INTRODUCTION
The Grb2-associated binding protein 2 (Gab2) is 74 kDa scaffolding protein that is involved in functional cellular signalling and cancer development [1–4]. It is composed of large disordered regions and various structural domains and docking sites that act as a platform for the assembly of several protein-protein interactions [5]. While lacking any enzymatic activity, Gab2 is a direct mediator of receptor tyrosine kinases and non-receptor tyrosine kinases, such as cytokine and G-protein–coupled receptors, transmitting and amplifying signals to downstream effectors [6–8]. In fact, upon stimulation, Gab2 activates and binds to several targets involved in signal transduction and has a role in a variety of cellular functions, such as proliferation, survival, migration, and differentiation [9–11].
Gab2 is implicated in a number of cancers of both solid and haematological origin. In particular, Gab2 is overexpressed in breast [12], gastric [13] and lung [14] cancers. Furthermore, its role is clearly established in haematological cancers, like juvenile myelomonocytic leukaemia [15], chronic myelogenous leukemia (CML) [16], acute leukaemia [17] and acute lymphoblastic leukaemia [18]. Importantly, whilst the functions of Gab2 are critical in the embryonic and postnatal development, its expression in a healthy mature cell is relatively suppressed. For these reasons, targeting Gab2 provides a promising route to develop new strategies for highly selective drug therapies in anticancer treatment. In fact, this protein acts at the head of signalling networks, by conveying inputs from membrane receptors to the cytoplasm, where it dictates the functions of multiple pathways.
From a structural perspective, the N-terminal part of Gab2 consists of a Pleckstrin Homology domain (of about 120 amino acids), which binds the lipid phosphatidylinositol-3, 4, 5-triphosphate of the plasma membrane and is critical for the membrane localization of the protein [5], whereas the remainder of the protein (whose total length is 676 amino acids) essentially lacks any defined secondary or tertiary structure, such that Gab2 may be classified as an intrinsically disordered protein (IDP).
The disordered regions of Gab2 have a fundamental functional relevance, by promoting the direct interaction with a number of cytoplasmic partners containing SH2 or SH3 domains [10, 19–21]. Typically, upon binding to their specific partners, the disordered regions of Gab2 encounter a disorder-to-order transition, as exemplified for example in [22]. Because of this complexity, being the IDPs typically elusive to a mechanistic characterizations [23], the detailed mechanisms of interactions between Gab2 and its physiological partners have remained elusive.
One of the critical physiological interactions involving Gab2 lies in the binding with Grb2 [19]. This recognition occurs between the C-terminal SH3 domain of the latter protein (Grb2 SH3C) and a region of Gab2 encompassing residues 503-524 (Gab2503-524) [24], which is predominantly disordered in isolation whereas it adopts a polyproline type II fold upon binding (Figure 1) [24]. Interestingly, in its unbound state Gab2503-524 has a degree of propensity to adopt conformations resembling the Grb2-bound state [22]. Whilst the structural features of Gab2503-524 has been described both in its free state and when bound the Grb2 SH3C [24], little is known about the mechanism of recognition between these two elements. Characterising this mechanism of recognition is of great importance for understanding both the molecular basis underlying the role of Gab2 in functional and pathological processes and for the general structural basis of protein-protein interactions by IDPs. In this work, we address this important open question by providing a detailed kinetic characterization on the recognition between Gab2503-524 and Grb2 SH3C. Binding kinetics, measured both in the μs and ms time range, were obtained by monitoring intrinsic fluorescence using both a stopped flow and temperature jump techniques. In agreement with what expected from a disordered system [25, 26], data reveal the presence of complex kinetics, which enabled the identification of at least two kinetic steps. We show that a comparison between the experiments performed by varying the concentration of Gab2503-524 or Grb2 SH3C reveals unambiguously that the reaction occurs via an induced fit type mechanism, whereby the folding of Gab2503-524 to a polyproline type II structure is subsequent to the formation of an initial encounter complex, despite its partial character of polyproline II when in the unbound state. Furthermore, by performing an alanine scanning of the four proline residues of Gab2503-524, we observe that the association rate constant between the two molecules correlate with the stability of the polyproline type II structure, an observation that supports the view that the initial encounter complex, whilst not fully folded, contains some native-like secondary interactions, as also suggested by the NMR analysis of the isolated fragment [22].
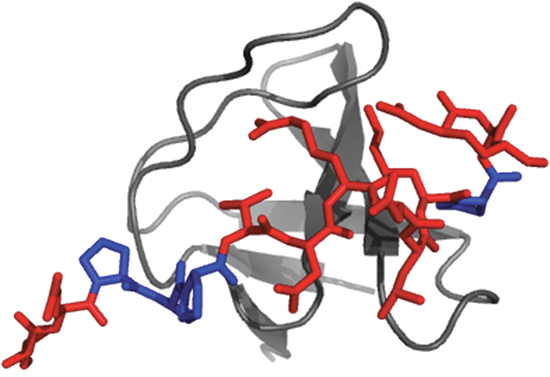
Figure 1: Cartoon representation of the Gab2-Grb2 SH3C complex. Three-dimensional structure of the complex between the C-terminal SH3 domain of the Grb2 protein (Grb2 SH3C) and a peptide representing the residues 503-524 of the Grb2-associated binding protein 2 (Gab2503-524). The SH3 domain is represented in gray while the peptide is represented in red. The four proline residues of Gab2503-524 are highlighted in blue. The picture was generated using the software PyMOL (pdb code: 2vwf). Crystallization conditions were 50mM HEPES pH 7.5, 50mM NaCl (peptide:protein 3:1 molar ratio); reservoir: 0.7 M tri-sodium citrate, 0.1 M Tris pH 8.5.
RESULTS AND DISCUSSION
Understanding the mechanism of binding between two molecules requires first to establish the simplest kinetic scheme describing the reaction adequately and then to infer the mechanistic details [27]. Accordingly, the rational of our experimental efforts to unveil the recognition between Gab2503-524 and Grb2 SH3C was focused first on unveiling the order of events and subsequently on understanding the main features of the reaction.
A folding after binding mechanism describes the recognition between Gab2503-524 and Grb2 SH3C
An intrinsically disordered protein, such as Gab2503-524, may explore a disorder-to-order transition upon binding to its physiological partner. In these cases, from a mechanistic perspective, it may be argued that the reaction could occur via two extreme mechanisms, namely conformational selection, formally equivalent to a Monod−Wyman−Changeaux model, [28] or folding after binding, equivalent to an induced fit model [29]. Although conformational selection requires the IDP to populate at least partly the folded state in the free form, i.e. as observed for Gab2503-524 [22], the pre-existence of ordered structures in IDPs does not necessarily commit to a conformational selection mechanism, and kinetic studies are needed to address these questions [30].
In an effort to understand the order of the events in the folding reaction of Gab2503-524 upon binding to Grb2 SH3C, we resorted to apply a kinetic test, originally introduced by Olson et al [31] and more recently applied on different protein systems [32–34]. This experimental strategy is essentially based on the comparison of the observed binding kinetics when performing pseudo-first order experiments with respect to each ligand. Thus, we performed relaxation binding experiments between Gab2503-524 and Grb2 SH3C by temperature jump. In particular, a constant concentration of Grb2 SH3C was mixed with different excess concentrations of Gab2503-524 and the resulting solution was subjected to a rapid increase of temperature of 9°C, from 16 to 25°C, using capacitor-discharge temperature-jump instrument. Similarly, we performed binding experiments by keeping the concentration of Gab2503-524 constant and varying the concentration of an excess of Grb2 SH3C. Binding was observed by monitoring the change in fluorescence of Grb2 SH3C, which is primarily ascribed to residues W35 and W36. A typical temperature jump trace is reported in Figure 2A. Under all the experimental conditions here considered, the binding time courses were consistent with a single exponential decay. While this observation, to a first approximation, may be consistent with a two-state mechanism, a detailed analysis of the observed rate constants, as shown below, revealed an additional level of complexity.
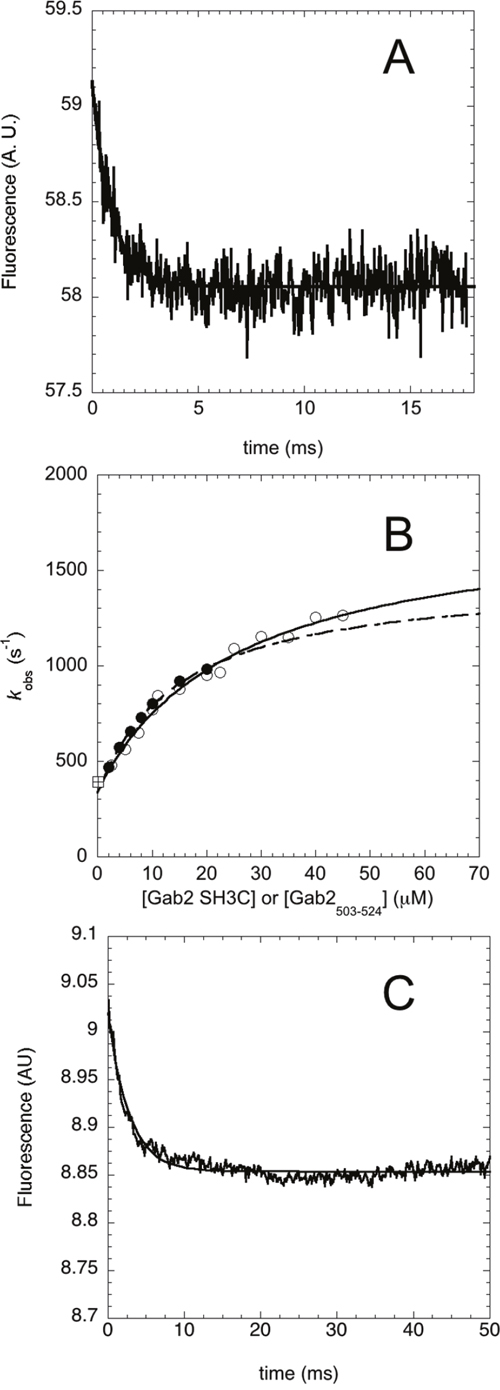
Figure 2: Binding kinetics between Grb2 SH3C and Gab2503-524 performed by using a temperature jump apparatus at 25°C. (A) Typical temperature jump time course obtained in a (un)binding experiment between Grb2 SH3C 4 μM and Gab2503-524 15 μM. Line is the best fit to a single exponential decay. (B) Plot of the observed relaxation rate constants as a function of reactant concentration. Pseudo-first-order binding experiments were performed by keeping the concentration of Grb2 SH3C at 4 μM and varying the concentration of Gab2503-524 (empty circles), and keeping the concentration of Gab2503-524 at 2 μM and varying the concentration of Grb2 SH3C (filled circles). In both cases the observed rate constants follow a hyperbolic behavior. The square symbol refer to the rate constant obtained from displacement experiments. (C) Displacement experiment. A pre-incubated complex of wild type Grb2 SH3C (10 μM) and Gab2503-524 (10 μM) was rapidly mixed with an excess of Grb2 SH3C variant W35Y/W36Y. The line is the best fit to an exponential decay.
A plot of the observed relaxation rate constants as a function of reactant concentration is reported in Figure 2B. The experiments performed under pseudo-first order conditions of the two different reactants provide data that are in both cases consistent with a hyperbolic dependence. This finding suggests that the observed kinetics is more complex than a simple two-state model, where a linear behavior would be expected, and at least a bimolecular and monomolecular step may be detected. For the nature of the reaction, we postulate these steps to be ascribed to a mechanism of binding and subsequent folding of Gab2503-524 into a polyproline type II structure. Since both experiments provide very similar values of observed rate constants, it may be concluded that the reaction occurs via an induced-fit type mechanism, whereby Gab2503-524 tends to recognize Grb2 SH3C in a relatively unstructured state and folding follows binding. In fact, it has been previously demonstrated that, due to its symmetry with respect to the reactants, only the induced fit scenario can account for a hyperbolic behavior when performing pseudo-first order experiments with respect to both ligands [31, 32, 34]. In contrast, in the case of conformational selection mechanism, an approximately linear dependence of the observed rate constant would have been observed when the concentration of Gab2503-524 was higher than the concentration of Grb2 SH3C.
In order to validate further the kinetic parameters obtained from pseudo-first-order experiments, we resorted to measure the dissociation rate constants with an additional method. In analogy to classical experiments on heme proteins [35], we carried out displacement experiments where a pre-incubated complex between two partners was challenged with an excess of a competing reactant. Thus, we performed displacement by producing a double mutant variant of Grb2 SH3C where the two fluorescent tryptophan residues were replaced with two tyrosines (W35Y/W36Y). Because W35Y/W36Y has a very different fluorescence yield compared to wild type Grb2 SH3C, a displacement between the pre-formed complex triggered by a rapid mix with an excess of W35Y/W36Y could be readily monitored. The observed dissociation rate constant was found insensitive to concentration of all the reactants and provided a value of koff = 350 ± 30 s-1. As shown in Figure 2, this value is consistent with what extrapolated from relaxation experiments.
Addressing the mechanism of recognition between Gab2503-524 and Grb2 SH3C by mutagenesis
It has been recently suggested that the proline residues of Gab2503-524 play a critical role both in tuning the preformed structure of the isolated protein and in modulating the affinity for Grb2 SH3C [22]. Thus, in an effort to understand directly the role of this position in the reaction mechanism, we carried out an alanine scanning of the four proline residues of Gab2503-524 and characterized the binding kinetics of the variants P510A, P511A, P512A and P519A. With the exception of P519, none of the other proline residues appear to participate directly to binding by establishing direct contacts with Grb2 SH3C. In order to compare the effect of mutagenesis more accurately, we performed these experiments by reducing the experimental temperature to 10°C and, therefore, measuring reaction kinetics using a stopped-flow apparatus. In these conditions, whilst we could not explore the full hyperbolic dependence of the observed rate constants (being too fast at high reactant concentrations for the stopped-flow apparatus), we could measure the apparent association and dissociation rate constant with higher precision. Importantly, control experiments with wild type Gab2503-524 show that the rate constants obtained with stopped-flow methodology were consistent with those obtained by temperature jump. Whilst for the variant P511A we could not measure any detectable binding, the observed pseudo-first-order rate constant for the P510A, P512A and P519A variant are reported in Figure 3. From the measured parameters a KD of 2.5 ± 0.2 μM, 2.5 ± 0.1 μM and 12 ± 1 μM could be calculated for P510A, P512A and P519A respectively.
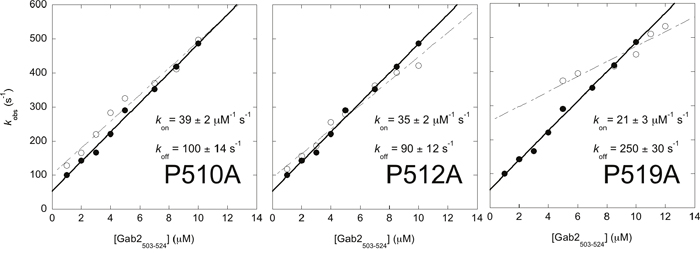
Figure 3: Alanine scanning of the proline residues of Gab2503-524 measured by using a stopped-flow apparatus at 10°C. The observed rate constants are plotted as a function of increasing Gab2503-524 concentration, while keeping Grb2 SH3C at a constant concentration of 0.5 μM. In each graph both Gab2503-524 wild-type (filled circles) and mutant (empty circles) are shown. Mutant P511A was not shown because it was not possible to measure any detectable binding. The observed rate constants follow a linear dependence from which the association (kon) and dissociation (koff) rate constants can be calculated. The calculated kon and koff for Gab2503-524 wild-type are 43 ± 1 μM-1 s-1 and 53 ± 7 s-1 respectively.
It is of interest to comment on the magnitude of the association rate constants of the different variants (Figure 3). In fact, earlier NMR analysis of the same variants suggested a clear ranking of WT > P510A > P512A > P519A > P511A in the tendency of populating preformed Grb2-bound like structures [22]. Accordingly, investigation of the data in Figure 3 suggests a similar trend when the association rate constant is taken into account. In particular, the lower the tendency of Gab2503-524 to explore pre-formed structure, the lower its rate constant of association to Grb2 SH3C. This finding appears to contrast our observation that the recognition between the two molecules occurs via a pure induced fit mechanism and suggests that some embryonic native-like secondary structure might be present in the initial encounter complex. On the basis of these observations, we suggest that, whilst the overall kinetics of binding is consistent with an induced fit scenario, at a molecular level a mixed mechanism may occur. More specifically, we propose that some pre-formed bound-like conformations are adopted by the disordered Gab2503-524, which confers some features of conformational selection. Thus, some native-like structure might be present already at the early stages of binding with Grb2 SH3C.
CONCLUSIONS
A comprehensive understanding of the binding between Gab2503-524 and Grb2 SH3C, a crucial interaction in normal cellular signalling for a number of cancer conditions, demands the full description of their mechanism of recognition. By performing a combined stopped-flow and temperature jump kinetic study, we successfully demonstrated that these two molecules bind through a initial encounter complex, whose characterization is potentially very important to the identification of molecules able to hijack this interaction and therefore to inhibit the aberrant functions of Gab2 in a number of solid and haematological cancers. Whilst the mechanism is formally similar to an induced fit scenario, the analysis of variants tuning the residual structure in Gab2503-524 indicates that the initial complex contains some embryonic bound-like secondary structure that funnels the binding energy landscape thereby effectively accelerating the formation of the macromolecular complex. Future work based on extensive site directed mutagenesis will shed light on the detailed structural features of such a metastable complex.
MATERIALS AND METHODS
Protein expression and purification
The sequence encoding for the Grb2 SH3C domain, subcloned in a pET28-b+ plasmid was used to transform E.coli cells (BL21). Expression of the protein was obtained growing bacterial cells in LB medium, at 37°C until they reached OD600 = 0.6-0.8 and then induced with IPTG at the concentration of 1mM at 37°C over night. Cells were then collected through centrifugation.
To purify the protein, pellet was resuspended in buffer TrisHCl 50mM, NaCl 0.5M, pH 7.5, with the addition of antiprotease tablet (Roche), then bacterial cells were lysed by sonication and centrifuged. The lysate was loaded onto a Nickel-charged Hi-trap Chelating column (GE Healthcare) equilibrated with buffer TrisHCl 50mM, NaCl 0.5M, 10mM Imidazole, pH 7.5 and eluted with a gradient from 10mM to 1M Imidazole using an ÄKTA-prime system. Fractions containing the purified protein were analysed through SDS-page and collected. Buffer was then exchanged with 25mM HEPES, 100mM potassium acetate, pH 7.5.
The Gab2503-524 peptide in its wild-type form (SRGSEIQPPPVNRNLKPDRKAK) and its variants were purchased from JPT, Germany. Peptide concentrations were measured by weighting 1 mg of the sample in an analytical balance and concentration was further confirmed by measuring absorbance at 205 nm.
Temperature-jump binding experiments
Kinetic experiments were performed using a PTJ-64 capacitor-discharge T-jump apparatus (TgK Scientific). Temperature was rapidly changed with a jump-size of 9°C, from 16°C to 25°C. The buffer used was 25mM HEPES, 100mM potassium acetate, pH 7.5, the excitation wavelength was 296 nm, and the fluorescence emission was collected using a 320-nm cut-off glass filter. Binding experiments were performed by keeping Grb2 SH3C concentration constant at 4 μM and varying the concentration of Gab2503-524 from 2.5 μM to 45 μM, and keeping Gab2503-524 concentration constant at 2 μM and varying the concentration of Grb2 SH3C from 2 μM to 20 μM. All the observed relaxation rate constants obtained and analysed were calculated from the average of 10 individual traces.
Stopped-flow binding experiments
Binding kinetics experiments were carried out on a single-mixing SX-18 stopped-flow instrument (Applied Photophysics); the excitation wavelength used was 280 nm and the fluorescence emission was collected using a 320-nm cut-off glass filter. Pseudo-first order binding experiments were performed mixing a constant concentration of Grb2 SH3C (0.5μM) versus increasing concentration of Gab2503-524, from 1μM to 10μM, in its wild-type form and its variants P510A, P511A, P512A, and from 1μM to 12μM for the P519A variant.
For all binding experiments the temperature was 10°C and the buffer used was 25mM HEPES, 100mM potassium acetate, pH 7.5. The observed rate constants were calculated from the average of 3-6 single traces. Association rate constants (kon) were calculated as the slope of a linear function fitting the observed rate constants versus ligand concentration, while dissociation rate constants (koff) were calculated either from extrapolation as the intercept of y-axis, or from separate displacement experiments.
Stopped-flow displacement experiments
The dissociation rate constant was measured by carrying out displacement experiments on a single-mixing SX-18 stopped-flow instrument (Applied Photophysics); the excitation wavelength used was 280 nm and the fluorescence emission was collected using a 320-nm cut-off glass filter. A pre-incubated complex of wild type Grb2 SH3C (10 μM) and Gab2503-524 (10 μM) was rapidly mixed with an excess of Grb2 SH3C variant W35Y/W36Y (100 μM and 200 μM). Displacement experiments were performed at 25°C, in presence of 25mM HEPES, 100mM potassium acetate, pH 7.5. The observed rate constants were calculated from the average of 5 single traces. Observed kinetics was consistent with a single exponential decay.
Author contributions
A.D.S and S.G. designed research; A.T. and D.B. performed research; A.T., D.B. and S.G. analysed data and all the authors revised the manuscript.
ACKNOWLEDGMENTS
Work partly supported by grants from the Italian Ministero dell’Istruzione dell’Università e della Ricerca (Progetto di Interesse ‘Invecchiamento’ to S.G.), Sapienza University of Rome (C26A155S48 and B52F16003410005 to S.G; C26N15J4A5 to A.T. and C26N15E4LB and B52F16001770005 to D.B.) and the Associazione Italiana per la Ricerca sul Cancro (Individual Grant - MFAG 2016, 18701 to S.G.). AT is a recipient of a post-doctoral fellowship from the Istituto Pasteur Italia – Cenci Bolognetti.
CONFLICTS OF INTEREST
Authors declare no conflicts of interest.
REFERENCES
1. Nishida K, Wang L, Morii E, Park SJ, Narimatsu M, Itoh S, Yamasaki S, Fujishima M, Ishihara K, Hibi M, Kitamura Y, Hirano T. Requirement of Gab2 for mast cell development and KitL/c-Kit signaling. Blood. 2002; 99:1866-9.
2. Nishida K, Yoshida Y, Itoh M, Fukada T, Ohtani T, Shirogane T, Atsumi T, Takahashi-Tezuka M, Ishihara K, Hibi M, Hirano T. Gab-family adapter proteins act downstream of cytokine and growth factor receptors and T- and B-cell antigen receptors. Blood. 1999; 93:1809-16.
3. Adams SJ, Aydin IT, Celebi JT. GAB2--a scaffolding protein in cancer. Mol Cancer Res. 2012; 10:1265-70.
4. Ding CB, Yu WN, Feng JH, Luo JM. Structure and function of Gab2 and its role in cancer. Mol Med Rep. 2015; 12:4007-14.
5. Simister PC, Feller SM. Order and disorder in large multi-site docking proteins of the Gab family--implications for signalling complex formation and inhibitor design strategies. Mol Biosyst. 2012; 8:33-46.
6. Kazi JU, Vaapil M, Agarwal S, Bracco E, Påhlman S, Rönnstrand L. The tyrosine kinase CSK associates with FLT3 and c-Kit receptors and regulates downstream signaling. Cell Signal. 2013; 25:1852-60.
7. Holgado-Madruga M, Emlet DR, Moscatello DK, Godwin AK, Wong AJ. A Grb2-associated docking protein in EGF- and insulin-receptor signalling. Nature. 1996; 379:560-4.
8. Gu H, Saito K, Klaman LD, Shen J, Fleming T, Wang Y, Pratt JC, Lin G, Lim B, Kinet JP, Neel BG. Essential role for Gab2 in the allergic response. Nature. 2001; 412:186-90.
9. Gu H, Griffin JD, Neel BG. Characterization of two SHP-2-associated binding proteins and potential substrates in hematopoietic cells. J Biol Chem. 1997; 272:16421-30.
10. Gu H, Pratt JC, Burakoff SJ, Neel BG. Cloning of p97/Gab2, the major SHP2-binding protein in hematopoietic cells, reveals a novel pathway for cytokine-induced gene activation. Mol Cell. 1998; 2:729-40.
11. Lynch DK, Daly RJ. PKB-mediated negative feedback tightly regulates mitogenic signalling via Gab2. EMBO J. 2002; 21:72-82.
12. Bentires-Alj M, Gil SG, Chan R, Wang ZC, Wang Y, Imanaka N, Harris LN, Richardson A, Neel BG, Gu H. A role for the scaffolding adapter GAB2 in breast cancer. Nat Med. 2006; 12:114-21.
13. Lee SH, Jeong EG, Nam SW, Lee JY, Yoo NJ, Lee SH. Increased expression of Gab2, a scaffolding adaptor of the tyrosine kinase signalling, in gastric carcinomas. Pathology. 2007; 39:326-9.
14. Xu XL, Wang X, Chen ZL, Jin M, Yang W, Zhao GF, Li JW. Overexpression of Grb2-associated binder 2 in human lung cancer. Int J Biol Sci. 2011; 7:496-504.
15. Mohi MG, Williams IR, Dearolf CR, Chan G, Kutok JL, Cohen S, Morgan K, Boulton C, Shigematsu H, Keilhack H, Akashi K, Gilliland DG, Neel BG. Prognostic, therapeutic, and mechanistic implications of a mouse model of leukemia evoked by Shp2 (PTPN11) mutations. Cancer Cell. 2005; 7:179-91.
16. Gu S, Chan WW, Mohi G, Rosenbaum J, Sayad A, Lu Z, Virtanen C, Li S, Neel BG, Van Etten RA. Distinct GAB2 signaling pathways are essential for myeloid and lymphoid transformation and leukemogenesis by BCR-ABL1. Blood. 2016; 127:1803-13.
17. Zatkova A, Schoch C, Speleman F, Poppe B, Mannhalter C, Fonatsch C, Wimmer K. GAB2 is a novel target of 11q amplification in AML/MDS. Genes Chromosomes Cancer. 2006; 45:798-807.
18. Wöhrle FU, Daly RJ, Brummer T. Function, regulation and pathological roles of the Gab/DOS docking proteins. Cell Commun Signal. 2009; 7:22.
19. Zhao C, Yu DH, Shen R, Feng GS. Gab2, a new pleckstrin homology domain-containing adapter protein, acts to uncouple signaling from ERK kinase to Elk-1. J Biol Chem. 1999; 274:19649-54.
20. Solomaha E, Szeto FL, Yousef MA, Palfrey HC. Kinetics of Src homology 3 domain association with the proline-rich domain of dynamins: specificity, occlusion, and the effects of phosphorylation. J Biol Chem. 2005; 280:23147-56.
21. Harkiolaki M, Lewitzky M, Gilbert RJ, Jones EY, Bourette RP, Mouchiroud G, Sondermann H, Moarefi I, Feller SM. Structural basis for SH3 domain-mediated high-affinity binding between Mona/Gads and SLP-76. EMBO J. 2003; 22:2571-82.
22. Krieger JM, Fusco G, Lewitzky M, Simister PC, Marchant J, Camilloni C, Feller SM, De Simone A. Conformational recognition of an intrinsically disordered protein. Biophys J. 2014; 106:1771-9.
23. Habchi J, Tompa P, Longhi S, Uversky VN. Introducing protein intrinsic disorder. Chem Rev. 2014; 114:6561-88.
24. Harkiolaki M, Tsirka T, Lewitzky M, Simister PC, Joshi D, Bird LE, Jones EY, O'Reilly N, Feller SM. Distinct binding modes of two epitopes in Gab2 that interact with the Sh3C domain of Grb2. Structure. 2009; 17:809-22.
25. Dogan J, Gianni S, Jemth P. The binding mechanisms of intrinsically disordered proteins. Phys Chem Chem Phys. 2014; 16:6323-31.
26. Kiefhaber T, Bachmann A, Jensen KS. Dynamics and mechanisms of coupled protein folding and binding reactions. Curr Opin Struct Biol. 2012; 22:21-9.
27. Fersht AR. Structure and mechanism in protein science. Freeman, New York, 1999.
28. Monod J, Wyman J, Changeux JP. On the nature of allosteric transitions: a plausible model. J Mol Biol. 1965; 12:88-118.
29. Koshland DE, Némethy G, Filmer D. Comparison of experimental binding data and theoretical models in proteins containing subunits. Biochemistry. 1966; 5:365-85.
30. Dogan J, Jemth P. Only kinetics can prove conformational selection. Biophys J. 2014; 107:1997-8.
31. Olson ST, Srinivasan KR, Bjork I, Shore JD. Binding of high affinity heparin to antithrombin III. Stopped flow kinetic studies of the binding interaction. J Biol Chem. 1981; 256:11073-9.
32. Gianni S, Dogan J, Jemth P. Distinguishing induced fit from conformational selection. Biophys Chem. 2014; 189:33-9.
33. Dosnon M, Bonetti D, Morrone A, Erales J, di Silvio E, Longhi S, Gianni S. Demonstration of a folding after binding mechanism in the recognition between the measles virus NTAIL and X domains. ACS Chem Biol. 2015; 10:795-802.
34. Gianni S, Walma T, Arcovito A, Calosci N, Bellelli A, Engstrom A, Travaglini-Allocatelli C, Brunori M, Jemth P, Vuister GW. Demonstration of long-range interactions in a PDZ domain by NMR, kinetics, and protein engineering. Structure. 2006; 14:1801-9.
35. Antonini E, Brunori M. Hemoglobin and myoglobin in their reactions with ligands. North-Holland Publ Co, Amsterdam, The Netherlands, 1971.

