INTRODUCTION
While breast cancer in Asia is characterized by a lower incidence rate than in the United States and Europe, [1, 2] it is still one of the leading causes of cancer death in Asia, particularly in Taiwan [3–5]. This disease is featured by its complexity due to the genetic heterogeneity of breast carcinomas [6]. Since a few decades ago, many gene-expression profiling studies of breast cancer have revealed the existence of four major subtypes differing markedly in prognosis: luminal-A, luminal-B, HER2-amplified, and basal-like [6], prevalence of which varies by racial/ethnic groups [7]. Among the Asian populations, prevalence of the luminal-A, luminal-B, HER2+/ER-, basal-like, and unclassified subtype has been shown to be 55-65%,, 10-20%, 10-15%, 10-15%, and 0-5%, respectively [8, 9]. In the current study, we focused on luminal-like breast cancer comprising the luminal-A and luminal-B subtypes, which are defined by the presence of hormonal receptor and absence of HER2 on the plasma membrane of tumor cells (i.e., HR+/HER2-) by the immunohistochemistry [10]. Several studies have suggested that luminal-like cancers tend to have the most favorable prognosis and longer-term survival when compared with the other subtypes [11, 12]. However, early-stage breast cancer patients with the luminal-like subtypes are commonly (up to 75%) overtreated with adjuvant chemotherapy despite that recent studies have indicated that adjuvant chemotherapy may not provide significant benefit in reducing risk of recurrence [13, 14].
To overcome this issue, several multigene panels, such as Oncotype DX®(ODx), MammaPrint® and EndoPredict® assay kits, have been developed to help clinical decision-making regarding adjuvant chemotherapy for patients with early-stage breast carcinomas. ODx (Genomic Health Inc., Redwood, CA) is a prognostic and predictive assay kit for women with HR+ and HER2- breast cancer. It is a 21-gene RT-PCR assay for 16 cancer-related and five housekeeping control genes with an aim to aid physicians and patients to determine the best course of treatment by predicting the risk of distant recurrence of breast cancer. The ODx assay produces a numerical recurrence score and places patients into three categories: low-, intermediate- and high-risk [15]. Another test (MammaPrint by Agendia BV, Amsterdam, Netherlands) is a microarray-based gene-expression profiling assay that can classify the risk of distant recurrence into two categories, low and high, by analyzing 70 genes of HR+ and node-negative patients that had not received adjuvant systemic therapy [16]. The third test, EndoPredict (Myriad Genetics, Salt Lake City, UT), is a 11-gene RT-PCR test that provides prognostic information regarding the risk of distant recurrence of breast cancer to patients with HR+ and HER2- tumors [17]. The assay measures the expression of eight cancer-related and three control genes, and classifies patients under endocrine therapy into low- or high-risk of distant recurrence.
Even though all such tools can help in making treatment decision on adjuvant chemotherapy for patients with early-stage breast carcinoma, none of them was originally developed for Asian patients even though Asian people may have different mechanisms in breast cancer due to multiple factors such as ethnic, environmental, and genetic variations [18–20]. To overcome such potential limitations, we developed an 18-gene classifier (18-GC) with tumor tissues obtained from Asian breast cancer patients and compared its performance with that of ODx in predicting distant metastasis in early-stage HR+/HER2- patients.
RESULTS
Patient characteristics
Two-hundred twenty-four HR+ and HER2- breast cancer patients were included in the study (Table 1) with 202 (90.2%) of the patients diagnosed with no metastasis during the development of their breast cancer and 185 (82.6%) treated with adjuvant chemotherapy. Among those receiving chemotherapy, 165 (89.2%) of the patients did not develop metastasis during their follow-up.
Table 1: Characteristics of patients with and without metastasis in HR+/HER2- invasive breast carcinomas (n=224)
Variable |
Absence of metastasis |
Presence of metastasis |
p-value |
|---|---|---|---|
n=202 |
n=22 |
||
n (%) |
n (%) |
||
Age |
|||
≦40 (n=48) |
38 (18.8) |
10 (45.5) |
0.011 |
>40 (n=176) |
164 (81.2) |
12 (54.5) |
|
T stage |
|||
T1 (n=110) |
106 (52.5) |
4 (18.2) |
0.003 |
T2-T3 (n=114) |
96 (47.5) |
18 (81.8) |
|
N stage |
|||
N0 (n=130) |
119 (58.9) |
11 (50.0) |
0.421 |
N1 (n=94) |
83 (41.1) |
11 (50.0) |
|
Chemotherapy |
|||
No (n=39) |
37 (18.3) |
2 (9.1) |
0.382 |
Yes (n=185) |
165 (81.7) |
20 (90.9) |
|
Radiotherapy |
|||
No (n=93) |
83 (41.1) |
10 (45.5) |
0.693 |
Yes (n=131) |
119 (58.9) |
12 (54.5) |
|
Hormone therapy |
|||
No (n=9) |
8 (4.0) |
1 (4.5) |
1.000 |
Yes (n=215) |
194 (96.0) |
21 (94.5) |
|
Lymph vessel invasion |
|||
Nil/ Minimal (n=182) |
166 (82.2) |
16 (72.7) |
0.263 |
Prominent (n=42) |
36 (17.8) |
6 (27.3) |
|
ER status |
|||
Negative (n=5) |
5 (2.5) |
0 (0.0) |
1.000 |
Positive (n=219) |
197 (97.5) |
22 (100.0) |
|
PR status |
|||
Negative (n=40) |
35 (17.3) |
5 (22.7) |
0.558 |
Positive (n=184) |
167 (82.7) |
17 (77.3) |
We found that two characteristics were significantly associated with the presence of metastasis in unadjusted analysis: a) an age at 40 or younger; b) a stage of T2-T3 (Table 1). Among the patients with no metastasis, 81.2% were over 40 years old, whereas only 54.5% of the patients with metastasis were categorized in the same age group (p=0.011). On the other hand, probability for patients with a T2-T3 tumor stage to develop metastasis is significantly higher than that for patients with a T1 tumor stage (81.8% with metastasis vs. 47.5% without metastasis for T2-T3 compared with 18.2% with metastasis vs. 52.5% without metastasis for T1, p=0.003).
Determination of the breakpoint score
For the breakpoint determination in the original panel of the 18 genes selected previously (Table 2), we compared the ROC curves with different cutoff points that stratify patients into two groups: low- or high-risk of recurrence. We selected a breakpoint score of 21 (Figure 1) since this score minimized the distance on the ROC curve to the left top edge of the diagram and produced a combined greatest sensibility (95.4%) and specificity (39.1%). By using this breakpoint, 80 (35.7%) of the 224 patients were classified as having low risk and 144 (64.3%) high risk.
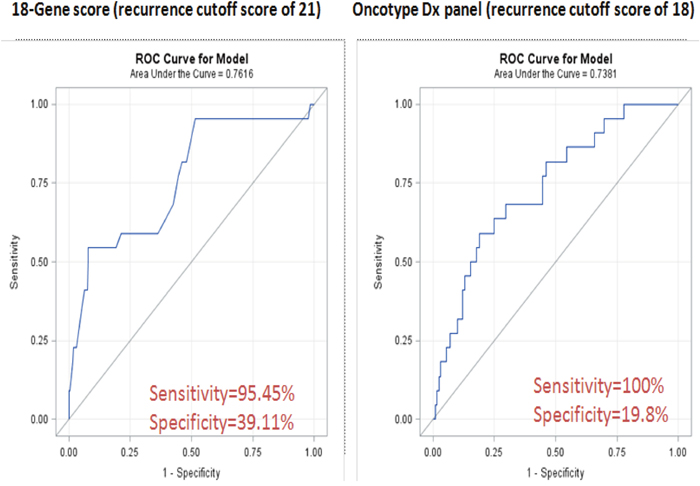
Figure 1: ROC curve analyses of the 18-GC and ODx.
Table 2: Genes selected for gene-expression profiling analysis
Gene symbol |
Gene title |
GenBank accession number |
|---|---|---|
TRPV6 |
Transient receptor potential cation channel, subfamily V, member 6 |
NM_018646 |
DDX39 |
DEAD (Asp-Glu-Ala-Asp) box polypeptide 39 |
NM_005804 |
BUB1B |
Budding uninhibited by benzimidazoles1 homolog beta (yeast) |
NM_001211 |
CCR1 |
Chemokine (C-C motif) receptor 1 |
NM_001295 |
STIL |
SCL/TAL1 interrupting locus |
NM_003035 |
BLM |
Bloom syndrome |
NM_000057 |
C16ORF7 |
Chromosome 16 open reading frame 7 |
NM_004913 |
PIM1 |
Pim-1 oncogene |
NM_002648 |
TPX2 |
TPX2, microtubule associated |
NM_012112 |
PTI1 |
Homo sapiens elongation factor 1-alpha 1 |
NM_001402 |
TCF3 |
Transcription factor 3 (E2A immunoglobulinenhancer binding factors E12/E47) |
NM_003200 |
CCNB1 |
Cyclin B1 |
NM_031966 |
DTX2 |
Deltex 2, E3 Ubiquitin Ligase |
NM_020892 |
ENSA |
Endosulfine alpha |
NM_004436 |
RCHY1 |
Ring Finger And CHY Zinc Finger Domain Containing 1, E3 Ubiquitin Protein Ligase |
NM_015436 |
NFATC2IP |
Nuclear Factor Of Activated T-Cells, Cytoplasmic, Calcineurin-Dependent 2 Interacting Protein |
NM_032815 |
OBSL1 |
Obscurin-like 1 |
NM_015311 |
MMP15 |
Matrix Metallopeptidase 15 (Membrane-Inserted) |
NM_002428 |
Patients classified in the intermediate-group by ODx were considered high risk when in comparison with the 18-gene panel. By using a breakpoint score of 21 in our gene panel assay, a total of 164 patients was classified in the same risk level as the ODx assay (73.2% concordance), indicating a significant agreement in the outcome predictions for individual patients (Table 3).
Table 3: Concordance between the 18-GC and ODx
Risk group |
18-gene classifier |
|||
|---|---|---|---|---|
Low (<21) |
High (≧21) |
Total |
||
Oncotype |
Low (<18) |
30 (75.0%) |
10 (25.0%) |
40 (17.9%) |
DX |
High (≧18) |
50 (27.2%) |
134 (72.8%) |
184 (82.1%) |
Total |
80 (35.7%) |
144 (64.3%) |
224 (100%) |
|
Recurrence rate and recurrence-free survival by distant metastasis
To evaluate the prognostic power of the 18-GC, we compared the status predicted by the 18-GC and the actual distant metastasis status (Table 4). Even though the calculated PPV at 14.6 is relatively low, the calculated NPV is relatively high at 98.8%, indicating that the 18-GC is relatively accurate in identifying patients that would not have distant metastasis in the end of clinical monitoring.
Table 4: Concordance between predictions by the 18-GC and the actual clinical distant metastasis outcomes
Distant metastasis |
Total |
|||
|---|---|---|---|---|
Risk group |
No |
Yes |
||
18-gene classifier |
Low (<21) |
79 (98.8%) |
1 (1.2%) |
80 (35.7%) |
High (>=21) |
123 (85.4%) |
21 (14.6%) |
144 (64.3%) |
|
Total |
202 (90.2%) |
22 (9.8%) |
224 (100%) |
|
In addition, we calculated the PPV and NPV for patients who was not treated by chemotherapy (n=39). Results showed that the PPV is at 10% and the NPV at 100%, meaning that the 18 GC is well precise for determining the risk if a patient will have distant metastasis, especially for patients classified as low-risk (Table 5).
Table 5: Concordance between predictions by the 18-GC and the actual clinical distant metastasis outcomes for patients without adjuvant chemotherapy
Distant metastasis |
Total |
|||
|---|---|---|---|---|
Risk group |
No |
Yes |
||
18-gene classifier |
Low (<21) |
19 (100%) |
0 (0%) |
19 (48.7%) |
High (>=21) |
18 (90%) |
2 (10%) |
20 (51.3%) |
|
Total |
37 (94.9%) |
2 (5.1%) |
39 (100%) |
|
We then performed a univariate and multivariate Cox proportional-hazards analysis for the factors of age, tumor stage, lymph node status, tumor grade, lymph vessel invasion, and risk classification by the 18-GC (Table 6). In the unadjusted model, we found that age at diagnosis, tumor stage, and classification by the 18-GC are significant prognostic factors of recurrence-free survival by distant metastasis. In the adjusted model, age at diagnosis, tumor stage and the 18-GC remain significantly associated with recurrence. Patients at age 40 or younger upon diagnosis had worse breast cancer-recurrence survival (HR=3.2; 95% CI = 1.4 – 7.4) than those older than 40. Similarly, patients with T2-T3 breast cancer (HR=3.3; 95% CI = 1.1 - 10.2) had worse prognosis than those with T1 breast cancer. A score equal to or higher than 21 by the 18-GC is a significant factor of shorter recurrence-free survival (HR=11.7; 95% CI = 1.5 - 89.9) after adjustment for other clinical and pathological variables.
Table 6: Cox proportional hazard models for recurrence-free survival
Variable |
Crude |
Adjusted |
||
|---|---|---|---|---|
HR (95% CI) |
P-value |
HR (95% CI) |
P-value |
|
Age |
||||
≦40 (n=48) |
3.2 (1.4 – 7.4) |
0.007 |
2.7 (1.1 – 6.3) |
0.025 |
>40 (n=176) |
1 (Reference) |
1 (Reference) |
||
T stage |
||||
T1 (n=110) |
1 (Reference) |
0.009 |
1 (Reference) |
0.039 |
T2-T3 (n=114) |
4.3 (1.4 – 12.6) |
3.3 (1.1 – 10.2) |
||
N stage |
||||
N0 (n=130) |
1 (Reference) |
0.439 |
1 (Reference) |
0.294 |
N1 (n=94) |
1.4 (0.6 – 3.2) |
0.6 (0.2 – 1.5) |
||
Tumor grade |
||||
1 (n=59) |
1 (Reference) |
0.396 |
1 (Reference) |
0.614 |
2 (n=106) |
2.2 (0.6 – 8.0) |
1.4 (0.4 – 5.0) |
||
3 (n=59) |
2.4 (0.6 – 9.4) |
0.8 (0.2 – 3.4) |
||
Lymph vessel invasion |
||||
Nil/ Minimal (n=182) |
1 (Reference) |
0.232 |
1 (Reference) |
0.298 |
Prominent (n=42) |
1.8 (0.7 – 4.5) |
1.7 (0.6 – 4.8) |
||
18-gene classifier |
||||
<21 (n=80) |
1 (Reference) |
0.015 |
1 (Reference) |
0.018 |
≧21 (n=144) |
12.1 (1.6 – 89.9) |
11.7 (1.5 – 89.9) |
||
Abbreviations: HR, hazard ratio; CI, confidence interval.
A forest plot of the HRs obtained from exploratory subgroup analyses for recurrence-free survival is shown in Figure 2. The results indicate that prominent lymph vessel invasions, tumor grade, and lymph node stage were no longer significant prognostic factors in some subgroups. A possible reason for this observation is that N1 patient usually would receive more adjuvant chemotherapy and radiotherapy than N0 patients. In contrast, the 18-gene panel classification, T2-T3 stage, and age at 40 or younger remain significant factors and might be important confounders for recurrence prognosis in breast cancer.
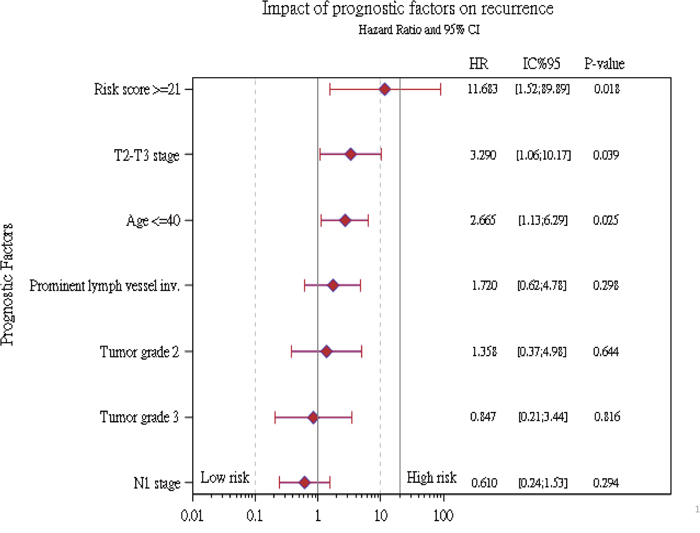
Figure 2: Summary of recurrence hazard ratios for different risk factors in subgroup analyses.
Recurrence-free survival rates were recorded at the 3-, 6-, and 9-year time points (Figure 3). It was found that patients classified as having low risk of recurrence by the 18-GC had high survival rates (0.960) even after nine years (Figure 3). In contrast, those classified as having high risk, the survival rates declined constantly over time and reached 0.772 at year nine (Figure 3). The survival probability of the low- and high-risk patients is plotted against the disease-free survival time.
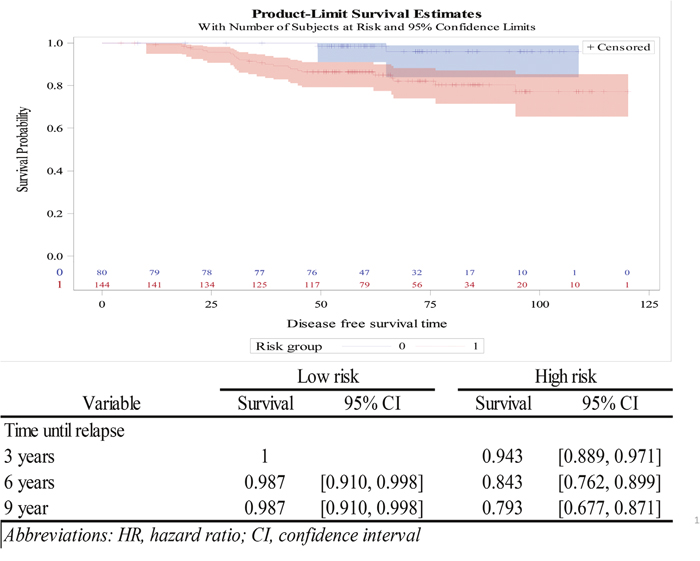
Figure 3: Survival plot analysis of low- and high-risk patients as determined by the 18-gene classifier.
DISCUSSION
Early-stage patients with the luminal-like subtypes of breast cancer are commonly overtreated with adjuvant chemotherapy [14]. To overcome this issue, genomic assays, including ODx, have been utilized to predict the recurrence risk of early-stage breast cancer. They can help patients to avoid the potential adverse toxicity from chemotherapy when the recurrence risk and treatment benefit is low [24]. On the other hand, such prognostic genomic assays should not miss to identify patients that are at high risk of recurrence and could benefit from chemotherapy [25].
Nevertheless, there may be differences in gene expression profiles among different ethnic groups. For example, ODx has been shown to overestimate the risk of recurrence in Asian patients [26]. Even in the intermediate-risk patients, the 10-year distant metastatic rate is 0%. Furthermore, Japanese women have been reported to have better survival according to the SEER data between 1973-1994 [27]. A recent study has also shown that the breast cancer-specific deaths in stage-I patients at year 7 in white, black, and Asian women are significantly different (hazard ratios of 1, 1.57 and 0.60, respectively) [18]. It may thus be important to have a prognostic kit specifically for Asian breast cancer patients.
To that end, based upon an Asian breast cancer patient cohort, we have developed a kit for expression profiling of a panel of 18 genes with an aim to predict distant recurrence risk in luminal-like breast cancer patients. To test the prognostic power of this 18-GC, we included 224 patients and analyzed the association of its prediction with the actual metastasis outcome retrospectively. It was found that this 18-GC is a significant independent prognostic factor of distant metastasis (Table 5; Figure 2). Further, at the breakpoint score of 21, this 18-gene panel classified 164 patients into the same risk group as did the 21-gene ODx panel, giving a concordance rate of 73%. Despite the relatively high concordance rate between the 18-GC and the ODx assay, it is noteworthy that 50 out 80 patients classified as having low risk by the 18-GC were classified as having high risk by ODx. These patients would be recommended to receive adjuvant chemotherapy if based on the risk scores assigned by ODx. However, from the retrospective clinical outcomes, it was found that patients assigned as having low risk of distance recurrence by the 18-GC had a high (98.8%) distant metastasis-free rate (Table 4) and a high probability of a long recurrence-free survival (Table 6 and Figure 3), which contrasts with the phenomenon observed among luminal-like (both the luminal-A and -B subtypes) breast cancer patients whose survival decreases constantly over time even though they had been classified in the low-risk group by ODx [28]. Even though a prospective study with a larger patient cohort is warranted, the new panel that we developed may serve as a good tool for breast cancer patients, especially those with an Asian ethnicity, to make a personalized and informed decision on whether chemotherapy should be performed.
MATERIALS AND METHODS
Patient selection
Retrospectively we included in this study a total of 224 luminal-like (HR+/HER2- ) and T1-3N0-1 breast cancer patients treated at Koo Foundation Sun Yat-Sen Cancer Center (KFSYSCC) in Taipei, Taiwan between 2005 and 2012, for evaluation of the 18-GC developed in our institute [21]. The institutional review board of KFSYSCC reviewed and approved the protocol and informed consent documents for the study. Eligible patients had invasive breast cancer; surgery as first treatment (mastectomy or breast-conserving surgery); a positive test result for estrogen or progesterone receptors (HR+); a negative test result for HER2 (HER2-); a few positive lymph nodes between 0 and 3. Patients with an N2, N3 or M1 stage and treated with pre-operative chemotherapy were excluded.
The 18-gene classifier
Development of the 18-GC has been reported previously [21]. Briefly, it was developed based on 135 breast cancer patients, including 112 patients treated in KFSYSCC, who developed no and 23 patients who developed LRR. Including a total of 18 recurrence-related genes, the 18-GC is a multifunctional gene panel that is associated with cell cycle and proliferation (DDX39, BUB1B, STIL, TPX2, CCNB1), oncogenic process (BLM, TCF3, PIM1, RCHY1, PTI1), inflammation and immune response (CCR1, NFATC2IP), cell-cell interaction (TRPV6, OBSL1, MMP15), apoptosis (C16ORF7, DTX2) and metabolism (ENSA) [21]. For risk classification, each gene was assigned a weight according to the Cox proportional hazards model to assemble the 18-gene scoring algorithm. With a range of risk scores between zero and 56, the breakpoint value of 21 was used to separate the low- from the high-risk category of distant recurrence. The algorithm of 18-GC is shown as below: [21]
18-gene score = 4 × TRPV6 + 3 × DDX39 + 8 × BUB1B + CCR1 + STIL + 3 × BLM + 11 × C16ORF7 + 4 × PIM1 + TPX2 + 2 × PTI1 + 2 × TCF3 + CCNB1 + DTX2 + 2 × ENSA + 5 × RCHY1 + 4 × NFATC2IP + OBSL1 + 2 × MMP15
Unlike ODx, we did not include an intermediate risk group because it is usually binary in clinic decision-making. By adopting the same statistical predictive model used by Paik et al [22], the raw recurrence score (Xi) is first calculated by using the following expression:
Xi = 0.47 × GRB7 group score - 0.34 × ER group score + 1.04 × proliferation group score + 0.10 × invasion group score + 0.05 × CD68 - 0.08 × GSTM1 - 0.07 × BAG.
The final recurrence score (Yi) was then calculated by transforming Xi using the following expression:
Prognostic factors
Along with the demographic and clinical variables previously identified with a prognostic value for distant recurrence (such as age at the diagnosis: ≦40 vs. >40 years old; tumor stage: T1 vs. T2-T3; lymph nodes: N0 vs. N1; tumor grade: grade 1 vs. grade 2 vs. grade 3; prominent vs. nil/focal lymph vessel invasion), the classification by our 18-GC (low vs. high risk) was included for analysis.
Statistical analysis
The demographic, clinical, and survival information were collected and analyzed among patients with and without metastasis. Crude and adjusted Cox analyses were used to compare patients in the low- and high-risk groups assigned by the 18-GC. ROC curve analyses were then performed to identify the optimal breakpoint [23]. We then evaluated sensitivity, specificity, accuracy, negative predictive value (NPV), positive predictive value (PPV), and the area under the curve (AUC) to determine how well the new 18-GC prediction model performs as compared with the ODx assay. All the statistical analyses (p < 0.05) were performed using SAS Software, version 9.4.
CONCLUSION
We have created an 18-GC for predicting the risk of distant recurrence in luminal-like breast cancer patients. Even though a study with a larger patient cohort conducted in a prospective way is warranted, the new 18-GC panel assay has the potential to become a good prognosis predictor for breast cancer patients, especially those of an Asian descent, to determine whether a given patient needs adjuvant chemotherapy.
Author contributions
Conceptualization: Skye Hung-Chun Cheng.
Data curation: Skye Hung-Chun Cheng, Tzu-Ting Huang, Yu-Hao Cheng, Chen-Fang Horng, Nicolas Pennarun.
Formal analysis: Skye Hung-Chun Cheng, Tzu-Ting Huang, Chen-Fang Horng, Nicolas Pennarun.
Funding acquisition: Skye Hung-Chun Cheng.
Investigation: Skye Hung-Chun Cheng, Tzu-Ting Huang, Yu-Hao Cheng, Nicolas Pennarun.
Methodology: Skye Hung-Chun Cheng, Tzu-Ting Huang, Chen-Fang Horng.
Project administration: Skye Hung-Chun Cheng, Chen-Fang Horng.
Resources: Skye Hung-Chun Cheng.
Software: Tzu-Ting Huang, Chen-Fang Horng.
Supervision: Skye Hung-Chun Cheng.
Validation: Tzu-Ting Huang.
Visualization: Yu-Hao Cheng.
Writing - original draft: Yu-Hao Cheng.
Writing - review & editing: Skye Hung-Chun Cheng, Jason Lei.
ACKNOWLEDGMENTS
The authors thank the multidisciplinary breast cancer team at the Koo Foundation Sun Yat-Sen Cancer Center (KF-SYSCC): (1) Patient care and sample collections: Drs. Chi-Min Chen, Ben-Long Yu, Tsung-Yen Cheng, and Tzu-Jung Tsai (Department of Surgery); Drs. Wan-Chen Tsai and Christopher KJ Lin (Department of Radiology); Drs. James J. Jian and Yu-Chen Tsai (Department of Radiation Oncology); (2) Data management and quality control: Yen-Chun Lin; (3) Tumor Bank: Theresa M. Tsai; and (4) Microarray Laboratory: Lih-Chian Wu and Dr. Kuo Jan Kao.
CONFLICTS OF INTEREST
The author (SHC) owns a patent relating to the content of this manuscript (Taiwan patent number: 104115832). None of the authors has any conflicts of interests in this research, either financial or non-financial.
FUNDING
This research was supported by the research fund of the Ministry of Science and Technology (NSC 102-3114-Y-368-001) of the Republic of China (Taiwan).
REFERENCES
1. Leong SP, Shen ZZ, Liu TJ, Agarwal G, Tajima T, Paik NS, Sandelin K, Derossis A, Cody H, Foulkes WD. Is breast cancer the same disease in Asian and Western countries? World J Surg. 2010; 34:2308-24.
2. Kim Y, Yoo KY, Goodman MT. Differences in incidence, mortality and survival of breast cancer by regions and countries in Asia and contributing factors. Asian Pac J Cancer Prev. 2015; 16:2857-70.
3. Binns C, Low WY, Lee MK. Breast cancer: an increasing public health problem in the Asia Pacific region. Asia Pac J Public Health. 2013; 25:364-7.
4. Chiang CJ, Lo WC, Yang YW, You SL, Chen CJ, Lai MS. Incidence and survival of adult cancer patients in Taiwan, 2002-2012. J Formos Med Assoc. 2016; 115:1076-88.
5. Shen YC, Chang CJ, Hsu C, Cheng CC, Chiu CF, Cheng AL. Significant difference in the trends of female breast cancer incidence between Taiwanese and Caucasian Americans: implications from age-period-cohort analysis. Cancer Epidemiol Biomarkers Prev. 2005; 14:1986-90.
6. Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature. 2012; 490:61-70.
7. Parise CA, Bauer KR, Brown MM, Caggiano V. Breast cancer subtypes as defined by the estrogen receptor (ER), progesterone receptor (PR), and the human epidermal growth factor receptor 2 (HER2) among women with invasive breast cancer in California, 1999-2004. Breast J. 2009; 15:593-602.
8. Telli ML, Chang ET, Kurian AW, Keegan TH, McClure LA, Lichtensztajn D, Ford JM, Gomez SL. Asian ethnicity and breast cancer subtypes: a study from the California Cancer Registry. Breast Cancer Res Treat. 2011; 127:471-8.
9. Lin CH, Liau JY, Lu YS, Huang CS, Lee WC, Kuo KT, Shen YC, Kuo SH, Lan C, Liu JM, Kuo WH, Chang KJ, Cheng AL. Molecular subtypes of breast cancer emerging in young women in Taiwan: evidence for more than just westernization as a reason for the disease in Asia. Cancer Epidemiol Biomarkers Prev. 2009; 18:1807-14.
10. Arvold ND, Taghian AG, Niemierko A, Abi Raad RF, Sreedhara M, Nguyen PL, Bellon JR, Wong JS, Smith BL, Harris JR. Age, breast cancer subtype approximation, and local recurrence after breast-conserving therapy. J Clin Oncol. 2011; 29:3885-91.
11. Sorlie T, Perou CM, Tibshirani R, Aas T, Geisler S, Johnsen H, Hastie T, Eisen MB, van de Rijn M, Jeffrey SS, Thorsen T, Quist H, Matese JC, et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc Natl Acad Sci U S A. 2001; 98:10869-74.
12. Weigelt B, Baehner FL, Reis-Filho JS. The contribution of gene expression profiling to breast cancer classification, prognostication and prediction: a retrospective of the last decade. J Pathol. 2010; 220:263-80.
13. Cheng SH, Yu BL, Horng CF, Tsai SY, Chen CM, Chu NM, Liu MC, Huang AT. In-Depth Evaluation of the AJCC 2010 Staging System for Luminal-like Breast Cancer --- An Analysis from a Free-Standing Cancer Hospital. Journal Cancer Research and Practice. 2013; 30:21-34.
14. Cheng SH, Yu BL, Horng CF, Tsai SY, Chen CM, Chu NM, Tsou MH, Lin CK, Shih LS, Liu MC. Long-term survival and stage I breast cancer subtypes. Journal of Cancer Research and Practice. 2016; 3:1-8.
15. Paik S, Tang G, Shak S, Kim C, Baker J, Kim W, Cronin M, Baehner FL, Watson D, Bryant J, Costantino JP, Geyer CE Jr, Wickerham DL, et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J Clin Oncol. 2006; 24:3726-34.
16. Cardoso F, van't Veer LJ, Bogaerts J, Slaets L, Viale G, Delaloge S, Pierga JY, Brain E, Causeret S, DeLorenzi M, Glas AM, Golfinopoulos V, Goulioti T, et al. 70-Gene Signature as an Aid to Treatment Decisions in Early-Stage Breast Cancer. N Engl J Med. 2016; 375:717-29.
17. Varga Z, Sinn P, Fritzsche F, von Hochstetter A, Noske A, Schraml P, Tausch C, Trojan A, Moch H. Comparison of EndoPredict and Oncotype DX test results in hormone receptor positive invasive breast cancer. PLoS One. 2013; 8:e58483.
18. Iqbal J, Ginsburg O, Rochon PA, Sun P, Narod SA. Differences in breast cancer stage at diagnosis and cancer-specific survival by race and ethnicity in the United States. JAMA. 2015; 313:165-73.
19. Chlebowski RT, Chen Z, Anderson GL, Rohan T, Aragaki A, Lane D, Dolan NC, Paskett ED, McTiernan A, Hubbell FA, Adams-Campbell LL, Prentice R. Ethnicity and breast cancer: factors influencing differences in incidence and outcome. J Natl Cancer Inst. 2005; 97:439-48.
20. Hausauer AK, Keegan TH, Chang ET, Clarke CA. Recent breast cancer trends among Asian/Pacific Islander, Hispanic, and African-American women in the US: changes by tumor subtype. Breast Cancer Res. 2007; 9:R90.
21. Cheng SH, Horng CF, Huang TT, Huang ES, Tsou MH, Shi LS, Yu BL, Chen CM, Huang AT. An Eighteen-Gene Classifier Predicts Locoregional Recurrence in Post-Mastectomy Breast Cancer Patients. EBioMedicine. 2016; 5:74-81.
22. Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, Baehner FL, Walker MG, Watson D, Park T, Hiller W, Fisher ER, Wickerham DL, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004; 351:2817-26.
23. Perkins NJ, Schisterman EF. The inconsistency of “optimal” cutpoints obtained using two criteria based on the receiver operating characteristic curve. Am J Epidemiol. 2006; 163:670-5.
24. Pittman J, Huang E, Dressman H, Horng CF, Cheng SH, Tsou MH, Chen CM, Bild A, Iversen ES, Huang AT, Nevins JR, West M. Integrated modeling of clinical and gene expression information for personalized prediction of disease outcomes. Proc Natl Acad Sci U S A. 2004; 101:8431-6.
25. Chen C, Dhanda R, Tseng WY, Forsyth M, Patt DA. Evaluating use characteristics for the oncotype dx 21-gene recurrence score and concordance with chemotherapy use in early-stage breast cancer. J Oncol Pract. 2013; 9:182-7.
26. Toi M, Iwata H, Yamanaka T, Masuda N, Ohno S, Nakamura S, Nakayama T, Kashiwaba M, Kamigaki S, Kuroi K; Japan Breast Cancer Research Group-Translational Research Group. Clinical significance of the 21-gene signature (Oncotype DX) in hormone receptor-positive early stage primary breast cancer in the Japanese population. Cancer. 2010; 116:3112-8.
27. Pineda MD, White E, Kristal AR, Taylor V. Asian breast cancer survival in the US: a comparison between Asian immigrants, US-born Asian Americans and Caucasians. Int J Epidemiol. 2001; 30:976-82.
28. Blows FM, Driver KE, Schmidt MK, Broeks A, van Leeuwen FE, Wesseling J, Cheang MC, Gelmon K, Nielsen TO, Blomqvist C, Heikkilä P, Heikkinen T, Nevanlinna H, et al. Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med. 2010; 7:e1000279.

