INTRODUCTION
The mesothelium is a monolayer of cells extending over the surface of the serosal cavities and organs. This layer facilitates free movement between tissues and organs whilst protecting them from infection and injury [1–3]. Mesothelioma is an aggressive cancer that develops in the mesothelium and is strongly associated with asbestos exposure [4]. The fibrous mineral erionite [5], carbon nanotubes [6, 7], genetic mutations [8], radiation [9] and Simian Virus 40 [10] have also been linked to mesothelioma.
The global incidence of mesothelioma has increased over the last decade and is predicted to peak sometime before 2030 [11]. Countries with the highest rates of mesothelioma include the USA and the UK, however Australia and Italy rank highly per capita [12]. Mesothelioma has a long latency period (20–40 years) following asbestos exposure [11]. Therefore, the new wave of predicted victims exposed through building demolition, renovation and repair [13] and in countries where asbestos is still mined and used (India, Vietnam, China, Russia, Zambia, Colombia and Kazakhstan) [14, 15], will likely see mesothelioma be a health burden beyond 2030.
Mesothelioma has a poor prognosis with a median survival of 4 to 14 months [12, 16]. Survival is influenced by histological subtype (epithelioid, biphasic, sarcomatoid) as epithelioid patients survive on average 9 months longer than sarcomatoid patients [12, 17, 18]. Cases of malignant pleural mesothelioma (MPM) are most commonly reported (80%) followed by malignant peritoneal mesothelioma (PMM) (20%) and rarely, mesothelioma of the pericardium [19–21] and testis [22].
Malignant pleural mesothelioma (MPM)
MPM consists of small tumour nodules that extend along the pleural surface. These tumours eventually enclose and invade the lung [15, 23]. Symptoms of MPM include chest tightness, pain and shortness of breath [24]. In 90% of patients, these symptoms will be caused by the presence of a pleural effusion (PE) [25].
PE, pleural thickening and pleural nodules are likely to be evident upon imaging of suspected cases [26, 27], however these findings do not distinguish MPM from other metastatic diseases [26, 28]. A cytological diagnosis of epithelioid MPM using PE samples is possible [29]. However, sarcomatoid cells are not often shed into the pleural space, therefore tissue demonstrating the malignant characteristics of invasion, cellular atypia and necrosis is required [30]. Collecting tissue by biopsy is invasive and can complicate disease management through the potential seeding of tumour cells [31]. To identify ‘at risk’ individuals and patients with early stage MPM, serum and PE markers have been investigated. The diagnostic accuracies reported for the more promising biomarkers mesothelin (MSLN) [32, 33], osteopontin [34, 35] and fibulin-3 [36, 37] are variable [31]. Therefore, there are currently no biomarkers that can be used alone for the accurate diagnosis of MPM.
Treatment of MPM requires multiple therapeutic modalities. Surgery (pleurectomy with or without decortication (PD) and extrapleural pneumonectomy (EPP)) can improve symptoms but rarely eradicate microscopic disease [38]. The regimen considered to be the standard of care for the palliation of MPM is the combination of pemetrexed with cisplatin [39, 40]. Compared to cisplatin alone, this combination improves patient survival by a few months [40]. Radiotherapy can also be given for palliative reasons [16, 41] but rarely improves survival outcomes [26, 42].
No new treatments for MPM exist outside the clinical trial setting [38, 39]. However, novel targets such as growth factors, apoptotic signalling pathways and various aspects of the immune system are being investigated. The immune checkpoint inhibitors cytotoxic T-lymphocyte-associated antigen 4 (CTLA-4) and programmed death 1 (PD-1) have become popular targets in recent times. Novel drugs targeting CTLA-4 and PD-1 have been approved for the treatment of melanoma and non-small cell lung cancer (NSCLC) and are being tested in ongoing trials for many cancers including MPM [39]. Other therapeutic options being investigated include the administration of oncolytic viruses, vaccination strategies to induce antigen-specific cell-mediated immune responses [43] and restoring down-regulated miRNA [44].
Malignant peritoneal mesothelioma (PMM)
PMM is often found as a diffuse tumour of the intestinal serosa or a large mass on the omentum or mesentery [23]. PMM presents with non-specific symptoms such as loss of appetite, nausea, vomiting, diarrhoea, constipation and ascites. Small bowel obstruction is a late feature [16].
Radiological findings that give the best indication of PMM include ascites associated with minimal soft tissue masses and preserved normal anatomy of the bowel and its mesentery [45, 46]. Laparotomy and laparoscopy with biopsy are the main diagnostic approaches used, however to ensure diagnostic accuracy, tissue samples need to be of adequate amount and quality [16]. Serum biomarkers for the less-invasive diagnosis of PMM have been investigated and potential targets include hyaluronic acid, osteopontin, mesothelin [16] and high mobility group box 1 (HMGB1) [47]. The identification of a robust biomarker for PMM is important as PMM has a high misdiagnosis rate and can easily be confused with ovarian cancer [48], diseases that affect the colon [49] and tuberculosis peritonitis [50].
The standard regimen used to treat PMM includes cytoreductive surgery (CRS) with hyperthermic intraperitoneal chemotherapy (HIPEC) using mitomycin C or cisplatin [16]. However, CRS plus HIPEC is not suitable for all patients, particularly those with disease outside the peritoneum or with poor health [51]. Whether radiotherapy improves patient survival remains unclear and adverse side effects such as adhesions and intestinal obstruction reduce the popularity of its use [16]. There are few clinical trials investigating novel avenues for the treatment of PMM, therefore new therapeutic targets are required.
Diagnosing mesothelioma can be challenging, invasive and lengthy. There is no way of identifying ‘at risk’ individuals who may benefit from early intervention. Current treatments are rarely curative and the lack of positive clinical trial results is a concern. Mesothelioma is likely to continue being a global health burden as the threat of asbestos exposure continues. If survival rates are to improve, novel treatment and diagnostic approaches are urgently needed.
MICRORNA
During the search for novel therapeutic and diagnostic targets for mesothelioma, the small but powerful gene regulators microRNA (miRNA) have become of interest. Cells produce miRNA in a multi-step process beginning in the nucleus (Figure 1) [52–55] where miRNA transcripts are cleaved by Drosha/DGCR8 to produce an intermediate structure of 60–70 nucleotides (nt) [52, 53, 56]. This precursor is transported into the cytoplasm by the Exportin-5 protein [53] where it is cleaved to form a duplex (21 to 25 nt) consisting of the mature miRNA and its complementary strand [57]. MiRNA associate with the argonaute protein to form the core of the miRNA Induced Silencing Complex (miRISC) [58] and guide the miRISC to complementary sites within target messenger RNA (mRNA) [52]. Once bound, the miRISC inhibits gene expression by initiating mRNA degradation and/or repressing translation [54, 59]. Cells can also produce miRNA through a number of non-canonical processes independent of Drosha and/or Dicer activity [60].
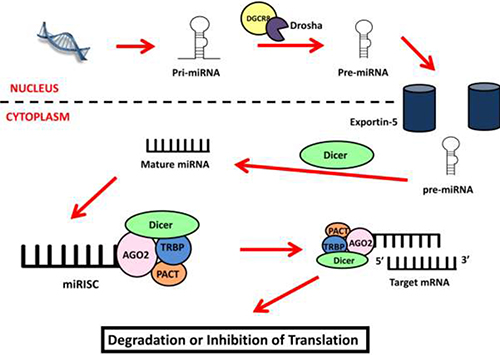
Figure 1: miRNA biogenesis. MiRNA biogenesis starts in the nucleus where the miRNA gene is transcribed into a primary miRNA transcript (pri-miRNA) and processed by Drosha/DGCR8 into the miRNA precursor (pre-miRNA). The precursor is exported into the cytoplasm by Exportin-5 where it is cleaved by Dicer to become the mature miRNA. This strand forms the miRISC complex with the AGO2, TRBP, PACT and Dicer proteins. The miRISC uses the miRNA as a guide to identify and bind to target mRNA causing the inhibition of target genes by inducing mRNA degradation or inhibiting translation.
MiRNA have extensive regulatory potential and the imprecise base pairing between miRNA and their targets allow miRNA to regulate a multitude of genes [61, 62]. Deviations in this regulation contribute to aberrant gene expression and influence disease biology [63]. MiRNA can function as oncogenes and/or tumour suppressors and are therefore potential targets for the development of novel treatments for cancer. MiRNA also have characteristics that make them attractive biomarkers including being stably expressed in tissue and fluid [64] and being easily measurable using techniques such as quantitative real-time PCR (qPCR) [65].
MIRNA IN MPM
Role of miRNA in MPM biology
MiRNA were first suggested to have biological roles in MPM by Guled and colleagues in 2009. The authors identified a number of miRNA that were expressed significantly different between MPM tissue and normal pericardium and the three MPM subtypes. The miRNA were predicted to target some of the more commonly affected genes in MPM including cyclin-dependent kinase inhibitor 2A (CDNK2A), neurofibromatosis type 2 (NF2), jun oncogene, hepatocyte growth factor and platelet derived growth factor. The miRNA were also located in chromosomal areas known to be deleted or gained in MPM [66]. A number of miRNA have now been identified as aberrantly expressed in MPM with a select few shown to regulate cell activity. Those with functional roles in MPM are described below and summarised in Table 1.
Table 1: MiRNA with biological roles in mesothelioma
miRNA |
Ref |
Expression in mesothelioma vs controls |
Samples Analysed |
MiRNA genomic location |
Chromosomal aberration in mesothelioma? |
Target gene regulated in mesothelioma |
Mesothelioma cell function regulated |
|---|---|---|---|---|---|---|---|
MPM |
|||||||
29c-5p |
[67] |
Lower |
8 MPM cell lines, LP9 mesothelial cells |
1q32.2 |
Yes [121] |
DNMT1, DNMT3A |
Proliferation, migration, invasion, colony formation, methylation |
31 |
[68] |
Lower |
8 MPM cell lines, LP9, primary mesothelial culture, Met-5A |
9p21.3 |
PPP6C |
Proliferation, migration, invasion, colony formation |
|
let-7a |
N/A |
2 EphrinA1 treated MPM cell lines |
22q13.31 |
Yes [121] |
RAS oncogenes |
Proliferation, migration |
|
let-7b |
[74] |
N/A |
2 Ursolic Acid treated MPM cell lines |
22q13.31 |
Yes [121] |
Twist |
Apoptosis, EMT |
16 |
Lower |
60 FFPE MPM tissues (46 Ep, 14 Bi), 23 FFPE normal pleura tissues, 6 MPM cell lines, Met-5A |
13q14.2 |
Yes [121] |
CCDN1, BCL-2, PD-L1 |
Proliferation, chemoresistance |
|
34 b/c |
Lower |
47 MPM tumours (32 Ep, 10 Bi, 4 Sa, 1 lymphohistiocytic), 10 non-neoplastic pleura, 6 MPM cell lines, 2 primary mesothelial cultures |
11q23.1 |
Yes [121] |
BCL-2 |
Proliferation, migration, invasion, resistance to radiotherapy |
|
126 |
[80] |
Lower |
29 FFPE MPM tissues, 5 MPM diagnostic biopsies, 14 matched non-neoplastic tissues, 5 pneumothorax benign reactive mesothelial tissues |
9q34.3 |
Yes [121] |
IRS1, PDK, ACL |
Mitochondrial metabolism, proliferation, autophagic flux |
1 |
Lower |
25 MPM tumours, 25 unmatched normal pleura tissue, 7 MPM cell lines |
20q13.33 |
Yes [121] |
PIM1 |
Proliferation, apoptosis, migration, invasion |
|
145 |
[86] |
Lower |
71 MPM tumours, 12 mesothelial benign cysts, 50 normal tissues, 3 MPM cell lines, primary mesothelial cell culture |
5q32 |
Yes [123] |
OCT4 |
Cell viability, clonogenicity, migration |
21-5p |
[89] |
N/A |
N/A |
17q23.1 |
Yes [121] |
MSLN |
Proliferation |
223 |
[90] |
Lower |
8 MPM cell lines (5 human, 3 mouse), primary mesothelial cell cultures, 26 MPM pleural effusions, 10 benign pleural effusions, 17 FFPE MPM tissues, 6 FFPE normal pleura |
Xq12 |
No |
STMN1 |
Migration |
302b |
[95] |
N/A |
2 EphrinA1 treated MPM cell lines |
4q25 |
Yes [121] |
MCL-1 |
Proliferation, apoptosis |
193a-3p |
[96] |
Lower |
120 MPM tissues (59 extrapleural pneumonectomy & 61 pleurectomy + decortication), 23 normal pleura, 10 MPM cell lines, Met-5A |
7q11.2 |
Yes [124] |
MCL-1 |
Proliferation, apoptosis, necrosis |
17-5p |
[97] |
Lower |
60 FFPE MPM tissues, 23 normal pleura, 7 MPM cell lines, Met-5A |
13q31.3 |
Yes [121] |
KCNMA1 |
Migration |
205 |
[99] |
N/A |
74 MPM tissues (21 Bi, 18 Sa, 35 Ep) |
1q32.3 |
Yes [121] |
ZEB1, ZEB2 |
EMT, invasion, migration |
PMM |
|||||||
34a |
[117] |
Lower |
45 PMM tissues, 5 PMM cell lines, 7 normal peritoneum |
1p36.22 |
Yes [125] |
c-MET, AKT |
Proliferation, apoptosis, invasion |
Ep – epithelioid, Bi – biphasic, Sa – sarcomatoid, N/A – not available, FFPE – formalin fixed paraffin embedded.
Pass and colleagues were the first to identify a miRNA with a functional role in MPM after miR-29c-5p was observed as downregulated in MPM cell lines compared to normal mesothelial controls. Transfection of the miR-29c-5p mimic in two MPM cell lines restored miR-29c-5p expression and caused an inhibition of cell proliferation, migration, invasion and colony formation. MiR-29c-5p was also suggested as a potential mediator of methylation in MPM after the expression of the DNA methyltransferases DNMT1 and DMT3A were reduced following miR-29c-5p overexpression. This study revealed that miR-29c-5p may be a tumour suppressor in MPM and is thus a potential therapeutic target [67].
MiR-31 is another downregulated miRNA in MPM caused by co-deletion of the miR-31 and CDKN2A genes from chromosome 9p21 [68]. Re-introducing miR-31 in MPM cells inhibited proliferation, migration, invasion and colony formation and reduced levels of protein phosphatase 6 (PPP6C). Aberrant PPP6C activity is associated with resistance to chemotherapy [69] and radiotherapy [70]. Therefore, loss of miR-31 in MPM may contribute to elevated PPP6C and promote chemoresistance. If this is true, miR-31 replacement therapy may be an effective tool to enhance tumour responses to chemotherapy.
Let-7 is a well-studied miRNA with a range of functions in cancer [71]. In MPM cell lines, let-7a was upregulated following activation of the Ephrin type A receptor by its ligand Ephrin A1. In turn, RAS family proto-oncogenes were suppressed causing an inhibitory effect on MPM cell growth [72]. In a subsequent study, nanoparticles packaged with let-7a inhibited MPM cell proliferation, migration and tumour growth [73]. Recently, another let-7 family member, let-7b, was shown to enhance the anti-tumour effect of ursolic acid in MPM cell lines. The overexpression of let-7b caused the cleavage of caspase-3 and PARP, the suppression of pAKT, B-catenin and Twist and the accumulation of cells in the sub-G1 phase of the cell cycle. The inhibition of let-7b in the same cell lines blocked the cytotoxicity of ursolic acid treatment and together, these results suggest that let-7b may regulate apoptosis and inhibit epithelial to mesenchymal transition (EMT) during ursolic acid treatment of MPM [74]. Increasing let-7a and b may be effective treatment strategies for MPM, either alone or in combination with novel chemotherapy agents.
MiRNA MPM therapeutic studies have recently advanced to clinical trial. MiR-16 is now the focus of a phase I trial that will be discussed in the following section of this review. The trial is based on work by Reid and colleagues who reported the downregulation of miR-15/16 in MPM tissue and cell lines in 2013. Reduced miR-15/16 was associated with increased levels of the target oncogenes CCND1 and Bcl-2 and re-expressing the miRNA in cell lines inhibited cell growth. Cell growth was inhibited most effectively following transfection of the miR-16 mimic. Restoring miR-16 also re-sensitised MPM cells to pemetrexed and gemcitabine and the intravenous administration of miR-16 in nanocells with epidermal growth factor receptor (EGFR) specific antibodies, inhibited tumour growth in mice [75]. The same laboratory has recently published results demonstrating that miR-16 is also a regulator of programmed death ligand 1 (PD-L1) in MPM and may therefore contribute to immune system evasion [76].
The miR-34 family are attractive targets for replacement therapy in MPM. The absence of miR-34b/c in MPM cells is caused by methylation and restoring miR-34b/c reverses malignant features such as migration, invasion, motility [77] and resistance to radiotherapy [78]. In normal mesothelial cells, reducing miR-34 induced cell proliferation, migration and invasion by up-regulating oncogenes such as C-MET and Bcl-2 [79]. These studies suggest that miR-34 has a role in the oncogenic transformation of mesothelial cells and the ongoing regulation of MPM biology.
Like miR-34, the expression of miR-126 in MPM is regulated by methylation [80] and also oxidative stress [81]. During such stress, miR-126 influences metabolic processes by altering mitochondrial function and inhibits malignant features such as cell growth, soft agar colony formation and tumour formation in mice [81]. MiR-126 potentially inhibits tumour progression through its ability to induce autophagic flux, thus supporting the idea that increased levels of autophagy may be protective in MPM [82].
MiR-1 was identified as a potential tumour suppressing miRNA in MPM when miR-1 was observed as downregulated in tumour samples compared to normal pleural mesothelium. Overexpressing miR-1 in MPM cells inhibited proliferation and induced apoptosis [83]. In a subsequent study, miR-1 was reported as downregulated in MPM cell lines and overexpressing miR-1 in two of these lines led to an inhibition of cell growth, invasion, migration and reduced levels the proto-oncogene PIM1 [84]. Recent evidence suggests that PIM1 is overexpressed in MPM and can influence MPM cell function [85]. Therefore, both miR-31 and PIM1 are potential targets for future MPM therapeutic studies.
MiR-145 is also downregulated in MPM cell lines and tumours and restoring miR-145 reduced proliferation, migration and invasion in vitro and inhibited tumour growth in mice. MiR-145 potentially exerts these tumour suppressive effects by regulating the transcription factor and stem cell marker OCT4 [86]. A recent study has shown that OCT4/SOX2 may be useful markers for identifying MPM cancer stem cell populations in vitro. Cells with high OCT4/SOX2 levels were resistant to chemotherapy, barely affected by re-expression of the NF2 tumour suppressor, and had a high-tumour initiating capability in vivo [87]. It has been hypothesised that this sub-population of cells is responsible for the poor response of MPM to treatment and important for tumour relapse. The role of miR-145 in the regulation of OCT4 in this MPM cell population will be important to investigate.
Identifying miRNA target genes is an important process for understanding how miRNA regulate cell function and disease biology. This can be done using results reported from previous studies, prediction software or affinity purification approaches. The “miR-CATCH” technique involves an affinity capture oligonucleotide that is used to co-purify a single target mRNA together with all endogenously bound miRNA [88]. This technique was combined with next generation sequencing to identify miRNAs that regulate the commonly upregulated gene in MPM MSLN. MiR-21-5p was identified as a candidate regulator of MSLN which was confirmed following miR-21-5p overexpression in a panel of MPM cell lines and the transformed mesothelial cell line MET-5A. The increased expression of miR-21-5p reduced MSLN expression and inhibited MPM cell proliferation, therefore uncovering another potential tumour suppressing miRNA in MPM [89].
MiR-223 was similarly identified by our laboratory as downregulated in MPM when miR-223 levels were found to be significantly lower in MPM cell lines, tissue and cells isolated from MPM PE compared to controls [90]. One target of miR-223 that is overexpressed in MPM is stathmin (STMN1) [91]. STMN1 is highly expressed in many malignancies and reducing STMN1 consistently inhibits cell growth, motility, invasion and the formation of metastasis in vivo. These processes are associated with changes in cell morphology and a decrease in microtubule stability [92]. We showed that re-expressing miR-223 in two MPM cell lines reduced STMN1 expression and MPM cell migration. We also showed that loss of miR-223 and overexpression of STMN1 in MPM could be due to aberrant c-JUN N-terminal kinase (JNK) signalling. Therefore, a potential tumour suppressive role for the JNK-miR-223-STMN1 axis was suggested and a novel role for JNK in MPM was revealed [90]. MiR-223 along with a number of other miRNA may also regulate tumour protein 53 (TP53) in MPM. These miRNA were identified as downregulated in tumours expressing the negative regulator of TP53 MDM2 [93]. Functional studies are required to validate these findings.
Myeloid cell leukaemia-1 (Mcl-1) is an anti-apoptotic protein regulated by a number of miRNA in MPM. Mcl-1 is overexpressed in MPM and is associated with the resistance of MPM cells to apoptosis [94]. Khodayari and colleagues recently determined that Mcl-1 was downregulated in MPM cells following the upregulation of miR-302b during ephrin-A1-mediated MPM cell growth inhibition. Transfecting MPM cells with miR-302b reduced Mcl-1 expression, cell and tumoursphere growth and induced apoptosis [95]. Mcl-1 is also regulated by miR-193a-3p in MPM and transfecting MPM cells with the miR-193a-3p mimic inhibited cell growth whilst inducing apoptosis and necrosis. MiR-193a-3p, delivered to MPM tumours in nanocells with EGFR antibodies, inhibited xenograft growth and induced tumour cell apoptosis in mice [96]. Using miRNA replacement therapy to target Mcl-1 in patients may prove to be an effective treatment for MPM.
The same laboratory recently published results supporting a role for miR-17-5p in regulating MPM cell migration [97]. An integrative approach was used to compare miRNA expression data from previous studies and mRNA gene expression datasets. Amongst the top enriched miRNA was the miR-17 family that was downregulated in MPM samples and significantly associated with the epithelioid subtype. The top enriched mRNA signalling pathways included genes linked to MPM cell migration. Some of these are regulated by miR-17 including KCNMA1 which encodes for the calcium-activated potassium channel subunit alpha 1 (KCa1.1) protein. In MPM cell lines, KCNMA1 and KCa1.1 were downregulated along with cell migration and invasion when these cells were transfected with the miR-17-5p mimic. Targeting KCa1.1 with the inhibitor paxilline also significantly inhibited MPM cell migration and colony formation. Therefore, inhibiting KCa1.1 using either the channel blocker paxilline or miR-17-5p replacement, may serve as novel treatments for MPM.
The morphologies of the different MPM subtypes are likely due to the different EMT stages [98]. During a study to explore the role of EMT in the three histological subtypes, Fassini et al., discovered that miR-205 was expressed significantly higher in epitheliod cells and tissue compared to both the biphasic and sarcomatoid subtypes. Therefore, loss of miR-205 correlated with a mesenchymal phenotype and a more aggressive tumour [99]. MiR-205 is a known regulator of EMT and maintains an epithelial phenotype by reducing ZEB1 and 2 and enhancing E-cadherin expression [100]. Transfecting miR-205 into MPM cell lines consistently reduced ZEB1 and 2 and cell migratory capability, thus suggesting a role for miR-205 in negatively regulating malignant features in MPM [99].
Most of the miRNA described above are downregulated in MPM and serve as potential tumour suppressors. This is a common phenomenon that has been reported in many malignancies. Interestingly, the genomic locations of the miRNA genes are associated with chromosomal aberrations that have been identified in MPM tumours and cells (Table 1). Therefore, chromosomal abnormalities are likely the cause of the global downregulation of miRNA in mesothelioma.
MiRNA replacement therapy for MPM
MiRNA are attractive therapeutic targets because of their powerful regulatory capabilities. Targeting multiple signalling pathways through a single miRNA may provide an effective way of combating drug resistance and improving tumour responses. Given that most miRNA are downregulated in MPM, strategies aimed at replacing miRNA in MPM may be therapeutically beneficial.
MiRNA replacement therapy for MPM has been an effective inhibitor of tumour growth in mice [73, 75, 81, 86, 96]. The most important development in moving this treatment forward to the clinic was the development of the miRNA delivery vehicles TargomiRs, by EnGeneIC. TargomiRs are minicells derived from asymmetric bacterial cell division that are loaded with miRNA mimics. They can be directed to malignant tissue using antibodies against specific tumour antigens [101, 102].
The TargomiRs used for the treatment of MPM in mice, loaded with the miR-16 mimic and conjugated with an anti-EGFR antibody [75], have now been intravenously administered to patients in a Phase I clinical trial MesomiR 1 (clinicaltrials.gov NCT02369198). Preliminary results indicate that this approach may have therapeutic benefit. All six patients enrolled in the initial stages completed the treatment regime. Four of the six showed stable disease and one patient had a partial response after eight weeks [44]. Ongoing trials will investigate the effects of increasing TargomiR dosage and compare TargomiR treatment to second or third-line chemotherapy.
MiRNA as diagnostic biomarkers for MPM
The lack of successful diagnostic and prognostic markers for MPM has encouraged researchers to investigate novel targets such as miRNA (summarised in Tables 2 and 3). Early research identified the miR-200 family as potential candidates for discriminating MPM from other cancers that invade the lung such as adenocarcinoma [103, 104]. A diagnostic assay based on the expression of miR-193-3p, miR-200c and miR-192 was developed reaching a sensitivity of 100% and a specificity of 94% in a blinded validation set of 68 samples from the lung and pleura [104]. A recent study comparing miRNA profiles in MPM tissue to non-neoplastic pleura using qPCR, identified a panel of four miRNA including miR-126, miR-143, miR-145 and miR-652 as significantly downregulated in MPM. These results were validated in a larger cohort and when the four miRNA were combined using logistic regression analysis, a high diagnostic accuracy (area under the curve (AUC)), as determined by receiver operator curve (ROC) analysis, of 0.96 was achieved [105]. Despite these promising results, these studies rely on miRNA expression in tissue. Preferably, a diagnostic test would measure miRNA in samples acquired in a less invasive way such as blood or urine.
Table 2: Potential diagnostic miRNA for MPM
miRNA |
Ref |
Source |
Cohort |
Number |
MPM Histological Subtype |
Statistical Measure |
|---|---|---|---|---|---|---|
200c, 141, 200b, 429 |
[102] |
Tissue |
1 |
15 MPM, 10 lung AD |
N/A |
AUC > 0.9 for each miRNA |
2 |
100 MPM, 32 lung AD |
32 U, 39 Ep, 19 Bi, 10 Sa |
||||
200c, 192, 193a-3p |
[104] |
Tissue |
1 |
29 MPM, 140 carcinomas |
22 Ep, 1 Bi, 6 Sa |
sensitivity 100%specificity 94% |
2 |
48 MPM, 136 carcinomas |
6 U, 29 Ep, 2 Bi, 7 Sa |
||||
3 |
14 MPM 49 carcinomas |
8 Ep, 4 Bi, 2 Sa |
||||
126, 143, 145, 652 |
[105] |
Tissue |
1 |
5 MPM, 5 matched diagnostic biopsies, 5 matched non-neoplastic pleura |
5 Ep |
AUC 0.96 for miRNA combined |
2 |
40 MPM, 12 matched diagnostic biopsies, 14 matched non-neoplastic pleura, 5 non-neoplastic reactive mesothelium |
27 Ep, 25 Bi |
||||
625-3p |
[106] |
Serum |
1 |
5 MPM, 3 healthy |
3 Ep, 2 Sa |
AUC 0.8 |
2 |
5 MPM, 14 healthy |
1 U, 9 Ep, 3 Bi, 2 Sa |
||||
3 |
30 MPM, 10 asbestosis |
1 U, 29 Ep, |
||||
Tissue |
4 |
18 MPM, 7 normal pericardium |
15 Ep, 3 Bi |
|||
103 |
[107] |
Cellular fraction of peripheral blood |
1 |
23 MPM, 17 asbestos exposed, 25 healthy |
3 U, 12 Ep, 7 Bi, 1 Sa |
AUC 0.75 - 0.87 |
126 |
[108] |
Tissue |
1 |
10 MPM 5 normal mesothelium |
9 Ep, 1 Sa |
AUC 0.7 |
2 |
27 MPM & adjacent normal tissue |
23 Ep, 3 Bi, 1 Sa |
||||
Serum |
3 |
44 MPM, 196 asbestos exposed, 50 healthy |
30 Ep, 8 Bi, 6 Sa |
sensitivity 60 - 73%, specificity 74% |
||
126, 132-3p |
[109] |
Plasma |
1 |
21 MPM, 21 asbestos exposed |
14 Ep, 4 Bi, 3 Sa |
AUC ~ 0.8 for each miRNA and combination |
2 |
22 MPM, 44 asbestos exposed |
4U, 14 Ep, 2 Bi, 2 Sa |
||||
197-3p, 1281, 32-3p |
[110] |
Serum |
1 |
10 MPM, 10 asbestos exposed, 10 healthy |
N/A |
AUC ~ 0.7 for each miRNA |
2 |
20 MPM, 15 asbestos exposed, 14 healthy |
N/A |
||||
126, 21 |
[111] |
Tissue |
1 |
40 FFPE benign pleura, 51 FFPE MPM |
34 Ep, 10 Bi, 75 Sa |
AUC 0.92 for miRNA combination |
Archived cytology samples |
2 |
24 Reactive mesothelium, 29 MPM |
29 Ep |
U – unknown, Ep – epithelioid, Bi – biphasic, Sa – sarcomatoid, N/A – not available, FFPE – formalin fixed paraffin embedded.
Table 3: Potential prognostic miRNA for MPM
miRNA |
Ref |
Source |
Cohort |
Number |
MPM Histological Subtype |
Expression change and survival |
|---|---|---|---|---|---|---|
29c-5p |
[67] |
Tissue |
1 |
37 MPM |
23 Ep, 14 other |
Higher expression = longer survival |
2 |
92 MPM |
58 Ep, 34 other |
||||
17-5p, 30c |
[112] |
Tissue |
1 |
24 MPM |
8 Ep, 8 Bi, 8 Sa |
Lower expression = longer survival in sarcomatoid MPM |
31 |
[113] |
Tissue |
1 |
25 FFPE MPM |
16 Ep, 4 Bi, 5 Sa |
Lower expression = longer survival in sarcomatoid MPM |
21-5p, 23a-3p, 30e-5p, 221-3p, 222-3p, 31-5p |
[114] |
Tissue |
1 |
64 EPP MPM |
47 Ep, 17 Bi |
Signature is associated with longer survival |
2 |
43 PD MPM |
25 Ep, 13 Bi, 5 Sa |
||||
Let-7c-5p, 151a-5p |
[115] |
Tissue |
1 |
52 FFPE MPM |
43 Ep, 8 Bi, 1 Sa |
Lower expression = longer survival |
2 |
16 fresh/frozen MPM |
11 Ep, 4 Bi, 1 Sa |
Ep – epithelioid, Bi – biphasic, Sa – sarcomatoid, FFPE – formalin fixed paraffin embedded, EPP – extrapleural pneumonectomy, PD –Pleurectomy with decortication.
Based on 90 miRNA previously associated with MPM, Kirschner et al., identified miR-625-3p as differentially expressed in the serum/plasma of MPM patients compared to controls. The increased levels of miR-625-3p in two independent cohorts suggest that this miRNA may be a promising diagnostic marker. In both cohorts the AUC reported for miR-625-3p was approximately 0.8 [106]. In the same year, Weber and colleagues reported potential miRNA markers in the cellular fraction of human peripheral blood of MPM patients, asbestos exposed and healthy individuals. MiR-103 was identified as a potential biomarker that could better discriminate MPM from healthy controls (AUC – 0.87) compared to MPM from asbestos exposed individuals (AUC – 0.75) [107].
Serum miRNA have also been analysed in MPM vs asbestos exposed vs healthy individuals. Santarelli and colleagues chose to analyse miR-126 as a potential biomarker for MPM in these groups and found that miR-126 could differentiate asbestos exposed from healthy individuals with a sensitivity of 60% and specificity of 74% and from MPM with a sensitivity of 73% and specificity of 73%. When evaluated in combination with MSLN, decreasing miR-126 and increasing MSLN were indicative of a higher risk of developing MPM [108]. Combining miR-126 with a recently discovered biomarker for MPM miR-132-3p, provided a potential diagnostic signature with an accuracy much higher than the accuracies of using either miR-132-3p or miR-126 alone to discriminate MPM patients from asbestos exposed individuals. The combination of these two miRNA could distinguish MPM from asbestos exposed samples with a sensitivity and specificity of 77% and 86% respectively [109].
Three novel serum miRNA biomarkers were recently identified when Bononi and colleagues compared miRNA in serum from MPM patients, ex-workers exposed to asbestos and healthy individuals. In this study, the miRNA identified included miR-197-3p, miR-1281 and miR-32-3p. MiR-197-3p and miR-32-3p were both expressed significantly higher in MPM patients compared to both asbestos exposed ex-workers and healthy individuals. MiR-1281 was expressed significantly higher in MPM and asbestos exposed ex-workers compared to healthy individuals. These miRNA were moderately good discriminators between the three sample cohorts with diagnostic accuracies (AUC) of around 0.7 [110]. Combining the three miRNA may diagnose MPM and predict individuals at risk of developing this disease more efficiently. This is yet to be determined.
Together, these studies demonstrate the potential of measuring miRNA to diagnose MPM and identify ‘at risk’ individuals. The results are encouraging as the diagnostic accuracies for most of the miRNA signatures are equivalent to or superior than the diagnostic accuracies of the current MPM serum biomarkers mesothelin (AOC 0.7–0.9) [33], osteopontin (AOC 0.83) [35] and fibulin-3 (AOC 0.8) [36, 37]. However, larger prospective studies are required to validate these findings.
Given that most MPM patients develop a PE, it is surprising that the diagnostic potential of miRNA within PE has not been thoroughly investigated. PE are routinely drained to alleviate discomfort [25] and are ideal samples to analyse during the diagnostic process. To date there is only one study that suggests measuring miRNA in PE may assist a diagnosis of MPM. Cappallesso and colleagues compared the miRNA profiles in MPM and reactive mesothelial (RM) archived histological samples prepared from PE. A combination of miR-126 and miR-21 could complement the cytological assessment of PE to differentiate MPM from RM with a sensitivity of 86% and a specificity of 87% [111]. Whilst the results of this study are promising, a subset of only 15 miRNA previously associated with MPM were analysed. Therefore, miRNA with higher diagnostic accuracies could have been overlooked.
MiRNA as prognostic biomarkers for MPM
The first study suggesting miRNA can be used to predict survival outcomes identified miR-29c-5p as an independent prognostic factor for time to disease progression as well as survival after surgical cytoreduction. Higher levels of miR-29c-5p predicted a more favourable prognosis [67]. Likewise, a more favourable outcome in sarcomatoid patients has been associated with reduced levels of miR-17-5p, miR-30c [112] and miR-31 [113].
Kirschner and colleagues reported a miRNA prognostic signature that could be used to predict survival outcomes in surgically resected MPM patients. A combination of six miRNA (miR-21-5p, miR-23a-3p, miR-30e-5p, miR-221-3p, miR-222-3p and miR-31-5p) provided a survival prediction accuracy of approximately 90% for MPM patients who had undergone EPP. When this signature was tested in an independent cohort of patients who had undergone palliative PD, survival was predicted at an accuracy of 72% [114]. Recently, a signature based on the expression of let-7c-5p and miR-151a-5p was identified in 52 MPM tumours as a potential tool for predicting survival. Higher levels of let-7c-5p and miR-151a-5p were associated with a poorer prognosis. This signature was validated in a second cohort of 16 fresh/frozen MPM tumours [115]. Correctly identifying patient prognosis using miRNA could allow for more intensive treatment after surgery to improve survival outcomes.
Whilst a number of miRNA have been identified as potential diagnostic and prognostic targets for MPM, there are discrepancies in the results reported between studies. Therefore, a systematic review and meta-analysis was recently undertaken to try and identify commonly reported miRNA in MPM. Because of the large differences in sample types and technologies used, comparing studies was difficult. The most consistent results were reported in blood and biopsy samples whereas cell line results varied greatly. Therefore, cell lines are best used only for functional assays [116].
Following an assessment of biomarker potential, a circulating miRNA signature based on the expression of miR-126, miR-103 and miR-625 in combination with MSLN was identified for distinguishing asbestos exposed individuals from MPM. Likewise the most consistently reported tissue specific miRNA (miR-16, miR-126, miR-143, miR-145, miR-192, miR-193, miR-200b, miR-203 and miR-652) were suggested to provide a MPM signature [116]. Validation studies are required to assess the clinical relevance of these signatures.
MIRNA IN PMM
Until recently, there was no information on the role of miRNA in PMM. The authors of the first study to address this chose to investigate miR-34a in diffuse malignant peritoneal mesothelioma (DMPM) [117]. MiR-34a was chosen as its aberrant expression has been associated with numerous malignancies and a liposomal nanoparticle formulated synthetic miR-34 (MRX34) recently entered a phase I trial [118]. MiR-34a was analysed in 45 DMPM tissues, seven normal peritoneum samples and five DMPM cell lines by qPCR. MiR-34a was significantly downregulated in DMPM samples compared to controls. Re-expressing miR-34a in DMPM cell lines inhibited cell proliferation and induced apoptosis, although at a variable extent across the five cell lines. The inhibitory effects were suggested to be caused by miR-34a regulating c-MET and AXL signalling. The delay in the induction of apoptosis seen in some of the cell lines following miR-34a overexpression was likely due to the activation of ERK1/2 and AKT. This cytoprotective mechanism was most prominent in the MP115 cell line that was derived from the more aggressive biphasic subtype tumour. Upregulating miR-34a also inhibited cell invasion, tumour growth in xenograft and orthotopic mouse models and influenced the tumour microenvironment by impairing the secretion of angiogenic factors [117]. MiR-34a is the first miRNA identified as a potential target for miRNA replacement therapy in PMM.
CONCLUSIONS
Mesothelioma is a fatal cancer induced by the presence of asbestos fibres. Diagnosis often occurs when the disease has reached an advanced stage and therapeutic modalities remain ineffective. Therefore, patients have a very poor prognosis and a reduced quality of life. The incidence of this disease is increasing as exposure to asbestos still occurs. A new wave of cases is a real concern.
In search of new diagnostic and therapeutic targets, mesothelioma research has evolved to include the analysis of the powerful gene regulators miRNA. A number of downregulated miRNA in mesothelioma, largely due to chromosomal aberrations, have been identified as regulators of oncogenic pathways. The re-expression of these miRNA in mesothelioma cells influences functions such as proliferation, migration, invasion, apoptosis, autophagy, methylation and chemoresistance. The development of TargomiRs is an exciting advancement in the field and a technology that can potentially be used to enhance the expression of any repressed miRNA in patients. It will be important to determine whether the replacement of multiple miRNAs or a combination of miRNAs with chemotherapy and other treatment modalities can enhance survival outcomes. There may also be an opportunity to personalise such a treatment for each patient’s miRNA tumour profile.
Potential diagnostic and prognostic miRNA have also been identified for mesothelioma, however it is important to note there is often a lack of reproducibility in the results across studies. This is most likely caused by differences in study design such as sample and control selection, sample cohort size, using inappropriate controls such as transformed cell lines and analysing only a selection of miRNA. In agreement with a recent commentary by Micollucci and colleagues [119], these issues can be overcome if a standardised approach for sample collection, storage and analysis is developed in a collaborative effort with large sample cohorts. Such an effort is already underway in the lung cancer field with Marzi et al., performing a series of tests to optimise miRNA quantification in serum from a cohort of more than 1000 patients. Variables such as patient fasting, haemolysis, RNA isolation protocol and data normalisation approaches were identified and controlled for and a standardised method for the analysis of miRNA in serum was suggested [120]. With the advancement of technology and an increase in collaborative efforts, novel miRNA diagnostic and therapeutic approaches for mesothelioma can be developed.
Author contributions
This manuscript was written by Kimberly Birnie. The majority of editing and restructuring was completed by Kimberly Birnie and Steven Mutsaers. All other authors contributed to reading, editing and restructuring several drafts.
CONFLICTS OF INTEREST
None.
FINANCIAL SUPPORT
This work was supported by grants from Dust Diseases Board N.S.W, Maurice Blackburn and Sir Charles Gairdner Hospital Research Committee. S Mutsaers was supported on a Cancer Council WA Research Fellowship.
REFERENCES
1. Mutsaers SE. The mesothelial cell. The International Journal of Biochemisty & Cell Biology. 2004; 36:9–16.
2. Mutsaers SE, Birnie K, Lansley S, Herrick SE, Lim CB, Prele CM. Mesothelial cells in tissue repair and fibrosis. Front Pharmacol. 2015; 6:113. https://doi.org/103389/fphar201500113.
3. Mutsaers SE. Mesothelial cells: their structure, function and role in serosal repair. Respirology. 2002; 7:171–91.
4. Robinson BW, Lake RA. Advances in malignant mesothelioma. N Engl J Med. 2005; 353:1591–603.
5. Rohl AN, Langer AM, Moncure G, Selikoff IJ, Fischbein A. Endemic pleural disease associated with exposure to mixed fibrous dust in Turkey. Science. 1982; 216:518–20.
6. Takagi A, Hirose A, Nishimura T, Fukumori N, Ogata A, Ohashi N, Kitajima S, Kanno J. Induction of mesothelioma in p53+/- mouse by intraperitoneal application of multi-wall carbon nanotube. J Toxicol Sci. 2008; 33:105–16.
7. Takagi A, Hirose A, Futakuchi M, Tsuda H, Kanno J. Dose-dependent mesothelioma induction by intraperitoneal administration of multi-wall carbon nanotubes in p53 heterozygous mice. Cancer Sci. 2012; 103:1440–4. https://doi.org/101111/j1349-7006201202318x.
8. Carbone M, Flores EG, Emi M, Johnson TA, Tsunoda T, Behner D, Hoffman H, Hesdorffer M, Nasu M, Napolitano A, Powers A, Minaai M, Baumann F, et al. Combined Genetic and Genealogic Studies Uncover a Large BAP1 Cancer Syndrome Kindred Tracing Back Nine Generations to a Common Ancestor from the 1700s. PLoS Genet. 2015; 11:e1005633. https://doi.org/101371/journalpgen1005633.
9. Goodman JE, Nascarella MA, Valberg PA. Ionizing radiation: a risk factor for mesothelioma. Cancer Causes Control. 2009; 20:1237–54. https://doi.org/101007/s10552-009-9357-4.
10. Pershouse MA, Heivly S, Girtsman T. The role of SV40 in malignant mesothelioma and other human malignancies. Inhal Toxicol. 2006; 18:995–1000.
11. Robinson BM. Malignant pleural mesothelioma: an epidemiological perspective. Ann Cardiothorac Surg. 2012; 1:491–6. https://doi.org/103978/jissn2225-319X20121104.
12. Bibby AC, Tsim S, Kanellakis N, Ball H, Talbot DC, Blyth KG, Maskell NA, Psallidas I. Malignant pleural mesothelioma: an update on investigation, diagnosis and treatment. Eur Respir Rev. 2016; 25:472–486. https://doi.org/101183/160006170063-2016.
13. Musk AW, de Klerk NH, Nowak AK. Asbestos exposure: challenges for Australian clinicians. Med J Aust. 2016; 204:48–9.
14. Kao SCH, Reid G, Lee K, Vardy J, Clarke S, Van Zandwijk N. Malignant mesothelioma. Internal Medicine Journal. 2010; 40:742–50. https://doi.org/10.1111/j.1445-5994.2010.02223.x.
15. Roe OD, Stella GM. Malignant pleural mesothelioma: history, controversy and future of a manmade epidemic. Eur Respir Rev. 2015; 24:115–131.
16. Cao S, Jin S, Cao J, Shen J, Hu J, Che D, Pan B, Zhang J, He X, Ding D, Gu F, Yu Y. Advances in malignant peritoneal mesothelioma. Int J Colorectal Dis. 2015; 30:1–10. https://doi.org/101007/s00384-014-2029-1.
17. Law MR, Hodson ME, Heard BE. Malignant mesothelioma of the pleura: relation between histological type and clinical behaviour. Thorax. 1982; 37:810–5.
18. Johansson L, Linden CJ. Aspects of histopathologic subtype as a prognostic factor in 85 pleural mesotheliomas. Chest. 1996; 109:109–14.
19. Godar M, Liu J, Zhang P, Xia Y, Yuan Q. Primary pericardial mesothelioma: a rare entity. Case Rep Oncol Med. 2013; 2013:283601. https://doi.org/101155/2013/283601.
20. Meyerhoff RR, Yang CF, Speicher PJ, Gulack BC, Hartwig MG, D'Amico TA, Harpole DH, Berry MF. Impact of mesothelioma histologic subtype on outcomes in the Surveillance, Epidemiology, and End Results database. J Surg Res. 2015; 196:23–32. https://doi.org/101016/jjss201501043.
21. Soeberg MJ, Leigh J, Driscoll T, Armstrong B, Young JM, van Zandwijk N. Incidence and survival trends for malignant pleural and peritoneal mesothelioma, Australia, 1982–2009. Occup Environ Med. 2016; 73:187–94. https://doi.org/101136/oemed-2015-103309.
22. Lowry SJ, Weiss NS. Geographic distribution of incidence of pericardial and paratesticular mesotheliomas in the USA. Cancer Causes Control. 2016; 27:1487–1489.
23. Inai K. Pathology of Mesothelioma. Environmental Health and Preventive Medicine. 2008; 13:60–4.
24. Boutin C, Schlesser M, Frenay C, Astoul P. Malignant pleural mesothelioma. European Respiratory Journal. 1998; 12:972–81.
25. Tsujimura T, Torii I, Sato A, Song M, Fukuoka K, Hasegawa S, Nakano T. Pathological and molecular biological approaches to early mesothelioma. International Journal of Clinical Oncology. 2012; 17:40–7. https://doi.org/10.1007/s10147-011-0369-1.
26. Kondola S, Manners D, Nowak AK. Malignant pleural mesothelioma: an update on diagnosis and treatment options. Ther Adv Respir Dis. 2016; 10:275–88. https://doi.org/101177/1753465816628800.
27. Finn RS, Brims FJ, Gandhi A, Olsen N, Musk AW, Maskell NA, Lee YC. Postmortem findings of malignant pleural mesothelioma: a two-center study of 318 patients. Chest. 2012; 142:1267–73.
28. Leung AN, Muller NL, Miller RR. CT in differential diagnosis of diffuse pleural disease. AJR Am J Roentgenol. 1990; 154:487–92.
29. Jamrozik E, de Klerk N, Musk AW. Asbestos-related disease. Internal Medicine Journal. 2011; 41:372–80. https://doi.org/10.1111/j.1445-5994.2011.02451.x.
30. Hjerpe A, Ascoli V, Bedrossian C, Boon M, Creaney J, Davidson B, Dejmek A, Dobra K, Fassina A, Field A, Firat P, Kamei T, Kobayashi T, et al. Guidelines for cytopathologic diagnosis of epithelioid and mixed type malignant mesothelioma. Complementary statement from the International Mesothelioma Interest Group, also endorsed by the International Academy of Cytology and the Papanicolaou Society of Cytopathology. Cytojournal. 2015; 12:26. https://doi.org/104103/1742-6413170726.
31. van der Bij S, Schaake E, Koffijberg H, Burgers JA, de Mol BAJM, Moons KGM. Markers for the non-invasive diagnosis of mesothelioma: a systematic review. Br J Cancer. 2011; 104:1325–33.
32. Creaney J, Robinson BWS. Serum and pleural fluid biomarkers for mesothelioma. Current Opinion in Pulmonary Medicine. 2009; 15:366–70. https://doi.org/10.1097/MCP.0b013e32832b98eb.
33. Pantazopoulos I, Boura P, Xanthos T, Syrigos K. Effectiveness of mesothelin family proteins and osteopontin for malignant mesothelioma. Eur Respir J. 2013; 41:706–15. https://doi.org/101183/0903193600226111.
34. Pass HI, Lott D, Lonardo F, Harbut M, Liu Z, Tang N, Carbone M, Webb C, Wali A. Asbestos exposure, pleural mesothelioma, and serum osteopontin levels. N Engl J Med. 2005; 353:1564–73.
35. Hu ZD, Liu XF, Liu XC, Ding CM, Hu CJ. Diagnostic accuracy of osteopontin for malignant pleural mesothelioma: a systematic review and meta-analysis. Clin Chim Acta. 2014; 433:44–8. https://doi.org/101016/jcca201402024.
36. Pass HI, Levin SM, Harbut MR, Melamed J, Chiriboga L, Donington J, Huflejt M, Carbone M, Chia D, Goodglick L, Goodman GE, Thornquist MD, Liu G, et al. Fibulin-3 as a blood and effusion biomarker for pleural mesothelioma. N Engl J Med. 2012; 367:1417–27. https://doi.org/101056/NEJMoa1115050.
37. Ren R, Yin P, Zhang Y, Zhou J, Zhou Y, Xu R, Lin H, Huang C. Diagnostic value of fibulin-3 for malignant pleural mesothelioma: A systematic review and meta-analysis. Oncotarget. 2016; 7:84851–9. https://doi.org/1018632/oncotarget12707.
38. Campbell NP, Kindler HL. Update on Malignant Pleural Mesothelioma. Semin Respir Crit Care Med. 2011; 32:102,10. https://doi.org/10.1055/s-0031-1272874.
39. Bonelli MA, Fumarola C, La Monica S, Alfieri R. New therapeutic strategies for malignant pleural mesothelioma. Biochem Pharmacol. 2016; 123:8–18. https://doi.org/ 10.1016/j.bcp.2016.07.012.
40. Vogelzang NJ, Rusthoven JJ, Symanowski J, Denham C, Kaukel E, Ruffie P, Gatzemeier U, Boyer M, Emri S, Manegold C, Niyikiza C, Paoletti P. Phase III Study of Pemetrexed in Combination With Cisplatin Versus Cisplatin Alone in Patients With Malignant Pleural Mesothelioma. Journal of Clinical Oncology. 2003; 21:2636–44. https://doi.org/10.1200/jco.2003.11.136.
41. Hoda MA, Klikovits T, Arns M, Dieckmann K, Zochbauer-Muller S, Geltner C, Baumgartner B, Errhalt P, Machan B, Pohl W, Hutter J, Eckmayr J, Studnicka M, et al. Management of malignant pleural mesothelioma-part 2: therapeutic approaches: Consensus of the Austrian Mesothelioma Interest Group (AMIG). Wien Klin Wochenschr. 2016; 128:618–26. https://doi.org/101007/s00508-016-1036-3.
42. Ung YC, Yu E, Falkson C, Haynes AE, Stys-Norman D, Evans WK. The role of radiation therapy in malignant pleural mesothelioma: a systematic review. Radiother Oncol. 2006; 80:13–8.
43. Bagia M, Nowak A. Novel Targeted Therapies and Vaccination Strategies for Mesothelioma. Current Treatment Options in Oncology. 2011; 12:149–62. https://doi.org/10.1007/s11864-011-0149-1.
44. Reid G, Kao SC, Pavlakis N, Brahmbhatt H, MacDiarmid J, Clarke S, Boyer M, van Zandwijk N. Clinical development of TargomiRs, a miRNA mimic-based treatment for patients with recurrent thoracic cancer. Epigenomics. 2016; 8:1079–85. https://doi.org/102217/epi-2016-0035.
45. Yan TD, Haveric N, Carmignani CP, Bromley CM, Sugarbaker PH. Computed tomographic characterization of malignant peritoneal mesothelioma. Tumori. 2005; 91:394–400.
46. Alexander HR Jr, Burke AP. Diagnosis and management of patients with malignant peritoneal mesothelioma. J Gastrointest Oncol. 2016; 7:79–86. https://doi.org/103978/jissn2078-68912015134.
47. Tabata C, Kanemura S, Tabata R, Masachika E, Shibata E, Otsuki T, Nishizaki T, Nakano T. Serum HMGB1 as a diagnostic marker for malignant peritoneal mesothelioma. J Clin Gastroenterol. 2013; 47:684–8. https://doi.org/101097/MCG0b013e318297fa65.
48. Taskin S, Gumus Y, Kiremitci S, Kahraman K, Sertcelik A, Ortac F. Malignant peritoneal mesothelioma presented as peritoneal adenocarcinoma or primary ovarian cancer: case series and review of the clinical and immunohistochemical features. Int J Clin Exp Pathol. 2012; 5:472–8.
49. Cao SB, Jin S, Cao JY, Shen J, Zhang JW, Yu Y. Colonic invasion induced by malignant peritoneal mesothelioma. International Journal of Colorectal Disease. 2014; 29:891–2. https://doi.org/10.1007/s00384-014-1875-1.
50. Yin WJ, Zheng GQ, Chen YF, Chen DQ, Sun NN, Yang YX, Sun XY, Kang LQ. CT differentiation of malignant peritoneal mesothelioma and tuberculous peritonitis. Radiol Med. 2016; 121:253–60. https://doi.org/101007/s11547-015-0609-y.
51. Turner K, Varghese S, Alexander HR Jr. Current concepts in the evaluation and treatment of patients with diffuse malignant peritoneal mesothelioma. J Natl Compr Canc Netw. 2012; 10:49–57.
52. Miyoshi K, Miyoshi T, Siomi H. Many ways to generate microRNA-like small RNAs: non-canonical pathways for microRNA production. Molecular Genetics and Genomics. 2010; 284:95–103. https://doi.org/10.1007/s00438-010-0556-1.
53. Kim NV. MicroRNA biogenesis: coordinated cropping and dicing. Nature Reviews/Molecular Cell Biology. 2005; 6:376–85.
54. Bartel DP. MicroRNAs: Genomics, biogenesis, mechanisms and function. Cell. 2004; 116:281–97.
55. Abdelfattah AM, Park C, Choi MY. Update on non-canonical microRNAs. Biomolecular concepts. 2014; 5:275–87. https://doi.org/10.1515/bmc-2014-0012.
56. Han J, Lee Y, Yeom KH, Nam JW, Heo I, Rhee JK, Sohn SY, Cho Y, Zhang BT, Kim VN. Molecular Basis for the Recognition of Primary microRNAs by the Drosha-DGCR8 Complex. Cell. 2006; 125:887–901.
57. Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K, Shiekhattar R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature. 2005; 436:740–4.
58. Schwarz DS, Hutvágner G, Du T, Xu Z, Aronin N, Zamore PD. Asymmetry in the Assembly of the RNAi Enzyme Complex. Cell. 2003; 115:199–208.
59. Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA translation and stability by microRNAs. The Annual Review of Biochemistry. 2010; 79:351–79.
60. Daugaard I, Hansen TB. Biogenesis and Function of Ago-Associated RNAs. Trends in Genetics. 2017; 33:208–19. https://doi.org/10.1016/j.tig.2017.01.003.
61. Ambros V. The functions of animal microRNAs. Nature. 2004; 431:350–5.
62. Olena AF, Patton JG. Genomic Organisation of microRNAs. Journal of Cell Physiology. 2010; 222:540–5.
63. Shah MY, Ferrajoli A, Sood AK, Lopez-Berestein G, Calin GA. microRNA Therapeutics in Cancer - An Emerging Concept. EBioMedicine. 2016; 12:34–42. https://doi.org/101016/jebiom201609017.
64. Etheridge A, Lee I, Hood L, Galas D, Wang K. Extracellular microRNA: A new source of biomarkers. Mutation Research/Fundamental and Molecular Mechanisms of Mutagenesis. 2011; 717:85–90.
65. Farazi TA, Spitzer JI, Morozov P, Tuschl T. miRNAs in human cancer. Journal of Pathology. 2011; 223:102–15.
66. Guled M, Lahti L, Lindholm PM, Salmenkivi K, Bagwan I, Nicholson AG, Knuutila S. CDKN2A, NF2, and JUN are dysregulated among other genes by miRNAs in malignant mesothelioma—A miRNA microarray analysis. Genes, Chromosomes and Cancer. 2009; 48:615–23. https://doi.org/10.1002/gcc.20669.
67. Pass HI, Goparaju C, Ivanov S, Donington J, Carbone M, Hoshen M, Cohen D, Chajut A, Rosenwald S, Dan H, Benjamin S, Aharonov R. hsa-miR-29c* Is Linked to the Prognosis of Malignant Pleural Mesothelioma. Cancer Research. 2009; 70:1916–24. https://doi.org/10.1158/0008-5472.can-09-3993.
68. Ivanov SV, Goparaju CMV, Lopez P, Zavadil J, Toren-Haritan G, Rosenwald S, Hoshen M, Chajut A, Cohen D, Pass HI. Pro-tumorigenic Effects of miR-31 Loss in Mesothelioma. Journal of Biological Chemistry. 2010; 285:22809–17. https://doi.org/10.1074/jbc.M110.100354.
69. MacKeigan JP, Murphy LO, Blenis J. Sensitized RNAi screen of human kinases and phosphatases identifies new regulators of apoptosis and chemoresistance. Nat Cell Biol. 2005; 7:591–600.
70. Mi J, Dziegielewski J, Bolesta E, Brautigan DL, Larner JM. Activation of DNA-PK by ionizing radiation is mediated by protein phosphatase 6. PLoS One. 2009; 4:e4395. https://doi.org/101371/journalpone0004395.
71. Balzeau J, Menezes MR, Cao S, Hagan JP. The LIN28/let-7 Pathway in Cancer. Front Genet. 2017; 8:31. https://doi.org/103389/fgene201700031.
72. Khodayari N, Mohammed KA, Goldberg EP, Nasreen N. EphrinA1 inhibits malignant mesothelioma tumor growth via let-7 microRNA-mediated repression of the RAS oncogene. Cancer Gene Ther. 2011; 18:806–16.
73. Lee HY, Mohammed KA, Kaye F, Sharma P, Moudgil BM, Clapp WL, Nasreen N. Targeted delivery of let-7a microRNA encapsulated ephrin-A1 conjugated liposomal nanoparticles inhibit tumor growth in lung cancer. Int J Nanomedicine. 2013; 8:4481–94. https://doi.org/102147/IJNS41782.
74. Sohn EJ, Won G, Lee J, Yoon SW, Lee I, Kim HJ, Kim SH. Blockage of epithelial to mesenchymal transition and upregulation of let 7b are critically involved in ursolic acid induced apoptosis in malignant mesothelioma cell. Int J Biol Sci. 2016; 12:1279–1288. https://doi.org/107150/ijbs13453.
75. Reid G, Pel ME, Kirschner MB, Cheng YY, Mugridge N, Weiss J, Williams M, Wright C, Edelman JJ, Vallely MP, McCaughan BC, Klebe S, Brahmbhatt H, et al. Restoring expression of miR-16: a novel approach to therapy for malignant pleural mesothelioma. Ann Oncol. 2013; 24:3128–35. https://doi.org/101093/annonc/mdt412.
76. Kao SC, Cheng YY, Williams M, Kirschner MB, Madore J, Lum T, Sarun KH, Linton A, McCaughan B, Klebe S, van Zandwijk N, Scolyer RA, Boyer MJ, et al. Tumour suppressor microRNAs contribute to the regulation of PD-L1 expression in malignant pleural mesothelioma. J Thorac Oncol. 2017; 12:1421–33. https://doi.org/10.1016/j.jtho.2017.05.024.
77. Kubo T, Toyooka S, Tsukuda K, Sakaguchi M, Fukazawa T, Soh J, Asano H, Ueno T, Muraoka T, Yamamoto H, Nasu Y, Kishimoto T, Pass HI, et al. Epigenetic Silencing of MicroRNA-34b/c Plays an Important Role in the Pathogenesis of Malignant Pleural Mesothelioma. Clinical Cancer Research. 2011; 17:4965–74. https://doi.org/10.1158/1078-0432.ccr-10-3040.
78. Maki Y, Asano H, Toyooka S, Soh J, Kubo T, Katsui K, Ueno T, Shien K, Muraoka T, Tanaka N, Yamamoto H, Tsukuda K, Kishimoto T, et al. MicroRNA miR-34b/c enhances cellular radiosensitivity of malignant pleural mesothelioma cells. Anticancer Res. 2012; 32:4871–5.
79. Tanaka N, Toyooka S, Soh J, Tsukuda K, Shien K, Furukawa M, Muraoka T, Maki Y, Ueno T, Yamamoto H, Asano H, Otsuki T, Miyoshi S. Downregulation of microRNA-34 induces cell proliferation and invasion of human mesothelial cells. Oncol Rep. 2013; 29:2169–74. https://doi.org/103892/or20132351.
80. Andersen M, Trapani D, Ravn J, Sorensen JB, Andersen CB, Grauslund M, Santoni-Rugiu E. Methylation-associated Silencing of microRNA-126 and its Host Gene EGFL7 in Malignant Pleural Mesothelioma. Anticancer Res. 2015; 35:6223–9.
81. Tomasetti M, Nocchi L, Staffolani S, Manzella N, Amati M, Goodwin J, Kluckova K, Nguyen M, Strafella E, Bajzikova M, Peterka M, Lettlova S, Truksa J, et al. MicroRNA-126 suppresses mesothelioma malignancy by targeting IRS1 and interfering with the mitochondrial function. Antioxid Redox Signal. 2014; 21:2109–25. https://doi.org/101089/ars20135215.
82. Tomasetti M, Monaco F, Manzella N, Rohlena J, Rohlenova K, Staffolani S, Gaetani S, Ciarapica V, Amati M, Bracci M, Valentino M, Goodwin J, Nguyen M, et al. MicroRNA-126 induces autophagy by altering cell metabolism in malignant mesothelioma. Oncotarget. 2016; 7:36338–52. https://doi.org/1018632/oncotarget8916.
83. Xu Y, Zheng M, Merritt RE, Shrager JB, Wakelee H, Kratzke RA, Hoang CD. miR-1 induces growth arrest and apoptosis in malignant mesothelioma. Chest. 2013; 144:1632–43. https://doi.org/101378/chest12-2770.
84. Amatya VJ, Mawas AS, Kushitani K, Mohi El-Din MM, Takeshima Y. Differential microRNA expression profiling of mesothelioma and expression analysis of miR-1 and miR-214 in mesothelioma. Int J Oncol. 2016; 48:1599–607. https://doi.org/103892/ijo20163358.
85. Mawas AS, Amatya VJ, Suzuki R, Kushitani K, Mohi El-Din MM, Takeshima Y. PIM1 knockdown inhibits cell proliferation and invasion of mesothelioma cells. Int J Oncol. 2017; 50:1029–1034. https://doi.org/103892/ijo20173863.
86. Cioce M, Ganci F, Canu V, Sacconi A, Mori F, Canino C, Korita E, Casini B, Alessandrini G, Cambria A, Carosi MA, Blandino R, Panebianco V, et al. Protumorigenic effects of mir-145 loss in malignant pleural mesothelioma. Oncogene. 2014; 33:5319–31. https://doi.org/101038/onc2013476.
87. Blum W, Pecze L, Felley-Bosco E, Wu L, de Perrot M, Schwaller B. Stem Cell Factor-Based Identification and Functional Properties of In Vitro-Selected Subpopulations of Malignant Mesothelioma Cells. Stem Cell Reports. 2017; 8:1005–17. https://doi.org/ 10.1016/j.stemcr.2017.02.005.
88. Vencken S, Hassan T, McElvaney NG, Smith SG, Greene CM. miR-CATCH: microRNA capture affinity technology. Methods Mol Biol. 2015; 1218:365–73. https://doi.org/101007/978-1-4939-1538-5_23.
89. De Santi C, Vencken S, Blake J, Haase B, Benes V, Gemignani F, Landi S, Greene CM. Identification of MiR-21-5p as a Functional Regulator of Mesothelin Expression Using MicroRNA Capture Affinity Coupled with Next Generation Sequencing. PLoS One. 2017; 12:e0170999. https://doi.org/101371/journalpone0170999.
90. Birnie KA, Yip YY, Ng DC, Kirschner MB, Reid G, Prele CM, Musk AW, Lee YC, Thompson PJ, Mutsaers SE, Badrian B. Loss of miR-223 and JNK Signaling Contribute to Elevated Stathmin in Malignant Pleural Mesothelioma. Mol Cancer Res. 2015; 13:1106–18. https://doi.org/101158/1541-7786MCR-14-0442.
91. Kim JY, Harvard C, You L, Xu Z, Kuchenbecker K, Baehner R, Jablons D. Stathmin is overexpressed in malignant mesothelioma. Anticancer Res. 2007; 27:39–44.
92. Belletti B, Nicoloso MS, Schiappacassi M, Berton S, Lovat F, Wolf K, Canzonieri V, D'Andrea S, Zucchetto A, Friedl P, Colombatti A, Baldassarre G. Stathmin activity influences sarcoma cell shape motility and metastatic potential. Molecular Biology of the Cell. 2008; 19:2003–13.
93. Walter RF, Vollbrecht C, Werner R, Wohlschlaeger J, Christoph DC, Schmid KW, Mairinger FD. microRNAs are differentially regulated between MDM2-positive and negative malignant pleural mesothelioma. Oncotarget. 2016; 7:18713–21. https://doi.org/1018632/oncotarget7666.
94. Soini Y, Kinnula V, Kaarteenaho-Wiik R, Kurttila E, Linnainmaa K, Paakko P. Apoptosis and expression of apoptosis regulating proteins bcl-2, mcl-1, bcl-X, and bax in malignant mesothelioma. Clin Cancer Res. 1999; 5:3508–15.
95. Khodayari N, Mohammed KA, Lee H, Kaye F, Nasreen N. MicroRNA-302b targets Mcl-1 and inhibits cell proliferation and induces apoptosis in malignant pleural mesothelioma cells. Am J Cancer Res. 2016; 6:1996–2009.
96. Williams M, Kirschner MB, Cheng YY, Hanh J, Weiss J, Mugridge N, Wright CM, Linton A, Kao SC, Edelman JJ, Vallely MP, McCaughan BC, Cooper W, et al. miR-193a-3p is a potential tumor suppressor in malignant pleural mesothelioma. Oncotarget. 2015; 6:23480–95. https://doi.org/10.18632/oncotarget.4346.
97. Cheng YY, Wright CM, Kirschner MB, Williams M, Sarun KH, Sytnyk V, Leshchynska I, Edelman JJ, Vallely MP, McCaughan BC, Klebe S, van Zandwijk N, Lin RC, et al. KCa1.1, a calcium-activated potassium channel subunit alpha 1, is targeted by miR-17-5p and modulates cell migration in malignant pleural mesothelioma. Mol Cancer. 2016; 15:44. https://doi.org/101186/s12943-016-0529-z.
98. Schramm A, Opitz I, Thies S, Seifert B, Moch H, Weder W, Soltermann A. Prognostic significance of epithelial-mesenchymal transition in malignant pleural mesothelioma. Eur J Cardiothorac Surg. 2010; 37:566–72. https://doi.org/101016/jejcts200908027.
99. Fassina A, Cappellesso R, Guzzardo V, Dalla Via L, Piccolo S, Ventura L, Fassan M. Epithelial-mesenchymal transition in malignant mesothelioma. Mod Pathol. 2012; 25:86–99. https://doi.org/101038/modpathol2011144.
100. Cao LY, Yang J, Fu XG, Lin YH, Lin F, Huang BY. [The MicroRNA miR-205 inhibits epithelial-messenchymal transition in HK-2 cells by down-regulating ZEB1 and ZEB2 expressions]. [Article in Chinese]. Nan Fang Yi Ke Da Xue Xue Bao. 2016; 36:1700–5.
101. MacDiarmid JA, Mugridge NB, Weiss JC, Phillips L, Burn AL, Paulin RP, Haasdyk JE, Dickson KA, Brahmbhatt VN, Pattison ST, James AC, Al Bakri G, Straw RC, et al. Bacterially derived 400 nm particles for encapsulation and cancer cell targeting of chemotherapeutics. Cancer Cell. 2007; 11:431–45.
102. MacDiarmid JA, Brahmbhatt H. Minicells: versatile vectors for targeted drug or si/shRNA cancer therapy. Curr Opin Biotechnol. 2011; 22:909–16. https://doi.org/101016/jcopbio201104008.
103. Gee GV, Koestlr DC, Christensen BC, Sugarbaker DJ, Ugolini D, Ivaldi GP, Resnick MB, Houseman E, Andres, Kelsey KT, Marsit CJ. Downregutaled microRNAs in the differential diagnosis of malignant pleural mesothelioma. International Journal of Cancer. 2010; 127:2859–69.
104. Benjamin H, Lebanony D, Rosenwald S, Cohen L, Giborri H, Barabash N, Ashkenazi K, Goren E, Meiri E, Morgenstern S, Perelman M, Barshack I, Goren Y, et al. A diagnostic assay based on microRNA expression accurately identifies malignant pleural mesothelioma. Journal of Molecular Diagnostics. 2010; 12:771–9.
105. Andersen M, Grauslund M, Ravn J, Sorensen JB, Andersen CB, Santoni-Rugiu E. Diagnostic potential of miR-126, miR-143, miR-145, and miR-652 in malignant pleural mesothelioma. J Mol Diagn. 2014; 16:418–30. https://doi.org/101016/jjmoldx201403002.
106. Kirschner MB, Cheng YY, Badrian B, Kao SC, Creaney J, Edelmann JJ, Armstrong NJ, Valley MP, Musk AW, Robinson BWS, McCaughan BC, Klebe S, Mutsaers SE, et al. Increased circulating miR-625-3p: a potential biomarker for patients with malignant pleural mesothelioma. Journal of Thoracic Oncology. 2012; 7:1184–91.
107. Weber DG, Johnen G, Bryk O, Jockel KH, Bruning T. Identification of miRNA-103 in the Cellular Fraction of Human Peripheral Blood as a Potential Biomarker for Malignant Mesothelioma "A Pilot Study". PLoS ONE. 2012; 7:e30221.
108. Santarelli L, Strafella E, Staffolani S, Amati M, Emanuelli M, Sartini D, Pozzi V, Carbonari D, Bracci M, Pignotti E, Mazzanti P, Sabbatini A, Ranaldi R, et al. Association of MiR-126 with Soluble Mesothelin-Related Peptides, a Marker for Malignant Mesothelioma. PLoS One. 2011; 6:e18232.
109. Weber DG, Gawrych K, Casjens S, Brik A, Lehnert M, Taeger D, Pesch B, Kollmeier J, Bauer TT, Johnen G, Bruning T. Circulating miR-132-3p as a Candidate Diagnostic Biomarker for Malignant Mesothelioma. Dis Markers. 2017; 2017:9280170. https://doi.org/101155/2017/9280170.
110. Bononi I, Comar M, Puozzo A, Stendardo M, Boschetto P, Orecchia S, Libener R, Guaschino R, Pietrobon S, Ferracin M, Negrini M, Martini F, Bovenzi M, Tognon M. Circulating microRNAs found dysregulated in ex-exposed asbestos workers and pleural mesothelioma patients as potential new biomarkers. Oncotarget. 2016; 7:82700–11. https://doi.org/1018632/oncotarget12408.
111. Cappellesso R, Nicole L, Caroccia B, Guzzardo V, Ventura L, Fassan M, Fassina A. Young investigator challenge: MicroRNA-21/MicroRNA-126 profiling as a novel tool for the diagnosis of malignant mesothelioma in pleural effusion cytology. Cancer Cytopathol. 2016; 124:28–37. https://doi.org/101002/cncy21646.
112. Busacca S, Germano S, De Cecco L, Rinaldi M, Comoglio F, Favero F, Murer B, Mutti L, Pierotti M, Gaudino G. microRNA signature of malignant meosthlelioma with potential diagnostic and prognostic implications. American Journal of Respiratory Cell and Molecular Biology. 2010; 42:312–9.
113. Matsumoto S, Nabeshima K, Hamasaki M, Shibuta T, Umemura T. Upregulation of microRNA-31 associates with a poor prognosis of malignant pleural mesothelioma with sarcomatoid component. Med Oncol. 2014; 31:303. https://doi.org/101007/s12032-014-0303-2.
114. Kirschner MB, Cheng YY, Armstrong NJ, Lin RC, Kao SC, Linton A, Klebe S, McCaughan BC, van Zandwijk N, Reid G. MiR-score: a novel 6-microRNA signature that predicts survival outcomes in patients with malignant pleural mesothelioma. Mol Oncol. 2015; 9:715–26. https://doi.org/101016/jmolonc201411007.
115. De Santi C, Melaiu O, Bonotti A, Cascione L, Di Leva G, Foddis R, Cristaudo A, Lucchi M, Mora M, Truini A, Tironi A, Murer B, Boldorini R, et al. Deregulation of miRNAs in malignant pleural mesothelioma is associated with prognosis and suggests an alteration of cell metabolism. Sci Rep. 2017; 7:3140. https://doi.org/101038/s41598-017-02694-0.
116. Micolucci L, Akhtar MM, Olivieri F, Rippo MR, Procopio AD. Diagnostic value of microRNAs in asbestos exposure and malignant mesothelioma: systematic review and qualitative meta-analysis. Oncotarget. 2016; 7:58606–37. https://doi.org/1018632/oncotarget9686.
117. El Bezawy R, De Cesare M, Pennati M, Deraco M, Gandellini P, Zuco V, Zaffaroni N. Antitumor activity of miR-34a in peritoneal mesothelioma relies on c-MET and AXL inhibition: persistent activation of ERK and AKT signaling as a possible cytoprotective mechanism. J Hematol Oncol. 2017; 10:19. https://doi.org/101186/s13045-016-0387-6.
118. Beg MS, Brenner AJ, Sachdev J, Borad M, Kang YK, Stoudemire J, Smith S, Bader AG, Kim S, Hong DS. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Invest New Drugs. 2017; 35:180–8. https://doi.org/101007/s10637-016-0407-y.
119. Micolucci L, Rippo MR, Olivieri F, Procopio AD. Progress of research on microRNAs with diagnostic value in asbestos exposure: A call for method standardization. Biosci Trends. 2017; 11:105–9. https://doi.org/105582/bst201601249.
120. Marzi MJ, Montani F, Carletti RM, Dezi F, Dama E, Bonizzi G, Sandri MT, Rampinelli C, Bellomi M, Maisonneuve P, Spaggiari L, Veronesi G, Bianchi F, et al. Optimization and Standardization of Circulating MicroRNA Detection for Clinical Application: The miR-Test Case. Clin Chem. 2016; 62:743–54. https://doi.org/101373/clinchem2015251942.
121. Jean D, Daubriac J, Le Pimpec-Barthes F, Galateau-Salle F, Jaurand MC. Molecular Changes in Mesothelioma With an Impact on Prognosis and Treatment. Archives of Pathology & Laboratory Medicine. 2012; 136:277–93. https://doi.org/10.5858/arpa.2011-0215-RA.
122. Cheng JQ, Jhanwar SC, Klein WM, Bell DW, Lee WC, Altomare DA, Nobori T, Olopade OI, Buckler AJ, Testa JR. Alterations and Deletion Mapping of 9p21-p22 in Malignant Mesothelioma. Cancer Research. 1994; 54:5547–51.
123. Minami D, Takigawa N, Kato Y, Kudo K, Isozaki H, Hashida S, Harada D, Ochi N, Fujii M, Kubo T, Ohashi K, Sato A, Tanaka T, et al. Downregulation of TBXAS1 in an iron-induced malignant mesothelioma model. Cancer Science. 2015; 106:1296–302. https://doi.org/10.1111/cas.12752.
124. Krismann M, Müller KM, Jaworska M, Johnen G. Severe Chromosomal Aberrations in Pleural Mesotheliomas with Unusual Mesodermal Features. The Journal of Molecular Diagnostics. 2000; 2:209–16. https://doi.org/10.1016/s1525-1578(10)60639-3.
125. Takeda M, Kasai T, Enomoto Y, Takano M, Morita K, Kadota E, Iizuka N, Maruyama H, Nonomura A. Genomic gains and losses in malignant mesothelioma demonstrated by FISH analysis of paraffin-embedded tissues. Journal of Clinical Pathology. 2011. https://doi.org/10.1136/jclinpath-2011–200208.

