INTRODUCTION
Equine infectious anemia virus (EIAV) is a member of the Lentivirus genus of the Retroviridae family. The Lentivirus genus also includes human immunodeficiency virus (HIV), simian immunodeficiency virus (SIV), feline immunodeficiency virus (FIV), bovine immunodeficiency virus (BIV), and Maedi-visnavirus (MVV) EIAV almost infected equids worldwide. It causes a persistent infection characterized by recurring febrile episodes associated with viremia, fever, thrombocytopenia, and wasting symptoms [1]. EIAV raised great concern among the international veterinary community due to the serious economic losses it caused, as well as the unique nature of this pathogen. In China, scientists successfully controlled the spread of EIAV, especially when an effective vaccine was widely applied. Here, we will focus on the control of EIAV in China.
EIAV introduction and epidemic in China
Reports and records of EIAV did not appear in China until the early 20th century. Japan spread EIAV to China during World War II. Due to an incomplete quarantine, horses with a latent infection were traded from the former Soviet Union and Mongolia, which also introduced EIAV to China. From 1954 to 1959, EIAV mainly occurred in Heilongjiang, Jilin, Liaoning, Inner Mongolia, Yunnan, Hebei, and the Shanxi province. EIAV prevalence in Shandong, Gansu, Jiangsu, and the Anhui Province was due to infected horses purchased from Xinjiang. EIAV prevalence in Hebei, Beijing, Tianjin, Shaanxi, and Henan provinces was caused by EIAV-latent horses purchased from Liaoning, Jilin, and Heilongjiang. The Chinese EIAV epidemic was started by horses traded from foreign countries, and gradually spread from north to south, eventually inducing a nationwide outbreak.
From 1954 to 1989, EIAV broke out in 23 provinces to varying degrees, contaminating 77% of the country’s total provinces (Figure 1) [2]. In the 1950s, EIAV was reported in only nine provinces: Heilongjiang, Inner Mongolia, Yunnan, Guizhou, Jilin, Guangdong, Guangxi, Hebei, and Shanxi. During the 1960s, the epidemic spread to 10 more provinces for a total of 19, including Liaoning, Beijing, Xinjiang, Shandong, Tianjin, Shaanxi, Henan, Gansu, Jiangsu, and Anhui. EIAV was reported in two more provinces in the 1970s, Qinghai and Sichuan, raising the total to 21 provinces. EIAV expanded into Ningxia and Hubei in the 1980s, capping the total at 23 provinces. Among all these regions, Heilongjiang, Liaoning, Jilin, Inner Mongolia, Hebei, Henan, Tianjin, Beijing, Shandong, Shanxi, Shaanxi, and Yunnan were the most infected [2]. The Chinese EIAV epidemic was stimulated by four factors: importing infected animals, frequent movement and trading of infected horses without effective control, no early detection method at early stage, and not sterilizing medical devices [2].
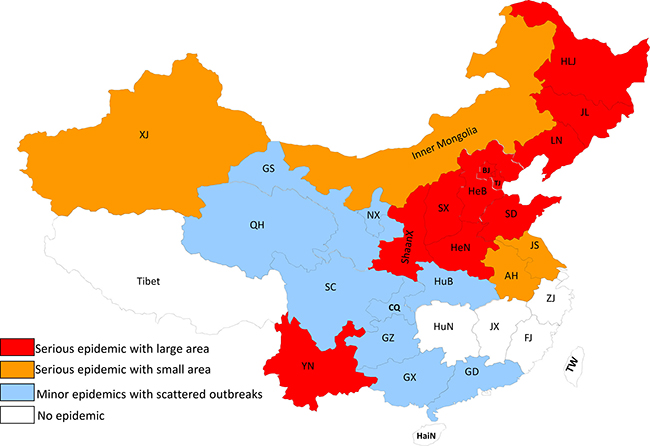
Figure 1: EIAV epidemic in China from 1954 to 1990. The EIAV epidemic state in China from 1954 to 1990 is summarized. The different colors represent epidemic degree of EIAV in different provinces. The jacinth represents serious epidemic with large area; the brown represents serious epidemic with small area; the light blue represents minor epidemics with scattered outbreaks.
EIAV control in China
EIAV prevention and control is accompanied by the development of detection technology. Because quarantine was the only control measure available in the early stages of EIAV, clinical comprehensive diagnosis (CCD) was widely applied [2]. EIAV could not be confirmed according to just one of the CCD methods, which included epidemiological investigation, clinical, blood, and pathology examination. CCD was used to screen possibly infected horses from 1954 to 1973 in China. EIAV-infected horses were quarantined or slaughtered to prevent spread of the virus. However, CCD could not detect latent infection which led to the reemergence of EIAV.
From 1974 to 1978, two serological methods dependent on antibody detection were developed to battle EIAV. The first was the complement-fixation (CF) test which detected specific antibodies that fixed complement when interacting with the corresponding antigen. Hemolysis was used as an indicator system to detect residual complement [2]. Another serological test based on agar gel immunodiffusion (AGID) [2]. EIAV-seropositive animals were either euthanized or kept inquarantine for the rest of their life, depending on local regulations. The use of these two tests controlled the EIAV infection in China. Not only were CCD, CF, and AGID time consuming and difficult to interpret, but some infected horses were misdiagnosed because some animals are positive for only one of the two serological tests. These disadvantages were overcome in 1979 when an enzyme-linked immunosorbent assay (ELISA) was developed to detect EIAV antibody [3], which was simple, quick, sensitive, and specific.
It is impossible to eradicate EIAV merely with quarantine measures. Antibody levels are not constant in the infected animal, so it is not possible to detect all the subclinical cases. An integrated control program including vaccination and quarantine was adopted. This not only eliminated most of the infected animals and reduced EIAV spread, but also enabled most of the healthy horses to obtain immunity. These measures accelerated the control and eradication of EIAV in China (Table 1; Figure 2).
Table 1: Statistics of EIAV control from 1976 to 1990 in China
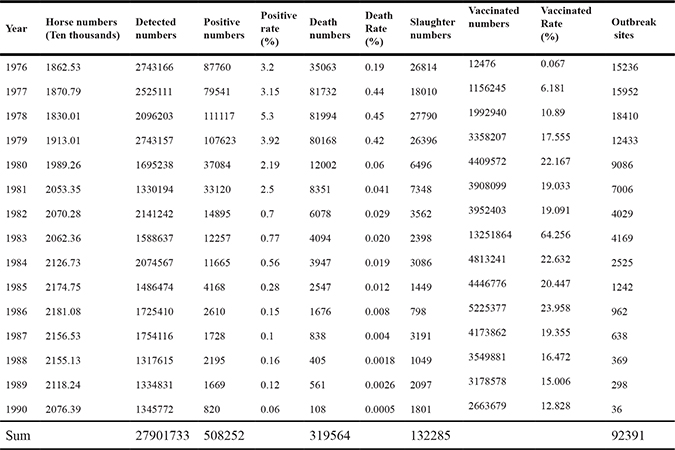
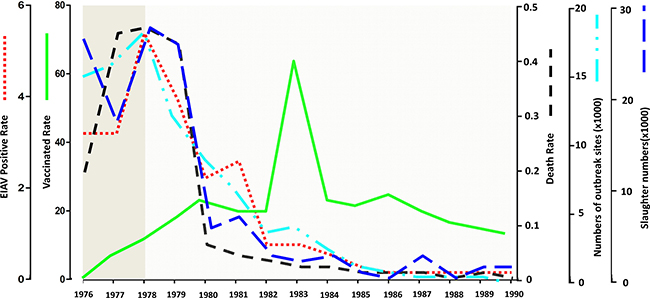
Figure 2: Statistics of EIAV control in China from 1976 to 1990. The EIAV control in China from 1976 to 1990 is summarized. The red dotted line represents EIAV positive rate, the green line represents vaccinated rate; the black dotted line represents death rate; the wathet dotted line represents numbers of outbreak sites; the deep blue dotted line represent slaughter numbers.
EIAV live attenuated vaccine development
For more than a century, the most successful vaccines were live attenuated vaccines which have been widely used to control the epidemics of many viruses, such as smallpox, polio, and measles [4, 5]. Since the 1960s, EIAV brought great losses to Chinese agriculture. After nearly 20 years of effort, Chinese scientists developed an effective live attenuated EIAV vaccine protected horses not only against homologous EIAV strains, but some heterogeneous virus strains as well [6–8]. The Chinese live attenuated EIAV vaccine is the only large-scale lenti-viral vaccine in the world, and serves as a model for the development of other lenti-virus vaccines [9].
Chinese scientists isolated a wild highly-pathogenic strain from horses in Liaoning, EIAVLN, and attempted to adapt this virus to several other animals, tissues, or cell lines. All these attempts failed, except in donkey. The virus experienced 117 passages (EIAVDV117) in donkey, with enhanced pathogenicity instead of attenuation. After 121 serial passages of EIAVDV117 strain on donkey leukocyte (EIAVDLV121), there was a progressive loss of virulence (Figure 3). EIAVDLV121 is an effective vaccine that can elicit protective immunity [2, 8]. This vaccine provided 85% protection for horse and 100% for donkey. However, it was costly and time consuming to isolate donkey leukocyte, so scientists successfully cultivatedEIAVDLV121 in fetal donkey dermal cells with 13 passages. This EIAVFDDV13 vaccine was as effective as EIAVDLV121 (Figure 3) [6]. Since 1976, this attenuated vaccine has been administered to over 70 million horses, mules, and donkeys, effectively controlling the EIAV epidemic in China.

Figure 3: Development of Chinese EIAV live attenuated vaccine. The development of Chinese EIAV live attenuated vaccine is summarized. EIAVLN:LN strain serial 16 passages in horse; EIAVDV117: LN strain serial 117 passages in donkey; EIAVDLV121:DV117 strain serial 121 passages in donkey leukocyte; EIAVFDDV13:DLV121 strain serial 13 passages in fetal donkey dermal cell.
Virulence determinant of EIAV
Previous studies utilizing chimeric proviruses in which parental viruses are acutely virulent or avirulent have identified critical regions that affect acute virulence. These data showed that U3 regions in the viral LTR, surface envelope protein, and the accessory S2 gene strongly affects EIAV virulence [10–12]. For the Chinese EIAV live attenuated vaccine, the LTR sequences of the different generations manifested stable genetic variations and mainly occurred in the transcriptional start site, the initial base of transactivation-response element (TAR), and the pre-mRNA cleavage site at the R-U5 boundary [13]. LTR sequence diversity was increased over passages in MDMs, and the genetic distances gradually increased between these MDMs-adapted EIAV strains to the parental virulent strain EIAVDV117 [14]. However, LTR was not the sole determinant of virulence for EAIV [15]. LTRs of the vaccine were variable, while EIAV LTRs of virulent strains were homologous [14]. LTRs from EIAV (DLA) showed higher Tat transactivated activity than LTRs from virulent strains.
By using chimeric clones of wild-type LTR and vaccine LTR, the main differences of Tat transactivated activity were mapped to the changes of R region, rather than U3 region [16]. For S2, reverse mutation of the virulence-associated S2 gene does not cause an attenuated equine infectious anemia virus strain to regain its pathogenicity [17]. Based on a full-length infectious clone of the EIAV vaccine strain, the envelope region supports the virulence and pathogenicity of EIAV [18, 19]. Furthermore, for the transmembrane envelope (TM), there was a high frequency of GP35 in vaccine strains rather than GP45 in virulent strain. This was because the vaccine strain had a high frequency of a premature stop codon, which generated a 154-residue truncation at the C-terminus. AC-terminal truncation of the transmembrane protein of the vaccine alters its in vitro replication and weakens its potential pathogenicity [20]. Cytoplasmic tail truncation of EIAV Env also increased cell necrosis [21].
Immune response induced by EIAV vaccine
Vaccinated horses can produce a significant humoral immune response. Two weeks after immunization, complement-fixation (CF) antibody and AGID antibody can be detected (Figure 4). Three weeks after immunization, the CF and AGID antibody sero-conversion rate were 100% and persisted to day 45. After 45 days, CF and AGID antibody positive rate began to decline. At 360 days, this rate was decreased to 22%. Neutralizing antibodies (Nab) appeared later than CF and AGID antibody. Nab can be detected 60 days after immunization. Nab levels peaked at day180, and the Nab sero-conversion rate was 100% and sustained at a high level over a long period with little variation (Figure 4). This indicated that Nab may promote vaccine-induced immunity [22].
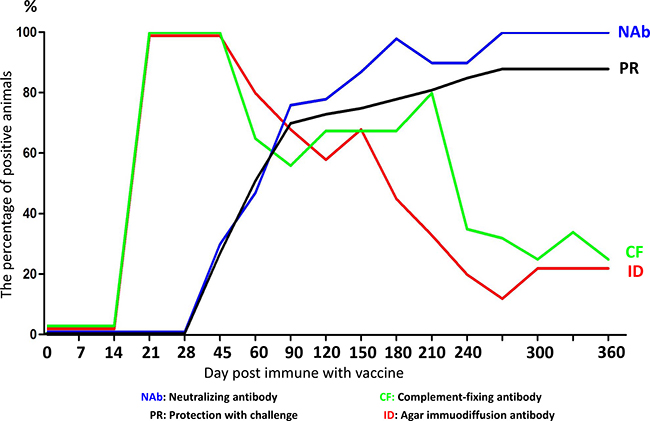
Figure 4: Correlation between antibody response and challenge protection. The horses were vaccinated and the correlation between antibody response and challenge protection was evaluated at different days post immune with vaccine. The blue line represents neutralizing antibody; the green line represents complement-fixing antibody; the black line represents protection with challenge; the red line represents agar immuodiffusion antibody.
The EIAV-specific cellular immune response was also evaluated. CD8 positive T cells increased after immunization and induced an EIAV-specific CTL response [7, 23, 24]. The vaccine group induced high levels of Th1 cytokines, and the EIAV-specific cytokines induced by the attenuated EIAV vaccine may contribute to the protective immune response against EIA disease [25, 26].
Furthermore, vaccine-induced TLR3 activation and INF beta production were higher than in the molecular clone strains [27]. Infection of equine monocyte-derived macrophages with an attenuated equine infectious anemia virus (EIAV) strain induces a strong resistance to infection by a virulent EIAV strain [28].
EIAV vaccine divergence and immune protection
The diverse immunogen composition of the EIAV vaccine may induce protective immunity [29–31]. Although the molecular clone strain and parental strain were replicated equally in vitro and in vivo, the molecular strain induced less protection and lower specific neutralizing production [32]. These findings indicated that diverse immunogen composition may stimulate the immune protection induced by the parental strain. In vivo evolution of the EIAV vaccine was investigated, and the envelope gp90 gene variation was associated with declining diversity after vaccination. This trend coincided with the maturation of immunity to EIAV [30]. Diverse immunogen composition may induce a high titer of neutralizing antibody.
EIAV and innate restriction factors
Innate restriction factors are host factors which could intervene against every step of the viral life cycle and inhibit viral replication [33, 34]. Many innate restriction factors have antiviral functions against EIAV infection. Fv1 was first reported in the 1970s and was homologous to gag gene of endogenous retrovirus (ERV), which was as a prototypic restriction factor protecting mice against murine leukemia virus (MLV) infection [35, 36]. Fv1 could also inhibit equine infectious anemia virus (EIAV) replication, indicating that Fv1 could have a broader antiviral activity [37].
Tetherin (also known as HM1.24, BST-2, or CD317) is a type II single-pass transmembrane protein with an unusual topology, including an N-terminal cytoplasmic tail (CT), a single transmembrane domain, an extracellular domain, and a C-terminal glycosylphosphatidylinositol (GPI) membrane anchor [38]. The host restriction factor of human tetherin can block the release of enveloped viruses and inhibit viral replication. Yin et al showed that the N-terminal domain of equine tetherin was shorter than human tetherin. Equine tetherin was localized on the cell surface and strongly blocked equine infectious anemia virus (EIAV) release. Moreover, EIAV envelope protein could neutralize the antiviral activity of equine tetherin [39].
Type I interferon (IFN) and the subsequent induction of interferon-stimulated genes (ISGs) promotes resisting viral infection. Viperin, also known as RSAD2, is an endoplasmicreticulum (ER) associated multifunctional protein with broad antiviral activity. Tang et al. demonstrated that equine viperin could distort the ER, inhibit viral Gag production and/or release, Env and Receptor, leading to reduced EIAV replication [40].
Schlafen (SLFN) family proteins are divided into three groups (Groups I, II, and III) based on size and structure. All these proteins contain a specific SLFN box upstream of the AAA domain, which has ATP-binding activity. Because of the RNA/DNA helicase function of the AAA domain, SLFNs have RNA structure-modelling activity and participate in RNA metabolism [41, 42]. Codon usage-based SLFN inhibition is a novel antiviral mechanism in innate immune response. Human schlafen11 (hSLFN11) could bind to transfer (t) RNAs and inhibit the changes of the tRNA pool, which would accommodate the specific codon usage of viruses during retroviral replication [43]. Equine schlafen 11 can restrict EIAV production and replication by a codon usage-dependent mechanism [44].
Adenosine deaminases that act on RNA (ADARs) are RNA-editing enzymes that catalyze the hydrolytic C6 deamination of adenosine (A) to produce inosine (I) in double-stranded RNA substrates [45–47]. ADAR1 is one member of the ADAR family that promotes viral evolution [48, 49] and stimulates EIAV replication and infectivity. ADAR1 may even contribute to the EIAV adaptation from horses to donkeys [50, 51].
EIAV evolution and the current control strategies
EIAV often results in a rapid, dynamic, three-stage disease process: acute, chronic, and long-term asymptomatic [52]. The initial acute phase is usually observed from 3 to 4 weeks post infection and is characterized by high levels of viremia and severe clinical symptoms. Most infected horses progress to chronic EIA, associated with repeated cycles of disease. With the irregular intervals of EIA cycle and the development of host immunity, most infected horses then become long-term asymptomatic. This makes EIAV an ideal model for understanding viral evolution in persistence and pathogenesis, providing better strategies for controlling EIA.
Because viral envelope variants were produced and selected sequentially, the virus could escape the host immune responses, leading to the cyclic nature of chronic EIA [53, 54]. Surface envelope glycoprotein gp90is the predominant site of viral variation of EIAV. Leroux et al showed that gp90 variation was independent of the number of disease cycles, viremia during chronic disease, and levels of virus replication during long-term asymptomatic infection. EIAV evolution can be associated with selection in target tissues, ongoing low levels of virus replication and be lack of abundant levels of plasma viremia [55].
The long terminal repeat (LTR) enhancer region of EIAV is one of the most variable regions in the EIAV genome [56, 57]. Up to 45% of the nucleotide positions within the LTR enhancer varied between the isolates [58]. EIAV LTR sequence variation influences the virus tropism. LTR enhancer changes are associated with EIAV adaptation in tissue culture [57]. Maury et al also indicated that due to a consequence of cell-specific selective pressures, genetic variants of EIAV LTR influenced cell tropism. However, there was no changes in the LTR enhancer over the course of a 3-year infection, implying the evolution of EIAV LTR was stable in vivo [59].
After using the attenuated vaccine EIAVFDDV13, China has successfully controlled and most areas have eradicated EIAV. However, EIA is still prevalent worldwide, especially in France, Slovenia, Ireland, Belgium, Italy, Japan, and Brazil [60–65]. Some of China’s experiences and measures can be adopted to battle EIA. First, the summer mosquito eradication must be conducted carefully. Second, regular quarantine is necessary. Serological examination is the main method for finding and euthanizing EIA-positive horses in a timely manner. Thirdly, all the horses should be immunized with an effective vaccine. Chinese EIAV attenuated vaccine EIAVFDDV13 is a very effective vaccine to control EIA.
Perspective
Developing an effective vaccine against lentivirus infections is still needed in both human and veterinarian medicine. EIA is still a severe lentivirus disease in horses of many other countries. In China, because many effective strategies were applied including the EIAV live attenuated vaccine, EIAV was successfully controlled and most areas have eradicated EIAV. While this serves as a model for controlling EIAV in other countries and battling other viral diseases, the attenuation mechanisms in these passages were not well understood. Understanding the attenuation mechanisms in the EIAV vaccine will provide further insight to controlling EIAV and other viruses.
ACKNOWLEDGMENTS AND FUNDING
This study was funded by National Natural Science Foundation of China (No.31602118) and the Fundamental Research Funds for the Central Universities (2016QNA6017). Many data in this article were cited from ‘Equine infectious anemia and its controlling in China, a document issued by the Chinese Ministry of Agriculture in 1997, which is written in Chinese.
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
REFERENCES
1. Leroux C, Cadore JL, Montelaro RC. Equine Infectious Anemia Virus (EIAV): what has HIV’s country cousin got to tell us? Veterinary Research. 2004; 35:485–512. https://doi.org/10.1051/vetres:2004020.
2. Feng JL. Enquine infectious anemia and its controlling in China. [Article In Chinese]. The Chinese Ministry of Agriculture. 1997; 4–10.
3. Dai YK, Ning XD, Shen RX, Wang ZY. Enzyme-linked immunosorbent assay (ELISA) for detection of equine infectious anemia viral antibody. The international symposium on immunity to equine infectious anemia. 1983: 231–9.
4. Ebert D. Experimental evolution of parasites. Science. 1998; 282:1432–5.
5. Hilleman MR. Vaccines in historic evolution and perspective: a narrative of vaccine discoveries. J Hum Virol. 2000; 3:63–76.
6. Dong JP, Wang ZQ, Peng DC, Shen RX. Studies on the equine infectious anemia fetal donkey dermal cell-adapted attenuated vaccine and its efficacy. Chinese Journal of Animal and Poultry Infectious Diseases. 1989: 77–83.
7. Lin YZ, Shen RX, Zhu ZY, Deng XL, Cao XZ, Wang XF, Ma J, Jiang CG, Zhao LP, Lv XL, Shao YM, Zhou JH. An attenuated EIAV vaccine strain induces significantly different immune responses from its pathogenic parental strain although with similar in vivo replication pattern. Antiviral Research. 2011; 92:292–304. https://doi.org/10.1016/j.antiviral.2011.08.016.
8. Shen RX, Xu ZD, He YS, Zhang SX. Development of a EIAV donkey leucocyte attenuated vaccine. China Agriculture Sciences. 1979; 12:1–15.
9. Liu L, Wan Y, Wu L, Sun J, Li H, Li H, Ma L, Shao Y. Broader HIV-1 neutralizing antibody responses induced by envelope glycoprotein mutants based on the EIAV attenuated vaccine. Retrovirology. 2010; 7:71. https://doi.org/10.1186/1742-4690-7-71.
10. Payne SL, Fuller FJ. Virulence determinants of equine infectious anemia virus. Curr HIV Res. 2010; 8:66–72.
11. Payne SL, Pei XF, Jia B, Fagerness A, Fuller FJ. Influence of long terminal repeat and env on the virulence phenotype of equine infectious anemia virus. J Virol. 2004; 78:2478–85.
12. Li F, Craigo JK, Howe L, Steckbeck JD, Cook S, Issel C, Montelaro RC. A live attenuated equine infectious anemia virus proviral vaccine with a modified S2 gene provides protection from detectable infection by intravenous virulent virus challenge of experimentally inoculated horses. J Virol. 2003; 77:7244–53.
13. Wei L, Fan X, Lu X, Zhao L, Xiang W, Zhang X, Xue F, Shao Y, Shen R, Wang X. Genetic variation in the long terminal repeat associated with the transition of Chinese equine infectious anemia virus from virulence to avirulence. Virus Genes. 2009; 38:285–8. https://doi.org/10.1007/s11262-008-0317-y.
14. Wang XF, Yang B, Han XE, Lin YZ, Jiang CG, Lv XL, Zhao LP, Zhou JH, Wang FL. Genetic evolution of equine infectious anemia virus LTR region during attenuation in vitro. Chinese Journal of Preventive Veterinary Medicine. 2010: 915–9.
15. Tu YB, Zhou T, Yuan XF, Qin HJ, Xue F, Sun CQ, Wang L, Wu DL, Peng JM, Kong XG, Tong GZ. Long terminal repeats are not the sole determinants of virulence for equine infectious anemia virus. Arch Virol. 2007; 152:209–18.
16. Zhou T, Yuan XF, Hou SH, Tu YB, Peng JM, Wen JX, Qiu HJ, Wu DL, Chen HC, Wang XJ, Tong GZ. Long terminal repeat sequences from virulent and attenuated equine infectious anemia virus demonstrate distinct promoter activities. Virus Res. 2007; 128:58–64. https://doi.org/10.1016/j.virusres.2007.04.005.
17. Gao X, Jiang CG, Wang XF, Lin YZ, Ma J, Han XE, Zhao LP, Shen RX, Xiang WH, Zhou JH. Reverse mutation of the virulence-associated S2 gene does not cause an attenuated equine infectious anemia virus strain to revert to pathogenicity. Virology. 2013; 443:321–8.
18. Liang H, He X, Shen RX, Shen T, Tong X, Ma Y, Xiang WH, Zhang XY, Shao YM. Combined amino acid mutations occurring in the envelope closely correlate with pathogenicity of EIAV. Arch Virol. 2006; 151:1387–403. https://doi.org/10.1007/s00705-005-0718-3.
19. Shen T, Liang H, Tong X, Fan X, He X, Ma Y, Xiang W, Shen R, Zhang X, Shao Y. Amino acid mutations of the infectious clone from Chinese EIAV attenuated vaccine resulted in reversion of virulence. Vaccine. 2006; 24:738–49. https://doi.org/10.1016/j.vaccine.2005.08.084.
20. Jiang CG, Gao X, Ma J, Lin YZ, Wang XF, Zhao LP, Hua YP, Liu D, Zhou JH. C-terminal truncation of the transmembrane protein of an attenuated lentiviral vaccine alters its in vitro but not in vivo replication and weakens its potential pathogenicity. Virus research. 2011; 158:235–45.
21. Meng Q, Li S, Liu L, Xu J, Liu Y, Zhang Y, Zhang X, Shao Y. Truncation of cytoplasmic tail of EIAV Env increases the pathogenic necrosis. Virus Res. 2008; 133:201–10. https://doi.org/10.1016/j.virusres.2008.01.004.
22. Shen RX. Development and use of an equine infectious anemia donkey leukocyte attenuated vaccine. The international symposium on immunity to equine infectious anemia. 1983; 20.
23. Zhang XY, Li HM, Liang H, Shen T, Ma Y, Xiang WH, Shen RX, Shao YM. [Induction of EIAV-specific cellular immune response by attenuated EIAV vaccine]. [Article in Chinese]. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi. 2006; 22:115–8.
24. Lin Y, Deng X, Shen N, Zhao L, Meng Q, Max J, Wang J, Shao Y, Zhou J. [A flow cytometric assay for the expression of interferon gamma in T lymphocytes and its application in the study of EIAV-induced immune response]. [Article in Chinese]. Wei Sheng Wu Xue Bao. 2008; 48:800–5.
25. Zhang XY, Wang Y, Liang H, Wei L, Xiang WH, Shen RX, Shao YM. Correlation between the induction of Th1 cytokines by an attenuated equine infectious anemia virus vaccine and protection against disease progression. Journal of General Virology. 2007; 88:998–1004. https://doi.org/10.1099/vir.0.82416-0.
26. Lin YZ, Cao XZ, Li L, Li L, Jiang CG, Wang XF, Ma J, Zhou JH. The pathogenic and vaccine strains of equine infectious anemia virus differentially induce cytokine and chemokine expression and apoptosis in macrophages. Virus Research. 2011; 160:274–82. https://doi.org/10.1016/j.virusres.2011.06.028.
27. Ma J, Wang SS, Lin YZ, Liu HF, Wei HM, Du C, Wang XF, Zhou JH. An attenuated EIAV strain and its molecular clone strain differentially induce the expression of Toll-like receptors and type-I interferons in equine monocyte-derived macrophages. Veterinary Microbiology. 2013; 166:263–9. https://doi.org/10.1016/j.vetmic.2013.06.005.
28. Ma J, Wang SS, Lin YZ, Liu HF, Liu Q, Wei HM, Wang XF, Wang YH, Du C, Kong XG, Zhou JH, Wang X. Infection of equine monocyte-derived macrophages with an attenuated equine infectious anemia virus (EIAV) strain induces a strong resistance to the infection by a virulent EIAV strain. Vet Res. 2014; 45:82. https://doi.org/10.1186/s13567-014-0082-y.
29. Wang XF, Jiang CG, Guo W, Xiang W, Lv XL, Zhao LP, Wang FL, Kong XG, Zhang XY, Shao YM, Zhou JH. [Comparison of proviral genomes between the Chinese EIAV donkey leukocyte-attenuated vaccine and its parental virulent strain]. [Article in Chinese]. Bing Du Xue Bao. 2008; 24:443–50.
30. Ma J, Jiang CG, Lin YZ, Wang XF, Zhao LP, Xiang WH, Shao YM, Shen RX, Kong XG, Zhou JH. In vivo evolution of the gp90 gene and consistently low plasma viral load during transient immune suppression demonstrate the safety of an attenuated equine infectious anemia virus (EIAV) vaccine. Archives of Virology. 2009; 154:867–73. https://doi.org/10.1007/s00705-009-0378-9.
31. Wei HM, Wang XF, Wang SS, Du C, Liu HF, Liu Q, Zhou JH. [The application of single-genome amplification and sequencing in genomic analysis of an attenuated EIAV vaccine]. [Article in Chinese]. Bing Du Xue Bao. 2012; 28:431–8.
32. Ma J, Shi N, Jiang CG, Lin YZ, Wang XF, Wang S, Lv XL, Zhao LP, Shao YM, Kong XG, Zhou JH, Shen RX. A proviral derivative from a reference attenuated EIAV vaccine strain failed to elicit protective immunity. Virology. 2011; 410:96–106. https://doi.org/10.1016/j.virol.2010.10.032.
33. Zheng YH, Jeang KT, Tokunaga K. Host restriction factors in retroviral infection: promises in virus-host interaction. Retrovirology. 2012; 9:112. https://doi.org/10.1186/1742-4690-9-112.
34. Malim MH, Bieniasz PD. HIV Restriction Factors and Mechanisms of Evasion. Cold Spring Harb Perspect Med. 2012; 2:a006940. https://doi.org/10.1101/cshperspect.a006940.
35. Lilly F. Fv-2: identification and location of a second gene governing the spleen focus response to Friend leukemia virus in mice. J Natl Cancer Inst. 1970; 45:163–9.
36. Pincus T, Rowe WP, Lilly F. Major Genetic Locus Affecting Resistance to Infection with Murine Leukemia Viruses .2. Apparent Identity to a Major Locus Described for Resistance to Friend Murine Leukemia Virus. Journal of Experimental Medicine. 1971; 133:1234–41. https://doi.org/10.1084/jem.133.6.1234.
37. Yap MW, Colbeck E, Ellis SA, Stoye JP. Evolution of the retroviral restriction gene Fv1: inhibition of non-MLV retroviruses. PLoS Pathog. 2014; 10:e1003968. https://doi.org/10.1371/journal.ppat.1003968.
38. Kupzig S, Korolchuk V, Rollason R, Sugden A, Wilde A, Banting G. BST-2/HM1.24 is a raft-associated apical membrane protein with an unusual topology. Traffic. 2003; 4:694–709. https://doi.org/10.1034/j.1600-0854.2003.00129.x.
39. Yin X, Hu Z, Gu QY, Wu XL, Zheng YH, Wei P, Wang XJ. Equine Tetherin Blocks Retrovirus Release and Its Activity Is Antagonized by Equine Infectious Anemia Virus Envelope Protein. Journal of Virology. 2014; 88:1259–70. https://doi.org/10.1128/Jvi.03148-13.
40. Tang YD, Na L, Zhu CH, Shen N, Yang F, Fu XQ, Wang YH, Fu LH, Wang JY, Lin YZ, Wang XF, Wang XJ, Zhou JH, et al. Equine Viperin Restricts Equine Infectious Anemia Virus Replication by Inhibiting the Production and/or Release of Viral Gag, Env, and Receptor via Distortion of the Endoplasmic Reticulum. Journal of Virology. 2014; 88:12296–310. https://doi.org/10.1128/Jvi.01379-14.
41. Neumann B, Zhao L, Murphy K, Gonda TJ. Subcellular localization of the Schlafen protein family. Biochem Biophys Res Commun. 2008; 370:62–6. https://doi.org/10.1016/j.bbrc.2008.03.032.
42. Mavrommatis E, Fish EN, Platanias LC. The Schlafen Family of Proteins and Their Regulation by Interferons. Journal of Interferon and Cytokine Research. 2013; 33:206–10. https://doi.org/10.1089/jir.2012.0133.
43. Li M, Kao E, Gao X, Sandig H, Limmer K, Pavon-Eternod M, Jones TE, Landry S, Pan T, Weitzman MD, David M. Codon-usage-based inhibition of HIV protein synthesis by human schlafen 11. Nature. 2012; 491:125–8. https://doi.org/10.1038/nature11433.
44. Lin YZ, Sun LK, Zhu DT, Hu Z, Wang XF, Du C, Wang YH, Wang XJ, Zhou JH. Equine schlafen 11 restricts the production of equine infectious anemia virus via a codon usage-dependent mechanism. Virology. 2016; 495:112–21. https://doi.org/10.1016/j.viro1.2016.04.024.
45. Bass BL. RNA editing by adenosine deaminases that act on RNA. Annual Review of Biochemistry. 2002; 71:817–46. https://doi.org/10.1146/annurev.biochem.71.110601.135501.
46. Samuel CE. Adenosine deaminases acting on RNA (ADARs) are both antiviral and proviral. Virology. 2011; 411:180–93. https://doi.org/10.1016/j.virol.2010.12.004.
47. Nishikura K. Functions and Regulation of RNA Editing by ADAR Deaminases. Annual Review of Biochemistry. 2010; 79:321–49. https://doi.org/10.1146/annurev-biochem-060208-105251.
48. Suspene R, Petit V, Puyraimond-Zemmour D, Aynaud MM, Henry M, Guetard D, Rusniok C, Wain-Hobson S, Vartanian JP. Double-Stranded RNA Adenosine Deaminase ADAR-1-Induced Hypermutated Genomes among Inactivated Seasonal Influenza and Live Attenuated Measles Virus Vaccines. Journal of Virology. 2011; 85:2458–62. https://doi.org/10.1128/Jvi.02138-10.
49. Ko NL, Birlouez E, Wain-Hobson S, Mahieux R, Vartanian JP. Hyperediting of human T-cell leukemia virus type 2 and simian T-cell leukemia virus type 3 by the dsRNA adenosine deaminase ADAR-1. Journal of General Virology. 2012; 93:2646–51. https://doi.org/10.1099/vir.0.045146-0.
50. Tang YD, Na L, Fu LH, Yang F, Zhu CH, Tang L, Li Q, Wang JY, Li Z, Wang XF, Li CY, Wang XJ, Zhou JH. Double-stranded RNA-specific adenosine deaminase 1 (ADAR1) promotes EIAV replication and infectivity. Virology. 2015; 476:364–71. https://doi.org/10.1016/j.virol.2014.12.038.
51. Tang YD, Zhang X, Na L, Wang XF, Fu LH, Zhu CH, Wang XJ, Zhou JH. Double-stranded-RNA-specific adenosine deaminase 1 (ADAR1) is proposed to contribute to the adaptation of equine infectious anemia virus from horses to donkeys. Archives of Virology. 2016; 161:2667–72. https://doi.org/10.1007/s00705-016-2951-3.
52. Montelaro RC, Ball JM, Ruslow KE. (1993). The Retroviridae, pp. 257–360.
53. Leroux C, Issel CJ, Montelaro RC. Novel and dynamic evolution of equine infectious anemia virus genomic quasispecies associated with sequential disease cycles in an experimentally infected pony. Journal of Virology. 1997; 71:9627–39.
54. Payne SL, Salinovich O, Nauman SM, Issel CJ, Montelaro RC. Course and extent of variation of equine infectious anemia virus during parallel persistent infections. J Virol. 1987; 61:1266–70.
55. Leroux C, Craigo JK, Issel CJ, Montelaro RC. Equine infectious anemia virus genomic evolution in progressor and nonprogressor ponies. Journal of Virology. 2001; 75:4570–83. https://doi.org/10.1128/Jvi.75.10.4570-4583.2001.
56. Lichtenstein DL, Craigo JK, Leroux C, Rushlow KE, Cook RF, Cook SJ, Issel CJ, Montelaro RC. Effects of long terminal repeat sequence variation on equine infectious anemia virus replication in vitro and in vivo. Virology. 1999; 263:408–17. https://doi.org/10.1006/viro.1999.9921.
57. Payne SL, La Celle K, Pei XF, Qi XM, Shao H, Steagall WK, Perry S, Fuller F. Long terminal repeat sequences of equine infectious anaemia virus are a major determinant of cell tropism. Journal of General Virology. 1999; 80:755–9.
58. Maury W, Perryman S, Oaks JL, Seid BK, Crawford T, McGuire T, Carpenter S. Localized sequence heterogeneity in the long terminal repeats of in vivo isolates of equine infectious anemia virus. Journal of Virology. 1997; 71:4929–37.
59. Maury W, Thompson RJ, Jones Q, Bradley S, Denke T, Baccam P, Smazik M, Oaks JL. Evolution of the equine infectious anemia virus long terminal repeat during the alteration of cell tropism. Journal of Virology. 2005; 79:5653–64. https://doi.org/10.1128/Jvi.79.9.5653-5664.2005.
60. Gaudaire D, Lecouturier F, Poncon N, Morilland E, Laugier C, Zientara S, Hans A. Molecular characterization of equine infectious anaemia virus from a major outbreak in southeastern France. Transbound Emerg Dis. 2017. https://doi.org/10.1111/tbed.12657.
61. Tigre DM, Brandao CFL, de Paula FL, Chinalia FA, Campos GS, Sardi SI. Characterization of isolates of equine infectious anemia virus in Brazil. Archives of Virology. 2017; 162:873–7. https://doi.org/10.1007/s00705-016-3172-5.
62. Kuhar U, Malovrh T. High genetic diversity of equine infectious anaemia virus strains from Slovenia revealed upon phylogenetic analysis of the p15 gag gene region. Equine Veterinary Journal. 2016; 48:205–10. https://doi.org/10.1111/evj.12404.
63. Quinlivan M, Cook F, Kenna R, Callinan JJ, Cullinane A. Genetic characterization by composite sequence analysis of a new pathogenic field strain of equine infectious anemia virus from the 2006 outbreak in Ireland. Journal of General Virology. 2013; 94:612–22. https://doi.org/10.1099/vir.0.047191-0.
64. Caij AB, Tignon M. Epidemiology and Genetic Characterization of Equine Infectious Anaemia Virus Strains Isolated in Belgium in 2010. Transboundary and Emerging Diseases. 2014; 61:464–8. https://doi.org/10.1111/tbed.12031.
65. Dong JB, Zhu W, Cook FR, Goto Y, Horii Y, Haga T. Identification of a novel equine infectious anemia virus field strain isolated from feral horses in southern Japan. Journal of General Virology. 2013; 94:360–5. https://doi.org/10.1099/vir.0.047498-0.

