INTRODUCTION
Esophageal cancer is one of the most common cancers worldwide, and carries a high mortality after diagnosis following the onset of symptoms [1]. Cancer of the esophagus occurs in two major histological forms, esophageal squamous cell carcinoma (ESCC) and esophageal adenocarcinoma (EAC). ESCC dominates in most parts of the world, especially in high-risk areas such as China, where it accounts for about 90% of the total esophageal cancer cases [2, 3]. Smoking and alcohol consumption are related with more than 90% of ESCC patients in the western countries [4, 5], but the role of smoking and alcohol consumption is less important in China. The risk factors for ESCC in China include poor nutrition, lack of fruit and vegetables, drinking hot beverages and opium [3, 6].
The peptidylarginine deiminase IV (PADI4 or PAD4) converts arginine residues at histone tails to citrulline [7]. PADI4 has been demonstrated to co-localize with cytokeratin, an intermediate filament protein that plays a role during cell differentiation and apoptosis [8–10]. In cancer, high PADI4 expression has been connected to tumor growth [11], as PADI4 was overexpressed in numerous malignant cancers, but not in healthy tissues [8]. Recent study using immunohistochemistry further verified a significant PADI4 expression in various malignancies, comprising esophageal squamous cancer cells [12]. Consistently, PADI4 level in the blood increased dramatically in the patients with various malignant tumors, but considerably declined after tumor excision surgery [12]. Notably, PADI4 can disrupt the apoptotic process via the citrullination of histone H3 in the promoter of p53-target genes [13]. Therefore, we postulated that PADI4 might play an important role in the carcinogenesis of the esophageal cancer.
Single nucleotide polymorphisms (SNPs) account for more than 90% genetic variations. Despite the evidence described above indicated a correlation between PADI4 and ESCC, few molecular epidemiological studies have explored the relationship between PADI4 SNPs and susceptibility of ESCC with inconsistent results [13]. In a small cohort of esophageal cancer patients (including ESCC and EAC), PADI4 rs10437048 and rs41265997 were found significantly associated with the risk of esophageal cancer [13]. To specifically examine the potential associations between genetic variants in PADI4 and ESCC risk, we studied the correlation with the tagging SNP strategy in a larger cohort of 629 subjects of ESCC and 686 controls.
RESULTS
Characteristics of the study population
Characteristics of cases and controls included in the study are summarized in Table 1. The cases and controls appeared to be adequately matched on age and sex as suggested by the χ2 test. As shown in Table 1, significant difference was detected on smoking status (p<0.001) between the cases and the controls, and drinking rate (p<0.001) was higher in ESCC patients than in control subjects.
Table 1: Distribution of selected demographic variables and risk factors in ESCC cases and controls
Variable |
Cases (n=629) |
Controls (n=686) |
p a |
||
|---|---|---|---|---|---|
n |
% |
n |
% |
||
Age (years) mean ± SD |
62.85 (±8.13) |
62.58 (±7.89) |
0.541 |
||
Age (years) |
0.155 |
||||
< 63 |
310 |
49.28 |
365 |
53.21 |
|
≥ 63 |
319 |
50.72 |
321 |
46.79 |
|
Sex |
0.185 |
||||
Male |
444 |
70.59 |
461 |
67.20 |
|
Female |
185 |
29.41 |
225 |
32.80 |
|
Tobacco use |
<0.001 |
||||
Never |
355 |
56.44 |
499 |
72.74 |
|
Ever |
274 |
43.56 |
187 |
27.26 |
|
Alcohol use |
<0.001 |
||||
Never |
428 |
68.04 |
526 |
76.68 |
|
Ever |
201 |
31.96 |
160 |
23.32 |
|
a Two-sided χ2 test and student t test; Bold values are statistically significant (p <0.05).
Associations between PADI4 tagging polymorphisms and risk of ESCC
The seven tagging SNPs were selected on the basis of their pairwise linkage disequilibrium (LD) with the r2 threshold of 0.8 and minor allele frequency (MAF) ≥0.05 to capture all the common SNPs. Among eligible SNPs, linkage disequilibrium analysis was performed in the Chinese Han population (https://www.ncbi.nlm.nih.gov/variation/tools/1000genomes/), and the SNP loci with moderate correlation were chosen for further analyses. The LD structure across the PADI4 genomic region was presented, and three blocks were defined (Figure 1). Next, we applied the “block-based” method, which exploits the principle of linkage disequilibrium observed within haplotype blocks, to search for tag SNPs. Several algorithms have been devised to partition chromosomal regions into haplotype blocks that are based on haplotype diversity, LD, four-gamete test and information complexity. We then used online database to predict the function of SNPs (http://www.regulomedb.org/) and selected seven tag SNPs for analysis (See Figure 1).
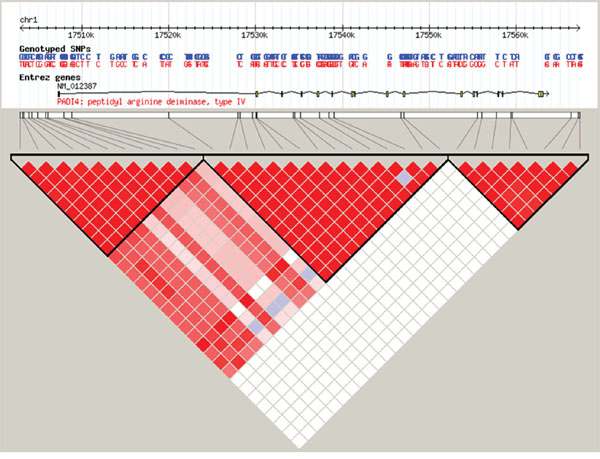
Figure 1: Linkage disequilibrium structure across the 50 kb region is represented, based on r2 coefficient calculated with the HapMap database. The middle panel shows the genomic structure of the human PADI4 gene. Exons are indicated by the vertical black bars. The genotyped tag SNPs are indicated with black bars. |D’| varies between 0 (no disequilibrium) and 1 (maximum disequilibrium), represented by shades of blue to white to pink to red. Blue:|D’| = 0 and red:|D’| = 1.
As shown in Table 2, the genotyping successful rates were ranging from 95.13% to 98.47%. In the control subjects, the genotype frequencies for these seven polymorphisms reached Hardy-Weinberg equilibrium (p-value for HWE, all p>0.05). The minor allele frequency (MAF) in our controls was comparable with the Chinese cohort in database for all seven SNPs loci.
Table 2: Primary information for PADI4 rs11203366, rs1886302, rs1635562, rs1635564, rs16825533, rs2240337, rs2477137 polymorphisms
Genotyped SNPs |
rs11203366 |
rs1886302 |
rs1635562 |
rs1635564 |
rs16825533 |
rs2240337 |
rs2477137 |
|---|---|---|---|---|---|---|---|
Ancestral Allele |
G |
T |
A |
C |
A |
G |
C |
Chromosome |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
Gene (ID) |
PADI4 (23569) |
PADI4 (23569) |
PADI4 (23569) |
PADI4 (23569) |
PADI4 (23569) |
PADI4 (23569) |
PADI4 (23569) |
Function |
Missense |
Intron region |
Intron region |
Intron region |
Intron region |
Intron region |
Intergene region |
Chr Pos (Genome Build 38.p7) |
17331039 |
17308901 |
17360325 |
17357031 |
17339386 |
17347727 |
17304110 |
Regulome DB Scorea |
No Data |
4 |
4 |
No Data |
4 |
5 |
4 |
TFBSb |
— |
Y |
— |
— |
— |
— |
Y |
nsSNP |
Y |
— |
— |
— |
— |
— |
— |
MAFc for Chinese in database |
0.256 |
0.268 |
0.354 |
0.232 |
0.061 |
0.073 |
0.146 |
MAF in our controls (n = 608) |
0.241 |
0.332 |
0.323 |
0.199 |
0.091 |
0.061 |
0.189 |
p value for HWEd test in our controls |
0.194 |
0.924 |
0.821 |
0.455 |
0.513 |
0.055 |
0.488 |
Genotyping methode |
LDR |
LDR |
LDR |
LDR |
LDR |
LDR |
LDR |
% Genotyping value |
96.42% |
96.80% |
96.34% |
95.13% |
98.47% |
95.13% |
98.47% |
bTFBS: Transcription Factor Binding Site (https://snpinfo.niehs.nih.gov/cgi-bin/snpinfo/snpfunc.cgi);
cMAF: minor allele frequency;
dHWE: Hardy–Weinberg equilibrium;
eLDR: ligation detection reaction
The genotype distributions of PADI4 SNPs in the cases and the controls are shown in Table 3. When the PADI4 rs2240337 G>A SNP GG homozygote genotype (AA) was used as the reference group, both the GA heterozygote genotype (AB) and the AA mutated homozygote genotype (BB) were associated with a significantly decreased risk of ESCC (AB vs. AA: adjusted OR = 0.52, 95% CI = 0.39-0.71, p<0.0001; BB vs. AA: adjusted OR = 0.30, 95% CI = 0.13-0.68, p = 0.004). Logistic regression analyses revealed that the PADI4 rs11203366 A>G, rs1886302 T>C, rs1635562 A>T, rs1635564 C>A, rs16825533 A>G, and rs2477137 C>A polymorphisms were not associated with the risk of ESCC. After the Bonferroni correction, for PADI4 rs2240337 G>A, the padj = 0.031 for GA vs. GG after adjusted for age, sex, smoking and drinking status. padj < 0.001 for AA vs. GG. None of the rest 6 SNPs, showed significant associations with ESCC in this study population (padj > 0.05 in all comparison models).
Table 3: Main effects of PADI4 SNPs on ESCC risk
Genotyped SNPs |
Genotyping |
AB vs. AA b Adjusted ORc (95% CI); p |
BB vs. AA Adjusted OR (95% CI); p |
p trend |
|
|---|---|---|---|---|---|
Case (n=629) (AA/AB/BB) a |
Control (n=686) (AA/AB/BB) |
||||
PADI4: rs11203366 A>G |
219/293/103 |
214/301/138 |
1.00 (0.78–1.29);0.985 |
0.77 (0.56–1.07);0.117 |
0.128 |
PADI4: rs1886302 T>C |
250/273/77 |
295/308/70 |
1.09 (0.86–1.39);0.487 |
1.37 (0.94–1.99);0.100 |
0.372 |
PADI4: rs1635562 A>T |
295/251/64 |
302/285/70 |
0.90 (0.71–1.15);0.406 |
0.91 (0.62–1.34);0.632 |
0.682 |
PADI4: rs1635564 C>A |
388/180/32 |
420/202/29 |
1.02 (0.80–1.31);0.860 |
1.22 (0.72–2.07);0.470 |
0.739 |
PADI4: rs16825533 A>G |
528/85/6 |
560/109/7 |
0.86 (0.63–1.18);0.349 |
0.97 (0.32–2.98);0.957 |
0.477 |
PADI4: rs2240337 G>A |
506/86/8 |
466/161/24 |
0.52 (0.39–0.71);<0.0001 |
0.30 (0.13–0.68);0.004 |
<0.0001 |
PADI4: rs2477137 C>A |
399/202/18 |
447/202/27 |
1.15 (0.90–1.47);0.256 |
0.76 (0.40–1.41);0.381 |
0.365 |
aAA/AB/BB means homozygote, heterozygote and mutated homozygote; b Bonferroni correction was performed to correct the p value (padj); For PADI4: rs2240337 G>A, the padj = 0.031 for GA vs. GG, padj < 0.001 for AA vs. GG, padj < 0.0001 for p trend. For the rest 6 SNPs, padj> 0.05 in all comparison models; Bold values are statistically significant (p <0.05); c Adjusted for age, sex, smoking and drinking status.
Associations between PADI4 rs2240337 polymorphism and pathologic character of ESCC
Furthermore, we analyzed the correlation between PADI4 rs2240337 G>A SNP and the clinic pathologic state. However, PADI4 rs2240337 G>A SNP did not correlate with clinical tumor stage (p = 0.215) or grade (p = 0.497) (Table 4).
Table 4: Distribution of clinic pathologic characters by PADI4 rs2240337 genotyping
Genotyping |
X2 |
p |
|||
|---|---|---|---|---|---|
AA |
AG |
GG |
|||
Pathologic grade |
|||||
1 |
4 |
22 (12.15%) |
155 (85.64%) |
3.38 |
0.496 |
2 |
4 |
53 (15.68%) |
281 (83.14%) |
||
3 |
0 |
11 (13.58%) |
70 (86.42%) |
||
Clinic stage |
|||||
1 |
3 |
14 (11.76%) |
102 (85.71%) |
8.34 |
0.215 |
2 |
1 |
42 (14.63%) |
244 (85.02%) |
||
3 |
2 |
25 (16.56%) |
124 (82.12%) |
||
4 |
2 |
5 (11.63%) |
36 (83.72%) |
||
Stratification analyses of seven polymorphisms and risk of ESCC
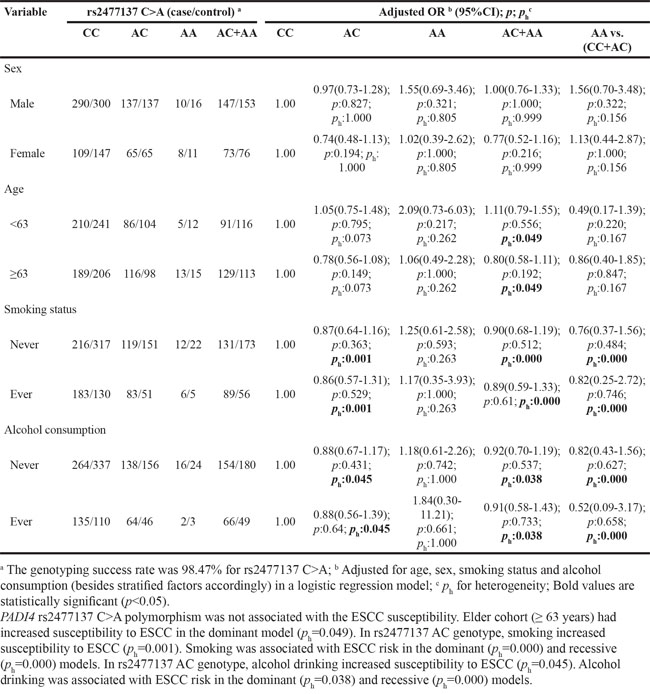
To further evaluate the effects of these seven SNPs on the risk of ESCC according to different age, gender, smoking and alcohol drinking status, stratification analyses were performed as shown in Table 5–11. We showed that smoking significantly increased ESCC risk in PADI4 rs11203366 AG/AA, rs1886302 CC/CT, rs1635562 AT, rs1635564 CA, rs2240337 AG and rs2477137 AC genotype. Alcohol drinking increased ESCC risk in PADI4 rs11203366 AG, rs1635562 AT, rs1635564 CA, rs2477137 AC, rs1886302 CT genotype. In younger cohort (<63 years), PADI4 rs16825533 AG genotype was associated with decreased risk of ESCC, while rs11203366 AA genotype was associated with increased risk of ESCC. In the non-drinking cohort, PADI4 rs11203366 AA variant was associated with increased risk of ESCC. PADI4 rs1886302 CC variant was associated with ESCC susceptibility in female cohort. In the non-alcohol drinking cohort, PADI4 rs1886302 CC and CT variants were associated with decreased risk of ESCC. In rs1635562 TT subgroup, elder people (≥63 years) were more susceptible to ESCC.
Table 5: Stratified analyses between PADI4 rs11203366 A>G polymorphism and ESCC risk by sex, age, smoking status and alcohol consumption
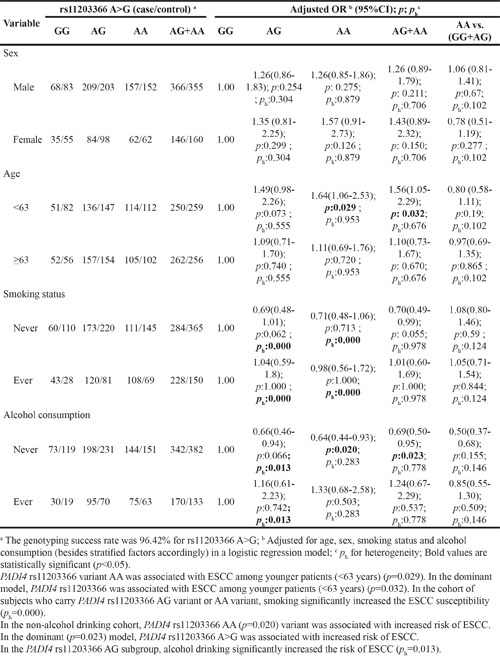
Table 6: Stratified analyses between PADI4 rs1886302 T>C polymorphism and ESCC risk by sex, age, smoking status and alcohol consumption
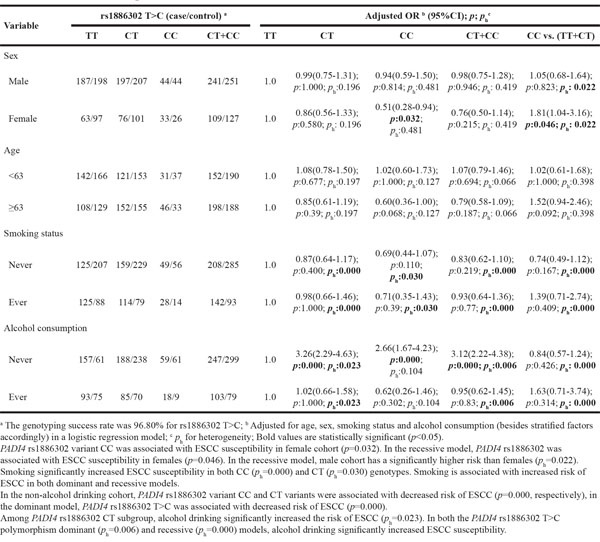
Table 7: Stratified analyses between PADI4 rs1635562 A>T polymorphism and ESCC risk by sex, age, smoking status and alcohol consumption
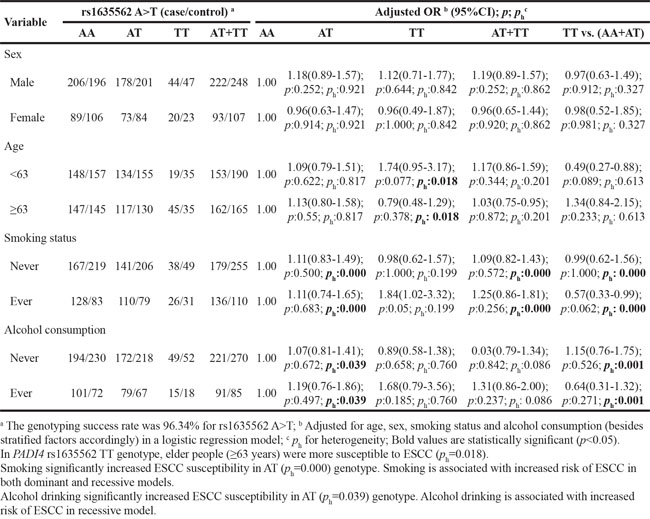
Table 8: Stratified analyses between PADI4 rs1635564 C>A polymorphism and ESCC risk by sex, age, smoking status and alcohol consumption
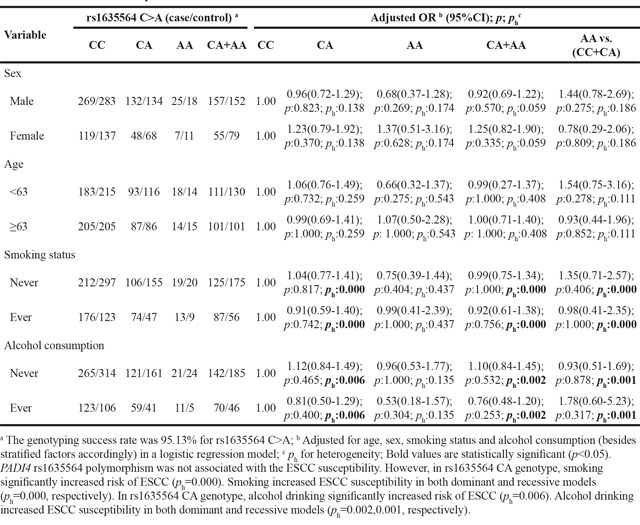
Table 9: Stratified analyses between PADI4 rs16825533 A>G polymorphism and ESCC risk by sex, age, smoking status and alcohol consumption
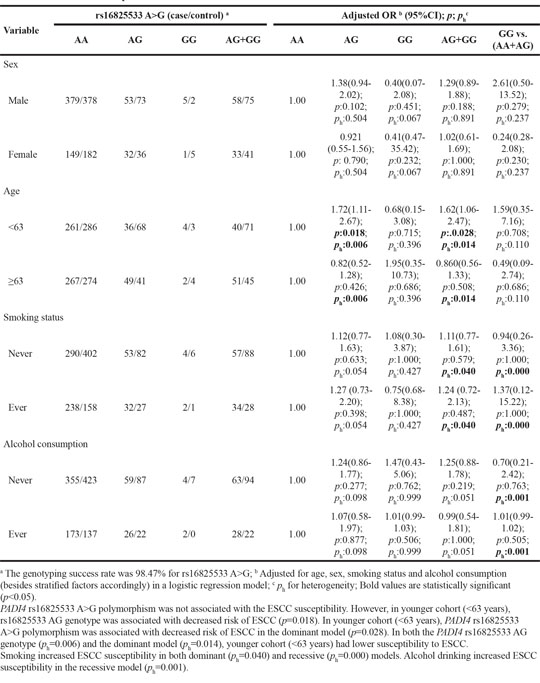
Table 10: Stratified analyses between PADI4 polymorphism rs2240337 G>A and ESCC risk by sex, age, smoking status and alcohol consumption
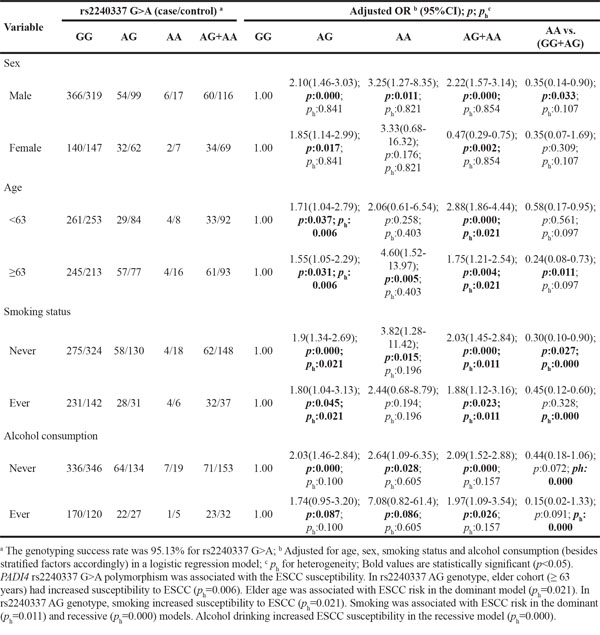
Table 11: Stratified analyses between PADI4 polymorphism rs2477137 and ESCC risk by sex, age, smoking status and alcohol consumption
Linkage disequilibrium analyses and association test
Linkage disequilibrium analyses in both controls and cases were conducted as shown in Table 12–13, there were correlations between these seven loci. Association test was performed using Haploview software (v 4.2), there were associations between these seven loci (Figure 2).
Table 12: Linkage disequilibrium analyses of PADI4 rs11203366, rs1886302, rs1635562, rs1635564, rs16825533, rs2240337, rs2477137 in control group
D‘: |
rs1886302 |
rs11203366 |
rs16825533 |
rs2240337 |
rs1635564 |
rs1635562 |
rs2477137 |
1 |
0.803 |
0.571 |
0.485 |
0.211 |
0.47 |
rs1886302 |
- |
0.764 |
0.509 |
0.508 |
0.211 |
0.209 |
rs11203366 |
- |
- |
0.978 |
0.525 |
0.445 |
0.28 |
rs16825533 |
- |
- |
- |
0.485 |
0.5 |
0.509 |
rs2240337 |
- |
- |
- |
- |
0.039 |
0.303 |
rs1635564 |
- |
- |
- |
- |
- |
0.836 |
r2: |
rs1886302 |
rs11203366 |
rs16825533 |
rs2240337 |
rs1635564 |
rs1635562 |
rs2477137 |
0.464 |
0.19 |
0.139 |
0.19 |
0.042 |
0.025 |
rs1886302 |
- |
0.367 |
0.051 |
0.097 |
0.022 |
0.011 |
rs11203366 |
- |
- |
0.124 |
0.065 |
0.065 |
0.03 |
rs16825533 |
- |
- |
- |
0.004 |
0.099 |
0.012 |
rs2240337 |
- |
- |
- |
- |
0 |
0.009 |
rs1635564 |
- |
- |
- |
- |
- |
0.084 |
D'>0, r2>0: There were linkage disequilibrium correlations among different loci; D'>0.7, r2>0.3: there were closer linkage disequilibrium correlation among different loci.
Table 13: Linkage disequilibrium analyses of PADI4 rs11203366, rs1886302, rs1635562, rs1635564, rs16825533, rs2240337, rs2477137 in case group
D‘: |
rs1886302 |
rs11203366 |
rs16825533 |
rs2240337 |
rs1635564 |
rs1635562 |
rs2477137 |
0.942 |
0.619 |
0.717 |
0.766 |
0.085 |
0.42 |
rs1886302 |
- |
0.658 |
0.687 |
0.653 |
0.18 |
0.214 |
rs11203366 |
- |
- |
1 |
0.748 |
0.245 |
0.251 |
rs16825533 |
- |
- |
- |
0.997 |
0.319 |
0.07 |
rs2240337 |
- |
- |
- |
- |
0.445 |
0.661 |
rs1635564 |
- |
- |
- |
- |
- |
0.879 |
r2: |
rs1886302 |
rs11203366 |
rs16825533 |
rs2240337 |
rs1635564 |
rs1635562 |
rs2477137 |
0.381 |
0.133 |
0.184 |
0.232 |
0.007 |
0.019 |
rs1886302 |
- |
0.341 |
0.072 |
0.072 |
0.015 |
0.011 |
rs11203366 |
- |
- |
0.123 |
0.078 |
0.023 |
0.019 |
rs16825533 |
- |
- |
- |
0.008 |
0.034 |
0 |
rs2240337 |
- |
- |
- |
- |
0.005 |
0.018 |
rs1635564 |
- |
- |
- |
- |
- |
0.089 |
D'>0, r2>0: There were linkage disequilibrium correlations among different loci; D'>0.7, r2>0.3: there were closer linkage disequilibrium correlation among different loci.
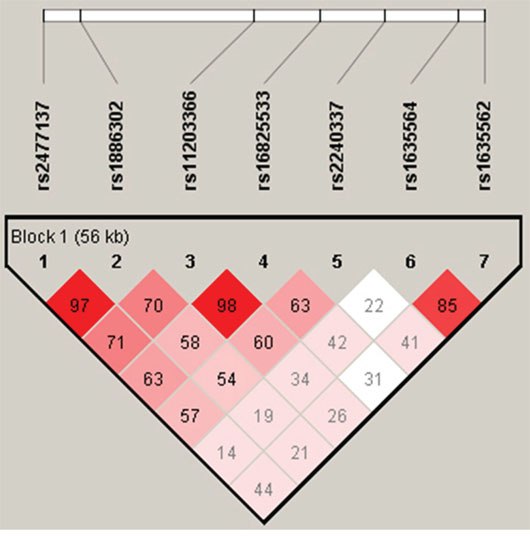
Figure 2: Association test of seven PADI4 SNPs (by Haploview Software, V 4. 2). There are associations between these seven loci.
Haplotype analyses of PADI4 polymorphisms and susceptibility to ESCC
As shown in Table 14, haplotype analyses showed that PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Crs1635564Ars1635562 was the most common haplotype in both groups (24.5% in controls, 25.5% in cases). The haplotype PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 frequency and PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 frequency were significantly lower in ESCC cases as compared with controls (0.019 vs. 0.036, p=0.007; 0.019 vs. 0.031, p=0.038, respectively), suggesting that both PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 and PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 haplotypes may be correlated with decreased susceptibility of ESCC (OR=0.491, 95%CI:0.290-0.831; OR=0.568, 95%CI:0.330-0.975, respectively). Haplotype PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 frequency was significantly higher in ESCC cases as compared with controls (0.073 vs. 0.049, p=0.042), suggesting that haplotype PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 genetic polymorphism may be correlated with increased susceptibility of ESCC (OR=1.435, 95%CI: 1.011-2.037).
Table 14: PADI4 haplotype frequencies (%) in cases and controls and risk of ESCC
Haplotypes |
Case (freq) |
Control (freq) |
Crude OR (95% CI) |
p |
|---|---|---|---|---|
PADI4 Ars2477137Crs1886302Grs11203366Ars16825533Ars2240337Crs1635564Ars1635562 |
63 (0.056) |
65 (0.055) |
0.964 [0.673~1.383] |
0.844 |
PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 |
22 (0.019) |
43 (0.036) |
0.491 [0.290~0.831] |
0.007 |
PADI4 Crs2477137Crs1886302Grs11203366Ars1635564Grs2240337Ars1635564Ars1635562 |
44 (0.040) |
28 (0.024) |
1.599 [0.989~2.585] |
0.054 |
PADI4 Crs2477137Crs1886302Grs11203366Ars1635564Grs2240337Crs1635564Ars1635562 |
56 (0.050) |
47 (0.040) |
1.209 [0.811~1.803] |
0.351 |
PADI4 Crs2477137Crs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 |
50 (0.044) |
50 (0.043) |
0.985 [0.658~1.474] |
0.941 |
PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 |
81 (0.073) |
58 (0.049) |
1.435 [1.011~2.037] |
0.042 |
PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Crs1635564Ars1635562 |
285 (0.255) |
288 (0.245) |
0.984 [0.807~1.201] |
0.877 |
PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Crs1635564Trs1635562 |
212 (0.190) |
216 (0.183) |
0.981 [0.789~1.219] |
0.859 |
PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Ars1635562 |
60 (0.054) |
63 (0.053) |
0.960 [0.665~1.385] |
0.827 |
PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 |
21 (0.019) |
37 (0.031) |
0.568 [0.330~0.975] |
0.038 |
Haplotypes were composited by PADI4 rs2477137, rs1886302, rs11203366, rs16825533, rs2240337, rs1635564 and rs1635562
All those frequency <0.03 were ignored in analysis
Haplotype PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 frequency was significantly lower in ESCC cases as compared with controls (0.019 vs. 0.036, p=0.007), suggesting that haplotype PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 genetic polymorphism may be correlated with decreased susceptibility of ESCC (OR=0.491, 95%CI: 0.290-0.831).
Haplotype PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 frequency was significantly higher in ESCC cases as compared with controls (0.073 vs. 0.049, p=0.042), suggesting that haplotype PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 genetic polymorphism may be correlated with increased susceptibility of ESCC (OR=1.435, 95%CI: 1.011-2.037).
Haplotype PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 frequency was significantly lower in ESCC cases as compared with controls (0.019 vs. 0.031, p=0.038), suggesting that haplotype PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 genetic polymorphism may be correlated with a decreased susceptibility of ESCC (OR=0.568, 95%CI: 0.330-0.975).
Power calculation
The power calculation was performed by “Power and Sample Size Calculation” Software (http://biostat.mc.vanderbilt.edu/wiki/Main/PowerSampleSize). Based on the assumption that the type I error probability for a two sided test (α) equals 0.05, the probability of exposure in controls p0 is 0.0698 in rs2240337 in the Chinese Han population according to the NCBI project. In the current study, using ligation detection reaction method, the successful rates of genotyping all exceeded 95%. There were 1,200 alleles successfully genotyped. The ratio of control/case (m) equals 1.085, and the correlation coefficient for exposure between matched case and controls (f) is 2.058 in rs2240337. The power value is 1.000.
DISCUSSION
In this hospital-based case-control epidemiological study in a Chinese population, we investigated whether tagging SNPs in PADI4 were associated with risk of developing ESCC. We found that the PADI4 rs2240337 G>A SNP was significantly associated with decreased risk of ESCC after the Bonferroni correction. PADI4 rs11203366, rs1886302, rs1635562, rs1635564 and rs2477137 polymorphisms were implicated with altered susceptibility of ESCC according to age, gender, smoking and alcohol drinking stratification analyses.
Recently, PADI4 has emerged as a novel transcriptional corepressor [14–16]. This enzyme catalyzes the posttranslational modification of arginine residues (to form citrulline) in histones H2A, H3, and H4 at the estrogen-regulated pS2 promoter [15–17] and at the apoptosis-related gene promoters p21 and OKL38 [14, 18], thereby repressing gene transcription. Additionally, the histone deaminating activity of PADI4 has been shown to downregulate the expression of numerous p53-dependent genes, including p21, PUMA, and GADD45 [14, 18]. PADI4 is overexpressed in numerous malignant cancers (e.g., breast, metastatic carcinomas, colon, bladder, lung, ovarian, and many others). In parallel, under normal circumstances, PADI4 exists as an intracellular protein, but in patients with malignant tumors, PADI4 can be detected in the plasma [16]. The PADI4 in blood increased in the presence of tumor and decreased after the tumor excision [12]. These studies bolstered the pathogenic role of PADI4 during carcinogenesis. Furthermore, expression of PADI4 was detected in esophageal cancer, but not in normal tissues. Significantly, PADI4 levels were positively correlated with the pathological classification of esophageal cancer [13].
In the present study, seven PADI4 gene variations in Chinese population were tested and associations between these variations and outcomes in ESCC were explored. Of the seven SNPs, rs2240337 G>A was validated as an ESCC susceptibility locus, showing highly significant evidence both in heterozygote group (p<0.0001) and homozygote group (p<0.004). A previous study in a small cohort of patients with EC (83 cases and 67 controls, including ESCC and EAC) has reported that the PADI4 rs10437048 genotype was significantly associated with decreased risk of EC, whereas rs41265997 were significantly associated with increased risk of EC [13]. In comparison with the cohort comprising ESCC and EAC in their study, we specifically focused on the relationship between ESCC and PADI4 in a larger cohort from East China, the seemingly discrepancy with previous findings may be attributed to the distinctive genetic variants characteristics in ESCC rather than EAC. In addition, the pairwise LD tagging approach for tagging SNPs selection in this study could possibly miss some SNPs in LD with rs2240337which were also susceptibility loci for ESCC. Notably, the frequencies of genetic polymorphisms vary drastically among different ethnic cohorts.
Rs2240337 is located in the intron region of PADI4 gene. The functions of SNPs in intron regions have not been fully elucidated. One study showed that rs2240337 could influence the mRNA stability or maturation in vitro [19], while the association between this SNP and rheumatoid arthritis severity has also been reported [20]. As the sample size was limited in our study, the correlation between rs2240337 and the pathologic character of ESCC was not evident, further investigation is desirable to demonstrate the functional relevance of rs2240337 polymorphism in ESCC.
Smoking and alcohol drinking have emerged as widely acknowledged risk factors of ESCC. This notion was in line with our finding, although PADI4 rs11203366, rs1886302, rs1635562, rs1635564, rs16825533 and rs2477137 were not associated with the susceptibility to ESCC, smoking significantly increased ESCC risk in PADI4 rs11203366 AG/AA, rs1886302 CC/CT, rs1635562 AT, rs1635564 CA and rs2477137 AC genotype, while alcohol drinking increased ESCC risk in PADI4 rs11203366 AG, rs1635562 AT, rs1635564 CA, rs2477137 AC, rs1886302 CT genotype. Interestingly, despite the fact that rs2240337 SNP was associated with decreased risk of ESCC, smoking increased ESCC risk in PADI4 rs2240337 AG genotype as compared with non-smokers. Our findings exemplified the significance of the environment and genetic risk factors interact and both contribute to the carcinogenesis. Our study showed the haplotype PADI4 Ars2477137Crs1886302Grs11203366Grs16825533Grs2240337Ars1635564Ars1635562 and PADI4 Crs2477137Trs1886302Grs11203366Ars1635564Grs2240337Crs1635564Trs1635562 genetic polymorphism may be correlated with decreased susceptibility to ESCC, while haplotype PADI4 Crs2477137Trs1886302Ars11203366Ars1635564Grs2240337Ars1635564Ars1635562 genetic polymorphism may be correlated with increased susceptibility of ESCC, which indicated that single locus polymorphism might not significantly modify the susceptibility to cancer, the chain effect lying in different loci leads to a more profound impact on the risk of cancer.
Our study provides the evidence that polymorphism of PIDA4 rs2240337 G>A is associated with the altered susceptibility of ESCC. We acknowledge there are several limitations in this study. First of all, the study subjects were all recruited from several local medical centers within same area, which might not completely represent the general Chinese population, especially when diverse regional environmental factors existed. Secondly, the detailed information regarding cancer metastasis and survival were not provided as the follow-up study is still ongoing, which hindered analyses of the impact of these SNP polymorphisms on ESCC progression and prognosis. Further studies with more loci and large sample size are warranted to elucidate the effect of PADI4 SNPs on ESCC risk. Last but not least, refrained by the limited technical support, we have not evaluated the biological function of the SNP polymorphism in the carcinogenesis of ESCC in the current study. As rs2240337 is located in the intron region of PADI4 gene, therefore overexpression of wild type and mutant type PADI4 coding sequence does not work. We speculate that rs2240337 may cause an alternative RNA splicing on PADI4 mRNA, thereby regulating the PADI4 protein function. Further studies using an rs2240337 G>A mutation cell or mouse model are needed to clarify the mutant PADI4 function.
MATERIALS AND METHODS
Ethical approval of the study protocol
We have complied with the World Medical Association Declaration of Helsinki regarding ethical conduct of research involving human subjects and/or animals. The Review Board of Jiangsu University (Zhenjiang, China) approved this hospital-based case-control study. To be included in the study, all subjects provided written informed consent.
Patients and controls
Between October 2008 and June 2013, 629 subjects with ESCC were consecutively recruited from the Affiliated People’s Hospital of Jiangsu University and Affiliated Hospital of Jiangsu University (Zhenjiang, China). All cases of ESCC were diagnosed pathologically. The exclusion criteria were patients who previously had: cancer; any metastasized cancer; radiotherapy or chemotherapy. The 686 controls were patients without cancer and were matched to the cases with regard to age (±5 years) and sex. Most of the controls were admitted to the hospitals for the treatment of trauma. They were recruited from the two hospitals mentioned above during the same time period.
Trained interviewers, using a pre-tested questionnaire, questioned each subject personally to obtain information on demographic data (e.g., age, sex) and related risk factors (including tobacco smoking and alcohol consumption). After the interview, 2mL of venous blood was collected from each subject. Individuals who smoked one cigarette per day for >1 year were defined as “smokers”. Subjects who consumed more than three alcoholic drinks a week for >6 months were considered to be “alcohol drinkers”.
Isolation of DNA, SNPs selection and genotyping by ligation detection reaction
Blood samples were collected from patients using vacutainers and transferred to tubes lined with ethylenediamine tetra-acetic acid (EDTA). Genomic DNA was isolated from whole blood with the QIAamp DNA Blood Mini Kit (Qiagen, Berlin, Germany) as described [21].
To find tagging SNPs, we used a block-based tagging strategy using Haploview 4.2 software, according to the HapMap database (http://www.hapmap.org/, phase II Nov08, on NCBI B36 assembly, dbSNP b126; population: Chinese Han population). Seven PADI4 tagging SNPs were selected on the basis of Hardy-Weinberg equilibrium (HWE) p ≥ 0.05, call rate ≥ 95% and minor allele frequency ≥ 0.05. The samples were genotyped using the ligation detection reaction (LDR) method, with technical support from the Shanghai Biowing Applied Biotechnology Company [22]. For quality control, repeated analyses were done for 110 (11.73%) randomly selected samples with high DNA quality.
Statistical analyses
Differences in the distributions of demographic characteristics, selected variables, genotypes of the PADI4 variants, and the correlation between genotyping and pathologic state were evaluated using the χ2 test. The associations between the seven SNPs and risk of ESCC were estimated by computing the odds ratios (ORs) and their 95% confidence intervals (CIs) using logistic regression analyses for crude ORs and adjusted ORs when adjusting for age, sex, smoking and drinking status. The HWE was tested by a goodness-of-fit χ2 test to compare the observed genotype frequencies to the expected frequencies among the control subjects. The Bonferroni correction procedure was applied because of the number of comparisons. As multiple hypotheses are tested, the chance of a rare event increases, and the likelihood of incorrectly rejecting a null hypothesis (type I error) increases, the Bonferroni correction was therefore performed. All statistical analyses were performed with SPSS 23.0 Statistical Package (SPSS Inc., Chicago, IL).
Abbreviations
PADI4: peptidylarginine deiminase type 4; ESCC: esophageal squamous cell carcinoma; EC: esophageal cancer; SNP: Single nucleotide polymorphism; LD: linkage disequilibrium; OR: odds ratio; CI: confidential interval.
Author contributions
LW, HG and TL carried out the molecular genetic studies, selected the tagged SNPs, performed the statistical analysis and drafted the manuscript. HP, LL, YS, JZ, YS, WT, GD, SC, YF, HD, QW, JY recruited the patients and collected the samples. SC, JY and YF participated in the design and coordination of the study. LT and JY conceived of the study, and participated in its design and coordination. All authors read and approved the final manuscript.
ACKNOWLEDGMENTS
We appreciate all patients who participated in this study. We wish to thank Dr. Yiqun Chen (Biowing Applied Biotechnology Company, Shanghai, China) for technical support.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
This study was supported by the National Natural Science Foundation of China (81300037, 81000028, 81370001, 81570031, 81101889, 81472332, 81341006); the Key Research and Development Program of Jiangsu Province (BE2016714); the Natural Science Foundation of Jiangsu Province (BK2010333, BK2011481); the “333” Elitist Training Program, Jiangsu, China (BRA2013135); the “Six Talent Peaks” Training Program, Jiangsu, China (2015-WSN-117, 2014-WSN-078); the “Distinguished Medical Specialist” Program, Jiangsu, China; the “Innovative and Entrepreneurial Elite Team” Program (2016), Jiangsu, China, the “Young Medical Talents” Training Program (QNRC2016447), Jiangsu, China, the research funding of Zhongshan Hospital (2016ZSLC15) and the Zhenjiang Social Development Program (SH2013039).
REFERENCES
1. Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011; 61: 69-90. doi: 10.3322/caac.20107.
2. Gholipour C, Shalchi RA, Abbasi M. A histopathological study of esophageal cancer on the western side of the Caspian littoral from 1994 to 2003. Dis Esophagus. 2008; 21: 322-7. doi: 10.1111/j.1442-2050.2007.00776.x.
3. Tran GD, Sun XD, Abnet CC, Fan JH, Dawsey SM, Dong ZW, Mark SD, Qiao YL, Taylor PR. Prospective study of risk factors for esophageal and gastric cancers in the Linxian general population trial cohort in China. Int J Cancer. 2005; 113: 456-63. doi: 10.1002/ijc.20616.
4. Engel LS, Chow WH, Vaughan TL, Gammon MD, Risch HA, Stanford JL, Schoenberg JB, Mayne ST, Dubrow R, Rotterdam H, West AB, Blaser M, Blot WJ, et al. Population attributable risks of esophageal and gastric cancers. J Natl Cancer Inst. 2003; 95: 1404-13.
5. Kamangar F, Chow WH, Abnet CC, Dawsey SM. Environmental causes of esophageal cancer. Gastroenterol Clin North Am. 2009; 38: 27-57, vii. doi: 10.1016/j.gtc.2009.01.004.
6. Nasrollahzadeh D, Kamangar F, Aghcheli K, Sotoudeh M, Islami F, Abnet CC, Shakeri R, Pourshams A, Marjani HA, Nouraie M, Khatibian M, Semnani S, Ye W, et al. Opium, tobacco, and alcohol use in relation to oesophageal squamous cell carcinoma in a high-risk area of Iran. Br J Cancer. 2008; 98: 1857-63. doi: 10.1038/sj.bjc.6604369.
7. Hagiwara T, Nakashima K, Hirano H, Senshu T, Yamada M. Deimination of arginine residues in nucleophosmin/B23 and histones in HL-60 granulocytes. Biochem Biophys Res Commun. 2002; 290: 979-83. doi: 10.1006/bbrc.2001.6303.
8. Chang X, Han J. Expression of peptidylarginine deiminase type 4 (PAD4) in various tumors. Mol Carcinog. 2006; 45: 183-96. doi: 10.1002/mc.20169.
9. Oshima RG. Apoptosis and keratin intermediate filaments. Cell Death Differ. 2002; 9: 486-92. doi: 10.1038/sj/cdd/4400988.
10. Iwaya K, Mukai K. Accumulation of ubiquitin-conjugated cytokeratin fragments in tumor cells. Semin Cancer Biol. 2005; 15: 309-18. doi: 10.1016/j.semcancer.2005.04.006.
11. Slack JL, Causey CP, Thompson PR. Protein arginine deiminase 4: a target for an epigenetic cancer therapy. Cell Mol Life Sci. 2011; 68: 709-20. doi: 10.1007/s00018-010-0480-x.
12. Chang X, Han J, Pang L, Zhao Y, Yang Y, Shen Z. Increased PADI4 expression in blood and tissues of patients with malignant tumors. BMC Cancer. 2009; 9: 40. doi: 10.1186/1471-2407-9-40.
13. Chang X, Hou X, Pan J, Fang K, Wang L, Han J. Investigating the pathogenic role of PADI4 in oesophageal cancer. Int J Biol Sci. 2011; 7: 769-81.
14. Yao H, Li P, Venters BJ, Zheng S, Thompson PR, Pugh BF, Wang Y. Histone Arg modifications and p53 regulate the expression of OKL38, a mediator of apoptosis. J Biol Chem. 2008; 283: 20060-8. doi: 10.1074/jbc.M802940200.
15. Wang Y, Wysocka J, Sayegh J, Lee YH, Perlin JR, Leonelli L, Sonbuchner LS, McDonald CH, Cook RG, Dou Y, Roeder RG, Clarke S, Stallcup MR, et al. Human PAD4 regulates histone arginine methylation levels via demethylimination. Science. 2004; 306: 279-83. doi: 10.1126/science.1101400.
16. Cuthbert GL, Daujat S, Snowden AW, Erdjument-Bromage H, Hagiwara T, Yamada M, Schneider R, Gregory PD, Tempst P, Bannister AJ, Kouzarides T. Histone deimination antagonizes arginine methylation. Cell. 2004; 118: 545-53. doi: 10.1016/j.cell.2004.08.020.
17. Denis H, Deplus R, Putmans P, Yamada M, Metivier R, Fuks F. Functional connection between deimination and deacetylation of histones. Mol Cell Biol. 2009; 29: 4982-93. doi: 10.1128/MCB.00285-09.
18. Li P, Yao H, Zhang Z, Li M, Luo Y, Thompson PR, Gilmour DS, Wang Y. Regulation of p53 target gene expression by peptidylarginine deiminase 4. Mol Cell Biol. 2008; 28: 4745-58. doi: 10.1128/MCB.01747-07.
19. Suzuki A, Yamada R, Chang X, Tokuhiro S, Sawada T, Suzuki M, Nagasaki M, Nakayama-Hamada M, Kawaida R, Ono M, Ohtsuki M, Furukawa H, Yoshino S, et al. Functional haplotypes of PADI4, encoding citrullinating enzyme peptidylarginine deiminase 4, are associated with rheumatoid arthritis. Nat Genet. 2003; 34: 395-402. doi: 10.1038/ng1206.
20. Abd-Allah SH, el-Shal AS, Shalaby SM, Pasha HF, Abou el-Saoud AM, el-Najjar AR, el-Shahawy EE. PADI4 polymorphisms and related haplotype in rheumatoid arthritis patients. Joint Bone Spine. 2012; 79: 124-8. doi: 10.1016/j.jbspin.2011.07.006.
21. Wang L, Tang W, Chen S, Sun Y, Fan Y, Shi Y, Zhu J, Wang X, Zheng L, Shao A, Ding G, Liu C, Liu R, et al. N-acetyltransferase 2 polymorphisms and risk of esophageal cancer in a Chinese population. PLoS One. 2014; 9: e87783. doi: 10.1371/journal.pone.0087783.
22. Chen ZJ, Zhao H, He L, Shi Y, Qin Y, Shi Y, Li Z, You L, Zhao J, Liu J, Liang X, Zhao X, Zhao J, et al. Genome-wide association study identifies susceptibility loci for polycystic ovary syndrome on chromosome 2p16.3, 2p21 and 9q33.3. Nat Genet. 2011; 43: 55-9. doi: 10.1038/ng.732.

