INTRODUCTION
Prostate cancer (PCa) is one of the most frequently diagnosed malignancies in men resulting in more than 250,000 deaths annually [1, 2]. Although PCa is the most common non-cutaneous tumor in developed countries, its etiology remains poorly understood [3, 4]. Identifying risk factors for PCa is critical for developing improved therapeutic interventions and gaining better understanding of the biology of this disease. The major risk factors for prostate cancer include age, race and family history [5–7]. Nearly 42% of the known risk factors of PCa are hereditary and more than 40 single nucleotide polymorphisms (SNPs) are significantly associated with the risk of PCa [8–18].
The enzyme UGT2B15 belongs to a large group of UDP glucuronosyltransferases [19] of the endoplasmic reticulum that inactivates lipophilic C19 steroid androgens, such as dihydrotestosterone (DHT) into water soluble metabolites that can be excreted. UGT2B15 is expressed in many steroid-sensitive tissues like the prostate [20] and its altered expression is associated with the growth of hormone-refractory PCa [21].
The D85Y polymorphism in the UGT2B15 gene that results from a single-base-pair change of guanine to thymine leads to a variant with higher Vmax with reduced risk of PCa [22]. Since UGT2B enzymes are involved in steroid metabolism and excretion, they play an important role in prostate health. However, recent studies that have investigated the association between the uridine diphosphate-glucuronosyltransferase 2B gene polymorphism and prostate cancer have shown contradictory results [23–28]. Some studies reported that the polymorphism increased the risk of prostate cancer, whereas, others did not. Therefore, we systematically reviewed the available literature and performed a meta-analysis to evaluate the association of UGT2B15 D85Y gene polymorphism with prostate cancer risk.
RESULTS
Characteristics of included studies
The details of the literature review process performed in this study are shown in Figure 1. Initially, a total of 509 unduplicated studies were identified from multiple literature databases and ultimately, six that were in accordance with the eligibility criteria were selected in agreement by all reviewers. The data from the six studies is summarized in Table 1. The six studies included 817 prostate cancer patients and 1000 normal controls and all of them reported exclusion/inclusion criteria [23–28]. In addition, all of these studies tested for the D85Y polymorphism in the UGT2B15 gene by using restriction fragment length polymorphism (RFLP) analysis after polymerase chain reaction (PCR) amplification.
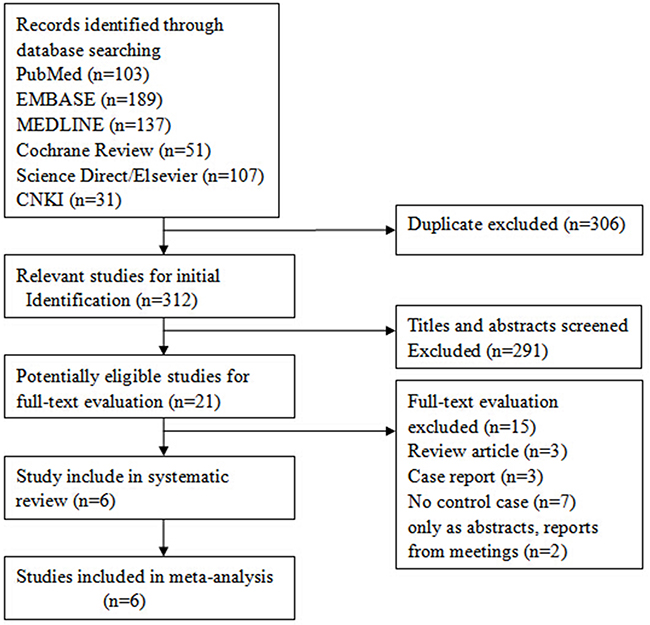
Figure 1: Flow diagram showing selection criteria of eligible studies.
Table 1: Characteristics of the included studies
Author |
Country |
Case |
Control |
Genotyping method |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
n |
D |
Y |
DD |
YD |
YY |
n |
D |
Y |
DD |
YD |
YY |
|||
Okugi H 2006 |
Japan |
102 |
139 |
65 |
50 |
39 |
13 |
117 |
137 |
97 |
33 |
71 |
13 |
PCR-RFLP |
Gsur A 2002 |
Australia |
190 |
179 |
201 |
40 |
99 |
51 |
190 |
187 |
193 |
47 |
93 |
50 |
PCR-RFLP |
Grant D J2013 |
USA |
100 |
111 |
73 |
32 |
47 |
13 |
297 |
275 |
319 |
68 |
139 |
90 |
PCR-RFLP |
Park J 2004 |
USA |
155 |
176 |
134 |
52 |
72 |
31 |
154 |
238 |
170 |
21 |
96 |
37 |
PCR-RFLP |
MacLeod SL 2007 |
USA |
64 |
89 |
39 |
26 |
37 |
1 |
64 |
73 |
55 |
12 |
49 |
3 |
PCR-RFLP |
Hajdinja T 2004 |
Slovenia |
206 |
195 |
217 |
47 |
101 |
58 |
178 |
148 |
208 |
28 |
92 |
58 |
PCR-RFLP |
Abbreviation: PCR-RFLP: polymerase chain reaction-restriction fragment length polymorphism
Meta-analysis
Based on the heterogeneity test results, the recessive model was analyzed by the fixed-effects model and the other (additive, dominant, co-dominant and allele) models were analyzed by the random-effects model. The meta-analysis revealed significant association between the UGT2B15 D85Y gene polymorphism and prostate cancer in all the genetic models (P<0.05). The combined ORs and 95% CIs were as follows: additive model (0.53, 0.32–0.88; Figure 2), co-dominant model (0.55, 0.35–0.86; Figure 3), dominant model (0.51, 0.33–0.79; Figure 4), recessive model (0.76, 0.60–0.96; Figure 5) and allele model (0.70, 0.55–0.89; Figure 6). The symmetric Begg’s funnel plots suggested that there was no publication bias in the meta-analysis (Figures 7A, 8A, 9A, 10A, 11A). The Egger’s regression test also indicated no evidence of publication bias in all the genetic models (P>0.05; Table 2). Further, we evaluated the sensitivity of the meta-analysis by omitting one study at a time and found that the combined ORs for the remaining studies remained consistent. Overall, no single study significantly changed the combined results indicating that the results were stable and reliable (Figures 7B, 8B, 9B, 10B, 11B).
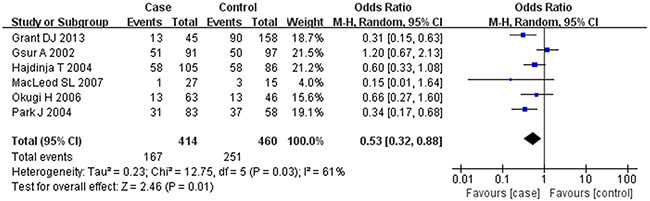
Figure 2: Forest plot showing the meta-analysis outcomes of the additive model.
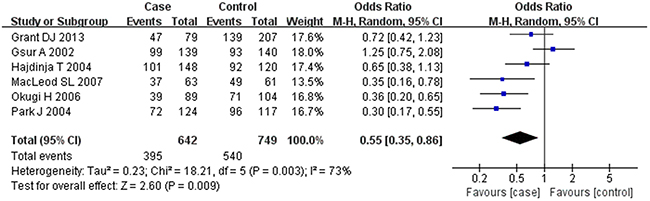
Figure 3: Forest plot showing the meta-analysis outcomes of the co-dominant model.
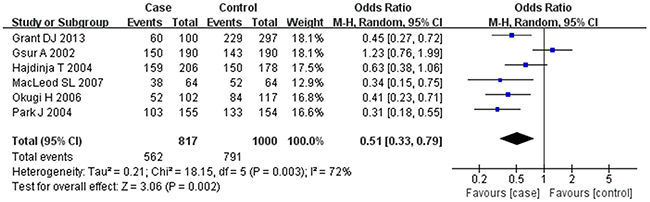
Figure 4: Forest plot showing the meta-analysis outcomes of the dominant model.
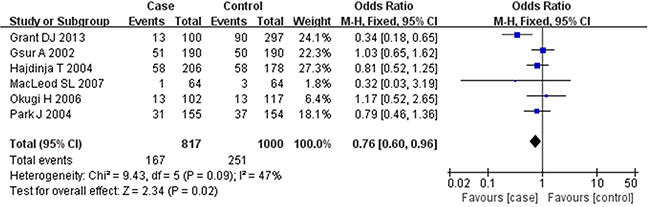
Figure 5: Forest plot showing the meta-analysis outcomes of the recessive model.
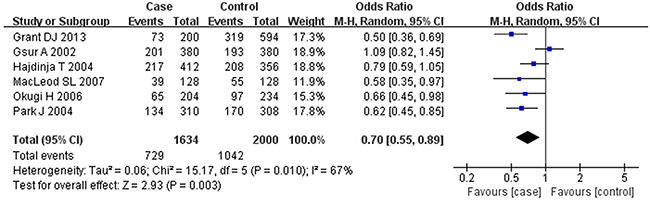
Figure 6: Forest plot showing the meta-analysis outcomes of the allele model.
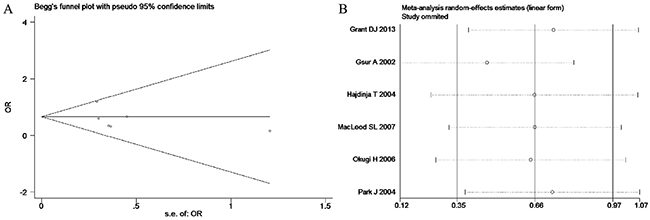
Figure 7: (A) Begg’s publication bias and (B) Sensitivity analysis plot of Additive model.
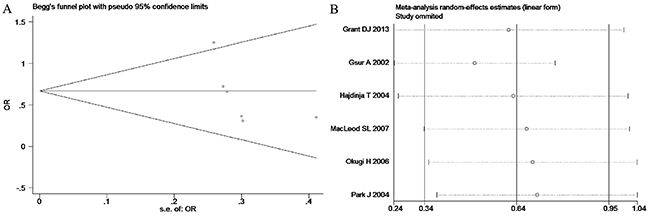
Figure 8: (A) Begg’s publication bias and (B) Sensitivity analysis plot of Co-dominant model.
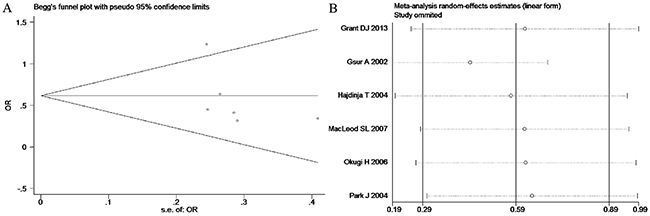
Figure 9: (A) Begg’s publication bias and (B) Sensitivity analysis plot of Dominant model.
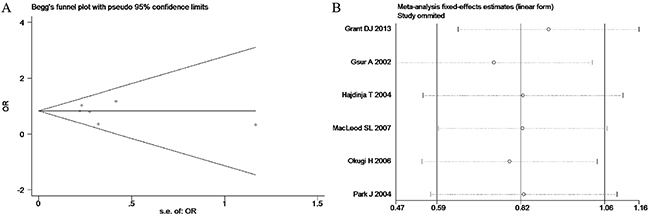
Figure 10: (A) Begg’s publication bias and (B) Sensitivity analysis plot of Recessive model.
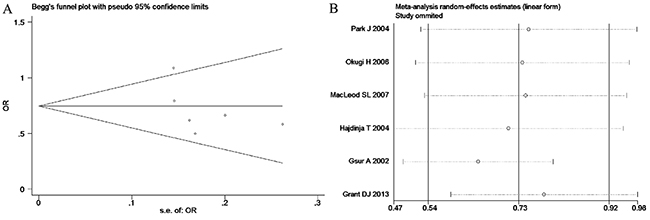
Figure 11: (A) Begg’s publication bias and (B) Sensitivity analysis plot of Allele model.
Table 2: Egger’s test of publication bias
Models |
Coeff. |
Std. Err. |
t |
P>|t| |
[95% Conf. Interval] |
|
|---|---|---|---|---|---|---|
Allele |
-3.36 |
2.88 |
-1.17 |
0.31 |
-13.46 |
5.89 |
Additive |
-1.09 |
1.36 |
-0.80 |
0.47 |
-4.88 |
2.69 |
Dominant |
-3.79 |
3.48 |
-1.09 |
0.34 |
-13.46 |
5.89 |
Recessive |
-0.51 |
1.06 |
-0.48 |
0.66 |
-3.45 |
2.43 |
Co-dominant |
-5.39 |
3.39 |
-1.59 |
0.19 |
-14.80 |
4.03 |
DISCUSSION
This study focused on six reports in the literature to analyze the association of the uridine diphosphate-glucuronosyltransferase 2B15 gene polymorphism with the risk of prostate cancer. Our results showed that all genetic models (additive, dominant, recessive, co-dominant and allele) demonstrated significant association between the UGT2B15 D85Y gene polymorphism and prostate cancer. Studies have shown that androgens play important roles in the growth, proliferation and progression of prostate cancer. Therefore, our study showed that the UGT2B15 gene product that is involved in androgen metabolism played a critical role. Specifically, the data indicated that UGT2B15 Y85 was protective against prostate cancer.
D85Y polymorphism was non-synonymous with UGT2B15 gene function [29–31]. Previous studies showed that S-oxazepam was a selective substrate for UGT2B15 and that the common UGT2B15 allozyme variant (D85Y) demonstrated altered S-oxazepam glucuronidation activity [30]. Further, the individual variability of oxazepam glucuronidation by human liver was determined in 35% of cases due to D85Y genotype and 14% of cases due to donor gender [31]. In addition, there was strong correlation between the genetic variants of UGT2B15 and sipoglitazar exposure with nearly two-thirds of the inter-subject variability in sipoglitazar plasma exposure attributed to the UGT2B15 genetic variation [32]. All these studies implied that UGT2B15 D85Y was a major determinant of S-oxazepam clearance and a useful in vivo probe for glucuronidation by UGT2B15 [33]. Similarly, other studies also demonstrated that the absence of the UGT2B15 protein during early gestation and relatively low expression during late gestation and the neonatal period were consistent with greater risks from some UGT2B15 substrates during these time periods. However, further studies are necessary to confirm these findings.
The UGT (UDP-glucuronosyltransferase) super- family of enzymes comprises endoplasmic reticulum-bound proteins that catalyze the glucuronidation of numerous compounds including drugs, carcinogens and hormones and the resultant inactive polar derivatives are excreted in bile and urine. The UGT1 enzymes on chromosome 2 are primarily involved in the conjugation of bilirubin, phenols, and estrogens. The larger UGT2B enzyme family (on chromosome 4) is grouped into two subfamilies with highly homologous members, namely UGT2A and UGT2B. The UGT2A genes are primarily expressed in olfactory epithelium and are involved in terminating odorant signaling. The UGT2B enzymes (UGT2B4, UGT2B7, UGT2B15, UGT2B17, and UGT2B28) are widely expressed in the liver and extra-hepatic steroid target tissues mediating the conjugation of diverse steroid substrates [34, 35]. Sequence divergence in the substrate-binding domains encoded by exons 1 and 2 determines the substrate specificity, whereas the remaining exons encode the conserved UDP-glucuronic acid-binding and the transmembrane domains [36]. Although the association of the UGT2B enzymes with serum steroid levels is unclear, they are important in the biotransformation of steroid hormones and few studies have suggested their association with the risk of prostate cancer [23–28].
As a member of the UGT2B enzyme family, UGT2B15 is a key enzyme for the inactivation of DHT, an active androgen in prostate cells. In the human UGT2B15 gene, a guanine-to-thymine SNP was identified that resulted in an amino acid change from aspartate (D) to tyrosine (Y) at position 85 [37]. Although both UGT2B15 (D85) and UGT2B15 (Y85) had similar substrate specificities, the Vmax of UGT2B15 (Y85) was twice that of UGT2B15 (D85) for C19 steroids, such as 5-androstane-3, 17-diol and DHT [38].
Levesque et al. reported that the Vmax of the enzyme resulting from the Y allele was approximately twice compared to the D allele, although no significant differences were evident in the Kmax of the two enzymes. These findings implied that the D enzyme inefficiently metabolized DHT, resulting in a high DHT concentration in the prostate tissue. Higher levels of DHT were associated with prostate cancer development [39, 40]. Our findings showed association of the D/D genotype with prostate cancer risk supporting the previous findings that associated high DHT with prostate cancer development.
However, there are few limitations in our study that need to be considered when interpreting the results of this meta-analysis. First, the sample size of each of the studies was relatively small with a total of 817 prostate cancer patients and 1000 normal controls investigated in the six studies. Second, several studies had to be excluded from the meta-analysis owing to lack of control data. Because of limited samples, subgroup analysis could not be performed and therefore, definitive conclusions could not be drawn regarding the clinical value of the UGT2B15 D85Y gene variant in prostate cancer.
In summary, the results of this meta-analysis suggest that the UGT2B15 D85Y gene variant is protective against prostate cancer. The plausible reason is that the variant has enhanced ability to metabolize DHT, thereby reducing the risk of prostate cancer by decreasing testosterone levels. However, studies with larger sample sizes are necessary to confirm this finding.
MATERIALS AND METHODS
Literature search
This meta-analysis was restricted to published studies that investigated the association between UGT2B15 D85Y gene polymorphism and prostate cancer risk. Two independent reviewers searched PubMed, EMBASE, Science Direct/Elsevier, MEDLINE CNKI, and the Cochrane Library from their inception until June 2016 without restrictions on the language of the report or the study type. The search terms combined text words and MeSH terms. For example, the search terms for the uridine diphosphate-glucuronosyltransferase 2B15 gene were “uridine diphosphate-glucuronosyltransferase 2 family, polypeptide B15” “UGT2B15”, “UDP glucuronosyltransferase 2 family”, “polypeptide B15” or “ugt2b15”; those for prostate cancer were “prostate cancer”, “prostatic neoplasms”, “cancer of prostate”, “cancer of the prostate”, “neoplasms, prostate”, “neoplasms, prostatic”, “prostate neoplasms”, “prostatic cancer” or “PCa”; and those for the polymorphism were “SNP”, “single-nucleotide polymorphism”, “polymorphism”, “variation” or “mutation.” All related articles and abstracts were retrieved. In addition, references cited within relevant reviews were retrieved manually. The search focused only on full articles for the meta-analysis.
Eligibility criteria
The inclusion criteria for the studies were: (1) Studies that tested the association of uridine diphosphate-glucuronosyltransferase 2B gene variants with prostate cancer; (2) The case groups were prostate cancer patients and healthy individuals were controls and (3) Genotyping for the D85Y SNP was conducted using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). The data extracted from relevant articles included eligible and genotyped cases with controls and the total numbers of cases and controls for each D85Y genotype.
The exclusion criteria were: (1)Studies that were case reports, abstracts, reports from meetings, review articles or duplicated previous publications (2) Studies that lacked a control population and (3) Studies that lacked data on genotype frequencies.
Study selection and validity assessment
Two independent reviewers screened the titles and abstracts of all citations obtained from the literature search and retrieved those that met the eligibility criteria. If the decision based on the title and abstract was ambiguous, the final decision was made after reviewing the full text. Disagreements were resolved by consensus between the two independent reviewers or decided by a third reviewer.
Data extraction and statistical analysis
The following data were extracted from the papers by the three reviewers: author names, year of publication, country, the numbers of genotype, genotyping methods, the numbers of cases and controls in each study regarding D85Y genotype and the outcomes. Quantitative meta-analysis was performed and analyzed by the two reviewers using Review Manager (RevMan) software (version 5.2; The Nordic Cochrane Centre, The Cochrane Collaboration, 2012, Copenhagen, Denmark) and Stata software (version 12.0; College Station, TX, USA).
The combined odds ratio (OR) and its 95% confidence interval (CI) were calculated. Heterogeneity or variation between studies was assessed using the P-value and the I-square statistic (I2) in the pooled analyses. If the P-value was less than 0.1 or the I2-value was greater than 50%, the summary estimate was analyzed in a random-effects model. Otherwise, a fixed-effects model was applied. The association between D85Y polymorphism in the UGT2B15 gene and prostate cancer risk was investigated in the allelic (Y vs. D), additive (YY vs. DD), dominant (YY and YD vs. DD), recessive (YY vs. YD and DD) and co-dominant (YD vs. DD) models, respectively, where D represents aspartic acid and Y represents tyrosine. Further, publication bias was detected by visual symmetry of the funnel plots, with asymmetry suggesting possible publication bias. It was also assessed by Begg’s and Egger’s test in the meta-analysis. A P-value of less than 0.05 indicated publication bias.
ACKNOWLEDGMENTS
This work was supported by the National Natural Science Foundation of China (Grant No. 81400761 and 81470989).
COMPETING INTERESTS
The authors declare no competing financial interests.
REFERENCES
1. Verzoni E, De Giorgi U, Derosa L, Caffo O, Boccardo F, Facchini G, Porcu L, De Vincenzo F, Zaniboni A, Chiuri VE, De Vivo R, Dinota A, Messina C, et al. Predictors of long-term response to abiraterone in patients with metastastic castration-resistantprostate cancer: a retrospective cohort study. Oncotarget. 2016; 7:40085-40094. doi: 10.18632/oncotarget.9485.
2. Stelloo S, Sanders J, Nevedomskaya E, de Jong J, Peters D, van Leenders GJ, Jenster G, Bergman AM, Zwart W. mTOR pathway activation is a favorable prognostic factor in human prostate adenocarcinoma. Oncotarget. 2016;7: 32916-24. doi: 10.18632/oncotarget.8767.
3. Jemal A, Center MM, DeSantis C, Ward EM. Global patterns of cancer incidence and mortality rates and trends. Cancer Epidemiol Biomarkers Prev. 2010; 19: 1893–1907.
4. Horwich A, Parker C, Bangma C, Kataja V. Prostate cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2010; 21: 129-133.
5. Carter BS, Bova GS, Beaty TH. Hereditary prostate cancer: epidemiologic and clinical features. J Urol. 1993; 150: 797-802.
6. Kupelian PA, Kupelian VA, Witte JS, Macklis R, Klein EA. Family history of prostate cancer in patients with localized prostate cancer: an independent predictor of treatment outcome. J Clin Oncol. 1997; 15: 1478-1480.
7. Bova GS, Partin AW, Isaacs SD. Biological aggressiveness of hereditary prostate cancer: long-term evaluation following radical prostatectomy. J Urol. 1998; 160: 660-663.
8. Gudmundsson J, Sulem P, Rafnar T. Common sequence variants on 2p15 and Xp11.22 confer susceptibility to prostate cancer. Nat Genet. 2008; 40: 281-283.
9. Eeles RA, Kote-Jarai Z, Giles GG. Multiple newly identified loci associated with prostate cancer susceptibility. Nat Genet. 2008; 40: 316-321.
10. Gudmundsson J, Sulem P, Gudbjartsson DF. Genome-wide association and replication studies identify four variants associated with prostate cancer susceptibility. Nat Genet. 2009; 41: 1122–1126.
11. Eeles RA, Kote-Jarai Z, Al Olama AA. Identification of seven new prostate cancer susceptibility loci through a genome-wide association study. Nat Genet. 2009; 41: 1116–1121.
12. Takata R, Akamatsu S, Kubo M. Genome-wide association study identifies five new susceptibility loci for prostate cancer in the Japanese population. Nat Genet. 2010; 42: 751–754.
13. Haiman CA, Chen GK, Blot WJ. Genome-wide association study of prostate cancer in men of African ancestry identifies a susceptibility locus at 17q21. Nat Genet. 2011; 43: 570-573.
14. Xu J, Mo Z, Ye D. Genomewide association study in Chinese men identifies two new prostate cancer risk loci at 9q31.2 and 19q13.4. Nat Genet. 2012; 44: 1231-1235.
15. Eeles RA, Olama AA, Benlloch S. Identification of 23 new prostate cancer susceptibility loci using the iCOGS custom genotyping array. Nat Genet. 2013; 45: 385-391.
16. Gudmundsson J, Sulem P, Manolescu A. Genome-wide association study identifies a second prostate cancer susceptibility variant at 8q24. Nat Genet. 2007; 39: 631–637.
17. Yeager M, Orr N, Hayes RB. Genomewide association study of prostate cancer identifies a second risk locus at 8q24. Nat Genet. 2007;39: 645-649.
18. Thomas G, Jacobs KB, Yeager M. Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet. 2008; 40: 310-315.
19. Tukey RH, Strassburg CP. Human UDP-glucuronosyltransferases: Metabolism, expression, and disease. Annu Rev Pharmacol Toxicol. 2000; 40: 581–616.
20. Barbier O, Lapointe H, Hum DW, Belanger A. Cellular localization of uridine diphosphoglucuronosyltransferase 2B enzymes in the human prostate by in situ hybridization and immunohistochemistry. J Clin Endocrinol Metab. 2000; 85: 4819-4826.
21. Kanaya J, Takashima M, Koh E, Namiki M. Androgenindependent growth in LNCaP cell lines and steroid uridine diphosphate-glucuronosyltransferase expression. Asian J Androl. 2003; 5: 9-13.
22. Levesque E, Beaulieu M, Green MD. Isolation and characterization of UGT2B15(Y85): A UDP-glucuronosyltransferase encoded by a polymorphic gene. Pharmacogenetics. 1997; 7: 317-325.
23. Grant DJ, Hoyo C, Oliver SD. Association of uridine diphosphate-glucuronosyltransferase 2B gene variants with serum glucuronide levels and prostate cancer risk. Genet Test Mol Biomarkers. 2013, 17: 3-9.
24. Park J, Chen L, Shade K. Asp85tyr polymorphism in the udpglucuronosyltransferase (UGT) 2B15 gene and the risk of prostate Cancer. J Urol. 2004; 171: 2484-2488.
25. Hajdinjak T, Zagradisnik B. Prostate cancer and polymorphism D85Y in gene for dihydrotestosterone degrading enzyme UGT2B15: Frequency of DD homozygotes increases with Gleason Score. Prostate. 2004; 59: 436-439.
26. Gsur A, Preyer M, Haidinger G. A polymorphism in the UDPGlucuronosyltransferase 2B15 gene (D85Y) is not associated with prostate cancer risk. Cancer Epidemiol Biomarkers Prev. 2002; 11: 497-498.
27. Okugi H, Nakazato H, Matsui H. Association of the polymorphisms of genes involved in androgen metabolism and signaling pathways with familial prostate cancer risk in a Japanese population. Cancer Detect Prev. 2006; 30: 262-268.
28. MacLeod S L, Nowell S, Plaxco J. An allele-specific polymerase chain reaction method for the determination of the D85Y polymorphism in the human UDP-glucuronosyltransferase 2B15 gene in a casecontrol study of prostate cancer. Ann Surg Oncol. 2000; 7: 777-782.
29. Ménard V, Eap O, Harvey M, Guillemette C, Lévesque E. Copy-number variations (CNVs) of the human sex steroid metabolizing genes UGT2B17 and UGT2B28 and their associations with a UGT2B15 functional polymorphism. Hum Mutat. 2009; 30:1310-9.
30. Court MH, Duan SX, Guillemette C, Journault K, Krishnaswamy S, Von Moltke LL, Greenblatt DJ. Stereoselective conjugation of oxazepam by human UDP-glucuronosyltransferases (UGTs): S-oxazepam is glucuronidated byUGT2B15, while R-oxazepam is glucuronidated by UGT2B7 and UGT1A9. Drug Metab Dispos. 2002; 30:1257–1265.
31. Court MH, Hao Q, Krishnaswamy S, Bekaii-Saab T, Al-Rohaimi A, von Moltke LL and Greenblatt DJ. UDP-glucuronosyltransferase (UGT) 2B15 pharmacogenetics: UGT2B15 D85Y genotype and gender are major determinants of oxazepam glucuronidation by human liver. J Pharmacol Exp Ther. 2004; 310: 656-665.
32. Stringer F, Scott G, Valbuena M, Kinley J, Nishihara M, Urquhart R. The effect of genetic polymorphisms in UGT2B15 on the pharmacokinetic profile of sipoglitazar, a novel anti-diabetic agent. Eur J Clin Pharmacol. 2013;69: 423-430.
33. He X, Hesse LM, Hazarika S, Masse G, Harmatz JS, Greenblatt DJ, Court MH. Evidence for oxazepam as an in vivo probe of UGT2B15: oxazepam clearance is reduced by UGT2B15 D85Y polymorphism but unaffected by UGT2B17 deletion. Br J Clin Pharmacol. 2009; 68: 721-730.
34. Turgeon D, Carrier JS, Hum DW, Bélanger A. Relative enzymatic activity, protein stability, and tissue distribution of human steroid-metabolizing UGT2B subfamily members. Endocrinology. 2001; 142: 778-787.
35. Levesque E, Turgeon D, Carrier JS. Isolation and characterization of the UGT2B28 cDNA encoding a novel human steroid conjugating UDP-glucuronosyltransferase. Biochemistry. 2001; 40: 3869-81.
36. Guillemette C, Harvey M, Bellemare J, Menard V. UGT genomic diversity: beyond gene duplication. Drug Metab Rev. 2010; 42: 24-44.
37. Levesque E, Beaulieu M, Green MD. Isolation and characterization of UGT2B15 (Y85): a UDP-glucuronosyltransferase encoded by a polymorphic gene. Pharmacogenetics. 1997; 7: 317-325.
38. Belanger A, Hum DW, Beaulieu M. Characterization and regulation of UDP-glucuronosyltransferases in steroid target tissues. J Steroid Biochem Mol Biol. 1998; 65: 301-310.
39. Ross R, Bernstein L, Judd H. Serum testosterone levels in healthy young black and white men. J Natl Cancer Inst. 1986; 76: 45-48.
40. Gann PH, Hennekens CH, Ma J, Longcope C, Stampfer MJ. Prospective study of sex hormone levels and risk of prostate cancer. J Natl Cancer Inst. 1996; 88: 1118-1126.

