INTRODUCTION
Pseudomonas putida is a non-fermentative Gram-negative bacillus belonging to the fluorescent group of the genus Pseudomonas [1]. It is an opportunistic human pathogen, responsible for nosocomial infections in immunocompromised patients and in those with catheter or indwelling devices. P. putida infection usually presents as bacteremia, urinary tract infection, or pneumonia [1, 2].
Worldwide, the increasing emergence of carbapenem-resistant bacteria, including Pseudomonas strains, has threatened the efficacy of carbapenems in the treatment of refractory infections [3]. Metallo-β-lactamases (MBLs) are a group of β-lactamases that can hydrolyze all β-lactams, including carbapenems, except aztreonam [3]. Genes encoding these MBLs are generally associated with integrons and transposons, and often coexist with genes conferring resistance to other classes of antibiotics [4]. This association results from co-selection under the pressure of multiple antibiotics, and leads to multidrug resistance (MDR) in Pseudomonas strains [4].
Production of MBLs, such as IMPs [5-7], VIMs [5, 8, 9], NDM-1 [10], and DIM-2 [11], has been identified to account for carbapenem resistance in P. putida. Previous studies of these MBL-producing P. putida strains have been confined to PCR detection and/or sequencing of bla genes and their genetic environments, except for that of the blaDIM-2-carrying plasmid p12969-DIM from a clinical MDR P. putida isolate, in which the plasmid was fully sequenced. The backbone of p12969-DIM was genetically closely related to blaIMP-9-carrying Pseudomonas aeruginosa plasmid pOZ176; however, the two plasmids contained dramatically different accessory regions, particularly those containing resistance genes [11, 12]. blaDIM-2 and blaIMP-9 in p12969-DIM and pOZ176 were embedded in integrons In1224 and In244, respectively, which were further associated with two different Tn21 subgroup transposons belonging to the Tn3 family [11, 12].
The current study presents the second fully-sequenced MBL-encoding plasmid, designated pSY153-MDR, from clinical P. putida. pSY153-MDR was a 468.2-kb mega plasmid, and carried blaIMP-45 (encoding carbapenem resistance) and armA (encoding aminoglycoside resistance), as well as additional markers involved in resistance to β-lactams, quinolones, macrolides, tetracyclines, amphenicols, quaternary ammonium compounds, sulphonamides, trimethoprim, and rifampicin. Twelve accessory modules, which especially included two novel MDR regions containing all of the above resistance genes, were inserted at different sites of the pSY153-MDR backbone.
RESULTS AND DISCUSSION
General features of P. putida SY153
PCR results demonstrated that strain SY153 harbored blaIMP, blaOXA, and armA, but none of the rest bla and 16S rRNA methylase genes screened for in this analysis, with the first two genes being confirmed as blaIMP-45 and blaOXA-1 by genomic sequencing. IMP-45 was closely related to IMP-9, with a single Ser214Gly variation, while the Carba NP test showed that SY153 had class B carbapenemase activity (data not shown). Strain SY153 was highly resistant to penicillins, cephalosporins, carbapenems, aminoglycosides, fluoroquinolones, tetracycline, trimethoprim/sulfamethoxazole, and chloramphenicol, but remained intermediately resistant to aztreonam. In addition, greatly elevated MIC values were observed for macrolides, rifampin, and nitrofurantoin, for which resistance breakpoints have not been established for P. putida (Supplementary Table 1). Repeated attempts at conjugation failed to transfer the blaIMP-45 marker from SY153 to E. coli J53 and P. aeruginosa PAO1.
Overview of pSY153-MDR
Plasmid pSY153-MDR had a closed circular DNA sequence, 468,170 kb in length, with a mean G+C content of 56.6%. There were 558 predicted open reading frames (ORFs), 68.6% of which encoded hypothetical proteins (Supplementary Figure 1). The molecular structure of pSY153-MDR could be divided into separate accessory modules that were defined as the acquired DNA regions associated with mobile elements, and the remaining backbone regions.
The pSY153-MDR backbone, 385 kb in length, was closely related (>98% nucleotide identity over >86% of the backbone regions) to five plasmids deposited in GenBank, namely pBM413 (a 423-kb blaIMP-45-harboring plasmid from P. aeruginosa; accession number CP016215), pOZ176 [12], pRBL16 (a 370-kb plasmid without antibiotic resistance genes from organic pollutant degradant Pseudomonas citronelloli; accession number CP015879), pTTS12 (a 584-kb solvent-resistance plasmid from P. putida; accession number CP015879) [13], and p12969-DIM [11] (last accessed December 1, 2016). repA (replication initiation protein of unknown incompatibility group) and parB2-parAB (partition) constituted the sole replication/stability system in pSY153-MDR. Together with pil (pilus assemble) and che (chemotaxis), this replication/stability system was found in all five plasmids. Two resistance loci, namely nfxB–mexCD–oprJ (resistance-nodulation-division (RND)-type multidrug efflux pump) and ter (tellurium resistance), were identified in the pSY153-MDR backbone. nfxB–mexCD–oprJ was also found in pBM413 and pDIM-12969, while the ter locus was also identified in plasmids pBM413, pRBL16, and pTTS12.
The accessory regions of pSY153-MDR were composed of four separate copies of each of the insertion sequence (IS) elements IS1491 and ISPpu29, single copies of IS1411 and ISPpu30, and two novel MDR regions, designated MDR-1 and MDR-2 (55.7 kb and 9.2 kb in length, respectively). These 12 accessory modules were inserted at different sites of the pSY153-MDR backbone, and all 10 individual IS elements were flanked by direct repeats (DRs; target site duplication signals of transposition). In contrast, only two accessory modules (an ISPpu23 element and a MDR region) were identified in p12969-DIM. Compared with p12969-DIM, pSY153-MDR had undergone much more massive insertions of foreign genetic contents, and showed a higher degree of genomic plasticity.
The MDR-1 region of pSY153-MDR was organized sequentially, as follows: the Tn1403–Tn1548 region, Tn6309, In48, and ISCfr1 (Figure 1). Either the MDR-1 region of pSY153-MDR or the MDR region of p12969-DIM was inserted immediately downstream of nfxB–mexCD–oprJ, indicating a “hotspot” for insertion of external genetic material in these two closely related plasmids.
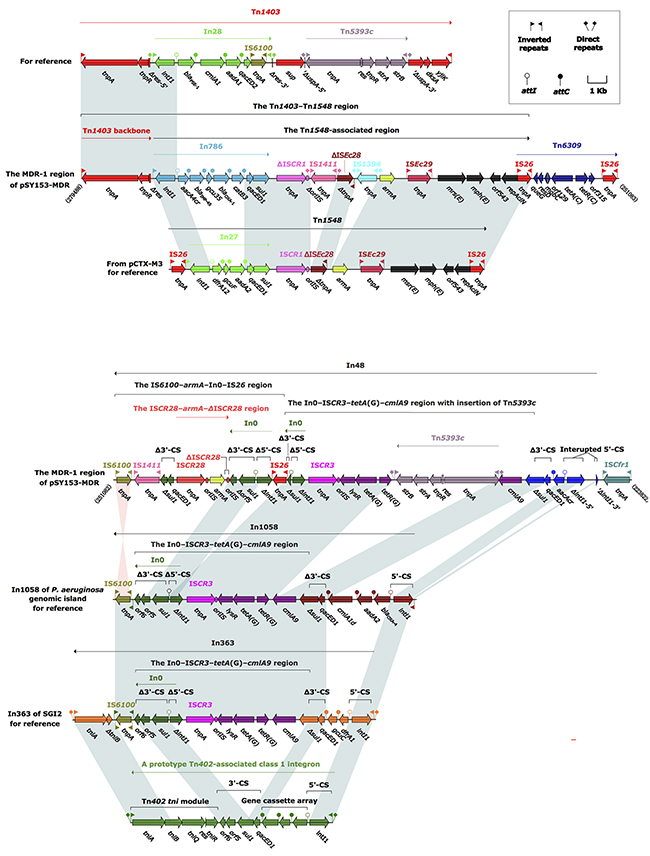
Figure 1: The MDR-1 region from pSY153-MDR, and comparison with related regions. Genes are denoted by arrows. Genes, mobile elements and other features are colored based on function classification. Shading denotes regions of homology (>95% nucleotide identity). Numbers in brackets indicate the nucleotide positions within plasmid pSY153-MDR. The accession numbers of Tn1403, Tn1548, In1058, and In363 for references are AF313472, AF550415, KJ463833, and AY963803, respectively.
The Tn1403–Tn1548 region and Tn6309 from the MDR-1 region
Tn1403 is a Tn3-family transposon initially identified in a clinical P. aeruginosa isolate in the United States in the 1970s [14]. It has a core backbone consisting of IRL (inverted repeat left), tnpA (transposase), tnpR (resolvase), res (resolution site), sup (sulfate permease), uspA (universal stress protein), dksA (RNA polymerase-binding transcription factor), yjiK (hypothetical protein), and IRR (inverted repeat right), with insertions of In28 and Tn5393c into res and dksA, respectively. Tn1403 and its close derivatives Tn6060, Tn6061, Tn6217, Tn6249, and Tn6286 in Pseudomonas often contain different insertions of foreign material (such as integrons and transposons), serving as important vehicles for transmission of resistance genes [11, 12, 15-17].
Tn1548, originally characterized in pCTX-M3 from Citrobacter freundii, is an IS26-flanked composite transposon without flanking DRs, and displays a modular structure consisting of IS26–In27–ISCR1–∆ISEc28–armA–ISEc29–msr(E)–mph(E) –orf543–repAciN–IS26 [18]. Tn1548 and its variants mainly differ from each other by insertion of distinct integrons or integron-like elements between IS26 and ISCR1, and are responsible for dissemination of the aminoglycoside resistance gene armA, the macrolide resistance operon msr(E)–mph(E), and other integron-borne resistance genes commonly found among Enterobacteriaceae and Acinetobacter species [19, 20].
The Tn1403–Tn1548 region of pSY153-MDR was composed of Tn1403 backbone remnant IRL–tnpAR–∆res, and a 19.0-kb Tn1548-associated region. Compared with Tn1548, the Tn1548-associated region of pSY153-MDR had undergone several evolutionary events: i) deletion of the IS26 at the 5’-flank; ii) replacement of In27 with In786; and iii) insertion of IS1411 and IS1394 upstream of ΔISEc28 and armA, respectively. In786 consisted of an inverted repeat at the integrase end (IRi), a 5’-conserved segment (5’-CS:intl1 (integrase)–attI1 (IntI1-recognizing recombination site)), a gene cassette array (GCA; organized as aacA4cr (aminoglycoside and quinolone resistance), blaIMP-45, gcu35 (unknown function), blaOXA-1 (β-lactam resistance), catB3 (chloramphenicol resistance), and a 3’-CS (qacED1 (quaternary ammonium compound resistance)–sul1 (sulfonamide resistance)), but lacked an inverted repeat at the tni end (IRt). The Tn1403–Tn1548 region was likely generated from homologous recombination between the Tn1403-like and Tn1548-like transposons, with In786 as the common component.
Tn6309 is an IS26-flanked composite transposon carrying class C tetracycline resistance module tetA(C) (class C tetracycline efflux protein)-tetR(C) (transcriptional repressor of tetA), and has been identified in genomic island Sm1-MDRGI from Stenotrophomonas maltophilia [21], and in sequenced plasmids pP10164-3 from Leclercia adecarboxylata P10164 [22], pB3 plasmids from Pseudomonas sp. GFP1 [23], pNDM-116-14 (accession number LN831184) from Vibrio cholerae 116-14, and pKZ3 from an uncultured bacterium [24]. Tn6309 from pP10164-3 and pB3 is bordered by 9-bp and 5-bp DRs, respectively, indicating that its mobilization into these two plasmids occurred via IS26-mediated replicative transposition. No DRs were associated with Tn6309 in pSY153-MDR and, moreover, Tn6309 and the Tn1548-associated region overlapped by one of their terminal IS26 elements, suggesting that the connection of Tn6309 and the Tn1548-associated region was promoted by IS26-mediated homologous recombination, rather than Tn6309 transposition.
In48 from the MDR-1 region
Class 1 integrons are frequently associated with the core transposition module tniABQ–res–tniR (designated tni) of Tn402, and display a prototype hybrid structure consisting of IRi, 5’-CS, GCA, 3’-CS, tni, and IRt [25].
In363 was initially characterized in the SGI2 resistance island from Salmonella enteric serovar Emerk [26], and is organized as follows: IRi, 5’-CS, GCA (dfrA1 (dihydrofolate resistance)–gcuC (unknown function)), Δ3’-CS (qacED1–Δsul1), the In0 (an empty class 1 integron)–ISCR3–tetA(G) (class G tetracycline resistance)–cmlA9 (chloramphenicol resistance) region, and IRi–IS6100–IRt–Δtni (ΔtniB–tniA)–IRt. In363 is bracketed by 5-bp DRs, indicating that its mobilization into SGI2 was a transposition event. In363 differed from the prototype class 1 integron by: i) insertion of the In0–ISCR3–tetA(G)–cmlA9 region within the 3’-CS of In363, likely resulting from homologous recombination based on the common 3’-CS region shared by In0 and ancestral In363; and ii) truncation of tni by the insertion of IS6100, generating the IRi–IS6100–IRt–Δtni–IRt structure.
In1058, a close variant of In363, was present in the blaVIM-2-carrying genomic island from a MDR P. aeruginosa isolate [27]. It carried a GCA consisting of blaOXA-4 (β-lactam resistance)–aadA2 (aminoglycoside resistance)–cmlA1d (chloramphenicol resistance). Other than their differing GCA contents, In1058 had lost the 3’-terminal IRt–Δtni–IRt region relative to In363.
Compared with In1058, In48 had at least five major modular differences: i) In48 carried a single-gene (aacA4cr) cassette; ii) IRi was deleted as a result of the connection of ISPpu31 to intI1; iii) intI1 was interrupted by a cryptic 617-bp sequence; iv) Tn5393c was inserted between tetA(G) and cmlA9 of the In0–ISCR3–tetA(G)–cmlA9 region; and v) the IS6100–armA–In0–IS26 region was inserted within sul1 of the In0–ISCR3–tetA(G)–cmlA9 region.
The IS6100–armA–In0–IS26 region was bordered by IS26 and IS6100, both of which belonged to the IS6 family, and possess almost identical 14-bp terminal inverted repeats. As such, this region might utilize a mechanism of replicative transposition for mobility similar to that used by the IS26-flanked composite transposons [28]. In addition, homologous recombination based on the common In0 sequence shared by the IS6100–armA–In0–IS26 region and the In0–ISCR3–tetA(G)–cmlA9 region might also promote the mobilization of the former region into In48.
The four copies of In0 found in In363, In1058, and In48 had an identical 37-bp attI1 site, which was the 5’-terminal segment of the intact attI1, and lacked the symmetrical structure. This would render these In0s incapable of capturing gene cassette(s) via site-specific recombination.
There were also two copies of armA in pSY153-MDR. One copy was located in the Tn1403–Tn1548 region that was a Tn1548 variant. The armA genes from Tn1548 and its variants are generally bordered by ∆ISEc28 and ISEc29 [19, 20]. The second copy of armA was flanked by two copies of ISCR28, displaying an ISCR28–armA–∆ISCR28 structure that represented a novel armA vehicle.
The MDR-2 region of pSY153-MDR
The MDR-2 region was composed of a class 1 integron, In1237, and two copies of IS1411, one intact and the other truncated (Figure 2). In1237 carried a GCA consisting of qnrVC1 (quinolones resistance)–gcu165 (unknown function)–arr2 (rifampicin resistance)–dfrA22e (dihydrofolate resistance). Notably, dfrA22e was a derivative of the reference dfrA22 gene (accession number HM173356), containing variations Pro3Arg, Leu5Ser, Lys29Thr, Asn62Ser, Ser73Gly, Arg114His, Asp137Asn, and Glu138Ala. This dfrA22e cassette was also found in In1218 from Aeromonas sobria from ornamental fish (accession number KT315928). The expression of the In1237 cassette array was driven by a sole PcW promoter. Compared with the prototype class 1 integron, In1237 had undergone two major changes: i) loss of IRi and truncation of intI1 through the connection of In1237 to ΔIS1411; and ii) replacement of tni by IS6100.
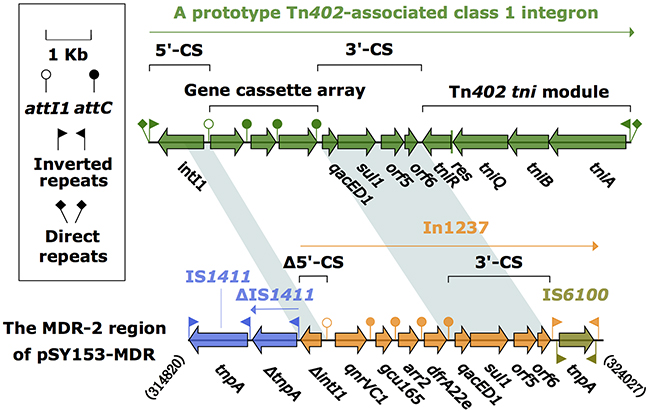
Figure 2: The MDR-2 region from pSY153-MDR, and comparison with related region. Genes are denoted by arrows. Genes, mobile elements and other features are colored based on function classification. Shading denotes regions of homology (>95% nucleotide identity). Numbers in brackets indicate the nucleotide positions within pSY153-MDR.
MATERIALS AND METHODS
Bacterial isolation and identification
The use of human specimens and all related experimental protocols was approved by the Committee on Human Research of the indicated institutions, and was carried out in accordance with the approved guidelines. Informed consent was obtained from the patient where indicated. Research involving biohazardous materials and all related procedures were approved by the Biosafety Committee of the National Institute for Communicable Disease Control and Prevention, Beijing, China. P. putida SY153 was isolated in 2012 from the urine specimen of a 68-year-old male patient with acute onset of cerebral infarction in a tertiary hospital in Sanya City, China. Bacterial species identification was performed by 16S rDNA gene amplification and sequencing [33]. The major plasmid-borne carbapenemase, extended-spectrum ß-lactamase, and 16S rRNA methylase genes were screened by polymerase chain reaction (PCR) assays [34, 35]. All PCR amplicons were sequenced on an ABI 3730 Sequencer (Applied Biosciences, Foster City, CA, USA), using the same primers as for PCR, according to the manufacturer’s instructions.
Plasmid conjugal transfer
Plasmid conjugal transfer experiments were carried out using sodium azide-resistant Escherichia coli J53 and colistin-resistant P. aeruginosa PAO1 (colistin MIC= 32 μg/mL) as the recipients, and SY153 as the donor. Aliquots (3 mL) of overnight culture of each donor and recipient strain were mixed, harvested, and resuspended in 80 μL of Brain Heart Infusion (BHI) broth (BD Biosciences). The suspensions were spotted on 1-cm2 hydrophilic nylon membrane filters with a 0.45-μm pore size (Millipore), which were then placed on BHI agar (BD Biosciences) plates and incubated at 25°C, 30°C, or 37°C for 12–18h. Bacteria were washed from the filter membranes and spotted onto Muller-Hinton (MH) agar plates containing 100 μg/mL sodium azide or 10 μg/mL colistin and 100 μg/mL ceftazidime for selection of blaIMP-45-positive E. coli or P. aeruginosa transconjugants.
Detection of carbapenemase activity
Activity of class A/B/D carbapenemases in bacterial cell extracts was determined using a modified CarbaNP test [34]. Briefly, 2 mL of bacterial cultures with an optical density at 600 nm of 1.0–1.4 were harvested, washed, and resuspended in 500 μL of 20 mM Tris-HCl (pH 7.8), lysed by sonication, and pelleted by centrifugation. Aliquots (50 μL) of the supernatants were individually mixed with 50 μL of solutions I–V, followed by incubation at 37°C for a maximum of 2 h. The substrates in solutions I-V consisted of 0.054% phenol red, 0.1mM ZnSO4 (pH 7.8), with or without 0.6 mg/μL imipenem, 0.8 mg/μL tazobactam and/or 3 mM EDTA (pH 7.8).
Antimicrobial susceptibility test
The antimicrobial susceptibility of the bacterial strains was determined by Etest (BioMérieux, Hazelwood, MO, USA), and interpreted as per Clinical and Laboratory Standards Institute guidelines [36].
Sequencing and annotation
Genomic DNA was isolated from SY153 using a Wizard Genomic DNA Purification Kit (Promega, Madison, WI, USA). The genome was sequenced using a Single Molecule Real Time technique on a PacBio platform (Tianjin Biochip Corporation, Tianjin, China). A total of 87,287 polymerase reads, with a mean read length of 11,530 bp, were generated, resulting in a total of 1,006,456,227 bases with a 104-fold average coverage. The DNA contigs were assembled using HGAP 2.0 [37]. Open reading frames and pseudogenes were predicted using RAST 2.0 [38], combined with BLASTP/BLASTN [39] searches against the UniProtKB/Swiss-Prot [40] and RefSeq [41] databases. Annotation of resistance genes, mobile elements, and other features was carried out using CARD [42], ResFinder [43], ISfinder [44], INTEGRALL [45]. Multiple and pairwise sequence comparisons were performed using MUSCLE 3.8.31 [46] and BLASTN, respectively. Gene organization diagrams were drawn in Inkscape (https://inkscape.org).
Nucleotide sequence accession number
The complete sequence of pSY153-MDR was submitted to GenBank under accession number KY883660.
CONCLUSIONblaIMP-45 genes have previously been documented in both chromosomally- and plasmid-located In786 sequences from P. aeruginosa [29-31] and S. maltophilia isolates [29]. To date, only one blaIMP-45-carrying plasmid, namely pBM413, from P. aeruginosa has been fully sequenced. Data presented in the current study showed that blaIMP-45 has spread to P. putida, and this is the first report of determination of a fully-sequenced plasmid, carrying In786-borne blaIMP-45 gene, from P. putida.
armA genes have been reported in three P. aeruginosa isolates [31, 32], and in two of these isolates, armA is embedded in the Tn1548-associated regions and coexists with blaIMP-45 on a single plasmid [31]. The current study is the first report of armA in P. putida.
Coexistence of a large number of antibiotic resistance genes accounts for the extensive drug resistance of strain SY153, which is likely to be a reservoir of antimicrobial resistance genes. The presence of these determinants also aids in the survival of strain SY153 under different antimicrobial selection pressures. All of the resistance genes contained on plasmid pSY153-MDR are clustered in the two MDR regions, which have very complex mosaic structures. These regions can be dissected into various integrons, transposons, and transposon-like elements. Further study on the evolution and diversification of pSY153-MDR and related plasmids, including those without antibiotic resistance genes, is needed.
Abbreviations
MDR: multidrug-resistant; MBLs: Metallo-β-lactamases; ORFs: open reading frames; DRs: direct repeats; IRL: inverted repeat left; IRR: inverted repeat right; IRi: inverted repeat at the integrase end; CS: conserved segment; GCA: gene cassette array; IRt: inverted repeat at the tni end; IS: insertion sequence; MIC: minimum inhibitory concentration; BHI: Brain Heart Infusion.
Author contributions
J.L., DS.Z. conceived the study and designed experimental procedures. M.Y., H.C., X.Z, X.C., and XF. Z performed the experiments. M.Y., J. F., Z. Z., DF. Z., and DS.Z. analyzed the data. H.C., X.Z, J.L. JX.L., and JG.X. contributed reagents and materials. M.Y., DS.Z., and J.L. wrote this manuscript.
CONFLICTS OF INTEREST
All authors declare that there are no conflicts of interest.
FUNDINGS
This work was supported by the National Natural Science Foundation of China (81501783) and the National Basic Research Program of China (2015CB554200).
REFERENCES
1. Carpenter RJ, Hartzell JD, Forsberg JA, Babel BS, Ganesan A. Pseudomonas putida war wound infection in a US Marine: a case report and review of the literature. J Infect. 2008; 56:234-240.
2. Yoshino Y, Kitazawa T, Kamimura M, Tatsuno K, Ota Y, Yotsuyanagi H. Pseudomonas putida bacteremia in adult patients: five case reports and a review of the literature. J Infect Chemother. 2011; 17:278-282.
3. Cornaglia G, Giamarellou H, Rossolini GM. Metallo-beta-lactamases: a last frontier for beta-lactams? Lancet Infect Dis. 2011; 11:381-393.
4. El Salabi A, Walsh TR, Chouchani C. Extended spectrum beta-lactamases, carbapenemases and mobile genetic elements responsible for antibiotics resistance in Gram-negative bacteria. Crit Rev Microbiol. 2013; 39:113-122.
5. Koh TH, Wang GC, Sng LH. IMP-1 and a novel metallo-beta-lactamase, VIM-6, in fluorescent pseudomonads isolated in Singapore. Antimicrob Agents Chemother. 2004; 48:2334-2336.
6. Gilarranz R, Juan C, Castillo-Vera J, Chamizo FJ, Artiles F, Alamo I, Oliver A. First detection in Europe of the metallo-beta-lactamase IMP-15 in clinical strains of Pseudomonas putida and Pseudomonas aeruginosa. Clin Microbiol Infect. 2013; 19:E424-427.
7. Docquier JD, Riccio ML, Mugnaioli C, Luzzaro F, Endimiani A, Toniolo A, Amicosante G, Rossolini GM. IMP-12, a new plasmid-encoded metallo-beta-lactamase from a Pseudomonas putida clinical isolate. Antimicrob Agents Chemother. 2003; 47:1522-1528.
8. Lombardi G, Luzzaro F, Docquier JD, Riccio ML, Perilli M, Coli A, Amicosante G, Rossolini GM, Toniolo A. Nosocomial infections caused by multidrug-resistant isolates of pseudomonas putida producing VIM-1 metallo-beta-lactamase. J Clin Microbiol. 2002; 40:4051-4055.
9. Bogaerts P, Huang TD, Rodriguez-Villalobos H, Bauraing C, Deplano A, Struelens MJ, Glupczynski Y. Nosocomial infections caused by multidrug-resistant Pseudomonas putida isolates producing VIM-2 and VIM-4 metallo-beta-lactamases. J Antimicrob Chemother. 2008; 61:749-751.
10. Bhattacharya D, Dey S, Kadam S, Kalal S, Jali S, Koley H, Sinha R, Nag D, Kholkute SD, Roy S. Isolation of NDM-1-producing multidrug-resistant Pseudomonas putida from a paediatric case of acute gastroenteritis, India. New Microbes New Infect. 2015; 5:5-9.
11. Sun F, Zhou D, Wang Q, Feng J, Feng W, Luo W, Liu Y, Qiu X, Yin Z, Xia P. Genetic characterization of a novel blaDIM-2-carrying megaplasmid p12969-DIM from clinical Pseudomonas putida. J Antimicrob Chemother. 2016; 71:909-912.
12. Xiong J, Alexander DC, Ma JH, Deraspe M, Low DE, Jamieson FB, Roy PH. Complete sequence of pOZ176, a 500-kilobase IncP-2 plasmid encoding IMP-9-mediated carbapenem resistance, from outbreak isolate Pseudomonas aeruginosa 96. Antimicrob Agents Chemother. 2013; 57:3775-3782.
13. Kuepper J, Ruijssenaars HJ, Blank LM, de Winde JH, Wierckx N. Complete genome sequence of solvent-tolerant Pseudomonas putida S12 including megaplasmid pTTS12. J Biotechnol. 2015; 200:17-18.
14. Stokes HW, Elbourne LD, Hall RM. Tn1403, a multiple-antibiotic resistance transposon made up of three distinct transposons. Antimicrob Agents Chemother. 2007; 51:1827-1829.
15. Roy Chowdhury P, Merlino J, Labbate M, Cheong EY, Gottlieb T, Stokes HW. Tn6060, a transposon from a genomic island in a Pseudomonas aeruginosa clinical isolate that includes two class 1 integrons. Antimicrob Agents Chemother. 2009; 53:5294-5296.
16. Di Pilato V, Pollini S, Rossolini GM. Tn6249, a new Tn6162 transposon derivative carrying a double-integron platform and involved with acquisition of the blaVIM-1 metallo-beta-lactamase gene in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2015; 59:1583-1587.
17. Coyne S, Courvalin P, Galimand M. Acquisition of multidrug resistance transposon Tn6061 and IS6100-mediated large chromosomal inversions in Pseudomonas aeruginosa clinical isolates. Microbiology. 2010; 156:1448-1458.
18. Galimand M, Sabtcheva S, Courvalin P, Lambert T. Worldwide disseminated armA aminoglycoside resistance methylase gene is borne by composite transposon Tn1548. Antimicrob Agents Chemother. 2005; 49:2949-2953.
19. Du XD, Li DX, Hu GZ, Wang Y, Shang YH, Wu CM, Liu HB, Li XS. Tn1548-associated armA is co-located with qnrB2, aac(6’)-Ib-cr and blaCTX-M-3 on an IncFII plasmid in a Salmonella enterica subsp. enterica serovar Paratyphi B strain isolated from chickens in China. J Antimicrob Chemother. 2012; 67:246-248.
20. Blackwell GA, Holt KE, Bentley SD, Hsu LY, Hall RM. Variants of AbGRI3 carrying the armA gene in extensively antibiotic-resistant Acinetobacter baumannii from Singapore. J Antimicrob Chemother. 2017; 72:1031-1039.
21. He T, Shen J, Schwarz S, Wu C, Wang Y. Characterization of a genomic island in Stenotrophomonas maltophilia that carries a novel floR gene variant. J Antimicrob Chemother. 2015; 70:1031-1036.
22. Sun F, Zhou D, Sun Q, Luo W, Tong Y, Zhang D, Wang Q, Feng W, Chen W, Fan Y, Xia P. Genetic characterization of two fully sequenced multi-drug resistant plasmids pP10164-2 and pP10164-3 from Leclercia adecarboxylata. Sci Rep. 2016; 6:33982.
23. Heuer H, Szczepanowski R, Schneiker S, Puhler A, Top EM, Schluter A. The complete sequences of plasmids pB2 and pB3 provide evidence for a recent ancestor of the IncP-1beta group without any accessory genes. Microbiology. 2004; 150:3591-3599.
24. Flach CF, Johnning A, Nilsson I, Smalla K, Kristiansson E, Larsson DG. Isolation of novel IncA/C and IncN fluoroquinolone resistance plasmids from an antibiotic-polluted lake. J Antimicrob Chemother. 2015; 70:2709-2717.
25. Chen Z, Fang H, Wang L, Sun F, Wang Y, Yin Z, Yang H, Yang W, Wang J, Xia P, Zhou D, Liu C. IMP-1 encoded by a novel Tn402-like class 1 integron in clinical Achromobacter xylosoxidans, China. Sci Rep. 2014; 4:7212.
26. Levings RS, Djordjevic SP, Hall RM. SGI2, a relative of Salmonella genomic island SGI1 with an independent origin. Antimicrob Agents Chemother. 2008; 52:2529-2537.
27. Perez F, Hujer AM, Marshall SH, Ray AJ, Rather PN, Suwantarat N, Dumford D 3rd, O’Shea P, Domitrovic TN, Salata RA, Chavda KD, Chen L, Kreiswirth BN, et al. Extensively drug-resistant pseudomonas aeruginosa isolates containing blaVIM-2 and elements of Salmonella genomic island 2: a new genetic resistance determinant in Northeast Ohio. Antimicrob Agents Chemother. 2014; 58:5929-5935.
28. Harmer CJ, Hall RM. IS26-mediated formation of transposons carrying antibiotic resistance genes. mSphere. 2016; 1:e00038-16.
29. Wu K, Wang F, Sun J, Wang Q, Chen Q, Yu S, Rui Y. Class 1 integron gene cassettes in multidrug-resistant Gram-negative bacteria in southern China. Int J Antimicrob Agents. 2012; 40:264-267.
30. Wang Y, Wang X, Schwarz S, Zhang R, Lei L, Liu X, Lin D, Shen J. IMP-45-producing multidrug-resistant Pseudomonas aeruginosa of canine origin. J Antimicrob Chemother. 2014; 69:2579-2581.
31. Janvier F, Otto MP, Jove T, Mille A, Contargyris C, Meaudre E, Brisou P, Plesiat P, Jeannot K. A case of multiple contamination with methylase ArmA-producing pathogens. J Antimicrob Chemother. 2017; 72:618-620.
32. Gurung M, Moon DC, Tamang MD, Kim J, Lee YC, Seol SY, Cho DT, Lee JC. Emergence of 16S rRNA methylase gene armA and cocarriage of bla(IMP-1) in Pseudomonas aeruginosa isolates from South Korea. Diagn Microbiol Infect Dis. 2010; 68:468-470.
33. Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA, Olsen GJ. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol. 2008; 74:2461-2470.
34. Chen Z, Li H, Feng J, Li Y, Chen X, Guo X, Chen W, Wang L, Lin L, Yang H, Yang W, Wang J, Zhou D, et al. NDM-1 encoded by a pNDM-BJ01-like plasmid p3SP-NDM in clinical Enterobacter aerogenes. Front Microbiol. 2015; 6:294.
35. Doi Y, Arakawa Y. 16S ribosomal RNA methylation: emerging resistance mechanism against aminoglycosides. Clin Infect Dis. 2007; 45:88-94.
36. CLSI. (2015). Performance standards for antimicrobial susceptibility testing: twenty-fifth informational supplement M100-S25. (Wayne, PA, USA: CLSI).
37. Chin CS, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods. 2013; 10:563-569.
38. Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Thomason JA 3rd, Stevens R, et al. RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep. 2015; 5:8365.
39. Boratyn GM, Camacho C, Cooper PS, Coulouris G, Fong A, Ma N, Madden TL, Matten WT, McGinnis SD, Merezhuk Y, Raytselis Y, Sayers EW, Tao T, et al. BLAST: a more efficient report with usability improvements. Nucleic Acids Res. 2013; 41:W29-33.
40. Boutet E, Lieberherr D, Tognolli M, Schneider M, Bansal P, Bridge AJ, Poux S, Bougueleret L, Xenarios I. UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Mol Biol. 2016; 1374:23-54.
41. O’Leary NA, Wright MW, Brister JR, Ciufo S, Haddad D, McVeigh R, Rajput B, Robbertse B, Smith-White B, Ako-Adjei D, Astashyn A, Badretdin A, Bao Y, et al. Reference sequence (RefSeq) database at NCBI: current status, taxonomic expansion, and functional annotation. Nucleic Acids Res. 2016; 44:D733-745.
42. Jia B, Raphenya AR, Alcock B, Waglechner N, Guo P, Tsang KK, Lago BA, Dave BM, Pereira S, Sharma AN, Doshi S, Courtot M, Lo R, et al. CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res. 2017; 45:D566-D573.
43. Zankari E, Hasman H, Cosentino S, Vestergaard M, Rasmussen S, Lund O, Aarestrup FM, Larsen MV. Identification of acquired antimicrobial resistance genes. J Antimicrob Chemother. 2012; 67:2640-2644.
44. Siguier P, Perochon J, Lestrade L, Mahillon J, Chandler M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006; 34:D32-36.
45. Moura A, Soares M, Pereira C, Leitao N, Henriques I, Correia A. INTEGRALL: a database and search engine for integrons, integrases and gene cassettes. Bioinformatics. 2009; 25:1096-1098.
46. Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004; 32:1792-1797.

