Introduction
Osteosarcoma, the most common primary malignant bone tumor, ranks as main cause of cancer-related death in children and adolescents. It frequently localizes to the distal femur and proximal tibia [1]. Decades ago, with surgery as the sole treatment, diagnosis of osteosarcoma often portended death. Patients had only a 15-20% chance of cure [2]. In the 1980s and 1990s, chemotherapy remarkably augmented survival rates; patients today have about 65-70% chance of 5-year relapse-free survival, if the disease is localized [3]. Yet other patients will relapse (with pulmonary metastasis) within five years and have undetectable metastatic disease at diagnosis. Pulmonary metastasis is the chief cause of death from osteosarcoma [4]. Patients with metastases have poor prognosis: long-term survival rate of 10-30% [5]. How to reduce metastasis in osteosarcoma is essential to osteosarcoma therapy. Thus, it is necessary to develop novel approaches for treating metastasis, meaning a need to clarify the molecular mechanisms underlying pathogenesis and progression.
Tumor metastasis is a multistep process. During metastasis, tumor cells can through secrete some proteins, growth factors and cytokines associated with the extracellular matrix (ECM) [6], some metastatic potential gene and protein can allow tumor cells to escape from the primary tumor, migrate, and invade surrounding tissues, enter the vasculature, circulate and reach secondary sites, extravasate, and finally establish metastatic foci [7, 8]. Connective tissue growth factor (CTGF/CCN2) is a member of the CCN family. CCN proteins modulate cell-ECM interactions and exhibit multi cellular functions such as regulation of cell division, apoptosis, adhesion, motility, and ion transport [9, 10]. CTGF is a secreted protein that acts as a multifunctional signaling modulator in various biologic or pathologic processes [11]. In cancer cell, many studies have report that CTGF enhanced tumor development and progression [12-14]. On the other hand, CTGF also promoted cancer cell migration and metastasis such as breast cancer [15], nasopharyngeal carcinoma [16], hepatocellular carcinoma [17], and melanoma [18], but negatively regulated in neuron-glioblastoma [19], and oral squamous cell carcinoma [20]. However, the role of CTGF in osteosarcoma metastasis has not yet been clarified.
One of the beginning step of metastasis is primary tumors invade locally through surrounding ECM or stromal cell layers. Matrix metalloproteinases (MMPs) are zinc-dependent endopeptidases and have been regarded as major critical molecules assisting tumor cells invasion [21]. Many studies linked MMPs with cancer invasion, migration or cancer cell-mediated tissue remodeling were the major positive correlations with MMP levels [8, 22]. MMPs have also been think one of promising therapeutic targets for osteosarcoma patients [23]. The function or expression of MMPs was regulated by many factors, including ECM molecules, growth factors, cytokines, and chemokines. Some of studies have demonstrated CCN proteins regulate MMPs protein expression [24, 25]. However, the role of CTGF in MMPs expression in human osteosarcoma is largely unknown.
MicroRNAs (miRNAs) are small noncoding RNAs that are 19 to 25 nucleotides in lengths, miRNAs regulate gene expression through binding to 3'untranslated regions (UTRs) of target mRNAs, either targeting the transcripts for degradation or blocking their translation [26]. There are more and more miRNAs have been involved in the regulation of cancer metastasis [27], and there have been considerable interest in the posttranscriptional regulation of MMPs via miRNAs [28]. Therefore, we hypothesized that miRNA may be involved in CTGF-mediated MMPs expression and tumor metastasis in human osteosarcoma. The results show that CTGF increases MMPs expression and subsequently promotes tumor metastasis in human osteosarcoma by down-regulating miR-519d through MEK and ERK pathway.
Results
CTGF enhances migration in human osteosarcoma
CTGF has been reported to promote cell migration and metastasis in variety of human cancer cells [29]. However, it is still not well-recognized whether CTGF stimulates tumor migration in osteosarcoma cells. First, we compared the protein expression of CTGF and migration ability in three osteosarcoma cell lines. Western blot revealed a higher level expression of CTGF in U-2 OS cells and lower level in MG-63 cells (Fig. 1A). U-2 OS cells were more migratory than MG-63 cells (Fig. 1B). Therefore, the migratory ability was positive corrected with CTGF expression in human osteosarcoma. To examine the link between CTGF expression and osteosarcoma migration, we established the CTGF-expressing plasmid (pcDNA3.1-CTGF). As shown in Fig. 1C and Supplementary Fig. S2B, the protein and mRNA expression were significant higher in CTGF-expressing cells (MG-63/CTGF and U-2 OS/CTGF) than in control cells (MG-63/vector and U-2 OS/vector). We also found the migration ability was dramatically increased in MG-63/CTGF cells (Fig. 1E). In contrast, shRNA-mediated knockdown of CTGF in U-2 OS/CTGF-shRNA cells and MG-63/CTGF-shRNA showed notably reduced CTGF expression, and the migration ability in U-2 OS cells (Fig. 1D&F; Supplementary Fig. S2A). On the other hand, incubation with recombinant human CTGF rescued CTGF-shRNA-reduced cell migration (Fig. 1F). These results indicate that CTGF promotes migration in human osteosarcoma cells.
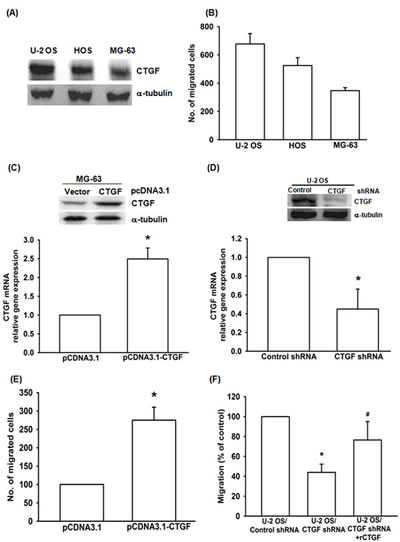
Figure 1: CTGF promotes migration in human osteosarcoma cells. (A&B) CTGF expression and migratory activity in indicated cells was examined by western blot and Transwell assay. (C&E) CTGF-overexpressing MG-63 cells were established, using pCDNA3.1 vector. Protein and mRNA expression of CTGF and migratory activity were examined by western blot, q-PCR and Transwell assay. (D) Protein and mRNA expression of CTGF in indicated cells were examined by western blot and q-PCR. (F) U-2 OS/CTGF shRNA cells were incubated with vehicle or recombine protein of CTGF (rCTGF; 50 ng/ml), cell migration examined by Transwell assay, results expressed as mean ± SEM. *, p < 0.05 as compared with controls, and #, p < 0.05 as compared with U-2 OS/CTGF shRNA group.
Repression of miR-519d by CTGF is critical in CTGF-mediated migration
MiRNA has been reported as an important regulator in cancer progression and metastasis [30]. In order to investigate miRNA differential expression in CTGF-overexpressed osteosarcoma cells, we used miRNome microRNA Profilers QuantiMir™ kit that contained 576 human miRNAs. Ten miRNAs were significantly up-regulated, and eight miRNAs were significantly down-regulated (Supplementary Table 1). Among the significantly regulated miRNAs, miR-519d was the most down-regulated in response to CTGF overexpression (Supplementary Table 1). We confirmed miR-519d expression using qRT-PCR, the miR-519d was down-regulated in MG-63/CTGF cells, and up-regulated in U-2 OS/CTGF-shRNA cells (Fig. 2A). In addition, directly applied CTGF in MG-63 cells and U-2 OS cells for 24 h reduced miR-519d expression in a concentration-dependent manner (Fig. 2B). To further verify the direct effect of miR-519d on cell motility, we transiently transfected the miR-519d mimic or inhibitor into osteosarcoma cells, and found miR-519d mimic but not inhibitor blocked the CTGF-induced cell migration (Fig. 2C). Taken together, CTGF increases osteosarcoma migration through down-regulating miR-519d.
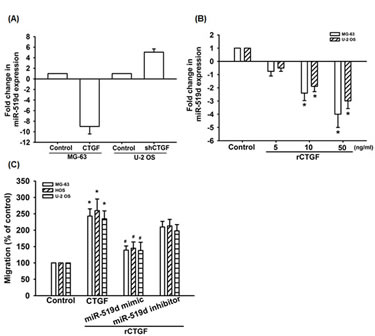
Figure 2: CTGF increases cell migration by suppressing miR-519d expression. (A) The miR-519d expression in indicated cells was examined by qRT-PCR. (B) MG-63 cells and U-2 OS cells were incubated with rCTGF (5-50 ng/ml) for 24 h, miR-519d expression examined by qRT-PCR. (C) Cells were transfected with miR-519d mimic or inhibitor for 24 h and cell migration examined by Transwell assay. Results are expressed as mean ± SEM. *, p < 0.05 as compared with controls, and #, p < 0.05 as compared with CTGF-treated control group.
MMP-2 and MMP-3 are the downstream target genes in CTGF-regulated miR-519d
To identify the mechanism of miR-519d-involved osteosarcoma migration, we searched for possible downstream genes using bioinformative screening analysis of miRNA target databanks: DIANA-mT, miRanda, miRDB, and Targetscan. From these four databanks overlapping between the predicted targets of miR-519d, the MMP-2 and MMP-3 were ranked as the most probable targets. We therefore hypothesized that both MMP-2 and MMP-3 were involved in CTGF-meidated migration. Transfection of cells with the MMP-2 or MMP-3 siRNA abolished CTGF-induced cell migration (Fig. 3A). Previous studies showed significant expression of MMP-1, -2, -3, -8, -9, and -13 in human osteosarcoma cells, and also indicating that these MMPs participated the progression and metastases in human osteosarcoma [31-34]. By using q-PCR and western blot, we found the mRNA and protein expression of MMP-2 and MMP-3 but not others were increased in MG-63/CTGF cells (Fig. 3B&C). In contrast, knockdown CTGF decreased MMP-2 and MMP-3 expression in U-2 OS/CTGF-shRNA cells (Fig. 3C&D). Furthermore, transfection with miR-519d mimic abolished CTGF-induced MMP-2 and MMP-3 expression (Fig. 3E). To examine whether miR-519d regulates the 3′UTR of MMP-2 and MMP-3, we constructed luciferase reporter vectors harboring the wild-type 3′UTR of the MMP-2 and MMP-3 mRNA (wt-MMP-2-3′UTR and wt-MMP-3-3′UTR) and vector containing mismatches in the predicted miR-519d binding site (mt-MMP-2-3′UTR and mt-MMP-3-3′UTR; Supplementary Fig. S1) and transfected these vectors into MG-63/CTGF and control cells. The results shown that CTGF increased luciferase activity in the wt-MMP-2-3′UTR and wt-MMP-3-3′UTR plasmid but not in the mt-MMP-2-3′UTR and mt-MMP-3-3′UTR (Fig. 3F). Taken together, these data demonstrated that miR-519d directly represses the MMP-2 and MMP-3 protein expression through binding to the 3′UTR of the human MMP-2 and MMP-3 gene.
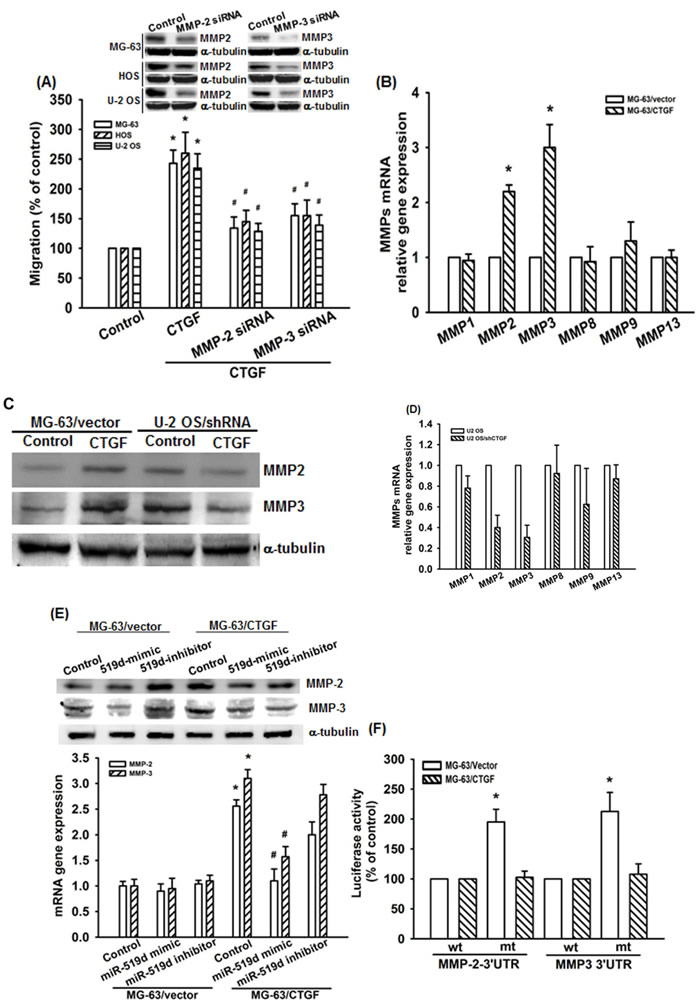
Figure 3: CTGF increases MMP-2 and MMP-3 expression and cell migration by down-regulating miR-519d. (A) Cells were transfected with MMP-2 or MMP-3 siRNA for 24 h. The MMP2 and MMP-3 expression and cell migration were examined by western blot and Transwell assay. (B-D) The mRNA and protein expression in indicated cells was examined by q-PCR and western blot. (E) Cells were transfected with miR-519d mimic or inhibitor for 24 h and MMPs expression was examined by western blot and q-PCR. (F) Cells were transfected with indicated luciferase plasmid for 24 h, and the relative luciferase activity was measured. Results are expressed as mean ± SEM. *, p < 0.05 as compared with control group, and #, p < 0.05 as compared with CTGF-treated control group.
MEK/ERK pathway is involved in CTGF-mediated migration and suppression of miRNA-519d
The MEK/ERK pathway has been reported to regulate MMPs expression and cancer metastasis [35-37]. We therefore examined whether the MEK/ERK is involved in CTGF-mediated cell migration and MMPs expression in osteosarcoma cells. Pretreatment of MG-63/CTGF cells with MEK inhibitors (PD98059 and U0126) or transfection with MEK1 and ERK2 mutant abolished CTGF-induced MMPs expression and cell migration (Fig. 4A-D). In addition, stimulation of cells with CTGF also promoted MEK and ERK phosphorylation time-dependently (Fig. 4E). Furthermore, MEK or ERK inhibitors and mutants reversed CTGF-inhibited miR-519d expression (Fig. 4F), indicating that CTGF increases MMPs and migration as well as suppresses miR-519d through MEK/ERK pathway.
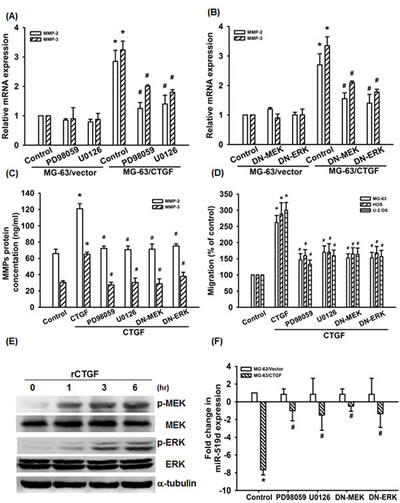
Figure 4: MEK/ERK pathway is involved in CTGF-induce migration and suppression of miRNA-519d. (A-D) Cells were pretreated with PD98059 (10 μM) and U0126 (10 μM) for 30 min or transfected with MEK1 and ERK2 mutant for 24 h, the MMPs expression and cell migration was examined by q-PCR, ELISA, and Transwell assay. (E) MG-63 cells were incubated with rCTGF (50 ng/ml) for indicated time intervals, MEK and ERK phosphorylation examined by western blot. (F) Cells were pretreated with PD98059 (10 μM) and U0126 (10 μM) for 30 min or transfected with MEK1 and ERK2 mutant for 24 h followed, the miR-519d expression was examined by qRT-PCR. Results expressed as mean ± SEM. *, p < 0.05 as compared with controls, and #, p < 0.05 as compared with CTGF-treated control gruop.
Knockdown of CTGF suppress lung metastasis in vivo
To determine whether knockdown of CTGF suppress lung metastasis in vivo, we established luciferase-expressing U-2 OS/Luc and U-2 OS/shCTGF-Luc cells. Cells were intravenously injected into SCID mice, and tumor metastasis was monitored by bioluminescence imaging. As shown in Fig. 5A, knockdown of CTGF significantly suppressed lung metastasis in the time course. Mice were sacrificed after 6 weeks injected, ex vivo imaging of the lungs derived from mice on day 42 showed a higher intensity and metastatic nodules in the U-2 OS/Luc versus U-2 OS/shCTGF-Luc group (Fig. 5B-D). Immunohistochemical staining also found expression of CTGF, MMP-2, and MMP-3 starkly decreasing in U-2 OS/shCTGF-Luc group (Fig. 5E). These indicates CTGF knockdown suppressing lung metastasis in vivo.
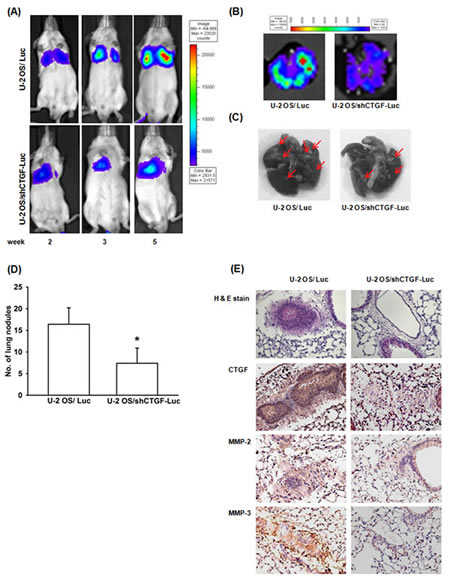
Figure 5: Knockdown of CTGF expression suppress lung metastasis in vivo. (A) U-2 OS/Luc or U-2 OS/shCTGF-Luc cells were injected into the lateral tail vein, and lung metastasis was monitored by bioluminescence imaging at indicated time intervals. (B-E) After 6 weeks mice were sacrificed, their lungs were excised and photographed, tumor colonies counted and stained with CTGF, MMP-2, and MMP-3. Results are expressed as mean ± SEM. *, p < 0.05 as compared with U-2 OS/Luc group.
Confirming significance of CTGF, MMPs, and miR-519d in clinical samples
Finally, we investigated clinical importance of CTGF, MMPs, and miR-519d in osteosarcoma patients. Expression of CTGF, MMP-2, and MMP-3 in osteosarcoma patients was significantly higher than in normal bone (Fig. 6A). Quantitative RT-PCR analysis of CTGF, MMP2, MMP-3, and miR-519d performed in osteosarcoma and normal samples, showed negative correlation between CTGF and miR-519d; positive correlation between CTGF and MMP-2, CTGF and MMP-3 (Fig. 6B-D). High CTGF expression was significantly associated with clinical stage and metastasis (Table 1); higher CTGF and lower miR-519d expression were linked with MMP-2 and MMP-3 expression, correlating with osteosarcoma development and metastasis.
Table 1: Correlation of CTGF expression with clinical status in osteosarcoma
Characteristic |
CTGF low (n=21) |
CTGF high (n=29) |
p-value |
Age (year) |
|||
< 24 |
7 |
12 |
0.5629 |
> 25 |
14 |
17 |
|
Gender |
|||
Female |
8 |
9 |
0.7635 |
Male |
13 |
20 |
|
Stage |
|||
IA |
4 |
2 |
0.0453* |
IIA |
9 |
7 |
|
IIB |
8 |
13 |
|
IVB |
0 |
7 |
|
Tumor status |
|||
T1 |
13 |
8 |
0.0152* |
T2-T3 |
8 |
21 |
|
Distant metastasis |
|||
M0 |
21 |
22 |
0.0152* |
M1 |
0 |
7 |
*,p <0.05
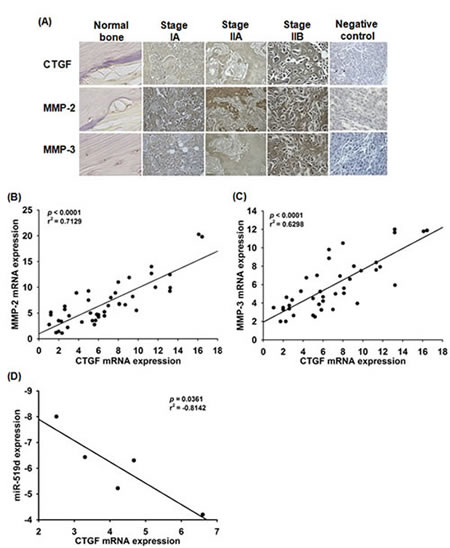
Figure 6: Clinical importance of CTGF, MMPs and miR-519d in osteosarcoma. (A) IHC stain of CTGF, MMP-2, and MMP-3 in normal versus osteosarcoma tissue. Correlation between CTGF/MMP-2 (B), CTGF/MMP-3 (C) and CTGF/mi-519d (D) in human osteosarcoma.
Discussion
Cancer is one of the leading causes of death worldwide [38]. Osteosarcoma is the most common primary bone tumor and with high morbidity that mainly occurs in children and adolescents. From a clinical perspective, primary osteosarcoma management can be successfully in most patients. But, the development of metastasis to the lung or other organs represents the most common cause of death [3]. Clear understanding of metastasis biology in osteosarcoma is necessary. This study characterized effect of CTGF on production of MMP-2 and MMP-3 in human osteosarcoma, which is responsible for subsequent increased migration and metastasis. Results indicated CTGF inducing upregulation of MMP-2 and MMP-3 expression in human osteosarcoma cells by down-regulating miR-519d through MEK and ERK pathway and contributing the tumor metastasis.
CTGF belongs to a family of secreted proteins that have a number of cellular functions, as well as pathologic processes [11]. In tumor metastasis, the double-sided sword effect of CTGF has been widely reported: CTGF promotes cancer growth or metastasis in chondrosarcoma, breast, and brain caners [10 , 39]; in contrast, it inhibits tumor metastasis in oral and lung cancers [40, 41]. However, the role of CTGF in osteosarcoma progression and metastasis remain unknown. Our in vitro data shown that overexpression of CTGF in osteosarcoma cells enhanced migration ability. On the other hand, knockdown of CTGF expression via CTGF-shRNA strongly decreased the cell migration. By using the IHC staining, we also found that the expression of CTGF in osteosarcoma tissues were correlates with tumor stage and metastasis. Finally, we used the animal model to confirm that down-regulation of CTGF decreased lung metastasis in vivo. To the best of our knowledge, this is the first study to demonstrate that CTGF promotes osteosarcoma metastasis in vitro and in vivo.
Newly identified small noncoding miRNAs, belong to a novel class that control gene expression by binding to complementary sequences in 3′UTRs of target mRNAs [42, 43]. Deregulated miRNA expression has been cited in human cancer and may affect multiple steps during metastasis [44]. It has been revealed that miR-21 [45], 125b [46], 143 [47], 20a [48], and 93 [49] regulates metastatic ability in osteosarcoma. Our study screened 576 miRNAs from miRNome microRNA Profilers QuantiMir™ kit and found miR-519d most down-regulated in response to CTGF overexpression; miR-519d reportedly suppresses tumor formation in human hepatocellular carcinoma, making it an attractive candidate biomarker for prediction of anticancer treatment response [50, 51]. In this study, transfection of cells with miR-519d mimic reduced CTGF-induced cell migration, meaning miR-519d can be an anti-metastatic gene.
Previous study demonstrated that MMPs functioning in many stages of cancer metastasis, including enhanced escape of survival in circulation, and extravasation at the secondary site [52]. CTGF has also been reported to regulate MMP-2 and MMP-3 expression [53, 54]. Our study found mRNA and protein expression of MMP-2 and MMP-3 increasing in MG-63/CTGF cells. Knockdown of MMP-2 or MMP-3 expression via transfection with MMP-2 or MMP-3 siRNA abolished CTGF-induced osteosarcoma cell migration activity, meaning both MMP-2 and MMP-3 involved in CTGF-mediated cell migration. By using miRNA target prediction (DIANA-mT, miRanda, miRDB, and Targetscan), we proved MMP-2 and MMP-3 most probable miR-519d targets; transfection of miR-519d-mimic strongly inhibited CTGF-induced MMP-2 and MMP-3 expression. We indicated miR-519d directly repressing MMP-2 and MMP-3 protein expression through binding to 3′UTR of human MMP-2 and MMP-3 genes, thus negatively regulating MMP-2 and MMP-3-mediated metastasis.
Metastasis is caused by cells wandering from a primary tumor, then colonizing other sites in the body. Most common features are described as invasion, intravasation or extravasation from the circulatory system, colonization and angiogenesis at distant sites. The latter is generally connected with poor prognosis of osteosarcoma patients. Development of anti-metastatic therapy could conceivably be useful in such cases. Our results point to high CTGF expression as significantly associated with clinical stage and metastasis. CTGF increases MMP-2/-3 expression and promotes metastasis by down-regulating miR-519d through MEK and ERK pathways; CTGF inhibition presents a new therapeutic target.
Materials and Methods
Materials
Anti-rabbit and anti-mouse IgG-conjugated horseradish peroxidase, rabbit polyclonal antibody specific for α-tubulin, and mouse monoclonal antibodies specific for CTGF, MMP-2, and MMP-3 were purchased from Biotechnology (Santa Cruz, CA). ON-TARGETplus siRNA of MMP2, MMP3, and control were purchased from Dharmacon Research (Lafayette, CO). PD98059, U0126 and human MMP-2 and MMP-3 enzyme-linked immunosorbent assay (ELISA) kits were purchased from Calbiochem (San Diego, CA). Recombinant human CTGF was purchased from PeproTech (Rocky Hill, NJ). MiRNome microRNA Profilers QuantiMir™ kit was from System Biosciences (SBI) (Mountain View, CA). MiR-519d mimic and inhibitor were purchased from Invitrogen (Carlsbad, CA). MEK1 dominant-negative mutant was a gift from Dr. W. M. Fu (National Taiwan University at Taipei). ERK2 dominant-negative mutant was a gift from Dr. M. Cobb (Southwestern Medical Center, Dallas, TX), all other chemicals purchased from Sigma-Aldrich (St. Louis, MO).
Cell culture
The human osteosarcoma cell lines (MG-63, U-2 OS, and HOS) were purchased from the American Type Cell Culture Collection (Manassas, VA, USA). MG-63 and HOS cells were maintained in Dulbecco`s Modified Eagle Medium (DMEM) and U-2 OS cells were maintained in McCoy’s 5A medium, which was supplemented with 20 mM HEPES and 10% heat-inactivated FCS, 2 mM glutamine, penicillin (100 U/ml), streptomycin (100 μg/ml) and 10% FBS at 37 °C with 5% CO2.
Patients and specimen preparation
The study protocol was approved by the Institutional Review Board of China Medical University Hospital, and all subjects gave informed written consent before enrollment. The specimens of tumor tissue or normal bone were obtained from patients who were had been diagnosed with osteosarcoma and had undergone surgical resection at China Medical University Hospital.
Migration assay
The migration assay was performed using Transwell (Costar, NY; pore size, 8 mm). Approximately 1×104 cells in 100 ml of serum-free medium were placed in the upper chamber, and 300 ml of the same medium was placed in the lower chamber. The plates were incubated for 24 h at 37°C in 5% CO2, cells were then fixed in 3.7% formaldehyde for 15 min and stained with 0.05% crystal violet in PBS for 15 min. Cells on the upper side of the filters were removed with cotton-tipped swabs, and the filters were washed with PBS. Cells on the underside of the filters were examined and counted under a microscope. Each clone was plated in triplicate for each experiment, and each experiment was repeated at least three times.
Immunohistochemical (IHC) staining
Human osteosarcoma tissue sections were deparaffinized with xylene and rehydrated through addition of ethanol. Endogenous peroxidase activity was blocked with 3% hydrogen peroxide in methanol for 10 min. Heat-induced antigen retrieval was carried out for all sections in 0.01 M sodium citrate buffer, pH 6 at 95°C for 25 min. Human CTGF, MMP-2, and MMP-3 antibodies were applied at a dilution of 1:200 and incubated at 4°C overnight. The antibody-binding signal was detected using the NovoLink Polymer Detection System (Leica Microsystems) and visualized with the diaminobenzidine reaction. The sections were counterstained with hematoxylin. The immunohistochemistry results were scored by taking into account the percentage of positive detection and intensity of the staining.
ELISA assay
Human osteosarcoma cells were cultured in 24-well plates. After reaching confluence, cells were changed to serum-free medium. Cells were then treated with CTGF alone for 24 h, or pretreated with pharmacological inhibitors or transfected with specific siRNA followed by stimulation with CTGF for 24 h. After treatment, the medium was removed and stored at -80°C. Then, MMP-2 and MMP-3 in the medium was determined using MMP-2 and MMP-3 ELISA kit according to the manufacturer’s protocol.
Luciferase activity assay
3’UTR regions of human MMP-2 and MMP-3 genes were amplified by PCR. The 3’UTR were cloned in pGL2-Control vector (Promega, Madison, WI, USA) downstream of the reporter gene. The predicted MMP-2 and MMP-3 binding sites, identified by the miRDB (http://mirdb.org/miRDB), were amplified by PCR and cloned in pGL2-Control vector upstream of the reporter gene (Supplementary Fig. S1). Mutant plasmids were generated using a QuickChange Site-Directed Mutagenesis kit (Stratagene, Cedar Creek, TX, USA).
Western blot analysis
Protein concentration was determined using the Thermo Scientific Pierce BCA Protein Assay Kit (Thermo Fisher Scientific Inc., USA). Proteins were resolved on SDS-PAGE and transferred to immobilon polyvinyl difluoride (PVDF) membranes. The blots were blocked with 4% BSA for 1 h at room temperature and incubated with the following primary antibodies for 1 h at room temperature. After 3 washes in tris-buffered saline with 0.05% Tween 20 (TBS-Tween), the blots were subsequently incubated with a donkey anti-rabbit or anti-mouse peroxidase-conjugated secondary antibody for 1 h at room temperature. The blots were visualized by enhanced chemiluminescence using Kodak X-OMAT LS film (Eastman Kodak, Rochester, NY). Quantitative data were obtained using a computing densitometer and ImageQuant software (Molecular Dynamics, Sunnyvale, CA).
Quantitative real-time PCR
Total RNA was extracted from osteosarcoma cells using a TRIzol kit (MDBio Inc., Taipei, Taiwan). The reverse transcription reaction was performed using 2 μg of total RNA that was reverse transcribed into cDNA using an oligo(dT) primer. Quantitative real-time polymerase chain reaction (q-PCR) analysis was carried out using TaqMan® one-step PCR Master Mix (Applied Biosystems, Foster City, CA, USA). Total complementary DNA (100ng/25μL reaction) was mixed with sequence-specific primers and TaqMan® probes according to the manufacturer’s instructions. Sequences for all target gene primers and probes were purchased commercially (β-actin was used as the internal control) (Applied Biosystems). Q-PCR assays were carried out in triplicate using a StepOnePlus sequence detection system. The cycling conditions were 10 min of polymerase activation at 95 °C, followed by 40 cycles at 95 °C for 15 s and 60 °C for 60 s.
For miRNAs detection, reverse transcription was performed using Mir-X™ miRNA First-Strand Synthesis and SYBR® qRT-PCR (Clontech Laboratories, Inc., CA, USA). U6 snRNA levels were used for normalization. The specific forward primer of miR-519d was as follows: 5’-CAAAGTGCCTCCCTTTAGAGTG-3’. Forward and reverse primers for U6 were 5’-CTCGCTTCGGCAGCACATATACT A-3’and 5’-ACGAATTTGCGTGTCATCCTTGCG-3’. The threshold was set above the non-template control background and within the linear phase of target gene amplification to calculate the cycle number at which the transcript was detected (denoted as CT).
In vivo metastasis model
Cells (U-2 OS/Luc or U-2 OS/shCTGF-Luc) were washed and resuspended in PBS. Subsequently, a unit suspension containing 5×106 cells in 100 µl PBS was injected into the lateral tail vein of 5-week-old severe combined immune deficient (SCID) mice. Lung metastasis was monitored using an in vivo imaging system (Xenogen IVIS imaging system). After 6 weeks, the mice were sacrificed by overdose with anesthetic. The lungs were removed, photographed, and counted tumor colonies. The lungs were then fixed in 10% formalin, embedded in paraffin and subsequently processed for IHC staining with CTGF, MMP-2 and MMP-3.
Statistics
Data are expressed as the mean ± standard error. The differences between groups were analyzed using the Student’s t-test or the χ2 test of variance. The difference was considered significant if the p value was less than 0.05.
Acknowledgments
This work was supported by grants from the National Science Council of Taiwan (NSC100-2320-B-039-028-MY3; NSC101-2320-B-715-002-MY3) and China Medical University Hospital (DMR-97-072). We thank Dr. W.M. Fu for providing the MEK1 mutant and Dr. M. Cobb for providing the ERK2 mutant.
Conflict of Interest Statement
The authors have no financial or personal relationships that could inappropriately influence this research.
References
1. Link MP, Goorin AM, Miser AW, Green AA, Pratt CB, Belasco JB, Pritchard J, Malpas JS, Baker AR, Kirkpatrick JA and et al. The effect of adjuvant chemotherapy on relapse-free survival in patients with osteosarcoma of the extremity. The New England journal of medicine. 1986; 314(25):1600-1606.
2. Hayden JB and Hoang BH. Osteosarcoma: basic science and clinical implications. Orthop Clin North Am. 2006; 37(1):1-7.
3. Osborne TS and Khanna C. A review of the association between osteosarcoma metastasis and protein translation. Journal of comparative pathology. 2012; 146(2-3):132-142.
4. Hattinger CM, Pasello M, Ferrari S, Picci P and Serra M. Emerging drugs for high-grade osteosarcoma. Expert Opin Emerg Drugs. 2010; 15(4):615-634.
5. Meyers PA. Muramyl tripeptide (mifamurtide) for the treatment of osteosarcoma. Expert Rev Anticancer Ther. 2009; 9(8):1035-1049.
6. Chen JC, Fong YC and Tang CH. Novel strategies for the treatment of chondrosarcomas: targeting integrins. BioMed research international. 2013; 2013:396839.
7. Geho DH, Bandle RW, Clair T and Liotta LA. Physiological mechanisms of tumor-cell invasion and migration. Physiology (Bethesda). 2005; 20:194-200.
8. Tang C-H. Molecular mechanisms of chondrosarcoma metastasis. BioMedicine. 2012; 2(3):92-98.
9. Dhar A and Ray A. The CCN family proteins in carcinogenesis. Experimental oncology. 2010; 32(1):2-9.
10. Chen PC, Cheng HC, Yang SF, Lin CW and Tang CH. The CCN family proteins: modulators of bone development and novel targets in bone-associated tumors. BioMed research international. 2014; 2014:437096.
11. Luft FC. CCN2, the connective tissue growth factor. Journal of molecular medicine. 2008; 86(1):1-3.
12. Chu CY, Chang CC, Prakash E and Kuo ML. Connective tissue growth factor (CTGF) and cancer progression. Journal of biomedical science. 2008; 15(6):675-685.
13. Lu H, Kojima K, Battula VL, Korchin B, Shi Y, Chen Y, Spong S, Thomas DA, Kantarjian H, Lock RB, Andreeff M and Konopleva M. Targeting connective tissue growth factor (CTGF) in acute lymphoblastic leukemia preclinical models: anti-CTGF monoclonal antibody attenuates leukemia growth. Annals of hematology. 2013.
14. Chang CC, Hsu WH, Wang CC, Chou CH, Kuo MY, Lin BR, Chen ST, Tai SK, Kuo ML and Yang MH. Connective tissue growth factor activates pluripotency genes and mesenchymal-epithelial transition in head and neck cancer cells. Cancer research. 2013; 73(13):4147-4157.
15. Chen PS, Wang MY, Wu SN, Su JL, Hong CC, Chuang SE, Chen MW, Hua KT, Wu YL, Cha ST, Babu MS, Chen CN, Lee PH, Chang KJ and Kuo ML. CTGF enhances the motility of breast cancer cells via an integrin-alphavbeta3-ERK1/2-dependent S100A4-upregulated pathway. Journal of cell science. 2007; 120(Pt 12):2053-2065.
16. Zhen Y, Ye Y, Yu X, Mai C, Zhou Y, Chen Y, Yang H, Lyu X, Song Y, Wu Q, Fu Q, Zhao M, Hua S, Wang H, Liu Z, Zhang Y, et al. Reduced CTGF expression promotes cell growth, migration, and invasion in nasopharyngeal carcinoma. PloS one. 2013; 8(6):e64976.
17. Xiu M, Liu YH, Brigstock DR, He FH, Zhang RJ and Gao RP. Connective tissue growth factor is overexpressed in human hepatocellular carcinoma and promotes cell invasion and growth. World journal of gastroenterology : WJG. 2012; 18(47):7070-7078.
18. Finger EC, Cheng CF, Williams TR, Rankin EB, Bedogni B, Tachiki L, Spong S, Giaccia AJ and Powell MB. CTGF is a therapeutic target for metastatic melanoma. Oncogene. 2013.
19. Romao LF, Mendes FA, Feitosa NM, Faria JC, Coelho-Aguiar JM, de Souza JM, Moura Neto V and Abreu JG. Connective tissue growth factor (CTGF/CCN2) is negatively regulated during neuron-glioblastoma interaction. PloS one. 2013; 8(1):e55605.
20. Yang MH, Lin BR, Chang CH, Chen ST, Lin SK, Kuo MY, Jeng YM, Kuo ML and Chang CC. Connective tissue growth factor modulates oral squamous cell carcinoma invasion by activating a miR-504/FOXP1 signalling. Oncogene. 2012; 31(19):2401-2411.
21. Fingleton B. Matrix metalloproteinases: roles in cancer and metastasis. Front Biosci. 2006; 11:479-491.
22. Hadler-Olsen E, Winberg JO and Uhlin-Hansen L. Matrix metalloproteinases in cancer: their value as diagnostic and prognostic markers and therapeutic targets. Tumour biology : the journal of the International Society for Oncodevelopmental Biology and Medicine. 2013; 34(4):2041-2051.
23. Yang J and Zhang W. New molecular insights into osteosarcoma targeted therapy. Current opinion in oncology. 2013; 25(4):398-406.
24. Maeta N, Osaki M, Shomori K, Inaba A, Kidani K, Ikeguchi M and Ito H. CYR61 downregulation correlates with tumor progression by promoting MMP-7 expression in human gastric carcinoma. Oncology. 2007; 73(1-2):118-126.
25. Hou CH, Chiang YC, Fong YC and Tang CH. WISP-1 increases MMP-2 expression and cell motility in human chondrosarcoma cells. Biochemical pharmacology. 2011; 81(11):1286-1295.
26. Iorio MV and Croce CM. MicroRNAs in cancer: small molecules with a huge impact. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2009; 27(34):5848-5856.
27. Nicoloso MS, Spizzo R, Shimizu M, Rossi S and Calin GA. MicroRNAs--the micro steering wheel of tumour metastases. Nature reviews Cancer. 2009; 9(4):293-302.
28. Li L and Li H. Role of microRNA-mediated MMP regulation in the treatment and diagnosis of malignant tumors. Cancer biology & therapy. 2013; 14(9).
29. Jacobson A and Cunningham JL. Connective tissue growth factor in tumor pathogenesis. Fibrogenesis & tissue repair. 2012; 5 Suppl 1:S8.
30. Garzon R, Marcucci G and Croce CM. Targeting microRNAs in cancer: rationale, strategies and challenges. Nature reviews Drug discovery. 2010; 9(10):775-789.
31. Ryu B, Li Y, Qian ZJ, Kim MM and Kim SK. Differentiation of human osteosarcoma cells by isolated phlorotannins is subtly linked to COX-2, iNOS, MMPs, and MAPK signaling: implication for chronic articular disease. Chemico-biological interactions. 2009; 179(2-3):192-201.
32. Minamitani K. Establishment of an osteoblastic osteosarcoma cell line and effects of cell culture conditions on secretion of matrix metalloproteinases from the cultured osteosarcoma cells. The Kurume medical journal. 2000; 47(2):115-124.
33. Korpi JT, Hagstrom J, Lehtonen N, Parkkinen J, Sorsa T, Salo T and Laitinen M. Expression of matrix metalloproteinases-2, -8, -13, -26, and tissue inhibitors of metalloproteinase-1 in human osteosarcoma. Surgical oncology. 2011; 20(1):e18-22.
34. Broadhead ML, Clark JC, Myers DE, Dass CR and Choong PF. The molecular pathogenesis of osteosarcoma: a review. Sarcoma. 2011; 2011:959248.
35. Tsubaki M, Satou T, Itoh T, Imano M, Ogaki M, Yanae M and Nishida S. Reduction of metastasis, cell invasion, and adhesion in mouse osteosarcoma by YM529/ONO-5920-induced blockade of the Ras/MEK/ERK and Ras/PI3K/Akt pathway. Toxicology and applied pharmacology. 2012; 259(3):402-410.
36. Ai X, Wu Y, Zhang W, Zhang Z, Jin G, Zhao J, Yu J, Lin Y, Zhang W, Liang H, Datta PK, Zhang M, Zhang B and Chen X. Targeting ERK pathway reduces liver metastasis of Smad4-inactivated colorectal cancer. Cancer biology & therapy. 2013; 14(11).
37. Kim S, Choi JH, Lim HI, Lee SK, Kim WW, Kim JS, Kim JH, Choe JH, Yang JH, Nam SJ and Lee JE. Silibinin prevents TPA-induced MMP-9 expression and VEGF secretion by inactivation of the Raf/MEK/ERK pathway in MCF-7 human breast cancer cells. Phytomedicine : international journal of phytotherapy and phytopharmacology. 2009; 16(6-7):573-580.
38. Yin M-c. Humans and cancer: An ongoing fight. BioMedicine. 2012; 2(3):83.
39. Tan TW, Lai CH, Huang CY, Yang WH, Chen HT, Hsu HC, Fong YC and Tang CH. CTGF enhances migration and MMP-13 up-regulation via alphavbeta3 integrin, FAK, ERK, and NF-kappaB-dependent pathway in human chondrosarcoma cells. J Cell Biochem. 2009; 107(2):345-356.
40. Chuang JY, Yang WY, Lai CH, Lin CD, Tsai MH and Tang CH. CTGF inhibits cell motility and COX-2 expression in oral cancer cells. Int Immunopharmacol. 2011; 11(8):948-954.
41. Chang CC, Shih JY, Jeng YM, Su JL, Lin BZ, Chen ST, Chau YP, Yang PC and Kuo ML. Connective tissue growth factor and its role in lung adenocarcinoma invasion and metastasis. J Natl Cancer Inst. 2004; 96(5):364-375.
42. Esquela-Kerscher A and Slack FJ. Oncomirs - microRNAs with a role in cancer. Nature reviews Cancer. 2006; 6(4):259-269.
43. Zhang Quan JK-T. Long noncoding RNAs and viral infections. BioMedicine. 2013; 3(1):34-42.
44. Chen PS, Su JL and Hung MC. Dysregulation of microRNAs in cancer. J Biomed Sci. 2012; 19:90.
45. Ziyan W, Shuhua Y, Xiufang W and Xiaoyun L. MicroRNA-21 is involved in osteosarcoma cell invasion and migration. Medical oncology. 2011; 28(4):1469-1474.
46. Liu LH, Li H, Li JP, Zhong H, Zhang HC, Chen J and Xiao T. miR-125b suppresses the proliferation and migration of osteosarcoma cells through down-regulation of STAT3. Biochemical and biophysical research communications. 2011; 416(1-2):31-38.
47. Osaki M, Takeshita F, Sugimoto Y, Kosaka N, Yamamoto Y, Yoshioka Y, Kobayashi E, Yamada T, Kawai A, Inoue T, Ito H, Oshimura M and Ochiya T. MicroRNA-143 regulates human osteosarcoma metastasis by regulating matrix metalloprotease-13 expression. Molecular therapy : the journal of the American Society of Gene Therapy. 2011; 19(6):1123-1130.
48. Huang G, Nishimoto K, Zhou Z, Hughes D and Kleinerman ES. miR-20a encoded by the miR-17-92 cluster increases the metastatic potential of osteosarcoma cells by regulating Fas expression. Cancer research. 2012; 72(4):908-916.
49. Montanini L, Lasagna L, Barili V, Jonstrup SP, Murgia A, Pazzaglia L, Conti A, Novello C, Kjems J, Perris R and Benassi MS. MicroRNA cloning and sequencing in osteosarcoma cell lines: differential role of miR-93. Cellular oncology. 2012; 35(1):29-41.
50. Hou YY, Cao WW, Li L, Li SP, Liu T, Wan HY, Liu M, Li X and Tang H. MicroRNA-519d targets MKi67 and suppresses cell growth in the hepatocellular carcinoma cell line QGY-7703. Cancer letters. 2011; 307(2):182-190.
51. Fornari F, Milazzo M, Chieco P, Negrini M, Marasco E, Capranico G, Mantovani V, Marinello J, Sabbioni S, Callegari E, Cescon M, Ravaioli M, Croce CM, Bolondi L and Gramantieri L. In hepatocellular carcinoma miR-519d is up-regulated by p53 and DNA hypomethylation and targets CDKN1A/p21, PTEN, AKT3 and TIMP2. The Journal of pathology. 2012; 227(3):275-285.
52. Deryugina EI and Quigley JP. Matrix metalloproteinases and tumor metastasis. Cancer metastasis reviews. 2006; 25(1):9-34.
53. Chintala H, Liu H, Parmar R, Kamalska M, Kim YJ, Lovett D, Grant MB and Chaqour B. Connective tissue growth factor regulates retinal neovascularization through p53 protein-dependent transactivation of the matrix metalloproteinase (MMP)-2 gene. The Journal of biological chemistry. 2012; 287(48):40570-40585.
54. Blaney Davidson EN, Vitters EL, Mooren FM, Oliver N, Berg WB and van der Kraan PM. Connective tissue growth factor/CCN2 overexpression in mouse synovial lining results in transient fibrosis and cartilage damage. Arthritis and rheumatism. 2006; 54(5):1653-1661.

