INTRODUCTION
Paclitaxel is a tubulin-targeting agents [1] commonly used in currently anti-cancer therapies. In the updated National Comprehensive Cancer Network (NCCN) guidelines (www.nccn.org/patients), paclitaxel-based adjuvant chemotherapy is still recommended as a first-line regiment for treating patients with triple-negative breast cancer (TNBC). Paclitaxel targets microtubules to interfere with the mitotic spindle, resulting in cell cycle arrest and ultimately apoptosis [2]. Although paclitaxel eliminates most tumor cells, the mechanisms causing resistance in residual cancer cells are unclear. Drug resistance is a crucial problem in cancer therapy that impacts mortality rates [3]. Recently, the combination of paclitaxel with carboplatin was shown to significantly increase the proportion of 315 TNBC patients achieving a pathological complete response (pCR): this therapy produced a pCR rate of 59% compared with 38% pCR in patients who did not receive carboplatin [4]. Moreover, a randomized phase II trial that enrolled 443 patients with stage II to III TNBC receiving standard paclitaxel therapy with or without carboplatin found that the combination resulted in a significantly higher pCR rate (54% vs. 41%) [5]. These findings indicate that the majority of TNBC patients are likely insensitive to paclitaxel treatment. Since patients suffered from chemotherapy, identification of useful biomarkers for predicting pCR in TNBC patients receiving paclitaxel-based chemotherapies is urgently needed.
Recent studies have demonstrated that an aberrant expression of β-tubulin subtype TUBB3 due to amino acid alteration at the paclitaxel-binding site predicts a poor response to paclitaxel-based chemotherapy and unfavorable prognosis in patients with metastatic gastric cancer [6], ovarian cancer [7] and non-small cell lung cancer [8, 9]. Although TUBB3 gene expression appeared to be an independent factor for breast cancer patients with luminal A/B subtypes, a Cox regression analysis of recurrence-free survival probability revealed that TUBB3 was not a useful prognostic factor for patients with HER2-enriched and basal-like (>75% triple-negative) breast cancers [10]. This result indicates that TUBB3 expression may be not a good biomarker for predicting paclitaxel response in these HER2-positive and triple-negative breast cancer patients. Even though alteration of β-tubulin subtypes may affect the anti-cancer effectiveness of paclitaxel, the discovery of new biomarkers other than TUBB3 to predict clinical outcomes in patients with HER2-positive and triple-negative breast cancers prior to receiving paclitaxel-based chemotherapy is likely valuable.
The 16 human Ovarian Tumour (OTU) family deubiquitinases (DUBs) are key regulators of the ubiquitin (Ub) code. Small OTU DUBs of the OTUD and OTUB subfamilies employ distinct mechanisms to achieve linkage specificity but the physiological roles of most members are unclear [11–13]. Accumulated evidence indicated that OTUD7B regulates inflammation and NF-κB signaling, T-cell activation and epidermal growth factor receptor (EGFR) trafficking [14–16]. IHere we find that OTUD7B were predominantly upregulated in paclitaxel-resistant MDA-MB436 TNBC cells after paclitaxel treatment and highly correlated with IC50 concentrations of paclitaxel in tested TNBC cell lines. Conversely, OTUD7B was dramatically downregulated in paclitaxel-sensitive HCC38 TNBC cells following treatment with paclitaxel. Intriguingly, compared to other OTUDs, OTUD7B transcript in tumors was significantly higher than that of normal tissues derived from patients with breast cancer. Moreover, OTUD7B upregulation was appeared to predict an increased risk for cancer recurrence/metastasis in breast cancer patients who may exhibit a poor pathologic complete response to paclitaxel chemotherapy. Finally, the in silico analysis showed that the loss of miR-1180 and the inhibition of inflammation-related pathways may be associated with OTUD7B-related mechanisms underlying paclitaxel resistance in TNBCs.
RESULTS
OTUD7B is predominantly upregulated following paclitaxel treatment in TNBC cells
To identify the molecules that are responsible for paclitaxel resistance in TNBCs, we analyzed transcriptional profiles of paclitaxel-sensitive HCC38 and paclitaxel-resistant MDA-MB436 with or without paclitaxel treatment for 24 hours at their respective 10-fold IC50 concentrations determined previously [17]. Then, we dissected 158 consensus genes with 1.5-fold changes after paclitaxel treatment in HCC38 and MDA-MB436 cells (Figure 1A, Supplementary Table 1). Our data showed that OTUD7B was upregulated but DSTNP2, destrin, actin depolymerizing factor pseudogene 2, and VCAN, a member of the aggrecan/versican proteoglycan family, were downregulated in MDA-MB436 cells (Figure 1B). An inversed effect on mRNA expression was detected in HCC38 cells (Figure 1B). However, mRNA levels from three independent experiments indicated that the transcriptional activity of OTUD7B was altered more dramatically upon paclitaxel treatment in both HCC38 and MDA-MB436 cells (Figure 1C) compared to DSTNP2 and VCAN.
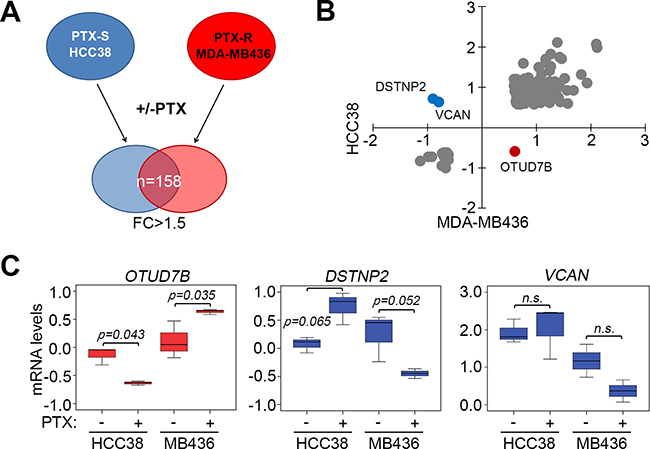
Figure 1: OTUD7B upregulation and DSTNP/VCAN downregulation predict a poor response to paclitaxel treatment in TNBC cells. (A) Flowchart of the procedure for identifying consensus genes with 1.5-fold change (FC) post-treatment with paclitaxel (PTX) at a concentration of 10 × IC50 for 24 hours in PTX-sensitive (PTX-S) HCC38 cells and PTX-resistant (PTX-R) MDA-MB436 cells. (B) Dotplot of mRNA levels (log2) of 158 consensus genes identified using the strategy shown in A. (C) mRNA levels of OTUD7B, DSTNP2 and VCAN in HCC38 and MDA-MB436 cells following treatment without or with paclitaxel at 10 × IC50 for 24 hours. Data from three independent experiments are shown as medians ± SD. Statistical differences were analyzed using one-way ANOVA with Tukey’s test.
Furthermore, we analyzed correlation between the IC50 concentrations of paclitaxel and the mRNA levels of OTUD7B, DSTNP2 and VCAN in a panel of TNBC cell lines BT-20, BT-549, HCC-1143, HCC-1806, HCC-1937, HCC-1954, HCC-30, HCC-70, Hs578T, MDA-MB-231, MDA-MB-436 and MDA-MB-468 after treatment with or without paclitaxel at the respective 10-fold IC50 concentrations. The data showed that correlations between the IC50 concentrations of paclitaxel and OTUD7B mRNA levels were marginal in the paclitaxel-untreated cells and significantly after treatment with paclitaxel in the tested TNBC cells (Figure 2A). Compared to untreated control cells, the correlation of OTUD7B mRNA levels with IC50 concentrations of paclitaxel appeared to be more significant in the tested TNBC cells (Figure 2A). In contrast, in both paclitaxel-treated and -untreated cells, IC50 concentrations of paclitaxel were inversely correlated with DSTNP2 (except in control cells) and VCAN mRNA levels (Figure 2B and 2C). In comparison with control cells, the mRNA levels of DSTNP2 and VCAN were determined respectively to significantly and marginally correlate with the IC50 concentrations of paclitaxel in the detected TNBC cells (Figure 2B and 2C). To validate these findings, we performed reverse transcription PCR (RT-PCR) experiment in order to determining the OTUD7B mRNA levels in a panel of TNBC cells (Figure 2D). Our data showed that the levels of OTUD7B transcript causally associated with the PTX IC50 concentrations in the detected TNBC cells (Figure 2E). Based on these findings, we subsequently focused on estimating the prognostic significance of OTUD7B in predicting the effectiveness of paclitaxel on clinical patients with breast cancer.
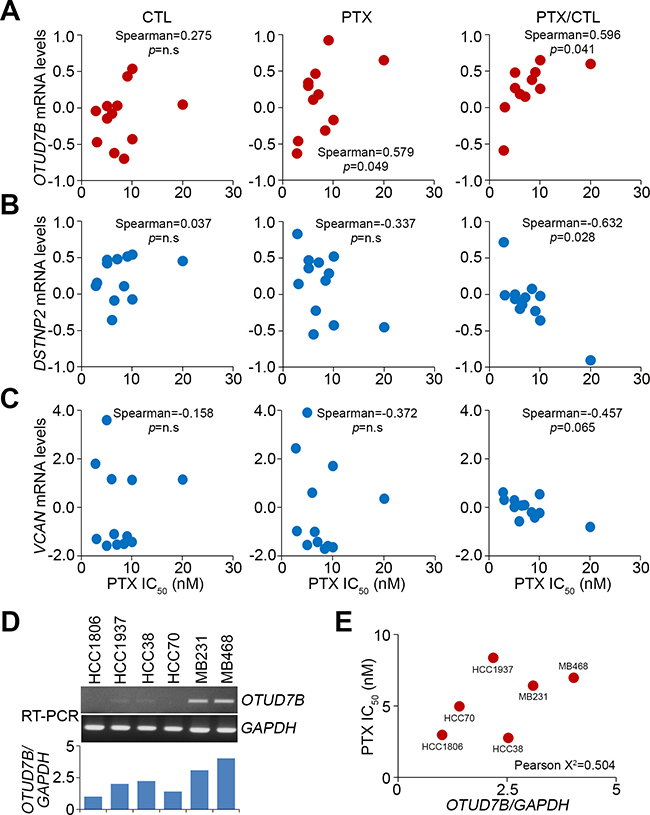
Figure 2: OTUD7B and DSTNP/VCAN positively and negatively correlate, respectively, with PTX IC50 concentrations in a panel of TNBC cell lines. (A–C) Correlations between OTUD7B, DSTNP and VCAN mRNA level and PTX IC50 concentration in tested TNBC cell lines. Correlations were assessed using Spearman’s test. The symbol “n.s” denotes non-significant results. Each dot in the dotplot indicates the median of mRNA levels from three independent experiments. (D) RT-PCR analysis for OTUD7B and GAPDH transcripts in TNBC cell lines (top). The levels of OTUD7B transcript were normalized by comparing with the respective GAPDH level in the tested cell lines and shown as ratios (bottom). (E) Correlates between the normalized OTUD7B levels and PTX IC50 concentrations in a panel of TNBC cell lines. Pearson’s correlation test was used to evaluate the statistical significance.
OTUD7B upregulation correlates with tumorigenesis and poor recurrence-free survival rates in clinical patients with TNBCs
To examine the association of OTUD7B expression with breast cancer development, we next analyzed the transcriptional profile of OTUD7B, as well as other OTUD subtypes, in TCGA breast invasive carcinoma (BRCA). The data showed that primary tumors expressed OTUD7B more abundantly than normal mammary epithelial tissues derived from patients with BRCA (Figure 3A). Moreover, among OTUD subtypes, the level of OTUD7B mRNA was detected to be much abundant in primary tumors compared to normal tissues derived from patients with breast cancer (Figure 3B). Accordingly, our data showed that the mRNA levels of OTUD7B in primary tumors are significantly higher than in normal adjacent tissues derived from BRCA patients (Figure 3C). Similar findings were also observed in paired normal adjacent tissues and primary tumors derived from patients with basal-like breast cancers, approximately 75% of which were TNBCs (Supplementary Figure 1).
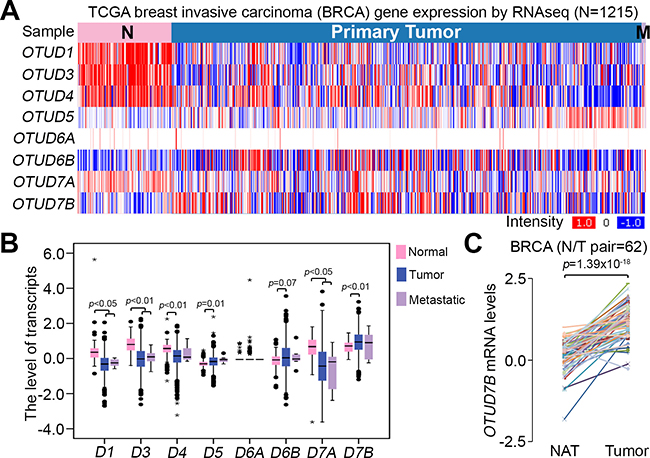
Figure 3: OTUD7B expression is upregulated in primary tumors compared to normal tissues derived from patients with breast cancer. (A) Heatmap for the transcription profile of OTUD subtypes in normal tissues, primary tumors and metastatic tumors derived from patients with breast invasive carcinoma (BRCA) using the TCGA database. (B) Boxplot of mRNA levels of OTUD subtypes in defined normal tissues (n = 113), primary tumors (n = 1061) and metastatic tumors (n = 7) from A. Data were analyzed by One-way ANOVA using Turkey’s test. (C) OTUD7B expression in paired normal adjacent tissues (NAT) and primary tumors derived from patients with BRCA. Data were analyzed using paired t-tests.
We next examined the in vivo tumor growth of TNBC cell lines HCC1806, HCC1937, HCC38, HCC70, MDA-MB231 and MDA-MB468 through the subcutaneous implantation into NOSCID mice for 4 weeks (Figure 4A). The data showed that tumor volumes of the tested TNBC cells post-implantation for 4 weeks are negatively correlated with OTUD7B mRNA levels (Figure 4B). Moreover, we detected the in vitro invasion ability of several TNBC cell lines HCC1806, HCC1937, HCC38 and MDA-MB231 by using trans-well cultivation (Figure 4C). Our results revealed that OTUD7B mRNA levels causally correlated with invasive ability of the tested TNBC cell lines (Figure 4D).
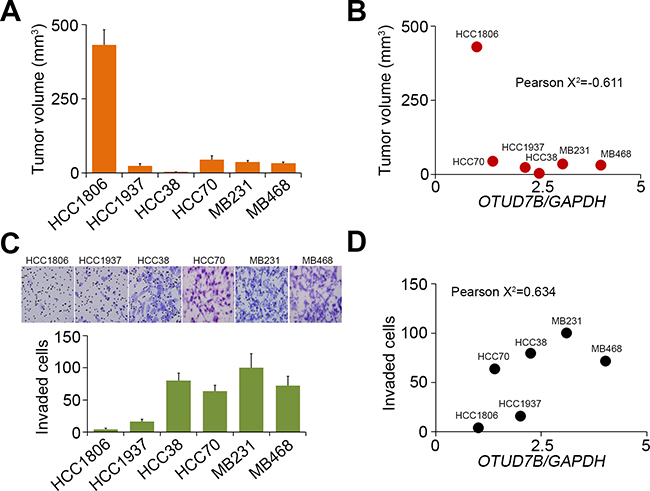
Figure 4: The correlations of OTUD7B transcriptional levels with the tumor growth, invasion and lung metastatic colonization abilities of TNBC cell lines. (A, B) Mice (n = 5) were subcutaneously implanted with breast cancer cell lines as indicated for 4 weeks. Tumor volume was measured from tumor-bearing mice and presented as mean ± SD (A). The correlation between OTUD7B transcriptional levels and the mean of tumor volume in each group was shown as dotplot (B). (C, D) The invaded cells in the invasion assay were stained with Giemsa regent (C, top). The data from triplicate experiments were presented as mean ± SD (C, bottom). The correlation between the normalized OTUD7B levels and invaded cell numbers of the tested cell lines was shown by dotplot (D). In (B, D), Pearson’s correlation test was used to evaluate the statistical significance.
Because chemotherapeutic efficacy is frequently assessed by recurrence-free survival (RFS) probability, we further estimated the prognostic significance of OTUD7B in predicting RFS rates in breast cancer patients. Using the SurvExpress database, we found that high-levels of OTUD7B mRNA expression significantly correlated with poor RFS rates, with a hazard ratio (HR) of 1.4 in 1901 breast cancer patients (Figure 5A). OTUD7B mRNA levels in the high-risk cohort were significantly upregulated compared to the low-risk cohort among enrolled breast cancer patients (Figure 5B). Similar results were also observed in the another cohort with 1574 breast cancer patients in SurvExpress database (Supplementary Figure 2). To investigate the predictive relevance among patients with different breast cancer subtypes, we performed further Kaplan-Meier analysis to compare RFS probability based on OTUD7B expression for patients represented in Figure 5A with luminal A/B, HER2-positive and basal-like breast cancers (Figure 5C–5F). Our data showed that OTUD7B upregulation was significantly associates with a poor RFS rate in patients with HER2-positive and basal-like breast cancers (Figure 5E and 5F); the worst HR was observed in patients with basal-like breast cancer compared to other subtypes (Figure 5F). These findings might implicate that OTUD7B severs as a useful biomarker to predict poor prognosis in patients with estrogen receptor-negative breast cancers.
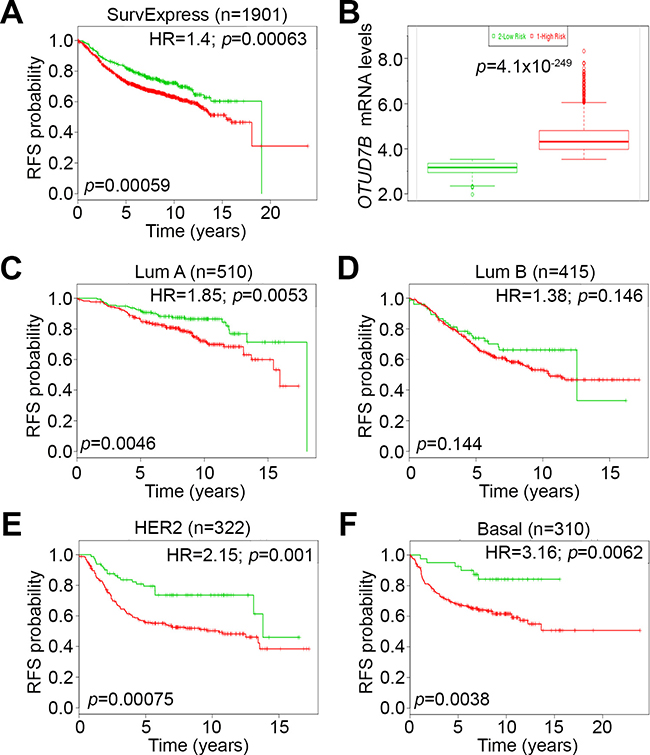
Figure 5: OTUD7B upregulation is associated with poor recurrence-free survival rates in breast cancer patients. (A) Kaplan-Meier analysis of recurrence-free survival (RFS) probability according to OTUD7B expression in breast cancer patients, performed using the SurvExpress database. HR denotes hazard ratio. (B) Boxplot of OTUD7B mRNA levels in low (green) and high (red)-risk cohorts in A. (C–F) Kaplan-Meier analysis of RFS probability according to OTUD7B expression in breast cancer patients with luminal A (Lum A), luminal B (Lum B), HER2 and basal-like (Basal) subtypes.
OTUD7B upregulation predicts a poor response to paclitaxel-based chemotherapy in breast cancer patients
We next evaluated the prognostic significance of OTUD7B in breast cancer patients who received paclitaxel-based chemotherapy. Using the K-M Plotter database, we found that OTUD7B upregulation reflected an unfavorable prognosis in terms of RFS probability in patients receiving pre-operative paclitaxel chemotherapy with unclassified (Figure 6A) and basal-like (Figure 6B) breast cancer. On the other hand, OTUD7B upregulation more significantly (p < 0.05) predicted a poor RFS probability in patients receiving post-operative paclitaxel chemotherapy with unclassified (Figure 6C) and basal-like (Figure 6D) breast cancer. Similar views were also observed in K-M analysis under the condition of distant metastasis-free survival probability using K-M Plotter database against patients receiving post-operative paclitaxel chemotherapy with unclassified (Supplementary Figure 3A) and basal-like (Supplementary Figure 3B) breast cancer. In comparison, we analyzed the transcriptional profile of OTUD7B in breast tumors derived from breast cancer patients receiving pre-operative paclitaxel-based chemotherapy. The data revealed that OTUD7B mRNA levels in breast tumors derived from patients with non-pathological complete response (pCR) were significantly higher than in patients with (Figure 6E). Similarly, among patients receiving post-operative paclitaxel-based chemotherapy, our results showed that OTUD7B mRNA levels in breast tumors derived from patients with non-pCR were predominantly elevated compared to those of patients with pCR (Figure 6F).
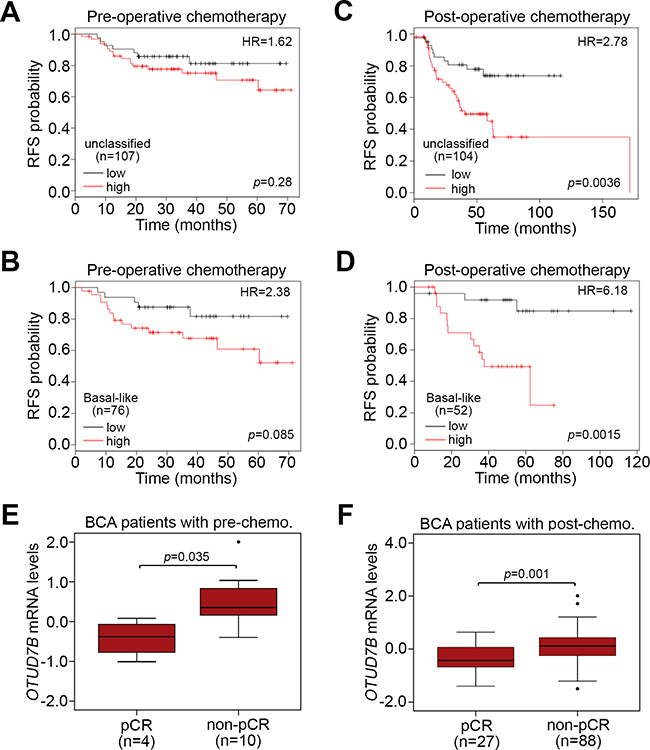
Figure 6: OTUD7B upregulation predicts a poor response to paclitaxel chemotherapy in breast cancer patients. (A, B) Kaplan-Meier analysis of RFS probability according to OTUD7B expression in patients who receive pre-operative paclitaxel-based chemotherapy with unclassified (A) or basal-like (B) breast cancer. (C, D) Kaplan-Meier analysis of RFS probability according to OTUD7B expression in patients who receive post-operative paclitaxel-based chemotherapy with unclassified (C) or basal-like (D) breast cancer. In A-D, Kaplan-Meier analyses were performed using the K-M Plotter database and HR denotes hazard ratio. (E, F) Boxplot of OTUD7B mRNA levels in tumor biopsy derived from BCA patients who received pre-operative paclitaxel-based chemotherapy (pre-chemo.) (E) or pre-operative paclitaxel-based chemotherapy (post-chemo.) (F). pCR denotes pathological complete response. Data were analyzed using a non-parametric Mann-Whitney U-test.
OTUD7B upregulation is associates with inhibition of inflammation-related pathways and activation of Let-7-mediated post-transcriptional regulation in paclitaxel-resistant TNBC cells
To ascertain the possible mechanism for OTUD7B upregulation-related paclitaxel resistance in TNBCs, we next used the TCGA database to analyze the transcriptional profile of OTUD7B and microRNA (miRNA)-1180 which has been shown to trigger post-transcriptional downregulation of OTUD7B [18] in primary tumors derived from patients with TNBC. We found that mRNA expression levels of OTUD7B and miR-1180 were inversely related in tested tumor tissues (Figure 7A). Moreover, we performed an in silico analysis using Ingenuity Pathway Analysis (IPA) software to computationally simulate possible activated or inhibited upstream regulators either in paclitaxel-sensitive HCC38 cells or paclitaxel-resistant MDA-MB436 cells after treatment with paclitaxel at the 10-folds IC50 concentrations for 24 hours. Our data showed that activation of Let-7 microRNA and LY294002, a pharmaceutical inhibitor for phosphatidylinositol 3-kinase (PI3K), was significantly predicted; conversely, inhibition of lipopolysaccharide (LPS) and toll-like receptor adaptor molecule 1 (TICAM1)-related signaling cascades was strongly predicted in MDA-MB436 cells following treatment with paclitaxel (Figure 7B, Supplementary Table 2). The inverse activity of these upstream regulators was computationally simulated in HCC38 cells (Figure 7B). Using the IPA database, we identified several Let-7-targeting genes in HCC38 and MDA-MB436 cells upon paclitaxel stimulation (Supplementary Figure 4A and 4B).
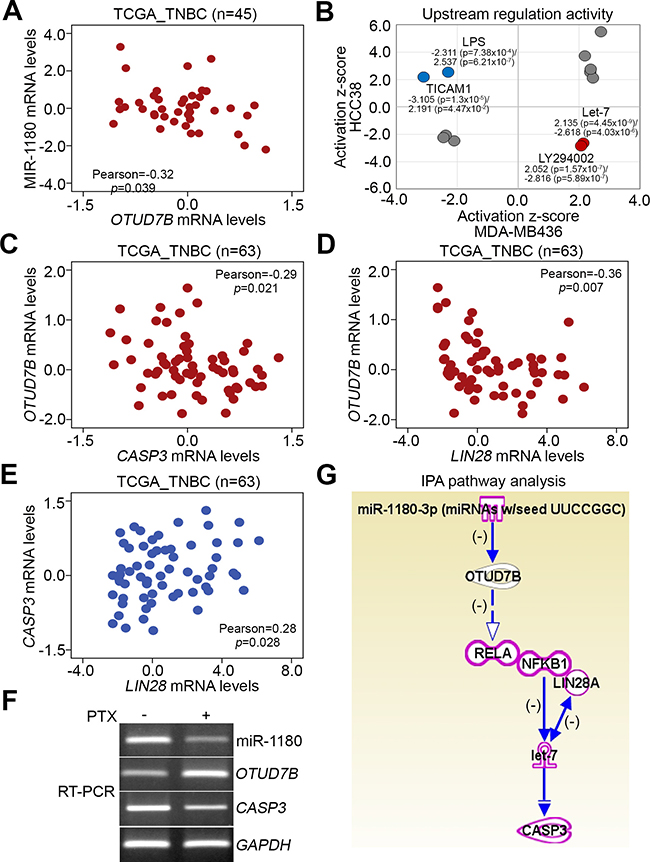
Figure 7: Possible mechanisms underlying paclitaxel resistance in TNBC cells. (A) Correlation between miR-1180 and OTUD7B mRNA levels in primary tumors derived from patients with TNBC using the TCGA database. Pearson’s correlation test was used to estimate the correlation. (B) In silico analysis of consensus upstream regulators that were possibly activated or inhibited after paclitaxel treatment at 10 × IC50 for 24 hours in HCC38 and MDA-MB436 cells, performed using Ingenuity Pathway Analysis software. (C–E) Correlations among OTUD7B, CASP3 and LIN28 mRNA levels in clinical tissues derived from patients with TNBC using TCGA database. In A, C, D and E, Pearson’s correlation test was used to estimate the statistical association. (F) RT-PCR analysis for primary miR-1180, OTUD7B, CASP3 and GAPDH transcripts derived from MDA-MB468 cells treated without or with PTX at 50 nM for 6 hours. (G) The computational simulation of pathway axis consisted with miR-1180, OTUD7B, NF-kB/LIN28, Let-7 and CASP3 by Ingenuity Pathway Analysis (IPA) software.
Since Let-7 microRNA has been shown to inhibit CASP3 expression, thereby promoting chemoresistance [19], and be negatively regulated by the NF-kB-Lin28 axis in breast cancer cells [20], we next analyzed the transcriptional profiles of OTUD7B, CASP3 and LIN28 in TNBC tissues. The data showed that OTUD7B expression was inversely correlated with CASP3 and LIN28 (Figure 7C and 7D), whereas mRNA levels of CASP3 and LIN28 (Figure 7E) were positively correlated in TNBC tissues. Moreover, RT-PCR analysis revealed that the mRNA levels of miR1180 and CASP3 are downregulated but OTUD7B transcript is upregulated upon PTX treatment in MDA-MB468 cells (Figure 7F). By using IPA software, the transcriptional inhibition of miR1180 on OTUD7B and the negative regulation of OTUD7B towards the NF-kB-Lin28 axis-inhibiting let-7 function on the transcriptional repression of CASP3 were computationally simulated (Figure 7G).
DISCUSSION
Resistance is a significant limitation to the effectiveness of cancer therapies. Here, we show that OTUD7B is upregulated in paclitaxel-resistant MDA-MB436 cells but downregulated in paclitaxel-sensitive HCC-38 cells after treatment with paclitaxel at 10-fold IC50 concentrations for 24 hours. Moreover, OTUD7B upregulation significantly predicts an unfavorable risk for cancer recurrence/metastasis and is associated with a poor response to paclitaxel-based chemotherapy in breast cancer patients. The molecular mechanism underlying OTUD7B upregulation-related paclitaxel resistance is likely associated with the downregulation of OTUD7B-targeting miR-1180 and LPS/PI3K-related signaling cascades and as well as the induction of miR-Let-7-mediated post-transcriptional inhibition of multiple genes in TNBC cells.
Previous studies have demonstrated that OTUD7B is a negative regulator for the NF-κB-related pathway through deubiquitination of TNF receptor-associated factor 3 (TRAF3) in mucosal immunity against infections [15]. Here, we find that paclitaxel-resistant MDA-MB436 cells suppress the LPS- and toll-like receptor adaptor molecule 1 (TICAM1)-related signaling pathways, which ultimately converge on the activation of NF-κB [21, 22]. Activation of the NF-kB-related pathway is known to be critical for mediating the cellular response to oxidative stress and inflammation [23–25]. Increased oxidative stress has been shown previously to affect potential paclitaxel cancercidal effectiveness in ovarian cancer [26] . Based on these findings, we conclude that negative regulation of the NF-κB-related pathway by upregulated OTUD7B is a key mechanism underlying paclitaxel resistance in TNBCs. In contrast, the NF-κB-related pathway was known to control cell growth by regulating the cell cycle machinery such as cyclins D1 [27, 28] which plays as a key element in mammary gland development and breast carcinogenesis [29]. Here we show that the OUTD7B upregulation inversely correlates with the in vivo tumor growth in a panel of TNBC cell lines even though the increased levels of OUTD7B transcript are extensively found in primary tumors in comparison with normal tissues derived from patients with breast cancer. Therefore, OTUD7B upregulation may suppress tumor growth via negatively regulating NF-κB-related pathway in TNBCs.
The PI3K/Akt pathway plays important roles in tumorigenesis and drug resistance in human cancers [30, 31]. A previous study has demonstrated that LY294002, a specific inhibitor of the PI3K/Akt pathway, induces resistant cells to become more sensitive to paclitaxel or docetaxel treatment in prostate cancer cells [30]. In contrast, inhibition of the LPS-triggered inflammation response appeared to suppress oxidative stress-mediated activation of the PI3K/Akt-NF-κB signaling axis in macrophages [32]. Interestingly, a recent report demonstrated that blockage of LPS-induced inflammatory responses inhibits the activity of the PI3K/Akt-NF-κB pathway but induces activation of the Nrf2 pathway [33], which is thought to be correlated with mechanism for chemoresistance in many cancer types [34–39], including breast cancer [40, 41]. In our simulation, the pharmaceutical inhibitory effect of LY294002 on PI3K/Akt pathway was activated upon paclitaxel stimulation in paclitaxel-resistant TNBC cells. Again, this finding supports the hypothesis that induction of OTUD7B-mediated inactivation of the NF-κB-related pathway plays a pivotal role in the mechanism for developing paclitaxel resistance in paclitaxel-resistant TNBC cells.
Previously, we have shown that induction of Let-7 microRNA inhibits the expression of caspase-3, a master caspase in mediating cell apoptosis, and thereby confers multidrug resistance in breast cancer cells [19]. Similar findings have also been reported in other cancer types [42]. Here, we show that Let-7 microRNA is computationally predicted to be highly active in paclitaxel-resistant TNBC cells following paclitaxel treatment. Since our previous report showed that caspase-3 downregulation is predominant in TNBCs with chemoresistance against paclitaxel-based chemotherapy [19], we believed that induction of Let-7 microRNA might be critical for protecting paclitaxel-resistant TNBC cells from paclitaxel-caused cell apoptosis by repressing caspase-3 expression. Induction of the inflammatory response mediated by NF-κB appeared to directly activate Lin28 transcription and rapidly reduce Let-7 microRNA levels, thereby promoting breast cancer tumor growth [20]. It has been known that Lin28 encodes RNA-binding protein that binds to Let-7 pre-microRNA and blocks the production of mature Let-7 microRNA [43, 44]. In contrast, activation of the NF-κB-related pathway due to increased oxidative stress was shown to enhance tumor proliferation but reduce resistance to paclitaxel treatment [26]. Here, we show that OTUD7B expression inversely correlates with both LIN28 and CASP3 expression in clinical tissues derived from patients with TNBCs. These results suggest that OTUD7B upregulation likely inactivates the NF-κB-Lin28 axis, which leads to the activation of Let-7 microRNA-mediated caspase-3 downregulation and eventually results in poor response to paclitaxel-induced cell apoptosis in paclitaxel-resistant TNBC cells.
In conclusion, OTUD7B upregulation is most likely due to miR-1180 downregulation and that this upregulation protects paclitaxel-resistant TNBC cells from paclitaxel-induced cell death by inhibiting NF-κB activity and thereby restraining Lin28-mediated suppression of maturation/activation of Let-7 microRNA. These effects may ultimately render the apoptotic pathway ineffective due to Let-7-triggered post-transcriptional inhibition of caspase-3 transcript. Our findings not only suggest that OTUD7B could be a prognostic biomarker for paclitaxel efficacy but also imply that targeting OTUD7B may be a new strategy to enhance the anti-cancer effectiveness of paclitaxel-based chemotherapy against breast cancer, particularly TNBCs.
MATERIALS AND METHODS
Cell lines and cell culture condition
Breast cancer cell lines MDA-MB-231 and MDA-MB468 were cultured in Leibovitz’s (L-15) medium (Gibco Life Technologies, Grand Island, NY, USA) supplemented with 10% foetal bovine serum (FBS, Invitrogen) and incubated at 37°C with free gas exchange with atmospheric air. Breast cancer cell lines HCC1806, HCC1937, HCC38 and HCC70 were cultured in RPMI-1640 medium (Gibco Life Technologies) with 10% FBS and incubated at 37°C with 5% CO2. BT-20 and Hs578t cells were cultured in Eagle’s Minimum Essential Medium (EMEM) and DMEM, respectively, with 10% FBS and incubated at 37°C with 5% CO2. All cell lines were obtained from American Type Culture Collection (ATCC). All cells were routinely authenticated on the basis of short tandem repeat (STR) analysis, morphologic and growth characteristics and mycoplasma detection.
Reverse transcription PCR (RT-PCR)
Total RNA was extracted from cells using TRIzol extraction kit (Invitrogen). Aliquots (5 μg) of total RNA were treated with M-MLV reverse transcriptase (Invitrogen) and then amplified with Taq-polymerase (Protech) using paired primers (for primary mR-1180, forward- CTGGTGCCCACCTCAGAGACGG and reverse-CAC AGCCACCAGGCTGAGCATG; for OTUD7B, forward- GAAGGAGAAGTCAAAGCGAGATCG and reverse-GC ATCACCTCCTGGCTATACTTGC; for CASP3, forward-CATGGAAGCGAATCAATGGACT and reverse-CTGTA CCAGACCGAGATGTCA; for GAPDH, forward-AGG TCGGAGTCAACGGATTTG and reverse-GTGATGG CATGGACTGTGGTC).
In vitro invasion assay
For invasion assay, Boyden chambers (Neuro Probe, Inc., USA) were used according to the manufacturer’s protocol. Briefly, polycarbonate membrane (8 μm pore size, 25 × 80 mm, Neuro Probe, Inc., USA) was pre-coated with 10 μg of human fibronectin (Sigma, MO, USA) on the lower side and matrigel on the upper side. Cells (1.5 × 104) were plated in the top chamber in 50 μl of starvation medium (0.1% FBS) containing drugs or DMSO. After 16 h, stationary cells were removed from the top side of the membrane, whereas migrated cells in the bottom side of the membrane was fixed in 100% methanol and stained with 10% Giemsa’s solution (Merck, Germany) for 1 hr. Invaded cells were counted under the light microscope (400x, ten random fields from each well). All experiments were performed in triplicates.
Microarray data processing
Transcriptional profiling results for HCC38 and MDA-MB436 cells treated with or without paclitaxel at 10× IC50 concentrations were obtained from GEO DataSets (accession no. GSE50832) on the NCBI website. Microarray data and related clinical data for GEO DataSet GSE22513 were also downloaded from NCBI website. Affymetrix DAT files were processed using the Affymetrix Gene Chip Operating System (GCOS) to generate .CEL files. Raw intensities in the .CEL files were normalized by robust multi-chip analysis (RMA), and fold-change analysis was performed using GeneSpring GX11 (Agilent Technologies). Relative mRNA expression levels were normalized by their median and presented as log2 values.
In silico analysis
Genes with a 1.5-fold-change threshold relative to control cells in HCC38 and MDA-MB436 cells after paclitaxel treatment were uploaded to the Ingenuity Pathway Analysis (IPA) website (Ingenuity Systems, www.ingenuity.com). Results of computational predictions for the activation or inhibition status of upstream regulators were then output as a text file. Consensus upstream regulators with significant z-scores from in silico analysis of paclitaxel-treated HCC38 and MDA-MB436 cells were analyzed in a PivotTable report and plotted as a dotplot in Microsoft Excel.
Animal studies
NOD/SCID mice were obtained from National Laboratory Animal Center in Taiwan and were maintained in compliance with the institutional guideline. All animal procedures were approved by the Institutional Animal Care and Use Committee at Academia Sinica, Taiwan. For the in vivo tumor growth, cells (5 × 105) suspended in 50 μL PBS were subcutaneously inoculated into the lateral dorsal skin of each mouse. Mice were humanely killed at the end of experiments and lungs were obtained for histological analysis.
Statistical analyses
SPSS 17.0 software (Informer Technologies, Roseau, Dominica) was used to analyze statistical significance. Paired t-tests were utilized to compare OTUD7B gene expression in cancer tissues and corresponding normal tissues. Spearman’s test was performed to estimate the association between OTUD7B mRNA and Paclitaxel IC50 concentrations in the panel of TNBC cell lines. Pearson’s correlation test was used to evaluate the association between OTUD7B and miR-1180 mRNA expression in basal-like breast cancer. Survival probabilities were determined by Kaplan-Meier analysis and log-rank tests. One-way ANOVA with Tukey’s test was used to estimate the difference in mRNA levels of OTUD7B, DSTNP2 and VCAN in HCC38 and MDA-MB436 cells after paclitaxel treatment. Mann-Whitney U-tests were used to analyze non-parametric data. P values < 0.05 in all analyses were considered statistically significant.
Author contributions
Conception and design: HWC and YFL. Acquisition of data: IJT. Analysis and interpretation of data: HWC, IJT and YFL. Writing, review and/or revision of the manuscript: HWC and YFL. Administrative, technical, or material support: IJT. All authors read and approved the final manuscript.
ACKNOWLEDGMENTS AND FUNDING
This study was supported by the Ministry of Science and Technology, Taiwan (MOST 105-2320-B-038-021-MY3 to HWC and MOST 104-2320-B-038-061-MY3 to YFL).
CONFLICTS OF INTEREST
The authors have no conflicts of interest to declare.
REFERENCES
1. Vyas DM, Kadow JF. Paclitaxel: a unique tubulin interacting anticancer agent. Prog Med Chem. 1995; 32:289–337.
2. Yusuf RZ, Duan Z, Lamendola DE, Penson RT, Seiden MV. Paclitaxel resistance: molecular mechanisms and pharmacologic manipulation. Curr Cancer Drug Targets. 2003; 3:1–19.
3. Baguley BC. Multiple drug resistance mechanisms in cancer. Mol Biotechnol. 2010; 46:308–316.
4. von MG, Schneeweiss A, Loibl S, Salat C, Denkert C, Rezai M, Blohmer JU, Jackisch C, Paepke S, Gerber B, Zahm DM, Kummel S, Eidtmann H, et al. Neoadjuvant carboplatin in patients with triple-negative and HER2-positive early breast cancer (GeparSixto; GBG 66): a randomised phase 2 trial. Lancet Oncol. 2014; 15:747–756.
5. Sikov WM, Berry DA, Perou CM, Singh B, Cirrincione CT, Tolaney SM, Kuzma CS, Pluard TJ, Somlo G, Port ER, Golshan M, Bellon JR, Collyar D, et al. Impact of the addition of carboplatin and/or bevacizumab to neoadjuvant once-per-week paclitaxel followed by dose-dense doxorubicin and cyclophosphamide on pathologic complete response rates in stage II to III triple-negative breast cancer: CALGB 40603 (Alliance). J Clin Oncol. 2015; 33:13–21.
6. Hwang JE, Hong JY, Kim K, Kim SH, Choi WY, Kim MJ, Jung SH, Shim HJ, Bae WK, Hwang EC, Lee KH, Lee JH, Cho SH, et al. Class III beta-tubulin is a predictive marker for taxane-based chemotherapy in recurrent and metastatic gastric cancer. BMC Cancer. 2013; 13:431.
7. Roque DM, Buza N, Glasgow M, Bellone S, Bortolomai I, Gasparrini S, Cocco E, Ratner E, Silasi DA, Azodi M, Rutherford TJ, Schwartz PE, Santin AD. Class III beta-tubulin overexpression within the tumor microenvironment is a prognostic biomarker for poor overall survival in ovarian cancer patients treated with neoadjuvant carboplatin/paclitaxel. Clin Exp Metastasis. 2013.
8. Edelman MJ, Schneider CP, Tsai CM, Kim HT, Quoix E, Luft AV, Kaleta R, Mukhopadhyay P, Trifan OC, Whitaker L, Reck M. Randomized phase II study of ixabepilone or paclitaxel plus carboplatin in patients with non-small-cell lung cancer prospectively stratified by beta-3 tubulin status. J Clin Oncol. 2013; 31:1990–1996.
9. Ohashi T, Yoshimasu T, Oura S, Kokawa Y, Kawago M, Hirai Y, Miyasaka M, Aoishi Y, Kiyoi M, Nishiguchi H, Honda M, Okamura Y. Class III Beta-tubulin Expression in Non-small Cell Lung Cancer: A Predictive Factor for Paclitaxel Response. Anticancer Res. 2015; 35:2669–2674.
10. Xu YC, Zhang FC, Li JJ, Dai JQ, Liu Q, Tang L, Ma Y, Xu Q, Lin XL, Fan HB, Wang HX. RRM1, TUBB3, TOP2A, CYP19A1, CYP2D6: Difference between mRNA and protein expression in predicting prognosis of breast cancer patients. Oncol Rep. 2015; 34:1883–1894.
11. Juang YC, Landry MC, Sanches M, Vittal V, Leung CC, Ceccarelli DF, Mateo AR, Pruneda JN, Mao DY, Szilard RK, Orlicky S, Munro M, Brzovic PS, et al. OTUB1 co-opts Lys48-linked ubiquitin recognition to suppress E2 enzyme function. Mol Cell. 2012; 45:384–397.
12. Mevissen TE, Hospenthal MK, Geurink PP, Elliott PR, Akutsu M, Arnaudo N, Ekkebus R, Kulathu Y, Wauer T, El OF, Freund SM, Ovaa H, Komander D. OTU deubiquitinases reveal mechanisms of linkage specificity and enable ubiquitin chain restriction analysis. Cell. 2013; 154:169–184.
13. Wiener R, Zhang X, Wang T, Wolberger C. The mechanism of OTUB1-mediated inhibition of ubiquitination. Nature. 2012; 483:618–622.
14. Hu H, Wang H, Xiao Y, Jin J, Chang JH, Zou Q, Xie X, Cheng X, Sun SC. Otud7b facilitates T cell activation and inflammatory responses by regulating Zap70 ubiquitination. J Exp Med. 2016; 213:399–414.
15. Hu H, Brittain GC, Chang JH, Puebla-Osorio N, Jin J, Zal A, Xiao Y, Cheng X, Chang M, Fu YX, Zal T, Zhu C, Sun SC. OTUD7B controls non-canonical NF-kappaB activation through deubiquitination of TRAF3. Nature. 2013; 494:371–374.
16. Pareja F, Ferraro DA, Rubin C, Cohen-Dvashi H, Zhang F, Aulmann S, Ben-Chetrit N, Pines G, Navon R, Crosetto N, Kostler W, Carvalho S, Lavi S, et al. Deubiquitination of EGFR by Cezanne-1 contributes to cancer progression. Oncogene. 2012; 31:4599–4608.
17. Dezso Z, Oestreicher J, Weaver A, Santiago S, Agoulnik S, Chow J, Oda Y, Funahashi Y. Gene expression profiling reveals epithelial mesenchymal transition (EMT) genes can selectively differentiate eribulin sensitive breast cancer cells. PLoS.One. 2014; 9:e106131.
18. Tan G, Wu L, Tan J, Zhang B, Tai WC, Xiong S, Chen W, Yang J, Li H. MiR-1180 promotes apoptotic resistance to human hepatocellular carcinoma via activation of NF-kappaB signaling pathway. Sci Rep. 2016; 6:22328.
19. Lin YF, Lai TC, Chang CK, Chen CL, Huang MS, Yang CJ, Liu HG, Dong JJ, Chou YA, Teng KH, Chen SH, Tian WT, Jan YH, et al. Targeting the XIAP/caspase-7 complex selectively kills caspase-3-deficient malignancies. J Clin Invest. 2013; 123:3861–3875.
20. Iliopoulos D, Hirsch HA, Struhl K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell. 2009; 139:693–706.
21. Cao L, Li R, Chen X, Xue Y, Liu D. Neougonin A Inhibits Lipopolysaccharide-Induced Inflammatory Responses via Downregulation of the NF-kB Signaling Pathway in RAW 264.7 Macrophages. Inflammation. 2016; 39:1939–1948.
22. Wu Y, Li W, Zhou C, Lu F, Gao T, Liu Y, Cao J, Zhang Y, Zhang Y, Zhou C. Ketamine inhibits lipopolysaccharide-induced astrocytes activation by suppressing TLR4/NF-kB pathway. Cell Physiol Biochem. 2012; 30:609–617.
23. Barham W, Chen L, Tikhomirov O, Onishko H, Gleaves L, Stricker TP, Blackwell TS, Yull FE. Aberrant activation of NF-kappaB signaling in mammary epithelium leads to abnormal growth and ductal carcinoma in situ. BMC Cancer. 2015; 15:647.
24. Zhong W, Qian K, Xiong J, Ma K, Wang A, Zou Y. Curcumin alleviates lipopolysaccharide induced sepsis and liver failure by suppression of oxidative stress-related inflammation via PI3K/AKT and NF-kappaB related signaling. Biomed Pharmacother. 2016; 83:302–313.
25. Spiga R, Marini MA, Mancuso E, Di FC, Fuoco A, Perticone F, Andreozzi F, Mannino GC, Sesti G. Uric Acid Is Associated With Inflammatory Biomarkers and Induces Inflammation Via Activating the NF-kappaB Signaling Pathway in HepG2 Cells. Arterioscler Thromb Vasc Biol. 2017; 37:1241–1249.
26. Mateescu B, Batista L, Cardon M, Gruosso T, de FY, Mariani O, Nicolas A, Meyniel JP, Cottu P, Sastre-Garau X, Mechta-Grigoriou F. miR-141 and miR-200a act on ovarian tumorigenesis by controlling oxidative stress response. Nat Med. 2011; 17:1627–1635.
27. Hinz M, Loser P, Mathas S, Krappmann D, Dorken B, Scheidereit C. Constitutive NF-kappaB maintains high expression of a characteristic gene network, including CD40, CD86, and a set of antiapoptotic genes in Hodgkin/Reed-Sternberg cells. Blood. 2001; 97:2798–2807.
28. Guttridge DC, Albanese C, Reuther JY, Pestell RG, Baldwin AS Jr. NF-kappaB controls cell growth and differentiation through transcriptional regulation of cyclin D1. Mol Cell Biol. 1999; 19:5785–5799.
29. Yu Q, Geng Y, Sicinski P. Specific protection against breast cancers by cyclin D1 ablation. Nature. 2001; 411:1017–1021.
30. Liu Z, Zhu G, Getzenberg RH, Veltri RW. The Upregulation of PI3K/Akt and MAP Kinase Pathways is Associated with Resistance of Microtubule-Targeting Drugs in Prostate Cancer. J Cell Biochem. 2015; 116:1341–1349.
31. West KA, Castillo SS, Dennis PA. Activation of the PI3K/Akt pathway and chemotherapeutic resistance. Drug Resist Updat. 2002; 5:234–248.
32. Zha L, Chen J, Sun S, Mao L, Chu X, Deng H, Cai J, Li X, Liu Z, Cao W. Soyasaponins can blunt inflammation by inhibiting the reactive oxygen species-mediated activation of PI3K/Akt/NF-kB pathway. PLoS One. 2014; 9:e107655.
33. Jayasooriya RG, Lee KT, Lee HJ, Choi YH, Jeong JW, Kim GY. Anti-inflammatory effects of beta-hydroxyisovalerylshikonin in BV2 microglia are mediated through suppression of the PI3K/Akt/NF-kB pathway and activation of the Nrf2/HO-1 pathway. Food Chem Toxicol. 2014; 65:82–89.
34. Rocha CR, Kajitani GS, Quinet A, Fortunato RS, Menck CF. NRF2 and glutathione are key resistance mediators to temozolomide in glioma and melanoma cells. Oncotarget. 2016; 7:48081–48092. http://doi.org/10.18632/oncotarget.10129.
35. Chen J, Solomides C, Simpkins F, Simpkins H. The role of Nrf2 and ATF2 in resistance to platinum-based chemotherapy. Cancer Chemother Pharmacol. 2017; 79:369–380.
36. Bao L, Wu J, Dodson M, Rojo de la Vega EM, Ning Y, Zhang Z, Yao M, Zhang DD, Xu C, Yi X. ABCF2, an Nrf2 target gene, contributes to cisplatin resistance in ovarian cancer cells. Mol Carcinog. 2017; 56:1543–1553.
37. Lu BC, Li J, Yu WF, Zhang GZ, Wang HM, Ma HM. Elevated expression of Nrf2 mediates multidrug resistance in CD133+ head and neck squamous cell carcinoma stem cells. Oncol Lett. 2016; 12:4333–4338.
38. Ryoo IG, Kim G, Choi BH, Lee SH, Kwak MK. Involvement of NRF2 Signaling in Doxorubicin Resistance of Cancer Stem Cell-Enriched Colonospheres. Biomol Ther Seoul. 2016; 24:482–488.
39. Xu Y, Luo Y, Wang ZY, Li X, Zheng P, Zhang TC. MRTF-A can activate Nrf2 to increase the resistance to doxorubicin. Oncotarget. 2017; 8:8436–8446. http://doi.org/10.18632/oncotarget.14246.
40. Zhong Y, Zhang F, Sun Z, Zhou W, Li ZY, You QD, Guo QL, Hu R. Drug resistance associates with activation of Nrf2 in MCF-7/DOX cells, and wogonin reverses it by down-regulating Nrf2-mediated cellular defense response. Mol Carcinog. 2013; 52:824–834.
41. Khatri R, Shah P, Guha R, Rassool FV, Tomkinson AE, Brodie A, Jaiswal AK. Aromatase Inhibitor-Mediated Downregulation of INrf2 (Keap1) Leads to Increased Nrf2 and Resistance in Breast Cancer. Mol Cancer Ther. 2015; 14:1728–1737.
42. Tsang WP, Kwok TT. Let-7a microRNA suppresses therapeutics-induced cancer cell death by targeting caspase-3. Apoptosis. 2008; 13:1215–1222.
43. Piskounova E, Viswanathan SR, Janas M, LaPierre RJ, Daley GQ, Sliz P, Gregory RI. Determinants of microRNA processing inhibition by the developmentally regulated RNA-binding protein Lin28. J Biol Chem. 2008; 283:21310–21314.
44. Viswanathan SR, Daley GQ, Gregory RI. Selective blockade of microRNA processing by Lin28. Science. 2008; 320:97–100.

