INTRODUCTION
Currently, cancers are a great challenge in the field of human health. Epidemiological data show that the global incidence rate for cancer amounts to approximately 14,000,000 per year and that mortality exceeds 8,200,000 [1, 2]. Early detection and diagnosis are critical for improving survival time and quality of life [3]. However, current clinical findings are mainly based on imaging analyses and are restricted by factors such as resolution. Additionally, tumors often cannot be diagnosed early [4, 5]. Fortunately, owing to their inherent sensitivity, tumor markers (especially molecular biomarkers) have revealed great potential in the early diagnosis of malignant tumors [6].
Long noncoding RNAs (lncRNA) are a type of noncoding RNA greater than 200 nucleotides in length [7]. In recent years, lncRNAs have been shown to play an important regulatory role in chromatin modification, X chromosome inactivation and transcription, translation, genetic imprinting, dosage compensation, and the regulation of protein activity and RNA alternative splicing [8–12]. LncRNAs play an important role in tumor incidence and development [13]. A variety of lncRNAs can be used as molecular tumor markers. These markers, together with clinical data, show potential value in the diagnosis and treatment of malignant tumors.
AFAP1-AS1 is a type of lncRNA encoded by the antisense strand of the AFAP1 gene. Our group and other research groups have found that AFAP1-AS1 plays an important role in the regulation of tumor incidence and development by promoting tumor cell metastasis [14–21]. We discovered that AFAP1-AS1 is closely correlated with clinical data - including survival time and TNM staging - in various cancers, such as nasopharyngeal carcinoma [14, 15] and lung cancer [16]. This demonstrates the potential value of AFAP1-AS1 as a new tumor marker. However, due to the restricted sample size, the credibility of previous studies remains doubtful. Thus, our study combined microarray data from the GEO database with results from several published studies in order to systemically review the prognostic value of AFAP1-AS1 in cancers and to determine the molecular mechanisms involved in the regulation of AFAP1-AS1 in tumorigenesis and development.
MATERIALS AND METHODS
Literature searching
Tumor microarray datasets based on the Affymetrix Human Genome U133 Plus 2.0 platform were obtained through independent searches of the GEO database by 3 reseachers (Wang, Mo, and Yang). We also retrieved all relevant published literature from Pubmed and Web of Science. The literature search was limited to studies published in the English language prior to July 2017. To increase the totality of the search, both mesh-terms and free words were used. Search terms were: ‘AFAP1-AS1’, ‘metastasis-associated lung adenocarcinoma transcript 1’, ‘long intergenic noncoding RNA’ or ‘lncRNA’ or ‘noncoding RNA’, ‘cancer’ or ‘carcinoma’ or ‘neoplasm’ and ‘prognosis’ or ‘survival’.
Literature selection
Results from the literature search were screened prior to analysis of datasets. Eligible studies met the following criteria: involved any type of human cancer; detected and analyzed AFAP1-AS1 expression in tissues; literature study involved the correlation between AFAP1-AS1 expression level and overall survival (OS) or disease free survival (DFS); literature provided survival curve or hazard ratio (HR) and the 95% confidence interval (CI). Literature was excluded based on the following criteria: experts’ opinions, letters, comments, case reports, reviews and meeting reports; the article was not found in full; survival rate data files were not presented in the literature; related survival data could not be gained from the literature; repeatedly published literature. When the same data subsets were published in more than one article, only the latest publication was included. All included studies were obtained as full text. Controversies regarding study selection were resolved via discussion with investigator Wang.
Data extraction
This process was carried out independently by six researchers and consensus was reached among all researchers for each dataset. For the 18 Affymetrix Human Genome U133 Plus 2.0 platform microarray datasets available in the GEO database (where both AFAP1-AS1 expression and the corresponding survival data was available), the following data were extracted: OS, RFS, lymph node metastasis, TNM stage, survival outcome, and AFAP1-AS1 expression value. For studies selected from Pubmed and Web of Science, the following items were extracted: author, publishing date, nationalities of patient groupings, sample size, expression level of AFAP1-AS1, length of follow-up, method of survival analysis, OR value and CI, OS, DFS, HR value and CI, RFS, lymphocyte metastasis, TNM staging, and distant metastasis. Screened studies were compiled and checked according to standards for meta-analyses. We then built the database and applied lnHR and SelnHR as a combination index. In cases where only the survival curve was provided, Engauge Digitizer 4.1 was applied to extract relevant data from the survival curve. We then performed calculations according to the method of Jayne F Tierney et al.
Quality assessment of the primary studies
Three researchers (Wang, Mo, and Yang) independently evaluated the quality of all selected studies according to the method of Steels et al. [22, 23] The final assessment is expressed as a percentage, where higher scores indicate higher literature quality.
Statistical approach
(1) We used a single factor cox analysis to calculate the HR value and its 95% CI, OS, and RFS from the gene microarray data. We used GraphPad Prism 5 to draw Kaplan-Meier survival curves based on AFAP1-AS1 expression levels. A ≥ 1.5-fold difference was used as the cut-off value for differentially expressed lncRNA and the false discovery ratio (FDR) was < 0.05. (2) Review Manager 5.3 was used to analyze the obtained data based on clinical indices including OS, DFS, and RFS. HR values from literature studies that acquired OS, RFS, and DFS were combined and heterogeneity was determined to be significant when I2 > 50% [4, 24]. A random effect model was adopted and sub-analyses were made when heterogeneity existed among study results. Otherwise, a fixed effect model was adopted to merge HR values and 95% CIs. Forest plots were applied to present calculation results. HR was calculated as the ratio of the prognoses in high AFAP1-AS1 cases to low AFAP1-AS1 cases. An HR > 1 demonstrated a poorer prognosis for the high AFAP1-AS1 expression group compared with the low AFAP1-AS1 expression group. (3) Publication biases were described through funnel plots for accessible OS and RFS data analysis. SelnHR was shown on the x-axis while lnHR was shown on the y-axis. Symmetry of the funnel plot was tested by linear regression models (Begge method and Eggs method) in STATA 12.0 to evaluate publication bias. Next, a sensitivity analysis was performed using STATA 12.0 to evaluate HR values for AFAP1-AS1 and OS extracted from single studies. Furthermore, OR values were combined and heterogeneity of clinical tumor diagnosis index data was analyzed using Review Manager 5.3. High AFAP1-AS1 expression was analyzed for correlation with TNM staging, lymphocyte metastasis, and distant metastasis across various cancer types. P < 0.05 indicated statistical significance of results.
RESULTS
Included studies and characteristics
As shown in Figure 1, we downloaded 20 Affymetrix Human Genome U133 Plus 2.0 datasets. We also found 10 relevant articles [14–21, 25–32]. Following the review process, 30 studies were chosen for the meta-analysis.
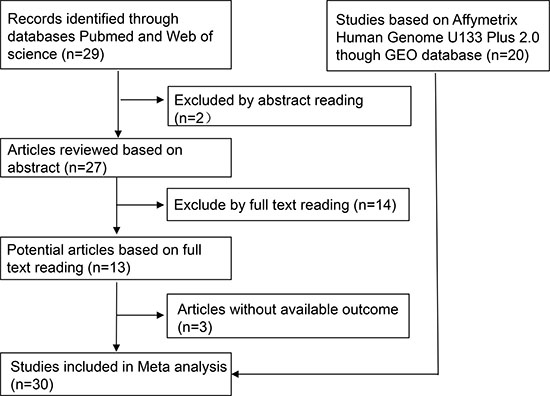
Figure 1: The flowchart of the meta-analysis. Researchers acquired 20 RNA microarray datasets from the GEO database. Of the 29 publications identified in initial research, the authors selected 10 publications for further analysis by reading abstracts and full texts. Thus, there were 30 studies included in this meta-analysis.
A total of 3482 patients were represented across these 30 studies, with a maximum sample size of 579 and a minimum sample size of 33. The mean sample size was 120. The most recent publication date was March 2017. The regions represented in the studies include China (9), America (9), Canada (5), Germany (2), Sweden (1), Japan (1), France (1), and Denmark (1). A total of 15 types of cancer were included in the meta-analysis (5 lung cancer, 4 breast cancer, 3 ovarian cancer, 3 DLBCL, 3 hepatocellular carcinoma, 1 colorectal cancer, 2 pancreatic cancer, 1 glioma, 1 nasopharyngeal carcinoma, 1 colon cancer, and 1 CN-AML).
All studies included a high AFAP1-AS1 expression group and a low AFAP1-AS1 expression group. 16 studies analyzed the expression level of AFAP1-AS1 by RT-PCR, while 19 utilized gene chips and 1 utilized fluorescence in situ hybridization (FISH). OS and RFS were estimated as a survival outcome in 76% (22) and 21% (6) of the studies, respectively. Associations between AFAP1-AS1 and clinical characteristics of cancers - TNM stage, lymph node metastasis, and distant metastasis - were estimated by 9, 5 and 4 studies, respectively.
GEO data analysis
We analyzed the relationship between AFAP1-AS1 and survival (OS/RFS). The Kaplan-Meier curve (Figure 2), as well as the HR and 95% CI (Table 1), were derived. The cut-off value for differentially expressed lncRNA was set at ≥ 1.5-fold, and the false discovery ratio (FDR) was < 0.05. We calculated the HR between AFAP1-AS1 and clinical characteristics of cancers (i.e., TNM stage, lymph node metastasis and distant metastasis) (Table 2).
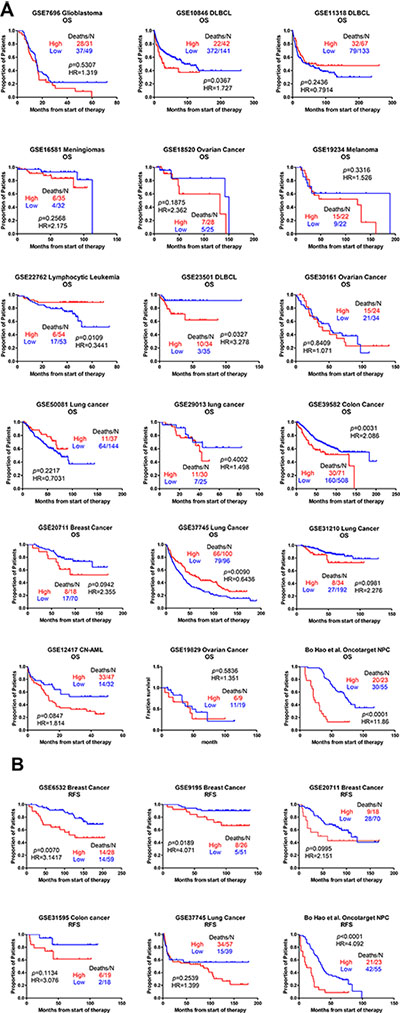
Figure 2: Kaplan-Meier curves relating AFAP1-AS1 expression to OS/RFS in different cancers. The relationship between AFAP1-AS1 and survival (A) OS, and (B) RFS. HR and 95% CIs were calculated in different cancers. The cut-off value for differentially expressed lncRNA was set at ≥ 1.5-fold difference and the false discovery ratio (FDR) was < 0.05.
Table 1: Characteristics of articles included in the meta-analysis
Study |
Year |
Region |
Tumor type |
Sample size |
Clinical stage of tumor |
Test methods of AFAP1-AS1 expression |
Survival outcome measures |
Survival analysis |
Reference |
|---|---|---|---|---|---|---|---|---|---|
Zhou et al. |
2016 |
China |
Esophageal Cancer |
162 |
TNM stage; distant metastasis; lymph node metastases |
qRTPCR |
OS |
COX analysis |
[28] |
Zhang et al. |
2016 |
China |
Hepatocellular Carcinoma |
78 |
TNM stage; lymph node metastases |
qRTPCR |
OS |
COX analysis |
[27] |
Wang et al. |
2016 |
China |
Colorectal Cancer |
52 |
TNM stage; distant metastasis |
qRTPCR |
OS |
COX analysis |
[25] |
Lu et al. |
2017 |
China |
Cholangiocarcinoma |
56 |
TNM stage; distant metastasis |
qRTPCR |
OS |
K-M curve |
[26] |
Ma et al. |
2016 |
China |
Gallbladder Cancer |
40 |
TNM stage; lymph node metastases |
qRTPCR |
None |
None |
[31] |
Ye et al. |
2016 |
China |
Pancreatic Cancer |
90 |
TNM stage; lymph node metastases |
qRTPCR |
OS |
K-M curve |
[21] |
Deng et al. |
2015 |
China |
Lung Cancer |
121 |
TNM stage; distant metastasis; lymph node metastases |
qRTPCR |
OS |
COX analysis |
[17] |
Bo et al. |
2015 |
China |
Nasopharyngeal Carcinoma |
112 |
TNM stage |
ISH |
OS RFS |
COX |
[30] |
Luo et al. |
2016 |
China |
Esophageal Cancer |
50 |
TNM stage |
qRTPCR |
None |
None |
[18] |
Table 2: Survival characteristics of studies based on Affymetrix Human Genome U133 Plus 2.0
Type of cancer |
GEO number |
Region |
No. of patients |
Outcome measure |
HR |
p value |
|---|---|---|---|---|---|---|
Lung Cancer |
GSE31210 |
Japan |
226 |
OS |
2.276 |
0.0981 |
DLBCL |
GSE11318 |
USA |
200 |
OS |
0.7914 |
0.2436 |
Glioblastoma |
GSE7696 |
Canada |
80 |
OS |
1.319 |
0.5307 |
DLBCL |
GSE10846 |
USA |
414 |
OS |
1.727 |
0.0367 |
Meningioma |
GSE16581 |
USA |
67 |
OS |
2.175 |
0.2568 |
Ovarian Cancer |
GSE18520 |
USA |
53 |
OS |
2.362 |
0.1875 |
Melanoma |
GSE19234 |
USA |
44 |
OS |
1.526 |
0.3316 |
Ovarian Cancer |
GSE19829 |
USA |
28 |
OS |
1.351 |
0.5836 |
Breast Cancer |
GSE20711 |
Canada |
88 |
OS |
2.355 |
0.0942 |
Lymphocytic Leukemia |
GSE22762 |
Germany |
107 |
OS |
0.344 |
0.0109 |
DLBCL |
GSE23501 |
USA |
69 |
OS |
3.278 |
0.0327 |
Ovarian Cancer |
GSE30161 |
USA |
58 |
OS |
1.071 |
0.8409 |
CN-AML |
GSE12417 |
Germany |
79 |
OS |
1.814 |
0.0847 |
Lung Cancer |
GSE37745 |
Sweden |
196 |
OS |
0.644 |
0.0090 |
Lung Cancer |
GSE29013 |
USA |
55 |
OS |
1.498 |
0.4002 |
Colon Cancer |
GSE39582 |
France |
579 |
OS |
2.086 |
0.0031 |
Lung Cancer |
GSE50081 |
Canada |
181 |
OS |
0.703 |
0.2217 |
Breast Cancer |
GSE6532 |
Canada |
87 |
RFS |
3.142 |
0.0070 |
Breast Cancer |
GSE9195 |
Canada |
77 |
RFS |
4.071 |
0.0189 |
Breast Cancer |
GSE20711 |
Canada |
88 |
RFS |
2.151 |
0.0995 |
Colon Cancer |
GSE31595 |
Denmark |
33 |
RFS |
3.076 |
0.1134 |
Lung Cancer |
GSE37745 |
Sweden |
96 |
RFS |
1.399 |
0.2539 |
Association between AFAP1-AS1 and survival in 14 types of cancers
In total, 22 studies used a Cox analysis to obtain the HR of OS. A significant association was found between AFAP1-AS1 and OS in cancer patients (pooled HR: 1.58; 95% CI: 1.21–2.21, Figure 3A). Significant heterogeneity existed between studies (Tau² = 0.36; Chi² = 95.68, df = 21 (P < 0.00001); I² = 78%), and there was no significant publication bias (Egg’s test P > |t| = 0.245 > 0.05, Begg’s test Pr > |z| = 0.401) (Figure 3B). We used both the fixed effect model and random effect model and found that the results did not differ between the two models.
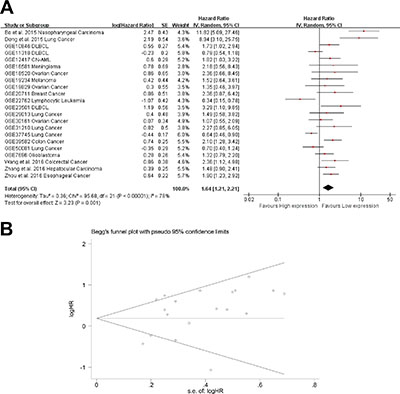
Figure 3: Meta-analysis of the independent role of AFAP1-AS1 in OS of different cancers. (A) Twenty-two studies used meta-analysis to identify the pooled HR of OS (pooled HR: 1.58; 95% CI: 1.21-2.21); (B) There was no significant publication bias for OS (Egg’s test P > |t| = 0.245 > 0.05, Begg’s test Pr > |z| = 0.401)
Due to the presence of heterogeneity, subgroups were analyzed for data source (Figure 4A) and region (Figure 4B). We found a significant association between AFAP1-AS1 and the OS of cancer patients in Asian countries (pooled HR: 3.17, 95% CI: 1.69–5.93). This association was not significant in western regions (pooled HR: 1.25, 95% CI: 0.94–1.67). AFAP1-AS1 was found to be significantly associated with the OS of cancer patients in data from published articles (pooled HR: 3.39, 95% CI: 1.64–6.99). The association was not significant for data from the GEO database (pooled HR: 1.29, 95% CI: 0.97–1.71). There was less significant heterogeneity across studies in the western subgroup (Tau² = 0.20; Chi² = 45.17, df = 15 (P < 0.0001); I² = 67%) as well as studies from the GEO database (Tau² = 0.20; Chi² = 47.22, df = 16 (P < 0.0001); I² = 66%). Greater heterogeneity existed in the Asian subgroup (Tau² = 0.46; Chi² = 24.61, df = 5 (P = 0.0002); I² = 80%) and in the data collected from published articles (Tau² = 0.55; Chi² = 24.59, df = 4 (P < 0.0001); I² = 84%).
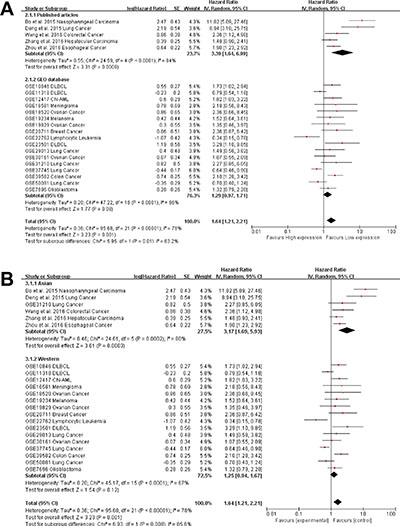
Figure 4: Subgroup analysis of independent regions and data sources in OS. (A) Subgroups were analyzed for the presence of heterogeneity based on the data source. The association between AFAP1-AS1 and OS of cancer patients was significant in the Asian population (pooled HR: 3.17, 95% CI: 1.69–5.93), but not significant in the western population (pooled HR: 1.25, 95% CI: 0.94–6.99) but not in data from the GEO database (pooled HR: 1.29, 95% CI: 0.97–.71).
To maximize clinical relevance, subgroups were analyzed based on tumor type. We found that AFAP1-AS1 was a risk factor in the prognosis of lung cancer (pooled HR: 1.54; 95% CI: 1.01–2.34), digestive system cancer (pooled HR: 1.87; 95% CI: 1.45–2.41) and nasopharyngeal carcinoma (HR: 11.82; 95% CI: 5.09–27.46). However, there was no significant association between AFAP1-AS1 and OS in lung cancer, ovarian cancer, or tumors of the nervous system. This result was strengthened by the low heterogeneity between the studies (Figure 5).
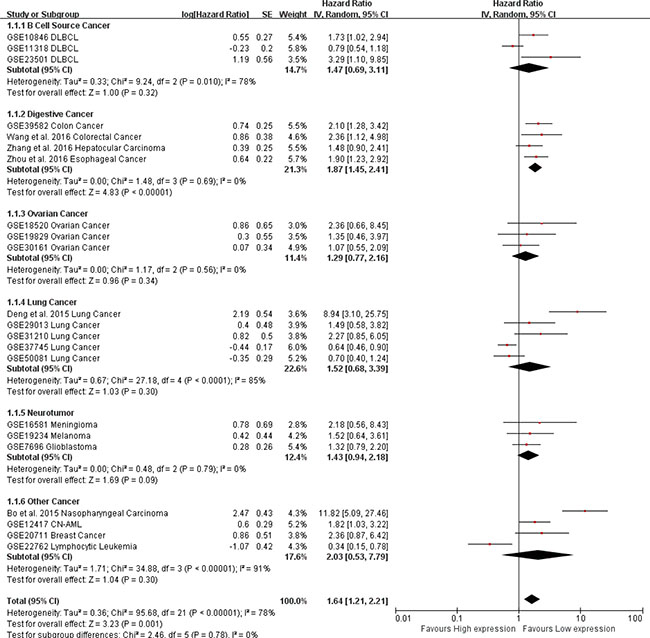
Figure 5: Subgroup analysis of tumor types in OS/RFS. AFAP1-AS1 was a risk factor in the prognosis of lung cancer (pooled HR: 1.54; 95% CI: 1.01–2.34), digestive system cancer (pooled HR: 1.87; 95% CI: 1.45–2.41) and nasopharyngeal carcinoma (HR: 11.82; 95% CI: 5.09–27.46).
The prognostic value of AFAP1-AS1 in RFS was evaluated in 6 studies. AFAP1-AS1 was significantly associated with RFS (pooled HR: 2.45, 95% CI: 1.76–3.42) (Figure 6A). There was no significant heterogeneity across the studies (Heterogeneity: Chi² = 6.96, df = 5 (P = 0.22); I² = 28%), nor was there significant publication bias (Begg’s test: Pr > |z| = 1.000, Egg’s test: P > |t| = 0.271 > 0.05) (Figure 6B). Three studies focused on breast cancer and a subgroup analysis found that AFAP1-AS1 was significantly associated with RFS in breast cancer (pooled HR = 2.90; 95% CI: 1.69–4.98; heterogeneity: Chi² = 0.76, df = 2 (P = 0.68); I² = 0%) (Figure 6C). Both fixed effect and random effect models were used to calculate the effect and the results were not markedly different between the two models.
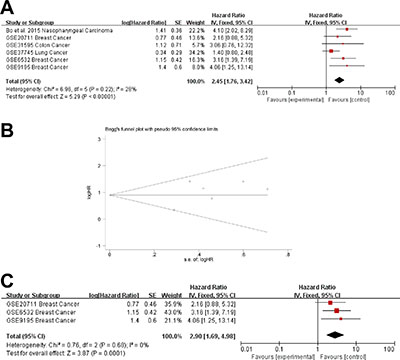
Figure 6: Meta-analysis of the independent role of AFAP1-AS1 in RFS of different cancers. (A) Six studies used meta-analysis to identify the pooled HR for RFS (pooled HR: 2.45, 95% CI: 1.76-3.42). (B) There was no significant publication bias for RFS (Begg’s test: Pr > |z| = 1.000, Egg’s test: P > |t| = 0.271 > 0.05). (C) AFAP1-AS1 was a risk factor for RFS in breast cancer (pooled HR = 2.90; 95% CI: 1.69–4.98).
Association between AFAP1-AS1 and the clinical characteristics of cancers
As shown in Table 3, 10 studies examined the association between AFAP1-AS1 and clinical characteristics in 6 types of cancer. 9 studies examined the association between TNM stage and AFAP1-AS1 in different cancers, including esophageal cancer (2), liver cancer (3), pancreatic cancer (1), lung cancer (2) and colorectal cancer (1). There was a significant association in esophageal cancer (pooled OR = 1.90; 95% CI: 1.01–3.57) and colorectal cancer (OR = 6.72; 95% CI: 1.92–23.58) but no significant association in lung cancer (pooled OR = 2.84; 95% CI: 0.12–65.10) and pancreatic cancer (OR = 2.15; 95% CI: 0.90–5.15). Association results for the two lung cancer studies were contradictory (Fixed effect pooled OR = 2.11; 95% CI: 1.25–3.55; Random effect pooled OR = 2.11; 95% CI: 0.78–5.66). There were 5 studies examining lymph node metastases in lung cancer (1), pancreatic cancer (1), liver cancer (1), esophageal cancer (1), and gallbladder cancer (1). We observed a significant association in lung cancer (OR = 2.85; 95% CI: 1.35–6.04), liver cancer (OR = 4.10; 95% CI: 1.43–11.75), pancreatic cancer (OR = 5.25; 95% CI: 2.09–13.20), and esophageal cancer (OR = 3.24; 95% CI: 1.70–6.17) but no significant association in gallbladder cancer (OR = 1.56; 95% CI: 0.44–5.53). There were 4 studies that examined distant metastases in different cancers, including lung cancer (1), liver cancer (1), colorectal cancer (1), and esophageal cancer (1). AFAP1-AS1 was significantly associated with distant metastasis in lung cancer (OR = 2.24; 95% CI: 1.08–4.67), colorectal cancer (OR = 7.50; 95% CI: 2.01–28.05), and esophageal cancer (OR = 2.98; 95% CI: 1.28–6.97) but not in liver cancer (OR = 3.33; 95% CI: 0.78–14.23). Subgroup analysis, sensitivity analysis and appraisal of publication bias were not performed due to the limited number and relative homogeneity of the studies.
Table 3: Results of meta-analysis of increased AFAP1-AS1 expression and clinical features in various cancers
Cancer types |
No. of studies |
No. of patients |
Pooled OR |
Heterogeneity |
|
|---|---|---|---|---|---|
Fixed |
Random |
I2 |
|||
TNM stage |
|||||
Liver Cancer |
3 |
290 |
2.11 [1.25, 3.55] |
2.11 [0.78, 5.66] |
68% |
Esophageal Cancer |
2 |
232 |
1.90 [1.01, 3.57] |
1.89 [1.01, 3.56] |
0% |
Pancreatic Cancer |
1 |
90 |
2.15 [0.90, 5.15] |
2.15 [0.90, 5.15] |
Not applicable |
Colorectal Cancer |
1 |
52 |
6.72 [1.92, 23.58] |
6.72 [1.92, 23.58] |
Not applicable |
Lung Cancer |
2 |
317 |
1.75 [0.98, 3.15] |
2.84 [0.12, 65.10] |
93% |
2.11 [1.56, 2.84] |
2.30 [1.30, 4.09] |
68% |
|||
Lymph node metastasis |
|||||
Gallbladder Cancer |
1 |
50 |
1.56 [0.44, 5.53] |
1.56 [0.44, 5.53] |
Not applicable |
Pancreatic Cancer |
1 |
90 |
5.25 [2.09, 13.20] |
5.25 [2.09, 13.20] |
Not applicable |
Liver Cancer |
1 |
78 |
4.10 [1.43, 11.75] |
4.10 [1.43, 11.75] |
Not applicable |
Esophageal Cancer |
1 |
162 |
3.24 [1.70, 6.17] |
3.24 [1.70, 6.17] |
Not applicable |
Lung Cancer |
1 |
121 |
2.85 [1.35, 6.04] |
2.85 [1.35, 6.04] |
Not applicable |
3.28 [2.24, 4.79] |
3.29 [2.24, 4.81] |
0% |
|||
Distant metastasis |
|||||
Lung Cancer |
1 |
121 |
2.24 [1.08, 4.67] |
2.24 [1.08, 4.67] |
Not applicable |
Liver Cancer |
1 |
56 |
3.33 [0.78, 14.23] |
3.33 [0.78, 14.23] |
Not applicable |
Colorectal Cancer |
1 |
52 |
7.50 [2.01, 28.05] |
7.50 [2.01, 28.05] |
Not applicable |
Esophageal Cancer |
1 |
162 |
2.98 [1.28, 6.97] |
2.98 [1.28, 6.97] |
Not applicable |
3.05 [1.89, 4.92] |
3.02 [1.86, 4.89] |
0% |
|||
DISCUSSION
The expression of the lncRNA AFAP1-AS1 has been shown by ourselves and others to be significantly upregulated in various tumor types. Additionally, high AFAP1-AS1 expression is closely correlated to a poor prognosis of cancer patients[ 16, 21, 27, 28, 30, 32]. In vitro experiments have shown that tumor cell characteristics, including proliferation and metastasis, can be inhibited by down-regulating AFAP1-AS1 expression. Further mechanistic studies found that AFAP1-AS1 was involved in tumorigenesis and development through the regulation of pathways including Rho/Rac [30] and PI3K-Akt [20]. On the other hand, as the antisense lncRNA of AFAP1, the second exon of AFAP1-AS1 is complementary to exons 14, 15, and 16 of AFAP1. Our group was the first to demonstrate that the specific knock-down of AFAP1-AS1 resulted in AFAP1 expression inhibition [30]. This lncRNA may be involved in the regulation of related proteins such as Src, thus affecting the cell motility, tumor invasion, and metastasis. (Figure 7)
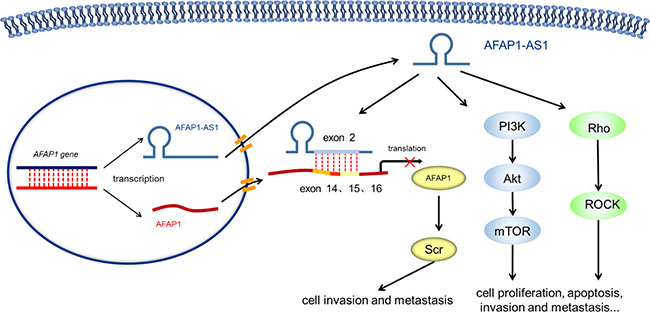
Figure 7: Molecular mechanisms of AFAP-AS1 in human cancer. AFAP1-AS1 is involved in tumorigenesis and development through the regulation of various pathways including Rho/Rac and PI3K-Akt. On the other hand, as the antisense lncRNA of AFAP1, the second exon of AFAP1-AS1 is complementary to exons 14, 15, and 16 of AFAP1. AFAP1-AS1 could inhibit AFAP1 protein translation, thus affecting cell motility, tumor invasion, and metastasis.
In our meta-analysis, we identified high expression of AFAP1-AS1 in various cancers, including 22 studies with OS data. By pooled HR values, AFAP1-AS1 was found to be an independent risk factor (pooled HR = 1.58; 95% CI: 1.21–2.21) of OS in tumor patients. In addition, 6 studies analyzed RFS and demonstrated that AFAP1-AS1 could be regarded as an independent prognostic biomarker of RFS (pooled HR = 2.45, 95% CI: 1.76–3.42).
Three previous studies found that high expression of AFAP1-AS1 correlated with TNM staging (III/IV vs. I/II) in esophageal cancer and colorectal cancer. Our study, which is based on 981 patients from five studies with TNM staging data, indicated that the correlation between high expression of AFAP1-AS1 and tumor TNM staging has a pooled OR value of 2.30 (95% CI: 1.30–4.09). However, this correlation may be possible in unstudied tumor types.
The correlation between AFAP1-AS1 expression and lymph node metastasis and distant metastasis was analyzed in five and four studies, respectively. AFAP1-AS1 was significantly associated with distant metastasis in lung cancer, colorectal cancer, and esophageal cancer but not in liver cancer. Additionally, AFAP1-AS1 was significantly associated with lymphocyte metastasis in lung cancer, liver cancer, pancreatic cancer, and esophageal cancer but not in gallbladder cancer. The pooled ORs (lymphocyte metastasis: OR = 3.28, 95% CI: 2.24–4.79, distant metastasis: HR = 3.05, 95% CI: 1.89–4.92) confirm the correlation between AFAP1-AS1 expression and lymphocyte metastasis and distant metastasis. Nevertheless, considering the inadequate sample size, our conclusions could change with the inclusion of more data.
In addition, our group recently found that AFAP1-AS1 expression was related to PD1 expression, indicating the possible participation of AFAP1-AS1 in the fascinating field of tumor immunology [15]. As the current star molecule in the field of tumor immunology, PD1 plays an important role in the immune tolerance of tumor cells [5, 33]. At present, anti-PD1 drugs (represented by Keytruda and Opdivo) have proven to be effective tumor therapies [34–38]. The close correlation between AFAP1-AS1 and PD1 indicates the potential of AFAP1-AS1 as a future target for immunological therapies and provides new avenues for treatment [39–41].
Nevertheless, there are many deficiencies in this meta-analysis. (1) Only English-language publications were included in this study, so any data demonstrating a correlation between AFAP1-AS1 and tumor prognosis in other languages were ignored. (2) Publications with positive results were more likely to get published than those with negative results, so data from negative as well as ongoing studies were underrepresented. Thus, the role of AFAP1-AS1 in tumor prognosis may be over-estimated. (3) The degree of AFAP1-AS1 expression varied between different publications, making a uniform analysis difficult. (4) Survival time was largely related to the therapeutic regimen, thus differences in treatment regimens influenced the calculation of HR values as well as study heterogeneity. (5) The variance in publication quality was one of the main factors influencing the analysis. (6) The differences between cancer sub-types were not considered in this analysis and were also rarely reported in most publications.
In conclusion, AFAP1-AS1 can act as a newfound independent biomarker for predicting cancer prognosis. However, more evidence is necessary to demonstrate the correlation between high AFAP1-AS1 expression levels and clinical indices such as TNM staging, lymphocyte metastasis, distant metastasis, and histological stage. Moreover, most patients in our study were Asian, thus our results do not reflect the global situation. Great heterogeneity remained in the study of the correlation between AFAP1-AS1 and clinical diagnosis. This can be explained by the application of different methods to classify high or low expression of AFAP1-AS1. Therefore, the conclusions from our study need to be further confirmed through multi-center randomized controlled trials with large sample sizes.
CONCLUSIONS
AFAP1-AS1 can serve as a novel molecular marker in different types of cancers. Further studies are necessary to demonstrate the correlation between high AFAP1-AS1 expression and clinical characteristics in order to improve the potential clinical benefits.
Abbreviations
lncRNAs: long noncoding RNAs; AFAP1-AS1: AFAP1 antisense RNA 1; HR: hazard ratio; OR: odds ratio; CI: Confidence interval; AFAP1: actin filament associated protein 1; OS: overall survival; RFS: recurrence-free survival; FISH: fluorescence in situ hybridization; Rho: rhodopsin; PI3K: phosphoinositide-3-kinase.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and material
Not applicable.
Author contributions
Y.W., Y.M., R.Z., X.Y., Y.Z., Y.X., W.Z., H.Z., and Y.D. collected the related publications and drafted the manuscript. Y.L., G.L., W.X. and Z.Z. participated in the design of the review and drafted the manuscript. All authors read and approved the final manuscript.
ACKNOWLEDGMENTS
We thank Dr. Mo Yang for the helpful discussion.
CONFLICTS OF INTEREST
The authors declare that they have no competing interests.
FUNDING
This work was supported in part by grants from the National Natural Science Foundation of China (81402009, 81572787, 81672683, 81702907 and 81772928), the Natural Science Foundation of Hunan Province (2016JC2035 and 2015JJ1022) and the Free Exploration Program of Central South University (ZY20170984 and 2017zzts374).
REFERENCES
1. Torre LA, Sauer AM, Chen MS Jr, Kagawa-Singer M, Jemal A, Siegel RL. Cancer statistics for Asian Americans, Native Hawaiians, and Pacific Islanders, 2016: Converging incidence in males and females. CA Cancer J Clin. 2016; 66:182–202.
2. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–132.
3. Siegel R, DeSantis C, Virgo K, Stein K, Mariotto A, Smith T, Cooper D, Gansler T, Lerro C, Fedewa S, Lin C, Leach C, Cannady RS, et al. Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin. 2012; 62:220–241.
4. Yu J, Zhang Y, Leung LH, Liu L, Yang F, Yao X. Efficacy and safety of angiogenesis inhibitors in advanced gastric cancer: a systematic review and meta-analysis. Journal of hematology & oncology. 2016; 9:111.
5. Lichtenegger FS, Krupka C, Haubner S, Kohnke T, Subklewe M. Recent developments in immunotherapy of acute myeloid leukemia. Journal of hematology & oncology. 2017; 10:142.
6. Gotwals P, Cameron S, Cipolletta D, Cremasco V, Crystal A, Hewes B, Mueller B, Quaratino S, Sabatos-Peyton C, Petruzzelli L, Engelman JA, Dranoff G. Prospects for combining targeted and conventional cancer therapy with immunotherapy. Nature reviews Cancer. 2017; 17:286–301.
7. Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, Bell I, Cheung E, Drenkow J, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007; 316:1484–1488.
8. Gong Z, Yang Q, Zeng Z, Zhang W, Li X, Zu X, Deng H, Chen P, Liao Q, Xiang B, Zhou M, Li X, Li Y, et al. An integrative transcriptomic analysis reveals p53 regulated miRNA, mRNA, and lncRNA networks in nasopharyngeal carcinoma. Tumour biology. 2016; 37:3683–3695.
9. Gong Z, Zhang S, Zhang W, Huang H, Li Q, Deng H, Ma J, Zhou M, Xiang J, Wu M, Li X, Xiong W, Li X, et al. Long non-coding RNAs in cancer. Science China Life sciences. 2012; 55:1120–1124.
10. Gong Z, Zhang S, Zeng Z, Wu H, Yang Q, Xiong F, Shi L, Yang J, Zhang W, Zhou Y, Zeng Y, Li X, Xiang B, et al. LOC401317, a p53-regulated long non-coding RNA, inhibits cell proliferation and induces apoptosis in the nasopharyngeal carcinoma cell line HNE2. PloS one. 2014; 9:e110674.
11. Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nature reviews Genetics. 2016; 17:47–62.
12. Wang Y, Mo Y, Gong Z, Yang X, Yang M, Zhang S, Xiong F, Xiang B, Zhou M, Liao Q, Zhang W, Li X, Li X, et al. Circular RNAs in human cancer. Molecular cancer. 2017; 16:25.
13. Fan C, Tang Y, Wang J, Xiong F, Guo C, Wang Y, Zhang S, Gong Z, Wei F, Yang L, He Y, Zhou M, Li X, et al. Role of long non-coding RNAs in glucose metabolism in cancer. Molecular cancer. 2017; 16:130.
14. Shi X, Zhang H, Wang M, Xu X, Zhao Y, He R, Zhang M, Zhou M, Li X, Peng F, Shi C, Shen M, Wang X, et al. LncRNA AFAP1-AS1 promotes growth and metastasis of cholangiocarcinoma cells. Oncotarget. 2017; 8:58394–58404. https://doi.org/10.18632/oncotarget.16880.
15. Tang Y, He Y, Shi L, Yang L, Wang J, Lian Y, Fan C, Zhang P, Guo C, Zhang S, Gong Z, Li X, Xiong F, et al. Co-expression of AFAP1-AS1 and PD-1 predicts poor prognosis in nasopharyngeal carcinoma. Oncotarget. 2017; 8:39001–39011. https://doi.org/10.18632/oncotarget.16545.
16. Zeng Z, Bo H, Gong Z, Lian Y, Li X, Li X, Zhang W, Deng H, Zhou M, Peng S, Li G, Xiong W. AFAP1-AS1, a long noncoding RNA upregulated in lung cancer and promotes invasion and metastasis. Tumour biology. 2016; 37:729–737.
17. Deng J, Liang Y, Liu C, He S, Wang S. The up-regulation of long non-coding RNA AFAP1-AS1 is associated with the poor prognosis of NSCLC patients. Biomedicine & pharmacotherapy. 2015; 75:8–11.
18. Luo HL, Huang MD, Guo JN, Fan RH, Xia XT, He JD, Chen XF. AFAP1-AS1 is upregulated and promotes esophageal squamous cell carcinoma cell proliferation and inhibits cell apoptosis. Cancer medicine. 2016; 5:2879–2885.
19. Lu X, Zhou C, Li R, Deng Y, Zhao L, Zhai W. Long Noncoding RNA AFAP1-AS1 Promoted Tumor Growth and Invasion in Cholangiocarcinoma. Cellular physiology and biochemistry. 2017; 42:222–230.
20. Guo JQ, Li SJ, Guo GX. Long Noncoding RNA AFAP1-AS1 Promotes Cell Proliferation and Apoptosis of Gastric Cancer Cells via PTEN/p-AKT Pathway. Digestive diseases and sciences. 2017; 62:2004–2010.
21. Ye Y, Chen J, Zhou Y, Fu Z, Zhou Q, Wang Y, Gao W, Zheng S, Zhao X, Chen T, Chen R. High expression of AFAP1-AS1 is associated with poor survival and short-term recurrence in pancreatic ductal adenocarcinoma. Journal of translational medicine. 2015; 13:137.
22. Steels E, Paesmans M, Berghmans T, Branle F, Lemaitre F, Mascaux C, Meert AP, Vallot F, Lafitte JJ, Sculier JP. Role of p53 as a prognostic factor for survival in lung cancer: a systematic review of the literature with a meta-analysis. The European respiratory journal. 2001; 18:705–719.
23. Xing X, Tang YB, Yuan G, Wang Y, Wang J, Yang Y, Chen M. The prognostic value of E-cadherin in gastric cancer: a meta-analysis. International journal of cancer. 2013; 132:2589–2596.
24. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. Bmj. 1997; 315:629–634.
25. Wang F, Ni H, Sun F, Li M, Chen L. Overexpression of lncRNA AFAP1-AS1 correlates with poor prognosis and promotes tumorigenesis in colorectal cancer. Biomedicine & pharmacotherapy. 2016; 81:152–159.
26. Lu X, Zhou C, Li R, Liang Z, Zhai W, Zhao L, Zhang S. Critical role for the long non-coding RNA AFAP1-AS1 in the proliferation and metastasis of hepatocellular carcinoma. Tumour biology. 2016; 37:9699–9707.
27. Zhang JY, Weng MZ, Song FB, Xu YG, Liu Q, Wu JY, Qin J, Jin T, Xu JM. Long noncoding RNA AFAP1-AS1 indicates a poor prognosis of hepatocellular carcinoma and promotes cell proliferation and invasion via upregulation of the RhoA/Rac2 signaling. International journal of oncology. 2016; 48:1590–1598.
28. Zhou XL, Wang WW, Zhu WG, Yu CH, Tao GZ, Wu QQ, Song YQ, Pan P, Tong YS. High expression of long non-coding RNA AFAP1-AS1 predicts chemoradioresistance and poor prognosis in patients with esophageal squamous cell carcinoma treated with definitive chemoradiotherapy. Molecular carcinogenesis. 2016; 55:2095–2105.
29. Wu W, Bhagat TD, Yang X, Song JH, Cheng Y, Agarwal R, Abraham JM, Ibrahim S, Bartenstein M, Hussain Z, Suzuki M, Yu Y, Chen W, et al. Hypomethylation of noncoding DNA regions and overexpression of the long noncoding RNA, AFAP1-AS1, in Barrett’s esophagus and esophageal adenocarcinoma. Gastroenterology. 2013; 144:956–966 e954.
30. Bo H, Gong Z, Zhang W, Li X, Zeng Y, Liao Q, Chen P, Shi L, Lian Y, Jing Y, Tang K, Li Z, Zhou Y, et al. Upregulated long non-coding RNA AFAP1-AS1 expression is associated with progression and poor prognosis of nasopharyngeal carcinoma. Oncotarget. 2015; 6:20404–20418. https://doi.org/10.18632/oncotarget.4057.
31. Ma F, Wang SH, Cai Q, Zhang MD, Yang Y, Ding J. Overexpression of LncRNA AFAP1-AS1 predicts poor prognosis and promotes cells proliferation and invasion in gallbladder cancer. Biomedicine & pharmacotherapy. 2016; 84:1249–1255.
32. He B, Zeng J, Chao W, Chen X, Huang Y, Deng K, Huang Z, Li J, Dai M, Chen S, Huang H, Dai S. Serum long non-coding RNAs MALAT1, AFAP1-AS1 and AL359062 as diagnostic and prognostic biomarkers for nasopharyngeal carcinoma. Oncotarget. 2017; 8:41166–41177. https://doi.org/10.18632/oncotarget.17083.
33. Arasanz H, Gato-Canas M, Zuazo M, Ibanez-Vea M, Breckpot K, Kochan G, Escors D. PD1 signal transduction pathways in T cells. Oncotarget. 2017; 8:51936–51945. https://doi.org/10.18632/oncotarget.17232.
34. Tu C, Zeng Z, Qi P, Li X, Yu Z, Guo C, Xiong F, Xiang B, Zhou M, Gong Z, Liao Q, Yu J, He Y, et al. Genome-Wide Analysis of 18 Epstein-Barr Viruses Isolated from Primary Nasopharyngeal Carcinoma Biopsy Specimens. Journal of virology. 2017; 91.
35. Wang M, Zhao J, Zhang L, Wei F, Lian Y, Wu Y, Gong Z, Zhang S, Zhou J, Cao K, Li X, Xiong W, Li G, et al. Role of tumor microenvironment in tumorigenesis. Journal of Cancer. 2017; 8:761–773.
36. Yu J, Liu Y, Guo C, Zhang S, Gong Z, Tang Y, Yang L, He Y, Lian Y, Li X, Deng H, Liao Q, Li X, et al. Upregulated long non-coding RNA LINC00152 expression is associated with progression and poor prognosis of tongue squamous cell carcinoma. Journal of Cancer. 2017; 8:523–530.
37. Tang Y, Wang J, Lian Y, Fan C, Zhang P, Wu Y, Li X, Xiong F, Li X, Li G, Xiong W, Zeng Z. Linking long non-coding RNAs and SWI/SNF complexes to chromatin remodeling in cancer. Molecular cancer. 2017; 16:42.
38. Xu K, Xiong W, Zhou M, Wang H, Yang J, Li X, Chen P, Liao Q, Deng H, Li X, Li G, Zeng Z. Integrating ChIP-sequencing and digital gene expression profiling to identify BRD7 downstream genes and construct their regulating network. Molecular and cellular biochemistry. 2016; 411:57–71.
39. Yang L, Tang Y, He Y, Wang Y, Lian Y, Xiong F, Shi L, Zhang S, Gong Z, Zhou Y, Liao Q, Zhou M, Li X, et al. High Expression of LINC01420 indicates an unfavorable prognosis and modulates cell migration and invasion in nasopharyngeal carcinoma. Journal of Cancer. 2017; 8:97–103.
40. Yu J, Liu Y, Gong Z, Zhang S, Guo C, Li X, Tang Y, Yang L, He Y, Wei F, Wang Y, Liao Q, Zhang W, et al. Overexpression long non-coding RNA LINC00673 is associated with poor prognosis and promotes invasion and metastasis in tongue squamous cell carcinoma. Oncotarget. 2017; 8:16621–16632. https://doi.org/10.18632/oncotarget.14200.
41. He B, Li W, Wu Y, Wei F, Gong Z, Bo H, Wang Y, Li X, Xiang B, Guo C, Liao Q, Chen P, Zu X, et al. Epstein-Barr virus-encoded miR-BART6–3p inhibits cancer cell metastasis and invasion by targeting long non-coding RNA LOC553103. Cell death & disease. 2016; 7:e2353.

