INTRODUCTION
There has been an epidemic rise in oropharyngeal cancer worldwide due to increasing rates of oral infection with human papillomavirus (HPV) [1, 2]. HPV-positive cancers typically arise in younger and healthier patients that consume less alcohol and tobacco than patients with HPV-negative head and neck squamous cell cancers (HNSCC). Although HPV-positive patients are much more likely to be cured of their disease [1, 3], an important challenge for these patients has been enduring the acute and chronic toxicities of therapy. In addition, HPV-positive tumors are distinct from a molecular perspective, with characteristic genetic, epigenetic and protein profiles [4]. Given these profound differences between HPV-positive and HPV-negative disease at the clinical and molecular level, it is logical that the development of any therapy is tailored specifically for the tumor HPV-status.
Cell lines are imperfect models of cancer [5, 6], however HNSCC cell lines appear to have a similar genomic landscape to primary tumors [7, 8]. More importantly, genomic markers of drug sensitivity in cell lines appear to generally correlate well with validated biomarkers in patient tumors [7, 9]. Thus, as cell lines can be easily and cheaply screened with large collections of drugs, they are invaluable tools to identify new agents with potent activity against tumor cells.
This study aimed to carry out a high-throughput drug screen of a large collection of compounds against HNSCC cell lines, with the objective of preliminary identification of potent and HPV-status selective agents suitable for further investigation.
RESULTS
High-throughput screening identified multiple drugs with high potency against HNSCC cell lines
The top 20 compounds with the highest activity from the kinase inhibitor and Tocris panels are reported in Figure 1A and 1B, respectively (List of compounds and mechanism of action can be found in Supplementary Table 1). This screen identified drugs with high potency that included a diverse range of chemotherapeutic and targeted agents, however several themes emerged (Supplementary Figure 1). Four inhibitors of the phosphoinositide 3 kinase (PI3K) pathway were identified (GDC0941, PIK-75, NVP-BEZ235, FAK Inhibitor 14), as were 3 drugs reported to affect various aspect of the cell cycle (MK 1775, TCS2312, NSC 146109). Three topoisomerase inhibitors were potent (Daunorubicin, Doxorubicin and SN 38). Multiple compounds affecting inflammatory pathways through inhibition of proteins such as NF-κB, IKKβ and IκBα were also identified (IMD0354, Pyrrolidinedithiocarbamate ammonium, MG 132 and Bay 11-7085). Cisplatin and carboplatin, chemotherapeutics in routine use head and neck cancer, did not meet our criteria for significant activity (B-score < -2) in the cell line panel at a dose of 4μM (Supplementary Figure 2). Complete tables of individual compound activity against each cell line from the kinase inhibitor and Tocris panels are provided in Supplementary Table 2.
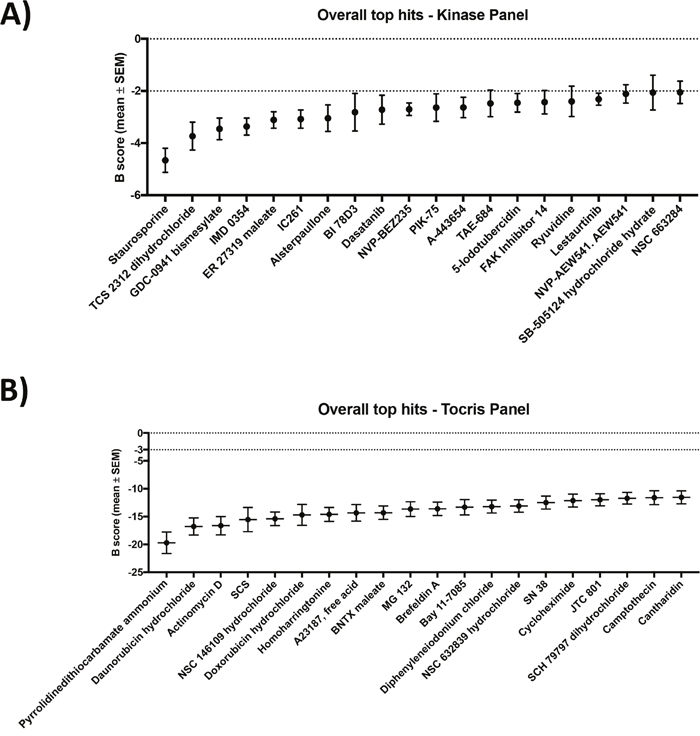
Figure 1: The top 20 most potent drugs and determined by the B-score in HNSCC cell lines from the Kinase inhibitor (A) and Tocris (B) panels in the high-throughput drug screen.
A subset of compounds demonstrated preferential activity against cell lines based on HPV-status
Thirty compounds from the kinase panel, and 27 compounds from the Tocris panel were found to be more potent against HPV-positive cell lines (Figure 2A shows top hits, full list in Supplementary Table 3). Among the identified hits, multiple compounds impacted cell cycle pathways (Supplementary Figure 1), including inhibition of cyclin-dependent kinases (CDK: SNS-032, Alvocidib), PI3K (PI-103, PIK90, Deguelin), polo-like kinase 1 (PLK1: GSK-461364, BI 2536), and checkpoint kinases (CHK: AZD-7762, SB 218078). Various Janus kinase (JAK) inhibitors were also seen in the HPV-positive preferential list (AZ-960, Merck-5, AT9283, Lestaurtinib). The compounds DIPPA hydrochloride, MVC 05-290, Way 629, Physostigmine and PNC 120596 that are known to act on neuro-receptors were demonstrated to be more potent in the HPV-positive cell lines.
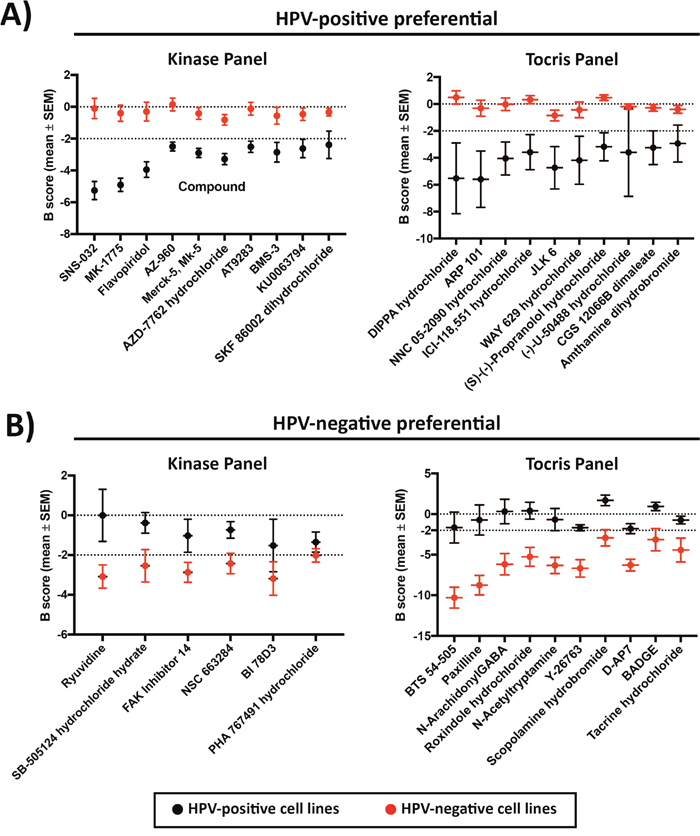
Figure 2: Top 10 drugs with (A) HPV-positive and (B) HPV-negative selective potency from Kinase inhibitor and Tocris compound panels.
Six compounds from the kinase inhibitor panel and 28 compounds from the Tocris panel were found to have higher activity in HPV-negative cell lines (Figure 2B shows top hits, full list in Supplementary Table 4). Three compounds impacting cellular proliferation and differentiation were identified (Supplementary Figure 1), including SB-505124, focal adhesion kinase (FAK) Inhibitor 14 and Aminopurvalanol A. From the Tocris panel, compounds impacting various aspects of cellular regulation and physiology were common hits against HPV-negative cell lines (Supplementary Figure 1), including Y-26763, D-AP7, BADGE, SKF 96365 hydrochloride, Olvanil, Embelin, GR 127935 hydrochloride, Demethylasterriquinone B1, Tyrphostin B44, Ro 90-7501, PALDA and OLDA.
Dose response curves confirm Ryuvidine and MK-1775 to be preferentially potent against HPV-negative and positive cell lines respectively
After the generation of multi-point dose response curves, Ryuvidine was selectively potent against HPV-negative cell lines, with an average IC50 of 1.562 μM ± 0.755 (mean ± standard deviation) in HPV-negative and 3.169 μM ± 1.305 in HPV-positive cell lines (Figure 3A, unpaired t-test p=0.003), confirming the findings in the high-throughput screen. MK-1775 showed significantly higher potency against HPV-positive cell lines with an average IC50 of 0.535 μM ± 0.4828 in HPV-positive and 3.647 μM ± 3.73 in HPV-negative cell lines (Figure 3B, unpaired t-test p=0.03), again showing compatible findings with the high-throughput drug screen. SNS-032, Flavopiridol, AZD-7762 and ARP-101 compounds did not show significantly different potency in cell lines stratified by HPV-status following validation studies across a range of concentrations (Supplementary Figure 3, unpaired t-test p>0.05). Dose-response studies of MK-1775, Ryuvidine, AZD7762 and Flavopiridol in control non-cancerous IMR90 cell line showed higher IC50 values (IC50 of >10 μM, 2.2 μM, 0.61 μM and >10 μM, respectively) compared to susceptible HNSCC cell lines, demonstrating the presence of a therapeutic window.
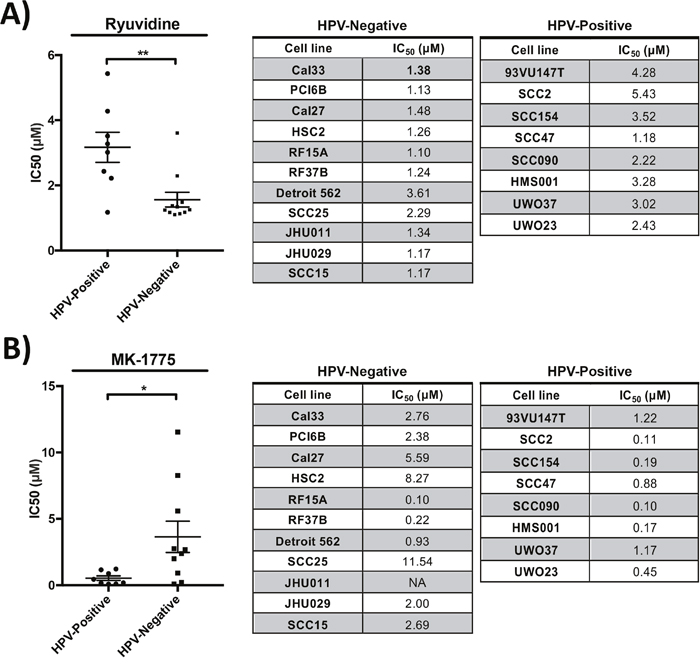
Figure 3: Drug potency validation studies revealed (A) Ryuvidine to be selective against HPV-negative cell lines (** p=0.003) and (B) MK-1775 to be selective against HPV-positive cell lines (* p=0.03).
DISCUSSION
To our knowledge, this is the first high-throughput drug screening study that focuses specifically on head and neck cancer cell lines. Our study of 1,433 compounds in 26 cell lines identified 57 and 34 compounds with preferential activity against HPV-negative and HPV-positive cell lines, respectively. Validation studies with six of these confirmed selective potencies in HPV-negative (Ryuvidine) and HPV-positive (MK-1775) cell lines. Given that HPV-positive and negative tumors are distinct from a molecular and prognostic perspective [10], this first step in the development of therapeutics specific to each disease type is critical to achieve the end goal of maximizing survival rates while minimizing toxicity.
Multiple studies and clinical trials have identified a superior response to chemotherapy, and better survival outcomes in HPV-positive HNSCCs compared to the HPV-negative tumors [3, 11–14]. Following these observations, treatment plans and clinical trials for HNSCC are increasingly being tailored based on tumor HPV status, particularly with the aim to de-intensify treatment for patients with HPV-positive tumors [15]. It is notable that despite the overall favourable outcomes in HPV-positive tumors, treatment failure still occurs in 20-40% of advanced disease [16]. As a result, there remains a necessity for further discovery and development of selective therapeutics in both HPV-positive and negative cohorts of HNSCCs.
Ryuvidine functions as an inhibitor of the CDK4 and SETD8 proteins [17, 18]. CDK4, and its close relative CDK6, interact with cyclin D and function in a pathway that is important for cell cycle progression. The CDK4/cyclin D complex impacts cell cycle progression by phosphorylating and inactivating retinoblastoma (Rb), a negative regulator of G1 exit [19]. Interestingly, Cyclin D1 is amplified frequently in HPV-negative HNSCC and p16, the negative regulator of the kinase complex, is frequently deleted [20]. Thus, Ryuvidine may reverse the effects of these mutations, leading to cell cycle arrest in HPV-negative HNSCC. In contrast, the Rb pathway is abrogated in HPV-positive cells as the HPV E7 oncoprotein binds Rb and targets it for degradation [21]. Thus, the drug would not be expected to have an effect in HPV-positive cells, which is consistent with our finding (Figure 3A). Further investigation will be required to clarify the mechanism behind the observed preferential potency of Ryuvidine against HPV-negative cell lines.
MK-1775, also known as AZD1775, is a WEE1 kinase inhibitor [22]. WEE1 kinase has been linked to G2-M cell cycle arrest in response to DNA damage. Inhibition of WEE1 kinase can therefore overcome G2 cell cycle arrest, which in the presence of DNA damage, can lead to mitotic catastrophe and ultimately cell death [22, 23]. As a result of this mechanism of action, WEE1 inhibitors have entered clinical trials as chemo-sensitizers of various tumors, such as breast cancer, leukemia and melanoma [24]. Previous studies have shown that MK-1775 can sensitize both HPV-positive and HPV-negative cell lines to cisplatin therapy, but through differing mechanisms; senescence [25] versus apoptosis [26] in HPV-negative and HPV-positive cell lines, respectively. Our validation study revealed MK-1775 to be independently potent in reducing cellular proliferation, with preferential activity in HPV-positive cell lines (Figure 3B). WEE1 kinase inhibition has been shown to selectively sensitize p53-deficient cells to exogenous DNA damage [25], and the higher susceptibility of HPV-positive cell lines to MK1775 may be due to the inhibition of p53 by the HPV E6 protein [27]. Further studies may be indicated to explore the potential of MK1775 compound as an HPV-specific chemotherapeutic agent.
Preliminary dose-response curve validation studies were able to validate the selectivity of 2 of the 6 tested compounds that were derived based on our high-throughput study. This is not surprising as the high-throughput study included only a single dose of each compound, with the purpose of preliminary identification of compounds with potential for further assessment. More of the identified compounds may show HPV-status specific activity against HNSCCs following validation. Furthermore, analyses of the available HNSCC cell lines have revealed common genetic alterations other than HPV-status that are also reflected in tumor populations [28]. These include amplifications in PIK3CA, EGFR and CCND2, deletions in CDKN2A, SMAD4 and NOTCH2, and mutations in TP53, CDKN2A, SMAD4 and PIK3CA [28]. Thus, the genetic characterization of our cell line panel paired with our high-throughput drug screen can potentially be used to identify genomic correlates of drug response as a further step towards providing personalize medicine for head and neck cancer patients.
High-throughput testing of large chemical compound libraries can serve as valuable resources in the context of a necessity for better chemotherapeutics in the field of head and neck oncology. In this study, we have provided the head and neck cancer research community with the results of a preliminary screen of the activity of over 1400 compounds against 26 HNSCC cell lines. Initial validation studies led to the identification of Ryuvidine and MK-1775 compounds to have preferential activity against HPV-negative and HPV-positive cell lines respectively. We hope that the provided datasets can serve as a valuable resource to lay the foundation for further investigation of compounds with therapeutic potential in the field of head and neck oncology.
MATERIALS AND METHODS
Cell lines and culture conditions
Twenty-eight HNSCC cell lines were obtained from established culture collections or from collaborators as listed in Supplementary Table 5. Short tandem repeat profiling was carried out as previously described [29] and compared to the literature to confirm the cell line identity (Supplementary Table 6). Cells were cultured in an incubator at 37°C with 5% CO2 in the medium specified in Supplementary Table 5.
High-throughput drug screening
A custom-made compound library from Tocris Bioscience comprised of 1,113 compounds and a kinase inhibitor library of 320 compounds were used for high throughput drug screening. The drug screen was carried out against 6 HPV-positive and 20 HPV-negative cell lines (Supplementary Table 5). High-throughput drug studies were carried out in the S.M.A.R.T Facility in the Samuel Lunenfeld Research Institute at Mount Sinai Hospital in Toronto, Ontario, Canada. Cells were robotically seeded at a density of 600 cells/well in 384-well plates and incubated for 24 hours prior to the addition of drugs using an automated drug pinning device (Beckman Multimek, Mississauga, Ontario). Cells were treated with either DMSO (control) or a single dose of each drug at a concentration of 4 μM, and cells were subsequently incubated for 48 hours based on methodology used in previous high-throughput studies [30, 31]. Cell viability was measured indirectly using the AlamarBlue reagent (ThermoFisher Scientific).
Statistical analysis
Fluorescence intensity derived from the high-throughput screen were normalized using B scores as per recommendation of Malo et. al. [32]. For cell lines with two replicates, the average B score per drug was calculated. In each of the drug panels, the most potent drugs were identified as those with an average B score below -2. Drugs were then sorted based on the highest B score to the lowest within that stratification scheme. To identify drugs with preferential activity based on HPV-status, Student’s t-tests were utilized to compare the average B score of HPV-positive cell lines to that of HPV-negative cell lines for each drug. These p-values were then controlled for local false discovery rates using the fdrtool package version 1.2.15 [33] in the R statistical environment (version 3.4.0). The statistical comparison was considered significant if the corrected p-value after correction by local false discovery rate (LFDR) was lower than 0.05. Drugs with significant preferential activity were then isolated by having a B score of lower than -2 in one disease type (ie HPV+ or HPV-) and a B score of higher than -2 in the opposite disease type, then sorted by the mean difference in the two arms of comparison (from high to low).
Generation of dose response curves
Six drugs (Ryuvidine, MK-1775, SNS-032, Flavopiridol, AZD-7762 and ARP-101) that demonstrated high differential activity in either HPV-positive or negative cell lines and mechanisms of action that were deemed to be interesting in HNSCC were selected for further analysis. Dose response curves were generated, and drug validation studies for these compounds were tested in 8 HPV-positive and 11 HPV negative cell lines (cell lines listed in Supplementary Table 5). Cells were seeded in 96-well plates at 3000 cells/well. After 24 hours compounds were added in a concentration series of 0.01, 0.03, 0.1, 0.3, 1, 3, 10, 30 and 100 μM. Three technical replicates were carried out in each cell line, per drug. Cells were incubated with the added drugs for 72 hours before assessing potency. Cell viability was measured indirectly using the PrestoBlue® Reagent (Thermofisher Scientific). To calculate the half-maximal inhibitory concentration (IC50) value for each cell line, normalized Relative Fluorescence Units (RFUs) of the drug-treated replicates were calculated as a percentage of the mean RFU of the vehicle control (DMSO-only) treatment replicates. IC50 values, defined as the concentration at which the normalized RFU reached 50%, were calculated by non-linear regression (Prism® 7 Graphpad Software, Inc). When a non-linear regression model could not be converged for a compound and cell line, no IC50 value was assigned and data was excluded from comparison analysis. An unpaired t-test was used to assess the significance of the differences in IC50 values between HPV-positive and HPV-negative cell lines for each of the compounds, and a p value of less than 0.05 was set as the significance limit. As a preliminary assessment of compound toxicity, we also tested 4 of the compounds (MK1775, Ryuvidine, AZD7762 and Flavopiridol) against the IMR90 non-cancerous cell line (ATCC CCL-186) over a dose range of 0.03 μM to 10 μM with the same methodology to generate dose-response curves and calculate IC50 values.
CONFLICTS OF INTEREST
The authors declare no potential conflicts of interest.
FUNDING
This study was funded by a Canadian Institutes for Health Research (CIHR) grant MOP 340674 to A.C.N. and P.C.B. PCB was supported by CIHR and Terry Fox Research New Investigator Awards and by the Ontario Institute for Cancer Research.
REFERENCES
1. Nichols AC, Palma DA, Dhaliwal SS, Tan S, Theuer J, Chow W, Rajakumar C, Um S, Mundi N, Berk S, Zhou R, Basmaji J, Rizzo G, et al. The epidemic of human papillomavirus and oropharyngeal cancer in a Canadian population. Curr Oncol. 2013; 20:212-9. https://doi.org/10.3747/co.20.1375.
2. Chaturvedi AK, Engels EA, Pfeiffer RM, Hernandez BY, Xiao W, Kim E, Jiang B, Goodman MT, Sibug-Saber M, Cozen W, Liu L, Lynch CF, Wentzensen N, et al. Human papillomavirus and rising oropharyngeal cancer incidence in the United States. J Clin Oncol. 2011; 29:4294-301. https://doi.org/10.1200/JCO.2011.36.4596.
3. Ang KK, Harris J, Wheeler R, Weber R, Rosenthal DI, Nguyen-Tan PF, Westra WH, Chung CH, Jordan RC, Lu C, Kim H, Axelrod R, Silverman CC, et al. Human papillomavirus and survival of patients with oropharyngeal cancer. N Engl J Med. 2010; 363:24-35. https://doi.org/10.1056/NEJMoa0912217.
4. Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature. 2015; 517:576-82. https://doi.org/10.1038/nature14129.
5. Gazdar AF, Girard L, Lockwood WW, Lam WL, Minna JD. Lung cancer cell lines as tools for biomedical discovery and research. J Natl Cancer Inst. 2010; 102:1310-21. https://doi.org/10.1093/jnci/djq279.
6. Hennessey PT, Ochs MF, Mydlarz WW, Hsueh W, Cope L, Yu W, Califano JA. Promoter methylation in head and neck squamous cell carcinoma cell lines is significantly different than methylation in primary tumors and xenografts. PLoS One. 2011; 6:e20584. https://doi.org/10.1371/journal.pone.0020584.
7. Barretina J, Caponigro G, Stransky N, Venkatesan K, Margolin AA, Kim S, Wilson CJ, Lehar J, Kryukov GV, Sonkin D, Reddy A, Liu M, Murray L, et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature. 2012; 483:603-7. https://doi.org/10.1038/nature11003.
8. Nichols AC, Black M, Yoo J, Pinto N, Fernandes A, Haibe-Kains B, Boutros PC, Barrett JW. Exploiting high-throughput cell line drug screening studies to identify candidate therapeutic agents in head and neck cancer. BMC Pharmacol Toxicol. 2014; 15:66. https://doi.org/10.1186/2050-6511-15-66.
9. Garnett MJ, Edelman EJ, Heidorn SJ, Greenman CD, Dastur A, Lau KW, Greninger P, Thompson IR, Luo X, Soares J, Liu Q, Iorio F, Surdez D, et al. Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature. 2012; 483:570-5. https://doi.org/10.1038/nature11005.
10. Gillison ML. Human papillomavirus-associated head and neck cancer is a distinct epidemiologic, clinical, and molecular entity. Semin Oncol. 2004; 31:744-54.
11. Bossi P, Orlandi E, Miceli R, Perrone F, Guzzo M, Mariani L, Granata R, Locati L, Fallai C, Cortelazzi B, Pilotti S, Scaramellini G, Gloghini A, et al. Treatment-related outcome of oropharyngeal cancer patients differentiated by HPV dictated risk profile: a tertiary cancer centre series analysis. Ann Oncol. 2014; 25:694-9. https://doi.org/10.1093/annonc/mdu004.
12. Kumar B, Cordell KG, Lee JS, Prince ME, Tran HH, Wolf GT, Urba SG, Worden FP, Chepeha DB, Teknos TN, Eisbruch A, Tsien CI, Taylor JM, et al. Response to therapy and outcomes in oropharyngeal cancer are associated with biomarkers including human papillomavirus, epidermal growth factor receptor, gender, and smoking. Int J Radiat Oncol Biol Phys. 2007; 69:S109-11. https://doi.org/10.1016/j.ijrobp.2007.05.072.
13. Posner MR, Lorch JH, Goloubeva O, Tan M, Schumaker LM, Sarlis NJ, Haddad RI, Cullen KJ. Survival and human papillomavirus in oropharynx cancer in TAX 324: a subset analysis from an international phase III trial. Ann Oncol. 2011; 22:1071-7. https://doi.org/10.1093/annonc/mdr006.
14. Fakhry C, Westra WH, Li S, Cmelak A, Ridge JA, Pinto H, Forastiere A, Gillison ML. Improved survival of patients with human papillomavirus-positive head and neck squamous cell carcinoma in a prospective clinical trial. J Natl Cancer Inst. 2008; 100:261-9. https://doi.org/10.1093/jnci/djn011.
15. Kelly JR, Husain ZA, Burtness B. Treatment de-intensification strategies for head and neck cancer. Eur J Cancer. 2016; 68:125-33. https://doi.org/10.1016/j.ejca.2016.09.006.
16. Swiecicki PL, Malloy KM, Worden FP. Advanced oropharyngeal squamous cell carcinoma: Pathogenesis, treatment, and novel therapeutic approaches. World J Clin Oncol. 2016; 7:15-26. https://doi.org/10.5306/wjco.v7.i1.15.
17. Ryu CK, Kang HY, Lee SK, Nam KA, Hong CY, Ko WG, Lee BH. 5-Arylamino-2-methyl-4,7-dioxobenzothiazoles as inhibitors of cyclin-dependent kinase 4 and cytotoxic agents. Bioorg Med Chem Lett. 2000; 10:461-4.
18. Blum G, Ibanez G, Rao X, Shum D, Radu C, Djaballah H, Rice JC, Luo M. Small-molecule inhibitors of SETD8 with cellular activity. ACS Chem Biol. 2014; 9:2471-8. https://doi.org/10.1021/cb500515r.
19. Giacinti C, Giordano A. RB and cell cycle progression. Oncogene. 2006; 25:5220. https://doi.org/10.1038/sj.onc.1209615.
20. Namazie A, Alavi S, Olopade OI, Pauletti G, Aghamohammadi N, Aghamohammadi M, Gornbein JA, Calcaterra TC, Slamon DJ, Wang MB, Srivatsan ES. Cyclin D1 amplification and p16(MTS1/CDK4I) deletion correlate with poor prognosis in head and neck tumors. Laryngoscope. 2002; 112:472-81. https://doi.org/10.1097/00005537-200203000-00013.
21. Munger K, Baldwin A, Edwards KM, Hayakawa H, Nguyen CL, Owens M, Grace M, Huh K. Mechanisms of human papillomavirus-induced oncogenesis. J Virol. 2004; 78:11451-60. https://doi.org/10.1128/JVI.78.21.11451-11460.2004.
22. Geenen JJJ, Schellens JHM. Molecular Pathways: Targeting the Protein Kinase Wee1 in Cancer. Clin Cancer Res. 2017; 23:4540-4. https://doi.org/10.1158/1078-0432.CCR-17-0520.
23. Wang Y, Decker SJ, Sebolt-Leopold J. Knockdown of Chk1, Wee1 and Myt1 by RNA interference abrogates G2 checkpoint and induces apoptosis. Cancer Biol Ther. 2004; 3:305-13.
24. Matheson CJ, Backos DS, Reigan P. Targeting WEE1 Kinase in Cancer. Trends Pharmacol Sci. 2016; 37:872-81. https://doi.org/10.1016/j.tips.2016.06.006.
25. Osman AA, Monroe MM, Ortega Alves MV, Patel AA, Katsonis P, Fitzgerald AL, Neskey DM, Frederick MJ, Woo SH, Caulin C, Hsu TK, McDonald TO, Kimmel M, et al. Wee-1 kinase inhibition overcomes cisplatin resistance associated with high-risk TP53 mutations in head and neck cancer through mitotic arrest followed by senescence. Mol Cancer Ther. 2015; 14:608-19. https://doi.org/10.1158/1535-7163.MCT-14-0735-T.
26. Tanaka N, Patel AA, Wang J, Frederick MJ, Kalu NN, Zhao M, Fitzgerald AL, Xie TX, Silver NL, Caulin C, Zhou G, Skinner HD, Johnson FM, et al. Wee-1 Kinase Inhibition Sensitizes High-Risk HPV+ HNSCC to Apoptosis Accompanied by Downregulation of MCl-1 and XIAP Antiapoptotic Proteins. Clin Cancer Res. 2015; 21:4831-44. https://doi.org/10.1158/1078-0432.CCR-15-0279.
27. Lechner MS, Laimins LA. Inhibition of p53 DNA binding by human papillomavirus E6 proteins. J Virol. 1994; 68:4262-73.
28. Li H, Wawrose JS, Gooding WE, Garraway LA, Lui VW, Peyser ND, Grandis JR. Genomic analysis of head and neck squamous cell carcinoma cell lines and human tumors: a rational approach to preclinical model selection. Mol Cancer Res. 2014; 12:571-82. https://doi.org/10.1158/1541-7786.MCR-13-0396.
29. Mundi N, Um S, Yoo J, Rizzo G, Black M, Pinto N, Palma DA, Fung K, MacNeil D, Mymryk JS, Barrett JW, Nichols AC. The control of anaplastic thyroid carcinoma cell lines by oncolytic poxviruses. Virus Res. 2014; 190:53-9. https://doi.org/10.1016/j.virusres.2014.07.009.
30. Prassas I, Paliouras M, Datti A, Diamandis EP. High-throughput screening identifies cardiac glycosides as potent inhibitors of human tissue kallikrein expression: implications for cancer therapies. Clin Cancer Res. 2008; 14:5778-84. https://doi.org/10.1158/1078-0432.CCR-08-0706.
31. Izrailit J, Berman HK, Datti A, Wrana JL, Reedijk M. High throughput kinase inhibitor screens reveal TRB3 and MAPK-ERK/TGFbeta pathways as fundamental Notch regulators in breast cancer. Proc Natl Acad Sci U S A. 2013; 110:1714-9. https://doi.org/10.1073/pnas.1214014110.
32. Malo N, Hanley JA, Cerquozzi S, Pelletier J, Nadon R. Statistical practice in high-throughput screening data analysis. Nat Biotechnol. 2006; 24:167-75. https://doi.org/10.1038/nbt1186.
33. Strimmer K. fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics. 2008; 24:1461-2. https://doi.org/10.1093/bioinformatics/btn209.

