INTRODUCTION
Breast cancer is a heterogeneous disease and the most common cancer among women with a drastically increasing rate in China [1]. Approximately 6% of women were initially diagnosed with metastatic breast cancer (MBC) and about 20% of patients would develop to MBC at an early stage [2]. Despite of significant improvements in the treatment of MBC during the last decade, it still remains an incurable disease with a median overall survival of 18-30 months [3]. Current therapy decision of MBC relies on clinical features, histological factors and well-defined biomarkers [4]. Effective chemotherapy drugs used in the treatment of various malignant tumors always result in drug resistance and toxicity. It was reported that many genetic polymorphisms were involved in metabolism enzyme function, drug resistance, toxicity and efficiency of chemotherapy [5–11]. Hunting for genetic markers to improve clinical outcome of MBC patients becomes a big challenge.
Glutathione S-transferase (GST) enzymes play an indispensable role in detoxifying chemotherapy drugs. They detoxify products of oxidation or alkylating drugs by directly combining to reactive compounds or drugs [12, 13]. GSTT1(glutathione S-transferase theta), located on chromosome 22, plays a role in human carcinogenesis. GSTP1(glutathione S-transferase pi), located on chromosome 11, prevents cells from carcinogen and cytotoxin. And GSTM1(glutathione S-transferase mu), located on chromosome 1, is involved in detoxification and drugs action [14, 15]. Studies have shown that GSTs gene polymorphisms might aid to identify high-risk individuals of chronic diseases such as type 2 diabetes mellitus (T2DM), hypertension and so on [16–18]. In addition, GSTs genetic variants have been reported to be involved in fluorouracil and platinum-based chemotherapy of various metastatic or advanced cancers, such as acute myeloid leukemia, gastrointestinal tumor, non-small cell lung cancer and prostate cancer [19–21].
For potential prognostic value, genetic polymorphism is also important for patients with MBC. Moreover, genetic polymorphism can be easily detected and applied to clinical application. Because genetic polymorphism is found to be strongly associated with chemotherapy efficacy and prognosis of breast cancer, it can be used to establish a refined model to predict prognosis of this disease [22]. Therefore, we performed a study in patients with previously untreated MBC to assess the impact of GSTM1 null/present, GSTT1 null/present, and GSTP1 rs1695 polymorphisms on the survival. The present study demonstrates that these genetic polymorphisms in MBC cancer patients have the potential to be developed as novel biomarkers for diagnosis and prognosis of MBC patients.
RESULTS
Patient characteristics and clinical outcomes
The distribution of demographic, treatment characteristics and clinical features of patients are presented in Table 1. By the time of the final analysis (December 2012), the median follow-up time of the patients was 35.8 months. One hundred and twenty-eight patients (75.3%) died and the median survival time was 21.6 months [95% confidence interval (CI), 18.6-24.6 months]. The 1-, 3-, and 5-year OS rates were 82. 4%, 65.9%, and 16.9%, respectively.
Table 1: Patient clinical and treatment characteristics
Characteristics |
N (%) |
|---|---|
Age at MBC diagnosis (years) |
|
Median |
50. 0 |
Range |
25. 0-74. 0 |
≥ 60 |
28 (16. 5) |
40-59 |
113 (66. 5) |
< 40 |
29 (17. 1) |
Menstruation status |
|
Post-menopausal |
71 (41. 8) |
Pre-menopausal |
99 (58. 2) |
Molecular subtype |
|
Luminal A |
36 (21. 2) |
Luminal B (HER-2 negative) |
12 (7. 1) |
HER-2 positive |
18 (10. 6) |
Triple-negative |
98 (57. 6) |
Unknown |
6 (3. 5) |
Adjuvant therapy |
|
No |
18 (10. 6) |
Only CT (± RT) |
119 (70. 0) |
CT + HT (± RT) |
33 (19. 4) |
Relapse-free interval |
|
Median (months) |
15. 2 |
≤ 2 years |
119 (70. 0) |
> 2 years |
51 (30. 0) |
No. of metastatic sites |
|
1 |
69 (40. 6) |
2 |
45 (26. 5) |
≥3 |
56 (32. 9) |
Metastatic site* |
|
Liver |
50 (29. 4) |
Lung |
75 (44. 1) |
Brain |
6 (3. 5) |
Lymph node |
108 (63. 5) |
Bone |
45 (26. 5) |
Chest wall |
32 (18. 8) |
Others |
38 (22. 4) |
Type of metastatic site |
|
Non-visceral |
51 (30. 0) |
Visceral |
119 (70. 0) |
Abbreviation: MBC, metastatic breast cancer; CT, chemotherapy; RT, radiotherapy; HT, hormone therapy; No., number.
* Number determined at MBC diagnosis includes metastasis to more than one site
Comparison of survival according to baseline characteristics of patients
To test whether various clinical characteristics contribute to overall survival (OS), patients were grouped according to age, menstruation status, molecular subtype, previous adjuvant treatment, RFI, number of metastatic sites, and type of metastatic site. According to univariate analysis, age, RFI, number of metastatic sites, and type of metastatic site significantly influenced patient prognosis (Table 2).
Table 2: Univariate and multivariate Cox-regression analyses for OS
Genotype |
Univariate analysis |
Multivariate analysis |
||
|---|---|---|---|---|
HR (95% CI) |
P |
HR (95% CI) |
P |
|
Age at MBC diagnosis |
||||
≥ 60 |
1. 00 |
|||
40-59 |
1. 90 (1. 12-3. 20) |
0. 017 |
||
< 40 |
2. 34 (1. 25-4. 37) |
0. 008 |
||
Menstruation status |
||||
Post-menopausal |
1. 00 |
|||
Pre-menopausal |
1. 28 (0. 90-1. 81) |
0. 176 |
||
Molecular subtype |
||||
Luminal A |
1. 00 |
|||
Luminal B (HER-2 negative) |
0. 92 (0. 45-1. 88) |
0. 817 |
||
HER-2 positive |
1. 30 (0. 82-2. 05) |
0. 271 |
||
Triple-negative |
1. 28 (0. 68-2. 40) |
0. 446 |
||
Adjuvant therapy |
||||
No |
1. 00 |
|||
Only CT (± RT) |
0. 70 (0. 40-1. 21) |
0. 201 |
||
CT + HT (± RT) |
0. 72 (0. 38-1. 36) |
0. 310 |
||
Relapse-free interval |
||||
≤ 2 years |
1. 00 |
1. 00 |
||
> 2 years |
0. 58 (0. 39-0. 86) |
0. 007 |
0. 56 (0. 38-0. 84) |
0. 005 |
No. of metastatic sites |
||||
1 |
1. 00 |
|||
2 |
1. 01 (0. 65-1. 56) |
0. 983 |
||
≥3 |
1. 63 (1. 09-2. 46) |
0. 019 |
||
Type of metastatic site |
||||
Non-visceral |
1. 00 |
1. 00 |
||
Visceral |
1. 80 (1. 20-2. 72) |
0. 005 |
1. 68 (1. 11-2. 55) |
0. 015 |
GSTT1 deletion |
1. 84 (1. 22-2. 76) |
0. 003 |
1. 92 (1. 26-2. 91) |
0. 002 |
GSTM1 deletion |
1. 43 (1. 00-2. 04) |
0. 046 |
1. 53 (1. 05-2. 23) |
0. 028 |
Abbreviation: OS, overall survival; MBC, metastatic breast cancer; CT, chemotherapy; RT, radiotherapy; HT, hormone therapy; No., number.
Effects of SNPs on OS
The allelic frequencies for multiple genes’ variants are summarized in Table 3. All observed genotype frequencies in patients were verified to be consistent with the HWE. Interestingly, the GSTT1, GSTM1 polymorphisms were significantly associated with patient survival. As shown in Table 3, patients with the present genotypes of GSTT1 and GSTM1 had 6.2 and 8.1 months longer survival (median OS, 23.4 and 28.2 months; 95% CI, 18.7-28.1 and 18.8-37.6 months, respectively) than those with the null genotypes (median OS, 17.2 and 20.1 months; 95% CI, 14.9-19.5 and 17.0-23.2 months, respectively; P = 0.003 and 0.046 for log-rank test; Figures 1 and 2). But GSTP1 rs1695 was not found associated with overall survival of previously untreated MBC in our study.
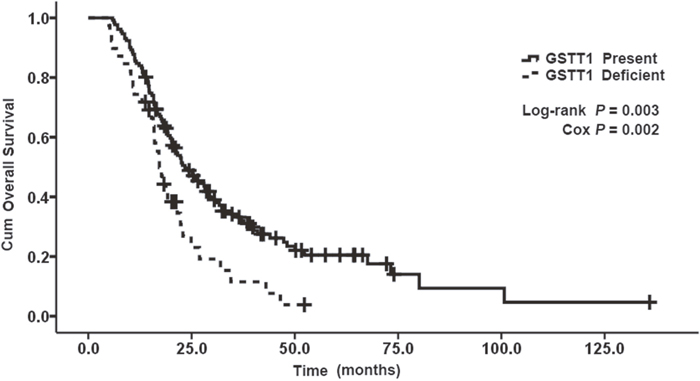
Figure 1: Kaplan–Meier curve demonstrating the overall survival (OS) of genotypes of GSTT1. The median OS was 23. 4 months (95% CI: 18. 7-28. 1) in present genotypes of GSTT1 and 17. 2 months (95% CI: 14. 9-19. 5) in null genotypes of GSTT1; p=0. 003.
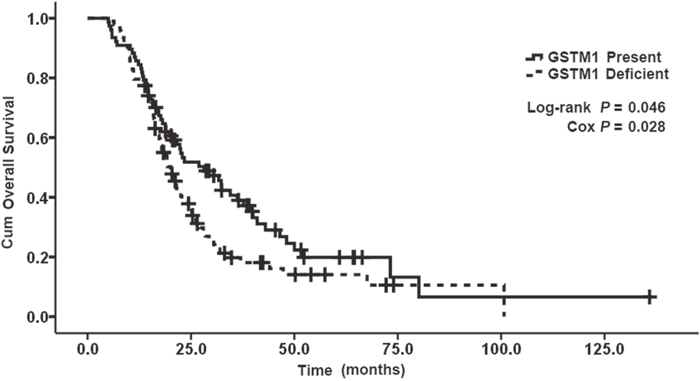
Figure 2: Kaplan–Meier curve demonstrating the overall survival (OS) of genotypes of GSTM1. The median OS was 28. 2 months (95% CI: 18. 8-37. 6) in present genotypes of GSTM1 and 20. 1 months (95% CI: 17. 0-23. 2) in null genotypes of GSTM1; p=0. 046.
Table 3: Associations between genotypes and OS
Genotype |
No. |
OS |
|||
|---|---|---|---|---|---|
Median (mo) |
P |
HR |
95%CI |
||
GSTT1 |
|||||
Present |
131 |
23. 4 |
1. 00 |
||
Null |
39 |
17. 2 |
0. 003 |
1. 84 |
1. 22-2. 76 |
GSTM1 |
|||||
Present |
77 |
28. 2 |
1. 00 |
||
Null |
93 |
20. 1 |
0. 046 |
1. 43 |
1. 00-2. 04 |
GSTP1 (rs1695) |
|||||
AA |
116 |
20. 4 |
1. 00 |
||
AG |
50 |
22. 9 |
0. 768 |
1. 06 |
0. 72-1. 57 |
GG |
4 |
50. 0 |
0. 292 |
0. 47 |
0. 12-1. 91 |
AG+GG |
54 |
24. 3 |
0. 992 |
1. 00 |
0. 68-1. 46 |
Abbreviation: No., number; mo, month; OS, overall survival; HR, hazard ratio; CI, confidence interval; NC, not calculated.
Multivariate analysis
In the multivariate Cox proportional hazards model, after adjustment for age, menstruation status, molecular subtype, previous adjuvant treatment, or number of metastatic sites, the prognostic significance of GSTT1, GSTM1 polymorphisms, RFI and type of metastatic site still existed. The hazard ratios (HRs) of patients with GSTT1 null genotype, GSTM1 null genotype, RFI > 2 years or visceral metastasis on OS were 1.92 (95% CI, 1.26-2.91), 1.53 (95% CI, 1.05-2.23), 0.56 (95% CI, 0.38-0.84) and 1.68 (95% CI, 1.11-2.55), respectively (Table 2).
DISCUSSION
As an incurable disease, MBC need systemic treatments which include chemotherapy, endocrine therapy, molecular therapy and immunotherapy. Some clinical characteristics are fundamental for therapy decision, such as lymph node metastasis, hormone receptors status, human epidermal growth factor receptor 2 (HER2) expression and types of metastatic site [23]. Molecular targeting therapies and immunotherapy have shown important and potential status in recent years for their remarkable effect and lower toxicity compared with the traditional chemotherapies. The discovery and use of agents targeted to ER, PR and HER2 have provided clinician with effective therapies. However, drug-resistance remains a crucial obstacle to tackle [24]. What is more, the potential of biomarker-based treatments improving target therapies, emphasized the requirement to find molecular markers involved in pathogenesis of breast cancer, which are the prognostic factors of therapeutic response and survival [25].
A number of prognostic factors have been shown to significantly predict the survival of patients with metastatic disease. These mainly include adjuvant chemotherapy, RFI, dominant metastatic site, menopausal status, receptor status, and multiple organ involvement [26]. In terms of the metastatic site, visceral like liver diffusion was reported to be a predictor of undesirable survival while non-visceral metastatic including only metastatic in bony skeleton or a single bone lesion can be considered as an indolent disease [27, 28]. Genetic polymorphisms involving in drug metabolism, DNA repair and apoptosis could alter the efficacy of chemotherapeutic regimens, and hence have effects on cancer progression.
In the present study, we examined the association of GSTs genetic polymorphisms and patient survival in a cohort of 170 patients with previously untreated MBC. We found that the null genotypes of GSTT1 and GSTM1 significantly contributed to poorer OS compared with the present genotypes, respectively. After adjusting for clinic-pathologic factors, the two genetic variants were still significantly associated with OS, showing that these polymorphisms were independent survival predictors. Additionally, RFI and type of metastatic site were also independently associated with OS of MBC patients in the cohort.
The glutathione-S-transferases (GSTs) make up a family of multifunctional enzymes with detoxification ability on electrophilic compounds [29]. In some previous clinical studies, the higher levels of GST enzymes in tumors are considered to reduce responses to chemotherapy and associated with a poorer survival in patients with carcinomas of the breast [30], stomach [31], esophagus [32], ovary [33, 34] and head and neck [35]. It has been identified that independent gene deletion are unable to express an active protein at both GSTM1 and GSTT1 [36, 37]. One might expect that a null genotype of enzyme would increase response to chemotherapy and improve clinical outcome [13]. Recently. Jian et al. [38] examined GST genotypes in 244 advanced non-small cell lung carcinoma patients, they found the null GSTM1 and the GG genotype of GSTP1 IIe105Val were correlated improved overall survival. Tatjana I. Djukic et al. [39] found patients with increased level of GSTT1 enzymes has the shorter mean life expectancy compared to null GSTT1.
Our results are in accordance with previous reports that null genotypes of GSTT1 and GSTM1 could be a poorer prognosis and showed a sluggish response to chemotherapy in various types of cancers [40–44]. To our knowledge, only one study has investigated the association between polymorphisms in the GST genes and OS in previously untreated patients with MBC [45]. Similar to our result, no associations were found between GSTP1 rs1695 and progression free survival [45]. It should be pointed out that, the results from early breast cancer (EBC) have been mixed. Sweeney et al. [46] and Ambrose et al. [47] found that the early stage patients with the low-activity GSTP1 Val/Val genotype and both null genotypes of GSTM1 and GSTT1 have better OS after chemotherapy. Whereas no effectiveness was reported for GSTP1 [48], GSTM1 [49], and GSTT1 [49] in other studies.
Thus, this is the first study to find the null genotypes of GSTT1 and GSTM1 significantly contribute to poorer OS compared with the present genotypes in MBC, which is quite different from the results in EBC setting [46, 47], but in accordance with the findings in other metastatic cancer types [40–44]. This may be partly because reduced GST activity leads to increased glutathione levels and elevated glutathione reduced the capability of DNA to bind to cytotoxic drugs such like platinum compounds [42, 50, 51], DNA-reactive metabolites of anthracyclines, and various alkylating agents. These most commonly used chemotherapeutic agents in the treatment of breast carcinoma, are substrates for GST-mediated glutathione conjugation [52, 53], more than that, the glutathione can protect DNA from damage and adduct formation by coupling [54]. That is why these enzymes are susceptible to chemotherapy. The function of GSTs extends beyond detoxification and chemosensitivity, as they have been found to play a critical role in kinase signaling [55, 56]. The function of cell signaling controlled will provide novel therapeutic targets of new drugs. It will provide the possibility to develop antagonists or agonists aiming at signaling pathway and exert positive biological effect. Additional, acquiring of GSTs genotyping from blood samples leads to personalized modality and better effectiveness. Therefore, the variants of GSTT1 and GSTM1 could be a novel and helpful predictive factor to identify specific MBC patients who may benefit from signaling pathway. As our study is the inclusion of untreated MBC, the identified patients may acquire higher response and lower toxicity as they accept targeting therapies earlier, at the same time, spare those patients unlikely to benefit from needless therapies and toxicity.
Several limitations of this study should be noted. The sample size of our study is relative moderate. And further research is necessary to choose patients according to genetic characteristics and find the optimized targeted treatment or tailored chemotherapy for patients with null genotypes of GSTT1 and GSTM1. Results of the presented study should be validated in prospective studies. And, due to possible ethnic differences, our results should be further verified in different ethnic populations to acquire more accurate and solid conclusions in the future.
In conclusion, we have reported for the first time that there were significant differences in the OS among previously untreated MBC patients with different GSTT1 and GSTM1 genotypes. Our results suggest that in addition to clinic-pathologic factors, genetic variants in GSTs might be suggestive factors in untreated MBC patients and further research is warranted.
MATERIALS AND METHODS
Patients
From March 2002 through November 2011, a total of 170 patients from Fudan University Shanghai Cancer Center (FUSCC) with previously untreated MBC were enrolled. Criteria for inclusion were as follows: female gender with histologically confirmed invasive ductal carcinoma, age of 18 to 70, with Eastern Cooperative Oncology Group (ECOG) performance status of 0 to 1, with adequate liver, renal function, and adequate bone marrow function. Exclusion criteria included: pretreatment of metastatic disease; more than one primary malignancy (except carcinoma in situ of the cervix or basal cell carcinoma of the skin with proper treatment); other serious complications/comorbidities that might affect survival.
Baseline information of these patients was collected and all specimens (blood samples) were obtained before treatment aiming at the metastatic disease. Classification of molecular subtypes and the clinic-pathology were based on the 2013 St. Gallen consensus [57]. Survival information was collected from hospital medical records and/or the follow-ups every 3 months. Each patient provided signed informed consent of using their DNA and clinical data. The study was approved by the Institutional Review Board of FUSCC.
SNP genotyping
DNA was collected from 5-mL blood sample from each patient. The polymorphisms of multiple genes including GSTM1 null/present, GSTT1 null/present, and GSTP1 rs1695 were performed by PCR-based RFLP methods, then applied DNA sequencing of the PCR products to further confirm the genotypes [58]. To make sure the accuracy of method and total reproducibility, 15% random samples were genotyped repeat by different people.
Statistical analysis
For each polymorphism, Pearson χ2 test was applied to test the Hardy-Weinberg equilibrium (HWE). OS was calculated from diagnosis of MBC to death. Survival distributions were analyzed by the Kaplan-Meier method and log-rank test was used to compare the survival analyses. Multivariate Cox proportional hazards models were applied to evaluate the effect of prognostic and clinical factors on OS, including age, molecular subtype, menstruation status, previous adjuvant treatment, relapse-free interval (RFI), number of metastatic sites, and type of metastatic site. Statistical significance was set at a level of 0.05 and all the statistical analyses were conducted using the SPSS software package (version 17. 0).
Abbreviations
GSTM1: glutathione S-transferase theta; GSTP1: glutathione S-transferase pi; GSTT1: glutathione S-transferase mu; MBC: metastatic breast cancer; OS: overall survival; HR: hazard ratio; GST: glutathione S-transferase; T2DM: type 2 diabetes mellitus; RFI: relapse-free interval; ER: estrogen receptor; PR: progestrone receptor; HER2: human epidermal growth factor receptor 2; FUSCC: Fudan University Shanghai Cancer Center; ECOG: Eastern Cooperative Oncology Group.
Author contributions
Conception and design: JZ, YW, XH and BW; Collection and assembly of data: JZ and YW; Data analysis and interpretation: JZ, YW and ZS; Pathological slides reviewing: JZ, YW, JC and LW; Manuscript writing: JZ, YW and ZW; Final approval of manuscript: JZ, YW, XH, BW, LW, SZ, JC and ZW.
ACKNOWLEDGMENTS
We’d like to show our sincere gratitude to all the faculty members in our department and all the patients and their families involved in the current study.
CONFLICTS OF INTEREST
No potential conflicts of interest were disclosed.
FUNDING
This research was supported by the grant from Shanghai Municipal Commission of Health and Family Planning (201640069) and Shanghai Natural Science Foundation (17ZR1405700).
REFERENCES
1. Li J, Zhang BN, Fan JH, Pang Y, Zhang P, Wang SL, Zheng S, Zhang B, Yang HJ, Xie XM, Tang ZH, Li H, Li JY, et al. A nation-wide multicenter 10-year (1999-2008) retrospective clinical epidemiological study of female breast cancer in China. BMC Cancer. 2011; 11:364. https://doi.org/10.1186/1471-2407-11-364.
2. Brewster AM, Hortobagyi GN, Broglio KR, Kau SW, Santa-Maria CA, Arun B, Buzdar AU, Booser DJ, Valero V, Bondy M, Esteva FJ. Residual risk of breast cancer recurrence 5 years after adjuvant therapy. J Natl Cancer Inst. 2008; 100:1179-83. https://doi.org/10.1093/jnci/djn233.
3. Mariani G. New developments in the treatment of metastatic breast cancer: from chemotherapy to biological therapy. Ann Oncol. 2005; 16:ii191-4. https://doi.org/10.1093/annonc/mdi719.
4. Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thurlimann B, Senn HJ; Panel Members. Strategies for subtypes--dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011; 22:1736-47. https://doi.org/10.1093/annonc/mdr304.
5. Pullarkat ST, Stoehlmacher J, Ghaderi V, Xiong YP, Ingles SA, Sherrod A, Warren R, Tsao-Wei D, Groshen S, Lenz HJ. Thymidylate synthase gene polymorphism determines response and toxicity of 5-FU chemotherapy. Pharmacogenomics J. 2001; 1:65-70.
6. Evans WE, Relling MV. Pharmacogenomics: translating functional genomics into rational therapeutics. Science. 1999; 286:487-91.
7. Nebert DW. Polymorphisms in drug-metabolizing enzymes: what is their clinical relevance and why do they exist? Am J Hum Genet. 1997; 60:265-71.
8. Bi N, Yang M, Zhang L, Chen X, Ji W, Ou G, Lin D, Wang L. Cyclooxygenase-2 genetic variants are associated with survival in unresectable locally advanced non-small cell lung cancer. Clin Cancer Res. 2010; 16:2383-90. https://doi.org/10.1158/1078-0432.CCR-09-2793.
9. Gurubhagavatula S, Liu G, Park S, Zhou W, Su L, Wain JC, Lynch TJ, Neuberg DS, Christiani DC. XPD and XRCC1 genetic polymorphisms are prognostic factors in advanced non-small-cell lung cancer patients treated with platinum chemotherapy. J Clin Oncol. 2004; 22:2594-601. https://doi.org/10.1200/JCO.2004.08.067.
10. Tibaldi C, Giovannetti E, Vasile E, Mey V, Laan AC, Nannizzi S, Di Marsico R, Antonuzzo A, Orlandini C, Ricciardi S, Del Tacca M, Peters GJ, Falcone A, et al. Correlation of CDA, ERCC1, and XPD polymorphisms with response and survival in gemcitabine/cisplatin-treated advanced non-small cell lung cancer patients. Clin Cancer Res. 2008; 14:1797-803. https://doi.org/10.1158/1078-0432.CCR-07-1364.
11. Yu D, Zhang X, Liu J, Yuan P, Tan W, Guo Y, Sun T, Zhao D, Yang M, Liu J, Xu B, Lin D. Characterization of functional excision repair cross-complementation group 1 variants and their association with lung cancer risk and prognosis. Clin Cancer Res. 2008; 14:2878-86. https://doi.org/10.1158/1078-0432.CCR-07-1612.
12. Salinas AE, Wong MG. Glutathione S-transferases--a review. Curr Med Chem. 1999; 6:279-309.
13. Hayes JD, Pulford DJ. The glutathione S-transferase supergene family: regulation of GST and the contribution of the isoenzymes to cancer chemoprotection and drug resistance. Crit Rev Biochem Mol Biol. 1995; 30:445-600. https://doi.org/10.3109/10409239509083491.
14. Vasieva O. The many faces of glutathione transferase pi. Curr Mol Med. 2011; 11:129-39.
15. Hollman AL, Tchounwou PB, Huang HC. The association between gene-environment interactions and diseases involving the human GST superfamily with SNP variants. Int J Environ Res Public Health. 2016; 13:379. https://doi.org/10.3390/ijerph13040379.
16. Gonul N, Kadioglu E, Kocabas NA, Ozkaya M, Karakaya AE, Karahalil B. The role of GSTM1, GSTT1, GSTP1, and OGG1 polymorphisms in type 2 diabetes mellitus risk: a case-control study in a Turkish population. Gene. 2012; 505:121-7. https://doi.org/10.1016/j.gene.2012.05.025.
17. Stoian A, Banescu C, Balasa RI, Motataianu A, Stoian M, Moldovan VG, Voidazan S, Dobreanu M. Influence of GSTM1, GSTT1, and GSTP1 polymorphisms on type 2 diabetes mellitus and diabetic sensorimotor peripheral neuropathy risk. Dis Markers. 2015; 2015:638693. https://doi.org/10.1155/2015/638693.
18. Pinheiro DS, Rocha Filho CR, Mundim CA, Junior Pde M, Ulhoa CJ, Reis AA, Ghedini PC. Evaluation of glutathione S-transferase GSTM1 and GSTT1 deletion polymorphisms on type-2 diabetes mellitus risk. PLoS One. 2013; 8:e76262. https://doi.org/10.1371/journal.pone.0076262.
19. Hung RJ, van der Hel O, Tavtigian SV, Brennan P, Boffetta P, Hashibe M. Perspectives on the molecular epidemiology of aerodigestive tract cancers. Mutat Res. 2005; 592:102-18. https://doi.org/10.1016/j.mrfmmm.2005.06.007.
20. McIlwain CC, Townsend DM, Tew KD. Glutathione S-transferase polymorphisms: cancer incidence and therapy. Oncogene. 2006; 25:1639-48. https://doi.org/10.1038/sj.onc.1209373.
21. Schnekenburger M, Karius T, Diederich M. Regulation of epigenetic traits of the glutathione S-transferase P1 gene: from detoxification toward cancer prevention and diagnosis. Front Pharmacol. 2014; 5:170. https://doi.org/10.3389/fphar.2014.00170.
22. Evans WE, Relling MV. Moving towards individualized medicine with pharmacogenomics. Nature. 2004; 429:464-8. https://doi.org/10.1038/nature02626.
23. Prat A, Pineda E, Adamo B, Galvan P, Fernandez A, Gaba L, Diez M, Viladot M, Arance A, Munoz M. Clinical implications of the intrinsic molecular subtypes of breast cancer. Breast. 2015; 24:S26-35. https://doi.org/10.1016/j.breast.2015.07.008.
24. Stopeck AT, Brown-Glaberman U, Wong HY, Park BH, Barnato SE, Gradishar WJ, Hudis CA, Rugo HS. The role of targeted therapy and biomarkers in breast cancer treatment. Clin Exp Metastasis. 2012; 29:807-19. https://doi.org/10.1007/s10585-012-9496-y.
25. Campos CZ, Losi Guembarovski R, de Oliveira CE, Banin Hirata BK, Vitiello GA, Dias FL, Hiroki CH, Watanabe MA, Mazzuco TL. Glutathione S-transferases deletions may act as prognosis and therapeutic markers in breast cancer. Clin Exp Med. 2017. https://doi.org/10.1007/s10238-017-0461-6.
26. Nicolini A, Giardino R, Carpi A, Ferrari P, Anselmi L, Colosimo S, Conte M, Fini M, Giavaresi G, Berti P, Miccoli P. Metastatic breast cancer: an updating. Biomed Pharmacother. 2006; 60:548-56. https://doi.org/10.1016/j.biopha.2006.07.086.
27. Koizumi M, Yoshimoto M, Kasumi F, Ogata E. Comparison between solitary and multiple skeletal metastatic lesions of breast cancer patients. Ann Oncol. 2003; 14:1234-40.
28. Sherry MM, Greco FA, Johnson DH, Hainsworth JD. Metastatic breast cancer confined to the skeletal system. An indolent disease. Am J Med. 1986; 81:381-6.
29. Coles BF, Kadlubar FF. Detoxification of electrophilic compounds by glutathione S-transferase catalysis: determinants of individual response to chemical carcinogens and chemotherapeutic drugs? Biofactors. 2003; 17:115-30.
30. Huang J, Tan PH, Thiyagarajan J, Bay BH. Prognostic significance of glutathione S-transferase-pi in invasive breast cancer. Mod Pathol. 2003; 16:558-65. https://doi.org/10.1097/01.MP.0000071842.83169.5A.
31. Nagashima F, Boku N, Ohtsu A, Yoshida S, Hasebe T, Ochiai A, Sakata Y, Saito H, Miyata Y, Hyodo I, Ando M. Biological markers as a predictor for response and prognosis of unresectable gastric cancer patients treated with irinotecan and cisplatin. Jpn J Clin Oncol. 2005; 35:714-9. https://doi.org/10.1093/jjco/hyi194.
32. Harpole DH Jr, Moore MB, Herndon JE 2nd, Aloia T, D’Amico TA, Sporn T, Parr A, Linoila I, Allegra C. The prognostic value of molecular marker analysis in patients treated with trimodality therapy for esophageal cancer. Clin Cancer Res. 2001; 7:562-9.
33. Yokoyama Y, Sato S, Fukushi Y, Sakamoto T, Futagami M, Saito Y. Significance of multi-drug-resistant proteins in predicting chemotherapy response and prognosis in epithelial ovarian cancer. J Obstet Gynaecol Res. 1999; 25:387-94.
34. Kigawa J, Minagawa Y, Kanamori Y, Itamochi H, Cheng X, Okada M, Oishi T, Terakawa N. Glutathione concentration may be a useful predictor of response to second-line chemotherapy in patients with ovarian cancer. Cancer. 1998; 82:697-702.
35. Shiga H, Heath EI, Rasmussen AA, Trock B, Johnston PG, Forastiere AA, Langmacher M, Baylor A, Lee M, Cullen KJ. Prognostic value of p53, glutathione S-transferase pi, and thymidylate synthase for neoadjuvant cisplatin-based chemotherapy in head and neck cancer. Clin Cancer Res. 1999; 5:4097-104.
36. London SJ, Yuan JM, Chung FL, Gao YT, Coetzee GA, Ross RK, Yu MC. Isothiocyanates, glutathione S-transferase M1 and T1 polymorphisms, and lung-cancer risk: a prospective study of men in Shanghai, China. Lancet. 2000; 356:724-9. https://doi.org/10.1016/S0140-6736(00)02631-3.
37. Park SK, Yoo KY, Lee SJ, Kim SU, Ahn SH, Noh DY, Choe KJ, Strickland PT, Hirvonen A, Kang D. Alcohol consumption, glutathione S-transferase M1 and T1 genetic polymorphisms and breast cancer risk. Pharmacogenetics. 2000; 10:301-9.
38. Jia W, Sun JY, Jia KY, Liu XC. Role of GSTM1, GSTT1, and GSTP1 IIe105Val gene polymorphisms in the response to chemotherapy and overall survival of advanced non-small cell lung cancer. Genet Mol Res. 2016. https://doi.org/10.4238/gmr.15037668.
39. Djukic TI, Savic-Radojevic AR, Pekmezovic TD, Matic MG, Pljesa-Ercegovac MS, Coric VM, Radic TM, Suvakov SR, Krivic BN, Dragicevic DP, Simic TP. Glutathione S-transferase T1, O1 and O2 polymorphisms are associated with survival in muscle invasive bladder cancer patients. PLoS One. 2013; 8:e74724. https://doi.org/10.1371/journal.pone.0074724.
40. Davies SM, Robison LL, Buckley JD, Tjoa T, Woods WG, Radloff GA, Ross JA, Perentesis JP. Glutathione S-transferase polymorphisms and outcome of chemotherapy in childhood acute myeloid leukemia. J Clin Oncol. 2001; 19:1279-87. https://doi.org/10.1200/jco.2001.19.5.1279.
41. Howells RE, Redman CW, Dhar KK, Sarhanis P, Musgrove C, Jones PW, Alldersea J, Fryer AA, Hoban PR, Strange RC. Association of glutathione S-transferase GSTM1 and GSTT1 null genotypes with clinical outcome in epithelial ovarian cancer. Clin Cancer Res. 1998; 4:2439-45.
42. Voso MT, D’Alo F, Putzulu R, Mele L, Scardocci A, Chiusolo P, Latagliata R, Lo-Coco F, Rutella S, Pagano L, Hohaus S, Leone G. Negative prognostic value of glutathione S-transferase (GSTM1 and GSTT1) deletions in adult acute myeloid leukemia. Blood. 2002; 100:2703-7. https://doi.org/10.1182/blood.V100.8.2703.
43. Goekkurt E, Al-Batran SE, Hartmann JT, Mogck U, Schuch G, Kramer M, Jaeger E, Bokemeyer C, Ehninger G, Stoehlmacher J. Pharmacogenetic analyses of a phase III trial in metastatic gastroesophageal adenocarcinoma with fluorouracil and leucovorin plus either oxaliplatin or cisplatin: a study of the arbeitsgemeinschaft internistische onkologie. J Clin Oncol. 2009; 27:2863-73. https://doi.org/10.1200/JCO.2008.19.1718.
44. Boige V, Mendiboure J, Pignon JP, Loriot MA, Castaing M, Barrois M, Malka D, Tregouet DA, Bouche O, Le Corre D, Miran I, Mulot C, Ducreux M, et al. Pharmacogenetic assessment of toxicity and outcome in patients with metastatic colorectal cancer treated with LV5FU2, FOLFOX, and FOLFIRI: FFCD 2000-05. J Clin Oncol. 2010; 28:2556-64. https://doi.org/10.1200/JCO.2009.25.2106.
45. Bewick MA, Conlon MS, Lafrenie RM. Polymorphisms in manganese superoxide dismutase, myeloperoxidase and glutathione-S-transferase and survival after treatment for metastatic breast cancer. Breast Cancer Res Treat. 2008; 111:93-101. https://doi.org/10.1007/s10549-007-9764-8.
46. Sweeney C, McClure GY, Fares MY, Stone A, Coles BF, Thompson PA, Korourian S, Hutchins LF, Kadlubar FF, Ambrosone CB. Association between survival after treatment for breast cancer and glutathione S-transferase P1 Ile105Val polymorphism. Cancer Res. 2000; 60:5621-4.
47. Ambrosone CB, Sweeney C, Coles BF, Thompson PA, McClure GY, Korourian S, Fares MY, Stone A, Kadlubar FF, Hutchins LF. Polymorphisms in glutathione S-transferases (GSTM1 and GSTT1) and survival after treatment for breast cancer. Cancer Res. 2001; 61:7130-5.
48. Goode EL, Dunning AM, Kuschel B, Healey CS, Day NE, Ponder BA, Easton DF, Pharoah PP. Effect of germ-line genetic variation on breast cancer survival in a population-based study. Cancer Res. 2002; 62:3052-7.
49. Yang G, Shu XO, Ruan ZX, Cai QY, Jin F, Gao YT, Zheng W. Genetic polymorphisms in glutathione-S-transferase genes (GSTM1, GSTT1, GSTP1) and survival after chemotherapy for invasive breast carcinoma. Cancer. 2005; 103:52-8. https://doi.org/10.1002/cncr.20729.
50. Tew KD. Glutathione-associated enzymes in anticancer drug resistance. Cancer Res. 1994; 54:4313-20.
51. Ishikawa T, Ali-Osman F. Glutathione-associated cis-diamminedichloroplatinum(II) metabolism and ATP-dependent efflux from leukemia cells. Molecular characterization of glutathione-platinum complex and its biological significance. J Biol Chem. 1993; 268:20116-25.
52. Stearns V, Davidson NE, Flockhart DA. Pharmacogenetics in the treatment of breast cancer. Pharmacogenomics J. 2004; 4:143-53. https://doi.org/10.1038/sj.tpj.6500242.
53. Bosch TM, Meijerman I, Beijnen JH, Schellens JH. Genetic polymorphisms of drug-metabolising enzymes and drug transporters in the chemotherapeutic treatment of cancer. Clin Pharmacokinet. 2006; 45:253-85. https://doi.org/10.2165/00003088-200645030-00003.
54. Ryberg D, Skaug V, Hewer A, Phillips DH, Harries LW, Wolf CR, Ogreid D, Ulvik A, Vu P, Haugen A. Genotypes of glutathione transferase M1 and P1 and their significance for lung DNA adduct levels and cancer risk. Carcinogenesis. 1997; 18:1285-9.
55. Townsend DM, Tew KD. The role of glutathione-S-transferase in anti-cancer drug resistance. Oncogene. 2003; 22:7369-75. https://doi.org/10.1038/sj.onc.1206940.
56. Dang DT, Chen F, Kohli M, Rago C, Cummins JM, Dang LH. Glutathione S-transferase pi1 promotes tumorigenicity in HCT116 human colon cancer cells. Cancer Res. 2005; 65:9485-94. https://doi.org/10.1158/0008-5472.CAN-05-1930.
57. Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thurlimann B, Senn HJ; Panel Members. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013; 24:2206-23. https://doi.org/10.1093/annonc/mdt303.
58. Abbas A, Delvinquiere K, Lechevrel M, Lebailly P, Gauduchon P, Launoy G, Sichel F. GSTM1, GSTT1, GSTP1 and CYP1A1 genetic polymorphisms and susceptibility to esophageal cancer in a French population: different pattern of squamous cell carcinoma and adenocarcinoma. World J Gastroenterol. 2004; 10:3389-93.

