INTRODUCTION
Historically, natural products have been considered invaluable as a source of therapeutic agents. Resveratrol is a polyphenolic phytoalexin present in many plants, including grapevines, pines, peanuts, berries, and grapes [1, 2]. Red wine is made from grapes (most commonly Vitis vinifera) that are rich in resveratrol; this has garnered much attention regarding the “French paradox,” which refers to the association between the consumption of red wine and the relatively low incidence of coronary heart disease in the French population, despite their high intake of dietary fat [3]. Numerous studies have examined the effects of resveratrol in detail, including work on sirtuin genes and longevity [4–6]. In addition, it has been suggested that resveratrol possesses potential anti-cancer properties, such as the inhibition of tumor initiation and progression, by inducing cell cycle arrest and apoptosis [7].
microRNAs (miRNAs), a family of small non-coding RNAs, have been identified in various organisms, such as worms, flies, mice, humans, and plants. Several miRNAs are conserved among different species, indicating that these miRNAs may serve important functions such as the modulation of gene expression. It has become realized that miRNAs contribute to many human diseases, including cancers, and are generally dysregulated in cancers compared with their expression in normal tissues. Accordingly, miRNA expression profiles can be used to classify poorly differentiated tumors, and some miRNAs are connected to well-studied tumor-suppressive or oncogenic networks [8, 9]. In addition, many studies have demonstrated that miRNAs are contained in exosomes/extracellular vesicles (EVs); these vesicles, which are secreted from all types of cells, act as mediators of cell-to-cell communication [10]. Furthermore, miRNAs are present in body fluids, such as blood, serum/plasma, saliva, and urine [11]. Researchers are currently investigating the potential utility of miRNA-expression patterns as novel specific and sensitive diagnostic markers, as well as that of miRNA profiles in body fluids as indicators of health conditions [12]. For instance, early-stage breast cancer could be detected and diagnosed by examining the expression profiles of serum miRNAs [13]. Thus, if we could maintain the expression levels and profiles of miRNAs in the body, this strategy would likely support the prevention and treatment of diseases. Recently, resveratrol was shown to regulate miRNAs in some cancer cell lines, which shed new light on cancer biology [14–20]. However, the molecular mechanisms through which resveratrol, a miRNA-regulating natural compound, can contribute to cancer treatment are not completely understood.
A previous study reported that resveratrol induces some tumor-suppressive miRNAs and Argonaute 2 (AGO2) expression, which can result in long-term gene silencing [9]. Here we examined the functional mechanisms underlying the regulatory role of resveratrol by analyzing both mRNA and miRNA profiles in breast cancer cells. Resveratrol up-regulated mainly three miRNAs (miR-34a, miR-424, and miR-503) that are under the control of p53. This was found to lead to the suppression of heterogeneous nuclear ribonucleoprotein A1 (HNRNPA1), which is up-regulated in a wide variety of cancers, and to the inhibition of cell proliferation. Our results provide novel information about the regulatory effects of resveratrol regarding the maintenance of good health.
RESULTS
Identification of resveratrol-regulated miRNAs in breast cancer
A previous study reported that AGO2 up-regulation by resveratrol enhanced the expression of some tumor-suppressive miRNAs and increased its RNA interference activity [15]. However, the understanding of the complete molecular mechanisms by which resveratrol contributes to cancer prevention and treatment is inadequate. For example, it is unclear which miRNAs are mainly regulated by resveratrol and how resveratrol-regulated miRNAs control cancer cell proliferation by modulating target gene expression. To obtain insight into the functional mechanisms of resveratrol, as resveratrol inhibited the proliferation of MDA-MB-231-luc-D3H2LN breast cancer cells (Supplementary Figure 1), we comprehensively analyzed miRNA and mRNA profiles of cancer cells treated with or without resveratrol. miRNA analysis revealed transcriptome changes between resveratrol-treated cells and untreated cells, and that resveratrol induced 24 miRNAs and suppressed 22 miRNAs (Figure 1A and Supplementary Figure 2A). Of these miRNAs, we selected the eight most highly up-regulated miRNAs and validated their expression levels by quantitative reverse transcription-PCR (qRT-PCR; Figure 1B). Additionally, to identify resveratrol-regulated miRNAs that are commonly expressed in different types of cancer cell lines, we examined the eight miRNAs in MCF7 cancer cells. We observed that only three miRNAs (miR-34a, miR-503, and miR-424) were significantly increased by >1.5-fold in both cell lines (Figure 1B); thus, we decided to focus on these miRNAs for further studies.
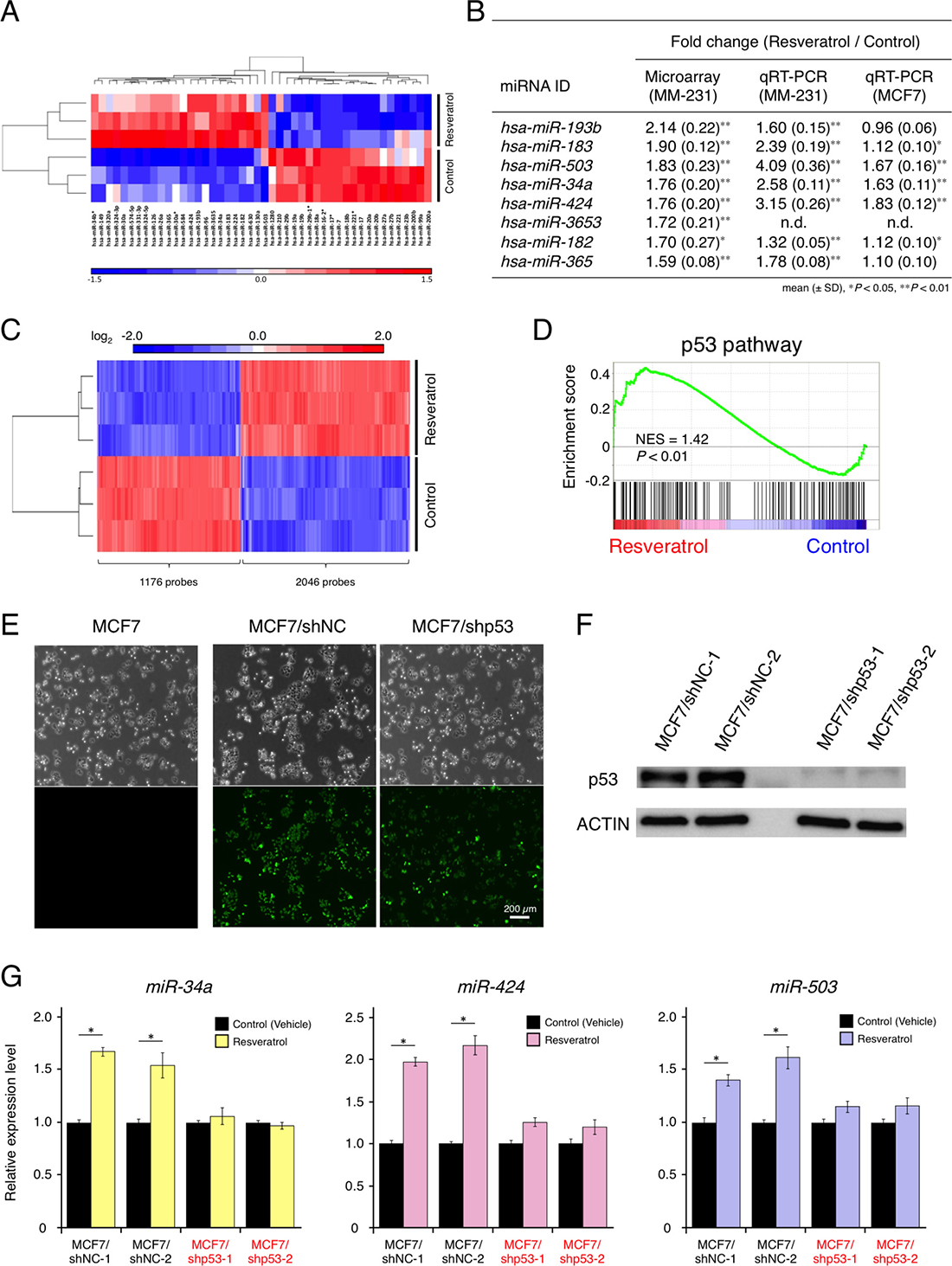
Figure 1: Effects of resveratrol on miRNA and mRNA profiles and resveratrol-regulated miRNAs in breast cancer. (A) Heat map showing miRNAs whose expression levels differed by >1.2-fold in MDA-MB-231-luc-D3H2LN cells, with or without resveratrol treatment (P < 0.1). Twenty-four miRNAs were up-regulated and 22 miRNAs were down-regulated by resveratrol treatment. (B) The eight most highly up-regulated miRNAs following resveratrol treatment, as determined by microarray analysis. miRNA expression was validated by qRT-PCR in MDA-MB-231-luc-D3H2LN and MCF7 cells (mean ± SD, *P < 0.05, **P < 0.01). n.d.: not determined (C) Heat map showing mRNAs that differed by >1.5-fold in MDA-MB-231-luc-D3H2LN cells, with or without resveratrol treatment (P < 0.01). A total of 2,046 mRNAs were up-regulated and 1,176 mRNAs were down-regulated by resveratrol treatment. (D) Gene set-enrichment analysis of the p53 pathway signature in MDA-MB-231-luc-D3H2LN cells treated with resveratrol, as compared with control cells (treated with vehicle). NES: normalized enrichment score (E) GFP signals in transgenic MCF7 cells expressing control non-target shRNA and TP53-knockdown shRNA. MCF7/shNC: MCF7/H1::non-target shRNA/CMV::copGFP and MCF7/shp53: MCF7/H1::TP53 shRNA/CMV::copGFP. (F) Immunoblot analysis of p53 expression in MCF7 cells expressing a control non-target shRNA and p53-knockdown shRNA. (G) The expression of resveratrol-regulated miRNAs in p53-knockdown MCF7 cells and control cells, with or without resveratrol treatment (mean ± SEM, *P < 0.05). Two independent clones were prepared for each cell line. The number of data points shown for each sample is four.
mRNA analysis suggested that resveratrol regulates a wide variety of genes in treated cells versus untreated cells and affects many biological pathways, such as the p53 pathway and apoptosis. We observed changes in p53-regulated gene-expression levels, which indicated that p53 activation by resveratrol contributed to apoptosis (Figure 1C, 1D, Supplementary Figure 2B–2E, and Supplementary Table 1). Intriguingly, miR-34a can be directly transactivated by p53, and miR-424 and miR-503 (which are transcribed together as the miR-424/503 cluster) are presumably under the control of p53 [21–24]. To investigate whether these miRNAs are regulated in the resveratrol/p53 pathway, we established two independent p53-knockdown MCF7 cell lines (Figure 1E and 1F), because MCF7 cells have wild-type p53 [25]. However, MDA-MB-231 cells contain mild mutations in p53 [26]. Resveratrol induced miR-34a, miR-503, and miR-424 expression in p53-expressing cells but not in p53-knockdown cells, suggesting that p53 is important for the regulatory function of resveratrol in miRNA induction (Figure 1G). These results showed that, in parallel with inhibiting breast cancer cell proliferation, resveratrol could mainly regulate miR-34a, miR-503, and miR-424 via the p53 pathway.
Identification of the target gene of resveratrol-regulated miRNAs and its clinical significance
To elucidate the mechanisms through which resveratrol-regulated miRNAs inhibit cancer cell proliferation, instead of searching target genes for each miRNAs, we investigated common targets of all three miRNAs. To this end, we integrated the results of in silico analysis using five target-prediction algorithms (miRecords, miRTarBase, TargetScan, miRanda, and PITA) with the transcriptome profile for genes that were down-regulated by >1.5-fold following resveratrol treatment (Figure 2A) [27–32]. Of the 1,176 down-regulated genes, we identified 187, 321, and 75 genes as putative direct target genes of miR-34a, miR-424, and miR-503, respectively. Venn diagram analysis revealed that 11 probe sets, consisting of eight gene symbols, were common candidates for the putative direct target genes of the three miRNAs (miR-34a, miR-424, and miR-503; Figure 2A and 2B). To narrow down the list of potential target genes, we performed gene ontology analysis and KEGG pathway analysis based on the mRNA microarray data. The findings showed that resveratrol treatment affected the HNRNPA1-related pathway, which alters the processing of capped intron-containing pre-mRNAs (Figure 2C) [33–35]. HNRNPA1 is up-regulated in a wide spectrum of cancers, such as breast cancer, colorectal cancer, lung cancer, and glioma [36–39]. These results led us to focus on HNRNPA1.
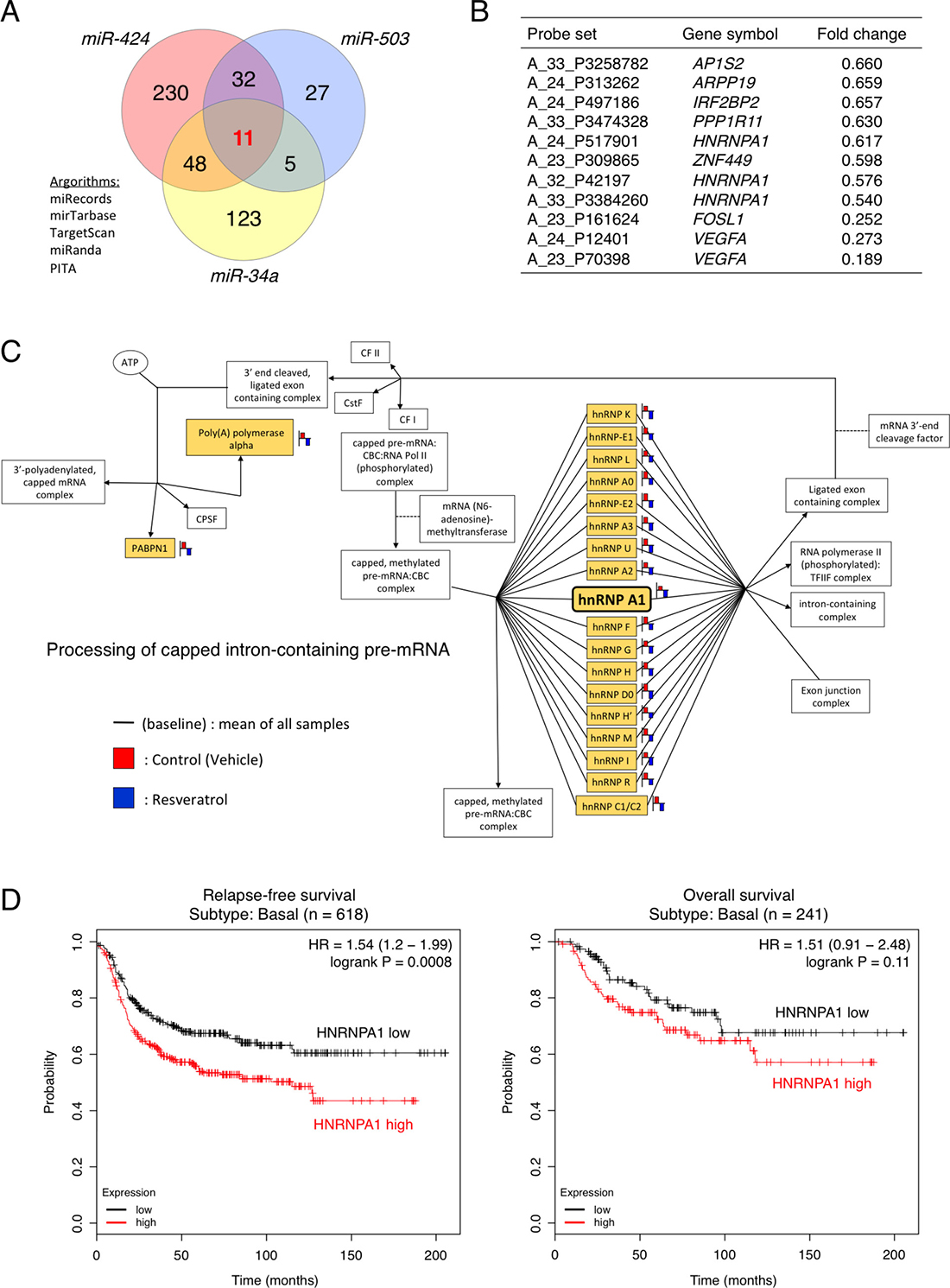
Figure 2: Clinical significance of HNRNPA1, a potential target gene of resveratrol-regulated miRNAs, in basal-like breast cancer. (A) Venn diagram depicting the overlap between predicted miR-34a, miR-424, and miR-503 targets. The target prediction algorithms miRecords, miRTarBase, TargetScan, miRanda, and PITA were chosen for whole-transcriptome analysis. (B) Common candidates for the putative direct target genes of miR-34a, miR-424, and miR-503 (C) KEGG pathway analysis of differentially expressed genes in the HNRNPA1-related pathway involved in processing capped intron-containing pre-mRNAs. The probe (gene symbol) on the arrays is represented as a yellow color box. (D) Left: Kaplan–Meier analysis of correlations between relapse-free survival in patients with basal-like breast cancer exhibiting high (above median value, n = 309) and low (below median value, n = 309) HNRNPA1 expression. Right: Kaplan–Meier analysis of correlations between overall survival of patients with basal-like breast cancer exhibiting high (above median value, n = 121) and low (below median value, n = 120) HNRNPA1 expression. The P value was determined by log-rank test. HR: hazard ratio.
Next, to evaluate the prognostic value of HNRNPA1 expression in basal-like breast cancer, we investigated the association between HNRNPA1 expression in breast cancer samples and disease progression (for other types of breast cancer, see Supplementary Figure 3) [40]. Intriguingly, Kaplan–Meier survival analysis demonstrated that patients with high HNRNPA1 expression had a significantly shorter relapse-free survival than patients with low HNRNPA1 expression (P = 0.0008 by log-rank test) and that patients with high HNRNPA1 expression had a shorter overall survival than patients with low HNRNPA1 expression (P = 0.11 by log-rank test; Figure 2D). In contrast, among potential target genes of miR-34a, miR-424, and miR-503, vascular endothelial growth factor A (VEGFA) and FOS-like 1 (FOSL1) expression did not exhibit clear correlations with poor prognosis in patients with basal-like cancer (Supplementary Figure 4). Therefore, the data above showed that, in basal-like breast cancer, HNRNPA1 was associated with poor clinical outcomes and could serve as a viable target for resveratrol/miRNAs in cancer treatment.
Resveratrol and resveratrol-regulated miRNAs suppress HNRNPA1 to mediate cell proliferation
As demonstrated above, HNRNPA1 could be highly relevant to cancer progression. However, it remained unclear whether HNRNPA1 expression is correlated positively or negatively with cell proliferation, miRNA expression, and resveratrol treatment. To determine the functional effects of HNRNPA1 on cell proliferation of basal-like breast cancer cells (MDA-MB-231-luc-D3H2LN cells), we knocked down HNRNPA1 using three independent siRNAs. siRNA transfection significantly reduced HNRNPA1 expression and cell proliferation (Figure 3A and 3B). This result suggested that HNRNPA1 promoted cancer cell proliferation and that suppressing HNRNPA1 expression inhibited cell proliferation.
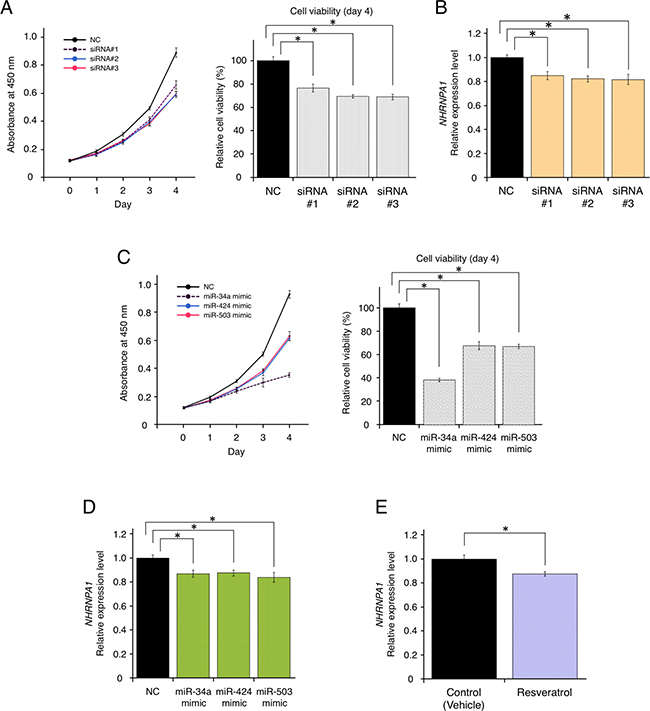
Figure 3: Resveratrol-mediated down-regulation of HNRNPA1 expression inhibited cancer cell proliferation. (A) Effects of HNRNPA1 siRNAs on the proliferation of MDA-MB-231-luc-D3H2LN cells (mean ± SEM). Left: cell viability at 0, 1, 2, 3, and 4 days after siRNA transfection. Right: cell proliferation normalized to that of the negative control (NC) at 4 days (*P < 0.05). The number of data points shown for each sample is six. (B) Effects of HNRNPA1 siRNAs on HNRNPA1 expression (mean ± SEM, *P < 0.05). The number of data points shown for each sample is four. (C) Effects of miR-34a, miR-424, or miR-503 mimics on the proliferation of MDA-MB-231-luc-D3H2LN cells (mean ± SEM). Left: cell viability at 0, 1, 2, 3, and 4 days after siRNA transfection. Right: cell proliferation normalized to that of the negative control (NC) at 4 days (*P < 0.05). The number of data points shown for each sample is six. (D) Effects of resveratrol-regulating miRNA (miR-34a, miR-424, and miR-503) mimics on HNRNPA1 expression (mean ± SEM, *P < 0.05). The number of data points shown for each sample is four. (E) Effects of resveratrol on HNRNPA1 expression (mean ± SEM, *P < 0.05). The number of data points shown for each sample is four.
Furthermore, we investigated whether resveratrol-regulated miRNAs (miR-34a, miR-424, and miR-503) and resveratrol could affect HNRNPA1 expression. After transfection with each miRNA mimic or non-target negative control miRNA mimic, we confirmed that overexpression of these miRNAs inhibited cell proliferation, demonstrating their effects as tumor-suppressive miRNAs (Figure 3C). We also observed that transfection of each miRNA significantly reduced HNRNPA1 expression (Figure 3D). Importantly, resveratrol treatment itself could down-regulate HNRNPA1 expression (Figure 3E). Taken together, these data suggested that the axis of resveratrol and resveratrol-regulated miRNAs functionally control the expression of HNRNPA1, which is involved in cancer cell proliferation.
Direct or indirect regulation of HNRNPA1 by resveratrol-regulated miRNAs
As shown in Figure 2A, using the target-prediction algorithms, HNRNPA1 was selected as a putative direct target of resveratrol-regulated miRNAs (miR-34a, miR-424, and miR-503). To elucidate the mechanism through which miR-34a, miR-424, and miR-503 regulate HNRNPA1, we prepared a luciferase reporter vector in which the 3′-untranslated region (UTR) of HNRNPA1 was cloned downstream of a luciferase gene (wild-type [WT]-HNRNPA1 3′-UTR vector). When the luciferase reporter WT-HNRNPA1 3′-UTR vector was co-transfected with miR-34a, miR-424, or miR-503 mimics, luciferase activity was significantly reduced by miR-424 and miR-503 mimics but not by the miR-34a mimic (Figure 4A). This result indicated that miR-34a could regulate HNRNPA1 without directly binding to the HNRNPA1 3′-UTR.
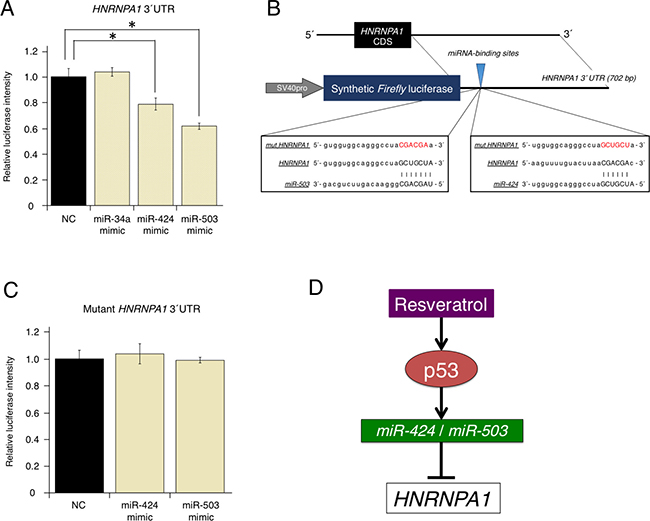
Figure 4: Direct regulation of oncogenic HNRNPA1 by the tumor-suppressive miRNAs, miR-424 and miR-503. (A) Effects of overexpressing resveratrol-regulated miRNA (miR-34a, miR-424, and miR-503) on WT-HNRNPA1 3′-UTR luciferase activity. Firefly luciferase activities were normalized to Renilla luciferase activities (mean ± SEM, *P < 0.05). Number of data points per sample was four. (B) Diagram of the luciferase reporter containing the WT and mutant HNRNPA1 3′-UTR regions. Sequences of miRNAs (miR-424 or miR-503), the miRNA-binding site, and the mutated site in the 3′-UTR of HNRNPA1 are shown. (C) Effects of overexpressing resveratrol-regulated miRNA (miR-424 and miR-503) on mutant HNRNPA1 3′-UTR luciferase activity. Firefly luciferase activities were normalized to Renilla luciferase activities (mean ± SEM, *P < 0.05). The number of data points shown for each sample is four. (D) Schematic representation of the molecular mechanisms through which resveratrol inhibits cancer cell proliferation via the p53/miRNA/HNRNPA1 axis.
We next prepared a mutant luciferase reporter (mut-HNRNPA1 3′-UTR vector) in which the putative binding site of miR-424 and miR-503 was changed (Figure 4B). In contrast to the WT-HNRNPA1 3′-UTR vector, when the mut-HNRNPA1 3′-UTR vector was used, mutations in the binding site abolished inhibition by miR-424 and miR-503 mimics (Figure 4C). These results strongly suggested that HNRNPA1 expression could be regulated directly by miR-424 and miR-503, but indirectly by miR-34a (Figure 4D).
DISCUSSION
It has long been considered that the consumption of healthy foods prevents or cures illness, whereas it has been shown that unhealthy eating habits are associated with increased risks for developing certain diseases, including cancer [41–44]. The therapeutic potential of naturally occurring products has attracted much attention in the past decade. Resveratrol, a multi-functional polyphenolic phytoalexin, has been known to exert anti-tumorigenic and anti-inflammatory effects, since it was demonstrated to inhibit carcinogenesis in mice [45]. Recently, it has been unveiled that some naturally occurring compounds, including resveratrol, regulate the expression of miRNAs in a favorable manner, which opened new avenues for cancer prevention and treatment. In breast cancer, some data have shown that resveratrol induces the expression of miR-663 and miR-744 in MCF7 cells and miR-200c and miR-141 in MDA-MB-231-luc-D3H2LN cells [14, 15]. However, it is still unclear how resveratrol controls cancer cell growth by regulating miRNA expression for the prevention of diseases. Here we showed that resveratrol controlled breast cancer cell proliferation by inducing tumor-suppressive miRNAs (miR-34a, miR-424, and miR-503) via the p53 pathway, which in turn suppressed HNRNPA1–the expression of which is associated with tumorigenesis and tumor progression.
miR-34a is a direct target of p53 and is known to promote cell cycle arrest in the G1 phase, induce senescence, and enhance apoptosis by directly repressing CDK4, CDK6, cyclin E2, E2F transcription factor 3 (E2F3), c-Myc, and BCL-2 [21–23, 46, 47]. Moreover, it was observed that resveratrol controls miR-34a expression in colon cancer and glioma cells [20, 48]. In breast cancer cells, consistent with previous studies, we showed that resveratrol induced miR-34a, suggesting that resveratrol could exert anticancer effects. miR-424 and miR-503, which are co-transcribed as the miR-424/503 cluster, are thought to belong to the extended miR-16 family based on the presence of the AGCAGC sequence in the 5′-end of the miRNA [49]. miR-16 has been identified as a tumor-suppressive miRNA and is deleted and/or downregulated in approximately 68% of cases of B-cell chronic lymphocytic leukemia, similar to results in many other cancers, strongly suggesting its important functions in tumor formation [50–54]. The seed sequences of miR-424 and miR-503 are similar to that of miR-16, indicating that they share target genes with miR-16 and act as tumor suppressors. Intriguingly, recent work in knockout mice has demonstrated that the miR-424/503 cluster is commonly lost in a subset of aggressive breast cancers, which promotes breast tumorigenesis in vivo [55]. These findings have provided evidence demonstrating that miR-424 and miR-503 act as tumor suppressors in breast cancer.
In this study, we found that resveratrol regulated the tumor-suppressive miRNAs miR-424 and miR-503 through the p53 pathway, in combination with miR-34a. These results support the therapeutic potential of resveratrol. Previous computational predictions suggested that various transcription factors, including runt-related transcription factor 1 (RUNX1), E2F3, SP3, yin yang 1 (YY1), nuclear factor erythroid 2-like 2 (NFE2L2), cAMP responsive element binding protein 1 (CREB1), activating transcription factor 2 (ATF2), upstream transcription factor 2 (USF2), ETS domain-containing protein (ELK1), CCAAT/enhancer binding protein beta (CEBPB), and homeobox A4 (HOXA4), can interact with the promoter region of miR-424/503 cluster genes [56]. Our data indicated that miR-424 and miR-503 expression was regulated by p53 in breast cancer; however, we have yet to elucidate whether these miRNAs were direct or indirect targets of p53.
Although we mainly focused on resveratrol-induced miRNAs in this research, it is also important to cast a spotlight on miRNAs that were decreased by resveratrol treatment. For example, some oncogenic miRNAs were down-regulated by resveratrol, such as miR-17, miR-17*, miR-18a, miR-19a, miR-19b, and miR-20a (miR-17-92 cluster). These miRNAs are transcriptionally repressed to varying degrees by p53 under normal conditions but, for instance, are up-regulated in human B-cell lymphoma and are amplified in malignant lymphoma [47, 57, 58]. Consistent with results showing that resveratrol controls tumor-suppressive miRNAs (miR-34a, miR-424, and miR-503) via the p53 pathway, p53-regulated oncogenic miRNAs were down-regulated by resveratrol, which suggests that such down-regulation could contribute to the inhibition of cancer cell growth and proliferation. It is reasonable to expect that balancing the expression of tumor-suppressive and oncogenic miRNAs is critical for its efficacy. Our results suggested that resveratrol could control tumor-suppressive and oncogenic miRNAs in a positive manner, but more studies are needed to fully understand these mechanisms.
HNRNPA1 is linked to tumorigenesis and tumor progression and has been shown to be up-regulated in a wide spectrum of cancers, such as breast cancer, colorectal cancer, lung cancer, and glioma [36–39]. It has been known that HNRNPA1 plays an important role in the modulation of gene expression by properly controlling the splicing of a wide variety of gene transcripts. Because numerous transcripts are targeted by HNRNPA1, it is currently difficult to determine which known or unknown target transcripts mainly mediate downstream cell growth and proliferation that is inhibited by resveratrol.
The mRNA transcripts of several genes, such as breast cancer 1 (BRCA1) and pyruvate kinase M (PKM), have been experimentally demonstrated to be alternatively spliced by HNRNPA1 [60]. As a tumor-suppressor gene, BRCA1 plays a pivotal role in cell-cycle regulation and DNA damage repair, and its defects result in defective cell-cycle checkpoints and genome instability [61]. Dysregulations of HNRNPA1 can lead to aberrant exon-skipping of BRCA1 transcripts and impair its function in cell-cycle regulation [62, 63]. Regarding PKM, the oncogenic transcription factor c-Myc promotes the expression of three HNRNP proteins (hnRNPI, hnRNPA1, and hnRNPA2), resulting in maintaining a high level of the embryonic isoform of PKM containing exon 10 (PKM2), as compared with the adult isoform of PKM containing exon 9 (PKM1) [39]. Because PKM2 plays an important role as a promoter of the Warburg effect, its overexpression is likely to be universal in many tumors and it is expected to be critical for cancer cell proliferation [59]. Although, after the treatment of resveratrol, it remains uncovered how the downstream genes of HNRNPA1 contribute to the phenotype observed in this research, aberrant expression of HNRNP genes, including HNRNPA1, has an impact on the regulation of pre-mRNA alternative splicing in cancer, suggesting that the maintenance of HNRNP levels is important for cancer prevention and treatment.
In this study, we focused on HNRNPA1 as a common target of resveratrol-regulated miRNAs (miR-34a, miR-424, and miR-503). As described above, HNRNPA1 could serve as a therapeutic target in breast cancer, consistent with the Kaplan–Meier analysis showing that high HNRNPA1 levels were associated with poor clinical outcomes in basal-like breast cancer. We also showed that HNRNPA1 expression was inhibited by resveratrol and its regulated miRNAs, and that HNRNPA1 is a new direct target of miR-424 and miR-503. These data shed new light on the molecular mechanisms through which resveratrol contributes to the prevention and treatment of cancer. Although miR-34a indirectly suppressed HNRNPA1, it directly represses key genes required for promoting cell-cycle arrest in the G1 phase, senescence, and apoptosis [64]. Because of the existence of many potential miR-34a target genes, we could not elucidate the relationship between the downstream genes and HNRNPA1, although this will be an important subject of future studies.
Notably, KEGG pathway analysis showed that resveratrol could exert broad effects on the HNRNPA1-related pre-mRNA splicing pathway. However, it is currently difficult to determine how resveratrol affects splicing regulation and sequential changes in gene expression. It will be of interest to examine whether resveratrol can regulate its target miRNAs (miR-34a, miR-424, and miR-503) and HNRNPA1 expression in other types of cancer. For therapeutic applications, future investigations will provide important information regarding the regulatory mechanisms of resveratrol.
For many years, a variety of studies have identified relationships between food and health, demonstrating the significance of a well-balanced diet for good health. Moreover, there is the possibility to prevent diseases and maintain good health if the expression levels and profiles of miRNAs in the body are appropriately maintained, as supported by the observation that changes in miRNA profiles in the bloodstream can be detected during the early stages of disease [20, 65]. It is particularly intriguing how foods and natural compounds substantially impact the expression of various miRNAs in the body, leading to the prevention of diseases. Thus, we believe that research and development will maintain or improve human health worldwide.
MATERIALS AND METHODS
Cell culture
MDA-MB-231-luc-D3H2LN cells (Xenogen) and MCF7 cells (American Type Culture Collection) were cultured in RPMI 1640 medium supplemented with 10% heat-inactivated fetal bovine serum and 1% antibiotic-antimycotic (Thermo Fisher Scientific) at 37° C in 5% CO2. Trans-resveratrol (Cayman Chemical) was used at concentrations ranging from 12.5–50 mM.
Cell proliferation assay (MTS assay)
Cells were seeded in 96-well plates at a density of 5,000 cells/well. The following day, the cells were treated with resveratrol. After 3 days in culture, cell viability was measured using the Cell Counting Kit-8 (Dojindo Laboratories) according to the manufacturer’s instructions. The absorbance at 450 nm was measured with a Synergy H4 microplate reader (BioTek Instruments). Experiments were performed in at least triplicate.
Expressing short-hairpin RNAs (shRNAs) via lentiviral transduction
Cell lines stably expressing a p53-specific shRNA (shp53) or a non-targeting control shRNA (shNC) were established by transducing cells with pLKO.1 lentiviral vectors encoding either shRNA, both of which were purchased from Sigma. The human p53 shRNA target sequence was as follows: 5′-GACUCCAGUGG UAAUCUAC-3′. The control target shRNA sequence was as follows: 5′-GAAAUGUACUGCGCGUGGAGAC-3′. In knockdown experiments, 293T cells were transduced with constructs expressing shp53 or shNC. MCF7 cells were infected with recombinant lentiviruses expressing shp53 or shNC.
Immunoblot analysis
Two days after resveratrol treatment, whole cell lysates were obtained using the M-PER mammalian protein-extraction reagent (Thermo Fisher Scientific). The proteins were resolved in a Mini-PROTEAN TGX Gel (4–12%, Bio-Rad) and electrotransferred onto a polyvinylidene fluoride membrane (Millipore). After blocking in Blocking One solution (Nacalai Tesque), the membranes were incubated for 1 h at room temperature with primary antibodies against p53 (sc-126; diluted 1:5000; Santa Cruz Biotechnology) and actin (MAB1501, diluted 1:5000; Millipore). The secondary antibody (horseradish peroxidase-conjugated anti-mouse IgG, NA931; GE Healthcare) was used at a dilution of 1:5000. The membranes were then exposed to ImmunoStar LD (Wako). The bound antibodies were visualized by chemiluminescence using the ECL Plus Western blotting detection system (GE HealthCare), and the signals were detected with a LAS-3000 LuminoImager (Fuji Film).
qRT-PCR analysis
Total RNA and miRNA were extracted from cells cultured with or without resveratrol treatment using the QIAzol and miRNeasy Mini Kit (Qiagen) according to the manufacturer’s protocol. cDNA was generated using the PrimeScript RT reagent Kit (TaKaRa) and the TaqMan® MicroRNA Reverse Transcript Kit (Applied Biosystems). TaqMan® probes (TaqMan® Gene Expression Assays and TaqMan® MicroRNA Assays) were purchased from Applied Biosystems. qRT-PCR reactions were performed with the TaqMan probes using the Premix Ex Taq™ (Probe qPCR) (TaKaRa) and TaqMan™ Universal PCR Master Mix, no AmpErase™ UNG (Thermo Fisher Scientific) on the StepOne® Real-Time PCR system (Applied Biosystems). Experiments were performed in at least triplicate. Expression levels of genes of interest were normalized to the expression of GAPDH and RNU44B.
Microarray analyses
For mRNA microarray analysis, total RNA (50 ng) was extracted using the QIAzol reagent (Qiagen) from MDA-MB-231-luc-D3H2LN cells, with or without exposure to 25 μM resveratrol for 2 days, and purified using the RNeasy Micro Kit (Qiagen). SurePrint G3 Human GE 8 × 60 K arrays (Agilent Technologies) were hybridized with cyanine 3 (Cy3)-labeled cRNA targets prepared from the RNA samples, according to the manufacturer’s instructions. Experiments were performed in biological triplicate.
For miRNA microarray analysis, total RNA (100 ng) was extracted using the QIAzol reagent (Qiagen) from MDA-MB-231-luc-D3H2LN cells, with or without exposure to 25 μM resveratrol for 2 days, and purified using the RNeasy Micro Kit (Qiagen). SurePrint G3 Human miRNA 8 × 60 K arrays (Agilent Technologies) were hybridized with 5′-cytidine bisphosphate-Cy3 (pCp-Cy3)-labeled cRNA targets prepared from the RNA samples according to the manufacturer’s instructions. Experiments were performed in biological triplicate. Each of the arrays was analyzed with the Microarray Scanner (Agilent Technologies). Gene-expression levels were calculated using Feature Extraction software, version 10.7.3.1 (Agilent Technologies). Normalized and log-transformed intensity values were analyzed using GeneSpring GX software, version 7.3.1 (Agilent Technologies). The intensity values were log2-transformed and imported into the Partek Genomics Suite 6.6 (Partek Incorporated, USA). One-way analysis of variance (ANOVA) was performed to identify differentially expressed genes. Unsupervised clustering and heat map generation were performed with sorted data sets by the Euclidean distance based on average linkage clustering, and principal component analysis mapping was conducted for all probe sets using Partek Genomics Suite 6.6. Gene set-enrichment analysis (http://software.broadinstitute.org/gsea/index.jsp) was performed to compare resveratrol-treated samples and control samples.
Transient assays
For siRNA transfections, pre-designed siRNAs targeting HNRNPA1 mRNA (IDs: s6710, s6712, and s223860) were purchased from Applied Biosystems. The Silencer® Select Negative Control #1 siRNA (Applied Biosystems) was used as a negative control. For miRNA transfections, synthetic hsa-miR-34a-5p (ID: MC11030), hsa-miR-424-5p (ID: MC10306), and hsa-miR-503-3p (ID: MC10378) were purchased from Applied Biosystems. The mirVana™ miRNA Mimic, Negative Control #1 (Applied Biosystems) was used as a negative control. Transfections were performed using the DharmaFECT transfection reagent (Thermo Fisher Scientific) according to the manufacturer’s protocol. A final concentration 25 nM in the supernatant was used when transfecting siRNAs or miRNAs. Experiments were performed in at least triplicate.
Luciferase reporter assays
For 3′-UTR luciferase reporter assays, pEZX-MT01 dual luciferase reporter plasmids containing the 3′-UTR of HNRNPA1, or the mutated 3′-UTR of HNRNPA1, as well as a control plasmid lacking the no 3′-UTR were purchased from GeneCopoeia. MCF7 cells were seeded into a 96-well plate at a density of 1.5 × 104 cells/well and co-transfected with the 100 ng of a vector and the 50 nM of a mirVana™ miRNA mimic or the mirVana™ miRNA inhibitor Negative Control (Thermo Fisher Scientific), using the DharmaFECT Duo transfection reagent (Thermo Fisher Scientific) for 24 h. The luciferase assays were performed using the Dual-Glo Luciferase assay system (Promega) according to the manufacturer’s instructions. Experiments were performed in at least triplicates.
Survival analysis
Overall survival in patients with basal-type, luminal A, luminal B, or HER2-positive breast cancer was stratified according to expression of the gene of interest and is presented as Kaplan–Meier plots (http://www.kmplot.com) [40].
Statistical analysis
The data are presented as the means ± SEM of at least three independent experiments. Statistical analyses of data containing more than two groups were performed using one-way ANOVA followed by Tukey’s honest significant difference test to account for multiple comparisons. P < 0.05 was considered statistically significant.
Author contributions
TO and KO designed and directed the study. KO designed and performed most of the experiments, analyzed data, and wrote the manuscript. KO and YY contributed to the data interpretation for microarray analyses. All authors critically reviewed the manuscript and approved the final version of the manuscript.
ACKNOWLEDGMENTS
We thank Ryotaro Fujimura, Norimitsu Yamagata, Hideto Yoshida, Ryosuke Matsuoka, Yasunobu Masuda, Masaaki Kunou, and Yoshinori Hamachiyo (Kewpie Corporation) for providing valuable comments on this paper. This work was supported in part by a Grant-in-Aid for Scientific Research on Priority Areas, Cancer, from the Ministry of Education, Culture, Sports, Science and Technology, Japan, and Japan Agency for Medical Research and Development (AMED).
CONFLICTS OF INTEREST
The authors have declared no conflicts of interests.
REFERENCES
1. Van Poppel H, Tombal B. Chemoprevention of prostate cancer with nutrients and supplements. Cancer Manag Res. 2011; 3:91–100.
2. Catalgol B, Batirel S, Taga Y, Ozer NK. Resveratrol: French paradox revisited. Front Pharmacol. 2012; 3:141.
3. Jasinski M, Jasinska L, Ogrodowczyk M. Resveratrol in prostate diseases - a short review. Cent European J Urol. 2013; 66:144–149.
4. Lippi G, Franchini M, Favaloro EJ, Targher G. Moderate red wine consumption and cardiovascular disease risk: beyond the “French paradox”. Semin Thromb Hemost. 2010; 36:59–70.
5. Howitz KT, Bitterman KJ, Cohen HY, Lamming DW, Lavu S, Wood JG, Zipkin RE, Chung P, Kisielewski A, Zhang LL, Scherer B, Sinclair DA. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature. 2003; 425:191–196.
6. Michan S, Sinclair D. Sirtuins in mammals: insights into their biological function. Biochem J. 2007; 404:1–13.
7. Scott E, Steward WP, Gescher AJ, Brown K. Resveratrol in human cancer chemoprevention - Choosing the ‘right’ dose. Mol Nutr Food Res. 2012; 56:7–13.
8. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, et al. MicroRNA expression profiles classify human cancers. Nature. 2005; 435:834–838.
9. Lujambio A, Lowe SW. The microcosmos of cancer. Nature. 2012; 482:347–355.
10. Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007; 9:654–659.
11. Vlassov AV, Magdaleno S, Setterquist R, Conrad R. Exosomes: current knowledge of their composition, biological functions, and diagnostic and therapeutic potentials. Biochim Biophys Acta. 2012; 1820:940–948.
12. Zen K, Zhang CY. Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev. 2012; 32:326–348.
13. Shimomura A, Shiino S, Kawauchi J, Takizawa S, Sakamoto H, Matsuzaki J, Ono M, Takeshita F, Niida S, Shimizu C, Fujiwara Y, Kinoshita T, Tamura K, et al. Novel combination of serum microRNA for detecting breast cancer in the early stage. Cancer Sci. 2016; 107:326–334.
14. Vislovukh A, Kratassiouk G, Porto E, Gralievska N, Beldiman C, Pinna G, El’skaya A, Harel-Bellan A, Negrutskii B, Groisman I. Proto-oncogenic isoform A2 of eukaryotic translation elongation factor eEF1 is a target of miR-663 and miR-744. Br J Cancer. 2013; 108:2304–2311.
15. Hagiwara K, Kosaka N, Yoshioka Y, Takahashi RU, Takeshita F, Ochiya T. Stilbene derivatives promote Ago2-dependent tumour-suppressive microRNA activity. Sci Rep. 2012; 2:314.
16. Yu YH, Chen HA, Chen PS, Cheng YJ, Hsu WH, Chang YW, Chen YH, Jan Y, Hsiao M, Chang TY, Liu YH, Jeng YM, Wu CH, et al. MiR-520h-mediated FOXC2 regulation is critical for inhibition of lung cancer progression by resveratrol. Oncogene. 2013; 32:431–443.
17. Liu P, Liang H, Xia Q, Li P, Kong H, Lei P, Wang S, Tu Z. Resveratrol induces apoptosis of pancreatic cancers cells by inhibiting miR-21 regulation of BCL-2 expression. Clin Transl Oncol. 2013; 15:741–746.
18. Tili E, Michaille JJ, Alder H, Volinia S, Delmas D, Latruffe N, Croce CM. Resveratrol modulates the levels of microRNAs targeting genes encoding tumor-suppressors and effectors of TGFbeta signaling pathway in SW480 cells. Biochem Pharmacol. 2010; 80:2057–2065.
19. Dhar S, Hicks C, Levenson AS. Resveratrol and prostate cancer: promising role for microRNAs. Mol Nutr Food Res. 2011; 55:1219–1229.
20. Kumazaki M, Noguchi S, Yasui Y, Iwasaki J, Shinohara H, Yamada N, Akao Y. Anti-cancer effects of naturally occurring compounds through modulation of signal transduction and miRNA expression in human colon cancer cells. J Nutr Biochem. 2013; 24:1849–1858.
21. He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y, Xue W, Zender L, Magnus J, Ridzon D, Jackson AL, Linsley PS, Chen C, et al. A microRNA component of the p53 tumour suppressor network. Nature. 2007; 447:1130–1134.
22. Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, Arking DE, Beer MA, Maitra A, et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007; 26:745–752.
23. Raver-Shapira N, Marciano E, Meiri E, Spector Y, Rosenfeld N, Moskovits N, Bentwich Z, Oren M. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell. 2007; 26:731–743.
24. Tarasov V, Jung P, Verdoodt B, Lodygin D, Epanchintsev A, Menssen A, Meister G, Hermeking H. Differential regulation of microRNAs by p53 revealed by massively parallel sequencing: miR-34a is a p53 target that induces apoptosis and G1-arrest. Cell Cycle. 2007; 6:1586–1593.
25. Lu X, Errington J, Curtin NJ, Lunec J, Newell DR. The impact of p53 status on cellular sensitivity to antifolate drugs. Clin Cancer Res. 2001; 7:2114–2123.
26. Olivier M, Eeles R, Hollstein M, Khan MA, Harris CC, Hainaut P. The IARC TP53 database: new online mutation analysis and recommendations to users. Hum Mutat. 2002; 19:607–614.
27. Xiao F, Zuo Z, Cai G, Kang S, Gao X, Li T. miRecords: an integrated resource for microRNA-target interactions. Nucleic Acids Res. 2009; 37:D105–110.
28. Hsu SD, Lin FM, Wu WY, Liang C, Huang WC, Chan WL, Tsai WT, Chen GZ, Lee CJ, Chiu CM, Chien CH, Wu MC, Huang CY, et al. miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res. 2011; 39:D163–169.
29. Chou CH, Chang NW, Shrestha S, Hsu SD, Lin YL, Lee WH, Yang CD, Hong HC, Wei TY, Tu SJ, Tsai TR, Ho SY, Jian TY, et al. miRTarBase 2016: updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Res. 2016; 44:D239–247.
30. Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005; 120:15–20.
31. Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E. The role of site accessibility in microRNA target recognition. Nat Genet. 2007; 39:1278–1284.
32. Betel D, Wilson M, Gabow A, Marks DS, Sander C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 2008; 36:D149–153.
33. Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000; 28:27–30.
34. Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016; 44:D457–462.
35. Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017; 45:D353–D361.
36. Pino I, Pío R, Toledo G, Zabalegui N, Vicent S, Rey N, Lozano MD, Torre W, García-Foncillas J, Montuenga LM. Altered patterns of expression of members of the heterogeneous nuclear ribonucleoprotein (hnRNP) family in lung cancer. Lung Cancer. 2003; 41:131–143.
37. Ushigome M, Ubagai T, Fukuda H, Tsuchiya N, Sugimura T, Takatsuka J, Nakagama H. Up-regulation of hnRNP A1 gene in sporadic human colorectal cancers. Int J Oncol. 2005; 26:635–640.
38. Li S, Xu H, Ding H, Huang Y, Cao X, Yang G, Li J, Xie Z, Meng Y, Li X, Zhao Q, Shen B, Shao N. Identification of an aptamer targeting hnRNP A1 by tissue slide-based SELEX. J Pathol. 2009; 218:327–336.
39. David CJ, Chen M, Assanah M, Canoll P, Manley JL. HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature. 2010; 463:364–368.
40. Györffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, Szallasi Z. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010; 123:725–731
41. Crozier A, Lean MEJ, McDonald MS, Black C. Quantitative Analysis of the Flavonoid Content of Commercial Tomatoes, Onions, Lettuce, and Celery. Journal of Agricultural and Food Chemistry. 1997; 45:590–595.
42. Willett WC. Diet, nutrition, and avoidable cancer. Environ Health Perspect. 1995; 103:165–170.
43. Sasazuki S, Inoue M, Iwasaki M, Sawada N, Shimazu T, Yamaji T, Tsugane S; JPHC Study Group. Combined impact of five lifestyle factors and subsequent risk of cancer: the Japan Public Health Center Study. Prev Med. 2012; 54:112–116.
44. Mayne ST, Playdon MC, Rock CL. Diet, nutrition, and cancer: past, present and future. Nat Rev Clin Oncol. 2016; 13:504–515.
45. Jang M, Cai L, Udeani GO, Slowing KV, Thomas CF, Beecher CW, Fong HH, Farnsworth NR, Kinghorn AD, Mehta RG, Moon RC, Pezzuto JM. Cancer chemopreventive activity of resveratrol, a natural product derived from grapes. Science. 1997; 275:218–220.
46. Tazawa H, Tsuchiya N, Izumiya M, Nakagama H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci U S A. 2007; 104:15472–15477.
47. Otsuka K, Ochiya T. Genetic networks lead and follow tumor development: microRNA regulation of cell cycle and apoptosis in the p53 pathways. Biomed Res Int. 2014; 2014:749724.
48. Wang H, Feng H, Zhang Y. Resveratrol inhibits hypoxia-induced glioma cell migration and invasion by the p-STAT3/miR-34a axis. Neoplasma. 2016; 63:532–539.
49. Caporali A, Emanueli C. MicroRNA-503 and the extended microRNA-16 family in angiogenesis. Trends Cardiovasc Med. 2011; 21:162–166.
50. Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, Rassenti L, Kipps T, Negrini M, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci U S A. 2002; 99:15524–15529.
51. Bottoni A, Piccin D, Tagliati F, Luchin A, Zatelli MC, degli Uberti EC. miR-15a and miR-16-1 down-regulation in pituitary adenomas. J Cell Physiol. 2005; 204:280–285.
52. Xia L, Zhang D, Du R, Pan Y, Zhao L, Sun S, Hong L, Liu J, Fan D. miR-15b and miR-16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells. Int J Cancer. 2008; 123:372–379.
53. Bonci D, Coppola V, Musumeci M, Addario A, Giuffrida R, Memeo L, D’Urso L, Pagliuca A, Biffoni M, Labbaye C, Bartucci M, Muto G, Peschle C, et al. The miR-15a-miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med. 2008; 14:1271–1277.
54. Bandi N, Zbinden S, Gugger M, Arnold M, Kocher V, Hasan L, Kappeler A, Brunner T, Vassella E. miR-15a and miR-16 are implicated in cell cycle regulation in a Rb-dependent manner and are frequently deleted or down-regulated in non-small cell lung cancer. Cancer Res. 2009; 69:5553–5559.
55. Rodriguez-Barrueco R, Nekritz EA, Bertucci F, Yu JY, Sanchez-Garcia F, Zeleke TZ, Gorbatenko A, Birnbaum D, Ezhkova E, Cordon-Cardo C, Finetti P, Llobet-Navas D, Silva JM. miR-424/503 is a breast cancer tumor suppressor whose loss promotes resistance to chemotherapy. Genes Dev. 2017; 31:553–566.
56. Schmeier S, MacPherson CR, Essack M, Kaur M, Schaefer U, Suzuki H, Hayashizaki Y, Bajic VB. Deciphering the transcriptional circuitry of microRNA genes expressed during human monocytic differentiation. BMC Genomics. 2009; 10:595.
57. Ota A, Tagawa H, Karnan S, Tsuzuki S, Karpas A, Kira S, Yoshida Y, Seto M. Identification and Characterization of a Novel Gene, C13orf25, as a Target for 13q31-q32 Amplification in Malignant Lymphoma. Cancer Research. 2004; 64:3087–3095.
58. Yan HL, Xue G, Mei Q, Wang YZ, Ding FX, Liu MF, Lu MH, Tang Y, Yu HY, Sun SH. Repression of the miR-17-92 cluster by p53 has an important function in hypoxia-induced apoptosis. EMBO J. 2009; 28:2719–2732.
59. David CJ, Manley JL. Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev. 2010; 24:2343–2364.
60. Jean-Philippe J, Paz S, Caputi M. hnRNP A1: the Swiss army knife of gene expression. Int J Mol Sci. 2013; 14:18999–19024.
61. Deng CX. BRCA1: cell cycle checkpoint, genetic instability, DNA damage response and cancer evolution. Nucleic Acids Res. 2006; 34:1416–1426.
62. Goina E, Skoko N, Pagani F. Binding of DAZAP1 and hnRNPA1/A2 to an exonic splicing silencer in a natural BRCA1 exon 18 mutant. Mol Cell Biol. 2008; 28:3850–3860.
63. Raponi M, Kralovicova J, Copson E, Divina P, Eccles D, Johnson P, Baralle D, Vorechovsky I. Prediction of single-nucleotide substitutions that result in exon skipping: identification of a splicing silencer in BRCA1 exon 6. Hum Mutat. 2011; 32:436–444.
64. Hermeking H. MicroRNAs in the p53 network: micromanagement of tumour suppression. Nat Rev Cancer. 2012; 12:613–626.
65. Otsuka K, Yamamoto Y, Matsuoka R, Ochiya T. Maintaining good miRNAs in the body keeps the doctor away?: Perspectives on the relationship between food-derived natural products and microRNAs in relation to exosomes/extracellular vesicles. Mol Nutr Food Res. 2018; 62:1700080.

