake A, Sato M, Komatsu R, Yamazaki F, Kimura I, Tsuchiya S, Kori H, Seo K, Yamaguchi Y, Matsuo M, Fustin JM, et al. Circadian regulation of intracellular G-protein signalling mediates intercellular synchrony and rhythmicity in the suprachiasmatic nucleus. Nat Commun. 2011; 2:327.
24. Zhang EE, Liu Y, Dentin R, Pongsawakul PY, Liu AC, Hirota T, Nusinow DA, Sun X, Landais S, Kodama Y, Brenner DA, Montminy M, Kay SA. Cryptochrome mediates circadian regulation of cAMP signaling and hepatic gluconeogenesis. Nat Med. 2010; 16:1152–6.
25. Rodriguez CI, Setaluri V. Cyclic AMP (cAMP) signaling in melanocytes and melanoma. Arch Biochem Biophys. 2014; 563:22–7.
26. Nürnberg B, Gudermann T, Schultz G. Receptors and G proteins as primary components of transmembrane signal transduction. Part 2. G proteins: structure and function. J Mol Med (Berl). 1995; 73:123–32.
27. Wettschureck N, Offermanns S. Mammalian G proteins and their cell type specific functions. Physiol Rev. 2005; 85:1159–204.
28. Albarran-Juarez J, Gilsbach R, Piekorz RP, Pexa K, Beetz N, Schneider J, Nürnberg B, Birnbaumer L, Hein L. Modulation of alpha2-adrenoceptor functions by heterotrimeric Galphai protein isoforms. J Pharmacol Exp Ther. 2009; 331:35–44.
29. Gohla A, Klement K, Piekorz RP, Pexa K, vom Dahl S, Spicher K, Dreval V, Haussinger D, Birnbaumer L, Nürnberg B. An obligatory requirement for the heterotrimeric G protein Gi3 in the antiautophagic action of insulin in the liver. Proc Natl Acad Sci U S A. 2007; 104:3003–8.
30. Nürnberg B. Classification and functional properties of Ga subunits, in: The Nature Reviews Drug Discovery GPCR Questionnaire Participants. The state of GPCR research in 2004. Nat Rev Drug Discov. 2004; 575:577–626.
31. Servillo G, Della Fazia MA, Sassone-Corsi P. Coupling cAMP signaling to transcription in the liver: pivotal role of CREB and CREM. Exp Cell Res. 2002; 275:143–54.
32. Li Y, Ross-Viola JS, Shay NF, Moore DD, Ricketts ML. Human CYP3A4 and murine Cyp3A11 are regulated by equol and genistein via the pregnane X receptor in a species-specific manner. J Nutr. 2009; 139:898–904.
33. Devanathan V, Hagedorn I, Köhler D, Pexa K, Cherpokova D, Kraft P, Singh M, Rosenberger P, Stoll G, Birnbaumer L, Piekorz RP, Beer-Hammer S, Ni eswandt B, Nürnberg B. Platelet Gi protein Gαi2 is an essential mediator of thrombo-inflammatory organ damage in mice. Proc Natl Acad Sci U S A. 2015; 112:6491–96.
34. Dizayee S, Kaestner S, Kuck F, Hein P, Klein C, Piekorz RP, Meszaros J, Matthes J, Birnbaumer L, Nürnberg B, Herzig S. Galphai2- and Galphai3-specific regulation of voltage-dependent L-type calcium channels in cardiomyocytes. PLoS One. 2011; 6:e24979.
35. Jiang M, Spicher K, Boulay G, Martin-Requero A, Dye CA, Rudolph U, Birnbaumer L. Mouse gene knockout and knockin strategies in application to alpha subunits of Gi/Go family of G proteins. Methods Enzymol. 2002; 344:277–98.
36. Köhler D, Devanathan V, Bernardo de Oliveira Franz C, Eldh T, Novakovic A, Roth JM, Granja T, Birnbaumer L, Rosenberger P, Beer-Hammer S, Nürnberg B. Galphai2- and Galphai3-deficient mice display opposite severity of myocardial ischemia reperfusion injury. PLoS One. 2014; 9:e98325.
37. Leiss V, Flockerzie K, Novakovic A, Rath M, Schonsiegel A, Birnbaumer L, Schurmann A, Harteneck C, Nürnberg B. Insulin secretion stimulated by l-arginine and its metabolite l-ornithine depends on Galphai2. Am J Physiol Endocrinol Metab. 2014; 307:E800-12.
38. Plummer NW, Spicher K, Malphurs J, Akiyama H, Abramowitz J, Nürnberg B, Birnbaumer L. Development of the mammalian axial skeleton requires signaling through the Galpha(i) subfamily of heterotrimeric G proteins. Proc Natl Acad Sci U S A. 2012; 109:21366–71.
39. Rudolph U, Finegold MJ, Rich SS, Harriman GR, Srinivasan Y, Brabet P, Boulay G, Bradley A, Birnbaumer L. Ulcerative colitis and adenocarcinoma of the colon in G alpha i2-deficient mice. Nat Genet. 1995; 10:143–50.
40. Wiege K, Ali SR, Gewecke B, Novakovic A, Konrad FM, Pexa K, Beer-Hammer S, Reutershan J, Piekorz RP, Schmidt RE, Nürnberg B, Gessner JE. Galphai2 is the essential Galphai protein in immune complex-induced lung disease. J Immunol. 2013; 190:324–33.
41. Wiege K, Le DD, Syed SN, Ali SR, Novakovic A, Beer-Hammer S, Piekorz RP, Schmidt RE, Nürnberg B, Gessner JE. Defective macrophage migration in Galphai2- but not Galphai3-deficient mice. J Immunol. 2012; 189:980–7.
42. Nürnberg B. Pertussis toxin as a Cell Biology Tool. In: Aktories K, editor. Bacterial toxins. London: Chapman & Hall. 1997: 33–45.
43. Aton SJ, Huettner JE, Straume M, Herzog ED. GABA and Gi/o differentially control circadian rhythms and synchrony in clock neurons. Proc Natl Acad Sci U S A. 2006; 103:19188–93.
44. O'Neill JS, Maywood ES, Chesham JE, Takahashi JS, Hastings MH. cAMP-dependent signaling as a core component of the mammalian circadian pacemaker. Science. 2008; 320:949–53.
45. Lai L, Yuan L, Chen Q, Dong C, Mao L, Rowan B, Frasch T, Hill SM. The Galphai and Galphaq proteins mediate the effects of melatonin on steroid/thyroid hormone receptor transcriptional activity and breast cancer cell proliferation. J Pineal Res. 2008; 45:476–88.
46. Mühlbauer E, Gross E, Labucay K, Wolgast S, Peschke E. Loss of melatonin signalling and its impact on circadian rhythms in mouse organs regulating blood glucose. Eur J Pharmacol. 2009; 606:61–71.
47. Venegas C, Garcia JA, Doerrier C, Volt H, Escames G, Lopez LC, Reiter RJ, Acuna-Castroviejo D. Analysis of the daily changes of melatonin receptors in the rat liver. J Pineal Res. 2013; 54:313–21.
48. Hardeland R, Madrid JA, Tan DX, Reiter RJ. Melatonin, the circadian multioscillator system and health: the need for detailed analyses of peripheral melatonin signaling. J Pineal Res. 2012; 52:139–66.
49. Dinet V, Ansari N, Torres-Farfan C, Korf HW. Clock gene expression in the retina of melatonin-proficient (C3H) and melatonin-deficient (C57BL) mice. J Pineal Res. 2007; 42:83–91.
50. Lopez-Molina L, Conquet F, Dubois-Dauphin M, Schibler U. The DBP gene is expressed according to a circadian rhythm in the suprachiasmatic nucleus and influences circadian behavior. EMBO J. 1997; 16:6762–71.
51. Bonaconsa M, Malpeli G, Montaruli A, Carandente F, Grassi-Zucconi G, Bentivoglio M. Differential modulation of clock gene expression in the suprachiasmatic nucleus, liver and heart of aged mice. Exp Gerontol. 2014; 55:70–9.
52. Kornmann B, Schaad O, Reinke H, Saini C, Schibler U. Regulation of circadian gene expression in liver by systemic signals and hepatocyte oscillators. Cold Spring Harb Symp Quant Biol. 2007; 72:319–30.
53. Mueller CR, Maire P, Schibler U. DBP, a liver-enriched transcriptional activator, is expressed late in ontogeny and its tissue specificity is determined posttranscriptionally. Cell. 1990; 61:279–91.
54. Ripperger JA, Shearman LP, Reppert SM, Schibler U. CLOCK, an essential pacemaker component, controls expression of the circadian transcription factor DBP. Genes Dev. 2000; 14:679–89.
55. Balsalobre A, Marcacci L, Schibler U. Multiple signaling pathways elicit circadian gene expression in cultured Rat-1 fibroblasts. Curr Biol. 2000; 10:1291–4.
56. Muchardt C, Li C, Kornuc M, Gaynor R. CREB regulation of cellular cyclic AMP-responsive and adenovirus early promoters. J Virol. 1990; 64:4296–305.
57. Stratmann M, Stadler F, Tamanini F, van der Horst GT, Ripperger JA. Flexible phase adjustment of circadian albumin D site-binding protein (DBP) gene expression by CRYPTOCHROME1. Genes Dev. 2010; 24:1317–28.
58. Bailey M, Silver R. Sex differences in circadian timing systems: implications for disease. Front Neuroendocrinol. 2014; 35:111–39.
59. Bur IM, Cohen-Solal AM, Carmignac D, Abecassis PY, Chauvet N, Martin AO, van der Horst GT, Robinson IC, Maurel P, Mollard P, Bonnefont X. The circadian clock components CRY1 and CRY2 are necessary to sustain sex dimorphism in mouse liver metabolism. J Biol Chem. 2009; 284:9066–73.
60. Hirao J, Nishimura M, Arakawa S, Niino N, Mori K, Furukawa T, Sanbuissho A, Manabe S, Nishihara M, Mori Y. Sex and circadian modulatory effects on rat liver as assessed by transcriptome analyses. J Toxicol Sci. 2011; 36:9–22.
61. Schwartz JB. The influence of sex on pharmacokinetics. Clin Pharmacokinet. 2003; 42:107–21.
62. Gachon F, Firsov D. The role of circadian timing system on drug metabolism and detoxification. Expert Opin Drug Metab Toxicol. 2011; 7:147–58.
63. Zmrzljak UP, Rozman D. Circadian Regulation of the Hepatic Endobiotic and Xenobitoic Detoxification Pathways: The Time Matters. Chem Res Toxicol. 2012; 25:811–24.
64. Lavery DJ, Lopez-Molina L, Margueron R, Fleury-Olela F, Conquet F, Schibler U, Bonfils C. Circadian Expression of the Steroid 15 alpha-Hydroxylase (Cyp2a4) and Coumarin 7-Hydroxylase (Cyp2a5) Genes in Mouse Liver Is Regulated by the PAR Leucine Zipper Transcription Factor DBP. Mol Cell Biol. 1999; 19:6488–99.
65. Takiguchi T, Tomita M, Matsunaga N, Nakagawa H, Koyanagi S, Ohdo S. Molecular basis for rhythmic expression of CYP3A4 in serum-shocked HepG2 cells. Pharmacogenet Genomics. 2007; 17:1047–56.
66. Luo G, Guenthner T, Gan LS, Humphreys WG. CYP3A4 induction by xenobiotics: biochemistry, experimental methods and impact on drug discovery and development. Curr Drug Metab. 2004; 5:483–505.
67. Schneider R, Linka RM, Reinke H. HSP90 affects the stability of BMAL1 and circadian gene expression. J Biol Rhythms. 2014; 29:87–96.
68. Le Lay J, Tuteja G, White P, Dhir R, Ahima R, Kaestner KH. CRTC2 (TORC2) Contributes to the Transcriptional Response to Fasting in the Liver but Is Not Required for the Maintenance of Glucose Homeostasis. Cell Metab. 2009; 10:55–62.
69. Nagata T, Redman RS, Lakshman R. Isolation of intact nuclei of high purity from mouse liver. Anal Biochem. 2010; 398:178–84.

