INTRODUCTION
Hepatocellular carcinoma (HCC) is the most common type of primary liver cancer [1, 2]. HCC ranks the fourth in the frequency of cancer in men [3]. Patients diagnosed at the early stages of HCC can be successfully cured; however, there is deficiency of effective methods for patients diagnosed at the advanced stages [4]. Therefore, to reduce the economic and social burden, screening methods for detecting HCC at an earlier stage are highly demanded.
In recent years, circular RNAs (circRNAs), a class of endogenous RNAs have been found to widely exist in mammalian cells, have been receiving extensive attention by researchers [5–7]. By interacting with microRNAs (miRNAs) or other molecules, circRNAs regulate gene expression at the transcriptional or post-transcriptional level [8]. Due to a lack of 3′ and 5′ ends and resistance to RNases, circRNAs might be used as potential biomarkers and treatment targets for cancers [6, 9]. Recently, researchers have reported that circRNAs are involved in carcinogenesis of several types of cancers, such as gastric cancer, colorectal cancer, ovarian cancer and HCC [6, 9–12]. Harboring miRNA response elements (MREs), some circRNAs act as competing endogenous RNAs (ceRNAs) to play their biological roles, especially in cancer. For examples, circHIPK3 regulates cell growth by sponging multiple miRNAs [13]; circITCH plays an inhibitory role in esophageal squamous cell carcinoma (ESCC) by suppressing the Wnt/β-catenin pathway through interacting with multiple miRNAs [14]; Cdr1 antisense locus (CDR1as) acts as an oncogene partly through targeting miR-7 in HCC [15]. However, the function of most circRNAs in HCC and their clinical significance remain unknown.
Hsa_circ_0005986 is one of the upregulated circRNAs, with fold change 5.841, in our microarray screen detected by comparing HCC tissues with adjacent non-tumorous tissues (GEO No. 94508: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE94508). Its gene is located at chr1: 14057494–14068652, and its associated-gene symbol is PRDM2 (PR/SET Domain 2). Via Arraystar’s home-made miRNA target prediction software based on TargetScan and miRanda [16], we found that hsa_circ_0005986 has hsa-miR-129-5p seed matches. And through checking the miRTarBase (http://mirtarbase.mbc.nctu.edu.tw/php/detail.php?mirtid=MIRT005412), we found Notch1 is one of validated target genes of hsa-miR-129-5p by luciferase reporter assay and microarray [17]. Several reports have shown that Notch1 plays an important role in HCC carcinogenesis and metastasis [18–21]. Hence, it could conceivably be hypothesized that hsa_circ_0005986 and Notch1mRNA may act as a pair of ceRNAs that are linked by miR-129-5p.
In this study, we first investigated the hsa_circ_0005986 expression characteristics in HCC tissues and cell lines, and explored the association between the hsa_circ_0005986 expression level and clinicopathological characteristics of patients with HCC. We then investigated whether hsa_circ_0005986 can act as a ceRNA for Notch1 through miR-129-5p.We found that hsa_circ_0005986 could directly interact with miR-129-5p through luciferase reporter assay, and regulate NOTCH1 mRNA expression by acting as a sponge for miR-129-5p. Finally, we knocked down the hsa_circ_0005986 expression level by using small interfering RNA (siRNA) and found that hsa_circ_0005986 affected not only cell cycle, but also cell proliferation by regulating the G0/G1 to S phase transition.
RESULTS
General characteristics of the subjects
There was a male dominance in studied patients. More than three-quarters (87.7 %) of patients suffering from HCC were men. The average age of HCC patients was 57.2 years. AFP negative patients accounted for a bit less than half (46.8%; Table 1).
Table 1: The relationship between hsa_circ_0005986 expression levels (ΔCt) in cancer tissues and clinicopathological factors of patients with HCC
Characteristics |
No. of patients |
Percent of patients (%) |
Mean ± SD |
P value |
|---|---|---|---|---|
Age (years) |
0.631 |
|||
≥ 50 |
62 |
76.5 |
11.27 ± 1.40 |
|
< 50 |
19 |
23.5 |
11.15 ± 0.85 |
|
Gender |
0.936 |
|||
Male |
71 |
87.7 |
11.25 ± 1.32 |
|
Female |
10 |
12.3 |
11.16 ± 1.09 |
|
Family history |
0.001 |
|||
Positive |
33 |
40.7 |
11.84 ± 1.27 |
|
Negative |
48 |
59.3 |
10.83 ± 1.14 |
|
Diabetes Mellitus |
0.156 |
|||
Yes |
13 |
14.5 |
10.78 ± 1.55 |
|
No |
68 |
85.5 |
11.33 ± 1.23 |
|
Diameter (cm) |
< 0.001 |
|||
≥ 5 |
35 |
16 |
11.80 ± 1.12 |
|
< 5 |
46 |
84 |
10.82 ± 1.26 |
|
Differentiation |
0.228 |
|||
Well |
16 |
19.8 |
11.24 ± 1.14 |
|
Moderate |
52 |
64.2 |
11.38 ± 1.38 |
|
Poor |
13 |
16 |
10.69 ± 0.95 |
|
Microvascular invasion |
0.026 |
|||
Positive |
25 |
31.2 |
11.70 ± 1.32 |
|
Negative |
55 |
68.8 |
11.01 ± 1.23 |
|
Cirrhosis |
0.763 |
|||
Yes |
48 |
60 |
11.18 ± 1.33 |
|
No |
32 |
40 |
11.26 ± 1.18 |
|
BCLC stage |
< 0.001 |
|||
A |
32 |
39.5 |
10.55 ± 0.98 |
|
B + C |
49 |
60.5 |
11.69 ± 1.27 |
|
TNM stage |
0.166 |
|||
I |
41 |
50.6 |
11.03 ± 1.20 |
|
II |
18 |
22.2 |
11.72 ± 1.49 |
|
III + IV |
22 |
27.2 |
11.24 ± 1.29 |
|
AFP |
0.690 |
|||
> 20 |
42 |
53.2 |
11.20 ± 1.36 |
|
≤ 20 |
37 |
46.8 |
11.31 ± 1.24 |
|
AKP |
||||
> 95 |
43 |
53.1 |
11.36 ± 1.23 |
0.397 |
≤ 95 |
38 |
46.9 |
11.11 ± 1.36 |
|
GGT |
0.410 |
|||
> 50 |
55 |
54.1 |
11.15 ± 1.31 |
|
≤ 50 |
45 |
45.9 |
11.39 ± 1.27 |
Hsa_circ_0005986 was downregulated in HCC tissues and HCC cell lines
The expression of hsa_circ_0005986 in HCC tissues was significantly lower than that in matched non-tumorous tissues (P < 0.001) (Figure 1A), and the average relative expression level of hsa_circ_0005986 was downregulated by 2.94-fold. Furthermore, the downregulated ratio of hsa_circ_0005986 reached 80.2% (65/81).
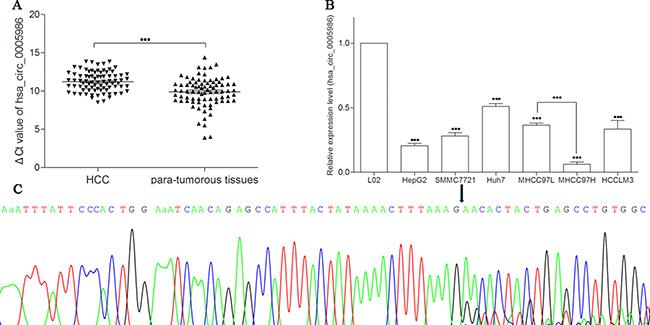
Figure 1: Detection of hsa_circ_0005986 expression by qRT-PCR. (A) Decreased expression of hsa_circ_0005986 between HCC tissues and para-tumorous tissues. Data are means ± SD. (B) Decreased expression of hsa_circ_0005986 between HCC cell lines and human normal hepatic cell line L02. Data are means ± SD. ***P < 0.001 (C) Sanger sequencing showed the back-splice junction sites.
To confirm the results of hsa_circ_0005986 in HCC tissues, we observed hsa_circ_0005986 levels in HCC cell lines (HepG2, SMMC7721, Huh7, MHCC97L, MHCC97H, and HCCLM3). The results showed that hsa_circ_0005986 levels in all detected HCC cell lines were significantly lower than those in normal hepatic cell line L02 (P < 0.001), and one interesting finding is the expression level in MHCC97L was significantly higher than that in MHCC97H (Figure 1B).
Sanger sequencing of the qRT-PCR products confirmed the back-splice junction sequence of hsa_circ_0005986 which was consistent with that from database (http://www.circbase.org/cgi-bin/listsearch.cgi) showed (Figure 1C).
Correlations between hsa_circ_0005986 expression level and clinicopathological parameters
The relationship between the expression level of hsa_circ_0005986 and clinicopathological factors of patients with HCC was analyzed. Low expression of hsa_circ_0005986 was significantly associated with chronic hepatitis B family history (P = 0.001), tumor diameters (P < 0.001), microvascular invasion (P = 0.026), Barcelona Clinic Liver Cancer staging system (BCLC) stage (P < 0.001) (Table 1).
Hsa_circ_0005986 and NOTCH1 mRNA are targeted by miR-129-5p
NOTCH1 is one of the validated targets of miR-129-5p (Figure 2C) [17]. Here, our study further verified the direct interaction between hsa_circ_0005986 and miR-129-5p via dual luciferase reporter assays (Figure 3).
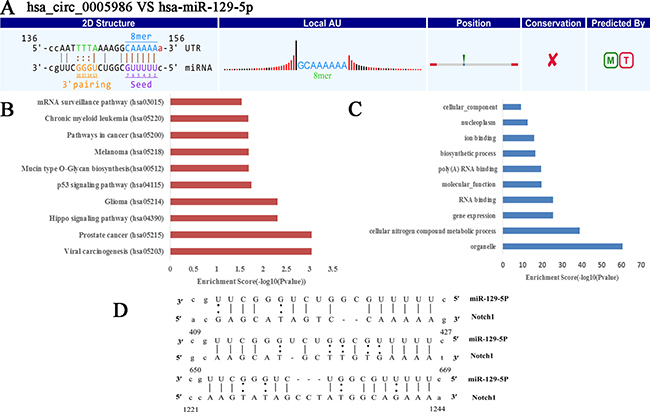
Figure 2: Prediction of hsa_circ_0005986-hsa-miR-129-5p-NOTCH1 interaction. (A) The interaction of hsa_circ_0005986-hsa-miR-129-5p was predicted based on TargetScan and miRanda. (B) The hsa_circ_0005986-miR-129-5p related KEGG pathway analysis. (C) The hsa_circ_0005986-miR-129-5p related GO analysis. (D) NOTCH1 is one of validated target genes of hsa-miR-129-5p (http://mirtarbase.mbc.nctu.edu.tw/php/detail.php?mirtid=MIRT005412). Detailed seed matches between hsa-miR-129-5p and Notch1mRNA.
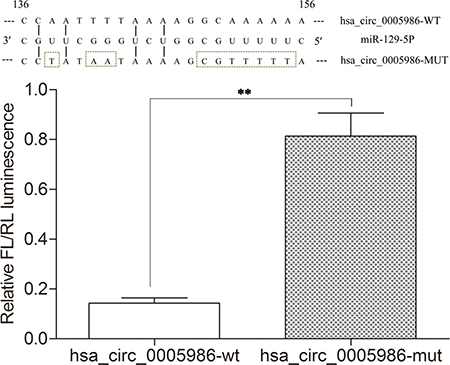
Figure 3: Results of dual luciferase reporter assays. Relative Firefly(FL)/Renilla(RL) luminescence mediated by luciferase plasmid harboring the wild-type or mutant hsa_circ_0005986. It indicates the direct interaction between hsa_circ_0005986 and miR-129-5p. Mean ± SD, n = 3; **P < 0.01.
To test whether hsa_circ_0005986 and Notch1 expression levels were affected by miR-129-5p, we increased the miR-129-5p level by transfection of its mimics into HepG2 and Huh7 cells. qRT-PCR analysis indicated that the transfection of miR-129-5p mimics decreased not only hsa_circ_0005986 levels but also Notch1mRNA levels in both HepG2 and Huh7 cells (Figure 4A).
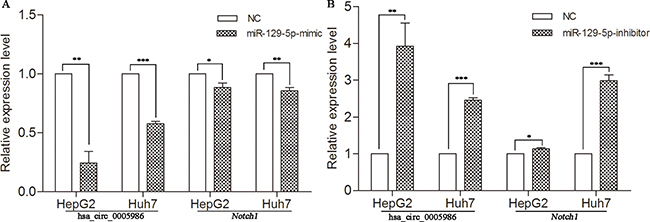
Figure 4: Expression of hsa_circ_0005986 and Notch1 mRNA in HepG2 and Huh7 cell lines transfected with miR-129-5p mimics (A) or inhibitors (B). Data are presented as mean ± SD, n = 3. NC, negative control. *P < 0.05, **P < 0.01, ***P < 0.001.
To decrease the miR-129-5p level, we transfected miR-129-5p inhibitors into HepG2 and Huh7 cells. Next, we utilized qRT-PCR analysis to reveal that miR-129-5p inhibitors increased both hsa_circ_0005986 and Notch1 abundance in two HCC cell lines (Figure 4B).
Effects of hsa_circ_0005986 downregulation on miR-129-5p and Notch1 expression
To verify that hsa_circ_0005986 and NOTCH1 are targets of miR-129-5p, we further sought to determine whether the downregulation of hsa_circ_0005986 would influence miR-129-5p and NOTCH1mRNA levels. If hsa_circ_0005986 functions as a ceRNA, its downregulation might free miR-129-5p. Therefore, we designed a siRNA against hsa_circ_0005986. The results showed that this not only effectively reduced the hsa_circ_0005986 level but also increased the miR-129-5p level in the HepG2 and Huh7 cells (Figure 5A, 5B). And at the same time, the level of NOTCH1mRNA decreased (Figure 5C).
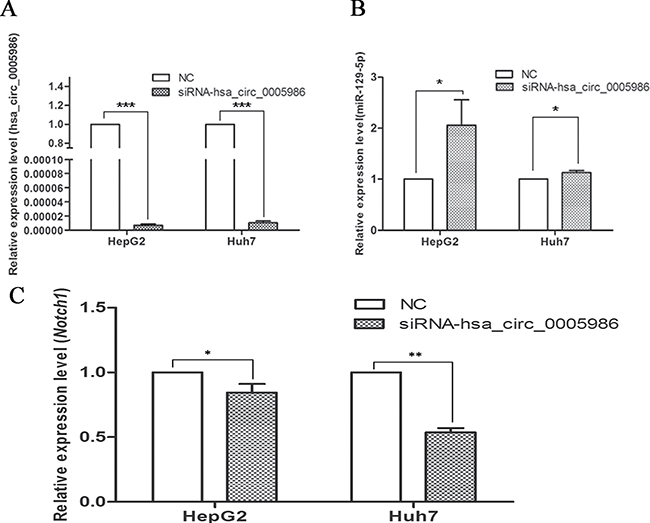
Figure 5: Expression of hsa_circ_0005986 (A), miR-129-5p (B) and NOTCH1 mRNA (C) in HepG2 and Huh7 cell lines after hsa_circ_0005986 knockdown. Data are presented as mean ± SD, n = 3. NC, negative control.*P < 0.05, **P < 0.01, ***P < 0.001.
Hsa_circ_0005986 regulates the cell cycle and cell proliferation
Because perturbations of the hsa_circ_0005986 level markedly affected NOTCH1mRNA expression, we decided to investigate the effects of interfering hsa_circ_0005986 on the cell proliferation and cell cycle.Hsa_circ_0005986 knockdown accelerated cell proliferation in HepG2 and Huh7 cell lines (Figure 6). Flow cytometry demonstrated that hsa_circ_0005986 downregulation promoted the G0/G1 to G2 phase transition in HepG2 and Huh7 cells (Figure 7).
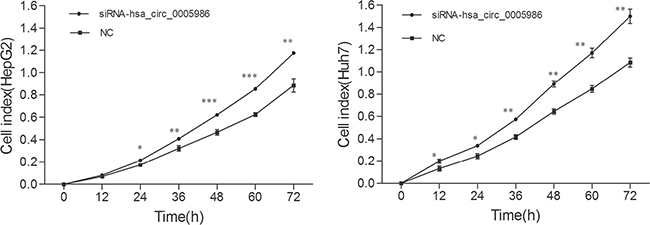
Figure 6: Growth curves of the HepG2 and Huh7 cell lines following by hsa_circ_0005986 knockdown. The proliferation assays were performed in E-Plate 96 using a Real-Time Cell Analyzer (RTCA). Data are presented as mean ± SD, n = 3. NC, negative control. *P < 0.05, **P < 0.01, ***P < 0.001.
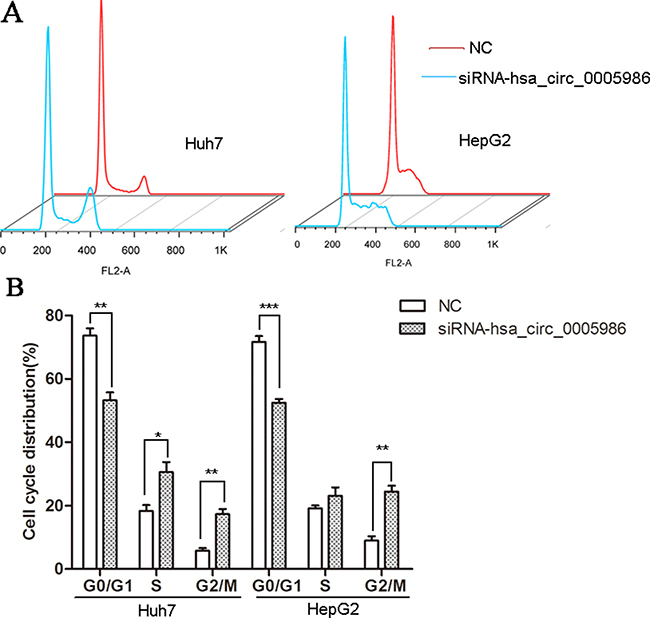
Figure 7: Cell cycle distributions in HepG2 and Huh7 cell lines following by hsa_circ_0005986 knockdown. (A) Representative original flow cytometry results. (B) Data are presented as mean ± SD, n = 3. NC, negative control. *P < 0.05, **P < 0.01, ***P < 0.001.
DISCUSSION
HCC with high morbidity and mortality is one of the prevalent cancers in the world [1, 22]. The carcinogenesis of HCC has been recognized as a complex process over many years; and its clinical diagnosis often happens at an advanced stage, generally missing the best time for radical treatments. So the early diagnosis is imperative for prognosis of HCC [23]. In our study, we found that the expression level of hsa_circ_0005986 in HCC was significantly lower than that of the para-tumorous tissues (Figure 1A). More importantly, for the first time, we found that the expression level of hsa_circ_0005986 was correlated with chronic hepatitis B family history, tumor diameters, microvascular invasion, and BCLC stage (Table 1). To further observe the diagnostic value of hsa_circ_0005986 in HCC, we compared its level in HCC cell lines, and found that hsa_circ_0005986 level in HCC cell lines (noninvasive HCC cell lines HepG2 and SMMC-7721, low invasiveness cell lines Huh7 and MHCC97L, and highly invasive cell lines MHCC97H and HCCLM3) were markedly lower than those in human normal hepatic cell line L02 (Figure 1B). It is interesting to note that the expression level of hsa_circ_0005986 in MHCC97L was higher than that in MHCC97H. MHCC97L and MHCC97H are derived from a subcutaneous xenograft of metastatic human HCC in nude mice, and show different metastatic power, especially in lung [24]. Nevertheless, hsa_circ_0005986 expression level is opposite to the result of our microarray study. The reason for this is that because of the limitation of microarray sample size and HCC is one of high heterogeneous tumors, the test results have certain false positives and false negatives, so it’s very important to expand the number of specimens, and use other methods (such as qRT-PCR) to verify.
Recently, circRNAs are known for being ubiquitous in cells [5, 25–26]. Thanks to improvements in high-throughput deep sequencing, circRNAs, once perceived as splicing errors, have been attracting more and more attention [7, 9, 27–28]. Sometimes, the expression levels of circRNAs can be more than ten times as high as those of their cognate linear isoforms [7, 29]. Because circRNAs are lack of the 5′cap and polyA tail, they can resist conventional RNA turnover and are more stable with a half-life more than 48 hours [25, 30, 31]. Intriguingly, scientists revealed that circRNAs can express in a complex tissue-specific, cell-type-specific or developmental-stage specific manner [9, 25, 32]. Taken together, we conclude that circRNAs could be incorporated into future clinical trials and become a kind of valuable biomarkers for precision medicine. For example, Chen et al. found that hsa_circ_0000190 harbored the diagnostic value in gastric cancer [33]. Zhang et al. disclosed that hsa_circ_101222 in combination with plasma protein endoglin strengthened the predictive value for pre-eclampsia [34].
As an enigmatic class of noncoding RNAs, circRNAs have been recently found to be associated with epigenetics and human disorders. Talhouarne et al. revealed that circular intronic RNAs can be passed on to offsprings in Xenopus oocytes and may be able to function in inheritance and epigenetics [35]. A circular isoform of antisense noncoding RNA in the INK4 locus (ANRIL) has been revealed to associate with the development of atherosclerosis [36]. CDR1 gene transcripts (also known as circular RNA sponge for miR-7, CiRS-7) abundantly express in brain, neuroblastoma lines, renal cell carcinoma lines etc., and influence many diseases including diabetes, prion disorders and cancers [37–38]. Du et al. demonstrated that the circular RNA circ-Foxo3 is associated with cell cycle progression through forming circ-Foxo3-p21-CDK2 ternary complex [39].
The study of ceRNA represents a new approach to examining complex post-transcriptional regulatory networks. ceRNA disturbances may be linked with many diseases. ceRNAs can not only be used to explain the biological mechanism of diseases, but can also be used in drug research [40]. As an indispensable member of ceRNA network, circRNAs have been paid more and more attention to by scientists [13–15, 41–47]. For example, cir-ITCH could act as sponge of miRNAs to enhance ITCH expression and thus suppress the activation of Wnt/β-catenin signaling in ESCC, lung cancer and colorectal cancer [14, 43–44]. In human HepG2 liver cells, Caiment et al. found that O6-methylguanine DNA methyltransferase (MGMT), one of DNA damage response enzymes, could interact with miR-181a-1-3p and a portion of circRNAs, and play roles in benzo[a]pyrene-induced carcinogenesis [11]. Through DIANA mirPath v.3, we identified has-miR-129-5p was closely related with a lot of cancer-related pathways, such as viral carcinogenesis, prostate cancer, hippo signaling pathway, p53 signaling pathway, pathways in cancer, PI3K-Akt signaling pathway, etc. (Figure 2B), and was involved with a lot of biological function processes such as organelle, cellular nitrogen compound, metabolic process, gene expression, etc. (Figure 2C). We focused on hsa_circ_0005986 and NOTCH1 in this study, because both of them are targets of miR-129-5p. Several reports have shown that miR-129-5p is linked to cancer, especially HCC [45, 46]. Based on these, by using luciferase reporter assay and qRT-PCR, we found that hsa_circ_0005986 could interact with miR-129-5p directly, that interference of the hsa_circ_0005986 level could influence miR-129-5p and Notch1 mRNA expression levels, and that hsa_circ_0005986 and Notch1 mRNA expression changed simultaneously. These all showed hsa_circ_0005986 could regulate NOTCH1 expression by acting as a sponge for miR-129-5p. Finally, we knocked down hsa_circ_0005986 expression by using siRNA and found that hsa_circ_0005986 affected cell proliferation by regulating the G0/G1 to S phase transition.
In summary, as one of circRNAs, hsa_circ_0005986 was lowly expressed in HCC. Its expression level was associated with chronic hepatitis B family history, tumor size, microvascular invasion and BCLC stage. One of the mechanisms underlying hsa_circ_0005986 influencing HCC carcinogenesis is that it regulated Notch1 expression through interacting with miR-129-5p. These indicated that hsa_circ_0005986 might not only be a potential biomarker for the diagnosis of HCC but might play important roles in carcinogenesis as well.
MATERIALS AND METHODS
Sample collection and ethics statement
Eighty-one HCC patients were enrolled in this study. All of them received surgeries at two cancer centers (Ningbo Lihuili Hospital and Ningbo No. 2 Hospital), from March 2013 to May 2016. Tissue samples were immediately soaked in RNAfixer Reagent (Bioteke, Beijing, China) and stored at −80°C till use. Histology was assessed independently by two experienced pathologists who were blinded to the clinical data.
This study was approved by the Human Research Ethics Committee from Ningbo University (IRB No.20100303). All patients signed the informed consent. Staging was determined by American Joint Committee on Cancer criteria and BCLC staging system. Patients with prior treatment of their tumor (such as trans-catheter arterial chemoembolization (TACE), ablation, radiotherapy, etc.) or with history of other solid tumors were excluded.
Cell culture
HCC cell lines (HepG2, SMCC7721, and Huh7) and human normal hepatic cell line L02 were purchased from the Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences (Shanghai, China). MHCC97L, MHCC97H and HCCLM3 were from the Liver Cancer Institute, Fudan University (Shanghai, China). All cell lines were cultured with RPMI 1640 Medium (Life Technologies, Carlsbad, CA, USA) containing 10% fetal bovine serum in a humidified atmosphere of 5% CO2.
RNA extraction
Total RNA was extracted from the frozen tissues and fresh cultured cells by using TRIzol (Invitrogen), based on the manufacturer’s instructions. After that, 10μl diethylpyrocarbonate (DEPC)-treated water was added to make the total RNA fully dissolved. Concentration and purity of total RNA samples were measured by using the SmartSpec Plus spectrophotometer (Bio-Rad, Hercules, CA, USA). The ratio of A260/A280 was used to indicate the purity of total RNA. Samples with value 1.8-2.0 were used for further experiments.
Reverse transcription
Following the manufacturer’s instructions, cDNA was generated by using the GoScript Rverse Transcription (RT) System (Promega, Madison, WI). In brief, 1 μl random primer, 1 μl oligo (dT)15 primer, 4 μl GoScript 5× reaction buffer, 2 μl MgCl2, 1 μl nucleotide mix, 0.5 μl recombinant RNasin ribonuclease, 1 μl GoScript reverse transcriptase, and 2 μg total RNA, were added in the system and then incubated at 42°C for 1h. RT reaction and no-template control were run at the same time.
Quantitative real-time PCR
Quantitative polymerase chain reaction (qPCR) was performed by using the GoTaq qPCR Master Mix (Promega, Madison, WI) on an Mx3005P real-time PCR System (Stratagene, La Jolla, CA, USA). A total of 5 μl cDNA product was used in a 25 μl reaction mixture which included 5.5μl DEPC-treated water, 1 μl upstream primer, 1 μl downstream primer, and 12.5 μl qPCR mix. Primers were designed with Primer3 (http://frodo.wi.mit.edu/), and synthesized by Sangon Biotech (Shanghai, China). Their sequences were as follows: for hsa_circ_0005986 (divergent primers) 5′-GAAACTGGCTGCGATATGTG-3′ (forward primer) and 5′-CACAGGCTCAGTAGTGTTCTTTAAA-3′ (reverse primer); for Notch1mRNA 5′-GAAACTGGCTG CGATATGTG-3′ (forward primer) and 5′-CACAGGCT CAGTAGTGTTCTTTAAA-3′ (reverse primer); for hsa-miR-129-5p 5′-CAACCTTACCTTTTTGCGGTC-3′ (forward primer) and 5′-TATGCTTGTTCTCGTCTCTG TGTC-3′ (reverse primer); for glyceraldehyde 3-phosphate dehydrogenase (GAPDH, reference gene1), 5′-TCGACAG TCAGCCGCATCTTCTTT-3′ (forward primer) and 5′-AC CAAATCCGTTGACTCCGACCTT-3′ (reverse primer); and for U6snRNA (reference gene2) 5′-ATTGGAA CGATACAGAGAAGATT-3′ (forward primer) and 5′-GG AACGCTTCACGAATTTG-3′ (reverse primer). Real-time PCR was done in triplicate. The amplification specific was confirmed by melting curve analysis. The data from qRT-PCR was analyzed by the ΔCt method. The ΔCt value was determined by subtracting the Ct value of reference gene from the Ct value of target gene. Larger ΔCt value indicates lower expression. Relative expression was calculated with the 2−ΔΔCt method. All results are expressed as the means ± SD of three independent experiments. All of assays were performed in a blinded fashion to clinical data.
Cloning and sequencing of qRT-PCR products
Based on the manufacturer’s instructions, the qRT-PCR product of hsa_circ_0005986 was first purified by using a UNIQ-10 PCR Product Purification Kit, and then cloned into the pUCm-T vector (Sangon Biotech, Shanghai, China). Finally, DNA sequencing was performed by Sangon Biotech Co., Ltd (Figure 1C).
Liver function and serological tumor marker analysis
Liver function including albumin (ALB), aspartate transaminase (AST), alanine aminotransferase (ALT), alkaline phosphatase (AKP), Gamma Glutamyl Transferase (GGT), and total bilirubin was measured by Olympus AU 2700 automatic biochemical analyzer with original kits (Olympus, Tokyo, Japan). Alpha-fetoprotein (AFP) was measured with an Elecsys 2010 machine (Roche Diagnostics, Basel, Switzerland).
Transient transfection
For the transfection of the miR-129-5p mimics, inhibitors and small interfering RNAs (siRNAs), HepG2 and Huh-7 cells (2 × 105) were seeded in 6-well plates. The following day, they were transfected with 120nM miR-129-5p mimics, inhibitors or siRNA using Lipofectamine 2000 Reagent (Life Technologies). The sequence of miR-129-5p mimic was 5′-CUUUUUGCGGUCUGGGCUUGC-3′ (sense) and 5′-AAGCCCAGACCGCAAAAAGUU-3′(antisense). The sequence of miR-129-5p inhibitor was GCAAGCCCAGACCGCAAAAAG. The sequences of three siRNA for hsa_circ_0005986 were 5′ - AACUUUAA AGAACACUACUGAGC-3′ (sense) and 5′ -UUAACUCG AAGCUGUCCUGTT-3′ (antisense); 5′ - ACUUUAAAGA ACACUACUGAG-3′ (sense) and 5′ - CUCAGUAGUGUU CUUUAAAGU -3′ (antisense); 5′ - AAACUUUAAAGA ACACUACUG-3′ (sense) and 5′ - CAGUAGUGUUCUUU AAAGUUU-3′ (antisense), and we selected 5′ - AACUUU AAAGAACACUACUGAGC-3′ (sense) and 5′ -UUAACU CGAAGCUGUCCUGTT-3′ (antisense) as the best one. The sequences of the negative control siRNAs were 5′ - AACUUUAAAGAACACUACUGAGC-3′ (sense) and 5′ - GCUCAGUAGUGUUCUUUAAAGUU-3′ (antisense). These sequences were synthesized by GenePharma Co., Ltd. (Shanghai, China).
Dual luciferase reporter assay
MiR-129-5p expression plasmid (pmirGLO) was purchased from GenePharma Co., Ltd. (Shanghai, China). The wild-type and mutant DNA sequences were synthesized by GenePharma Co., Ltd. and cloned into pmirGLO Firefly luciferase plasmid (GenePharma). The miR-129-5p MRE of hsa_circ_0005986 wild-type and mutant sequence showed in Figure 3 following Arraystar’s home-made miRNA target prediction (Figure 2A). HepG2 of 80% confluence in 24-well plates was transfected using Lipofectamine 2000 Reagent (Life Technologies) according to the manufacturer’s protocol. Firefly luciferase (FL) plasmid and miR-129-5p expression plasmid were cotransfected with pRL-TK Renilla luciferase (RL) vector (Promega, Madison, WI) for normalization. After 48 h, luciferase activity was measured using Dual-Glo Luciferase Assay System (Promega). For comparison, the FL activity was normalized with RL activity. All experiments were performed in triplicate.
Cell cycle analysis
The HepG2 and Huh7 cells were washed in PBS and fixed in 75% ice-cold ethanol at −20 °C overnight. After rehydrating with ice-cold PBS, the cells were stained with PI/RNase Staining Buffer (BD Biosciences, San Jose, CA,) and analyzed by flow cytometry on a FACSCalibur Flow Cytometer (BD Biosciences) using CellQuest Pro software.
Cell proliferation assays
The proliferation assays were performed in E-Plate 96 using a Real-Time Cell Analyzer (RTCA) (ACEA Biosciences, San Diego, CA) according to the manufacturer’s protocol.
Prediction for hsa_circ_0005986-miR-129-5p related pathways and GO analysis
The hsa_circ_0005986-miR-129-5p related pathway and gene ontology (GO) analysis were carried out based on DIANA-miRPath V.3 [48]. All of miRNA gene targets are experimentally validated (derived from TarBase7.0) [49]. P < 0.05 was used as the criterion for statistical significance.
Statistical analysis
Statistical analysis was performed according to the Statistical Product and Service Solutions (SPSS) 16.0 software package (IBM, Chicago, IL). All the graphs were plotted using GraphPad Prism 6.0 (GraphPad Software, La Jolla, CA). The data are presented as the means ± SD. Paired t test, independent-sample t test, one way analysis of variance (ANOVA), and two-tailed Student’s t-tests were flexibly used according to actual conditions. P value of 0.05 or less was considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
This study was supported by the Foundation for Hepatitis Prevention and Control Project of China (No. TQGB20150219), Zhejiang Medical Scientific Research Foundation (No.2017KY140), Regional Medical Center Project of Zhejiang Province (No. 201498), and Natural Science Foundation of Ningbo (No. 2015A10236). We would like to thank all our colleagues for their help in collecting specimens.
CONFLICTS OF INTEREST
None.
REFERENCES
1. Sun HZ, Song YL, Wang XY. Effects of different anesthetic methods on cellular immune and neuroendocrine functions in patients with hepatocellular carcinoma before and after surgery. J Clin Lab Anal. 2016; 30:1175–1182.
2. Bu Y, Jia QA, Ren ZG, Xue TC, Zhang QB, Zhang KZ, Zhang QB, You Y, Tian H, Qin LX, Tang ZY. The herbal compound Songyou Yin (SYY) inhibits hepatocellular carcinoma growth and improves survival in models of chronic fibrosis via paracrine inhibition of activated hepatic stellate cells. Oncotarget. 2015; 6:40068–80. doi: 10.18632/oncotarget.5313.
3. Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016; 66:115–132.
4. Li J, Li J, Bao Y, Pan K, Lin X, Liu X, Wang H, Xu Y, Luo X, Li H, Duan C. Low frequency of peripheral lymphocyte in chronic hepatitis B patients predicts poor progression to hepatocellular carcinoma. J Clin Lab Anal. 2016; 30: 208–215.
5. Li P, Chen S, Chen H, Mo X, Li T, Shao Y, Xiao B, Guo J. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015; 444:132–136.
6. Shao Y, Li J, Lu R, Li T, Yang Y, Xiao B, Guo J. Global circular RNA expression profile of human gastric cancer and its clinical significances. Cancer Med. 2017; 6:1173–1180. doi: 10.1002/cam4.1055.
7. Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013; 495:333–338.
8. Sen R, Ghosal S, Das S, Balti S, Chakrabarti J. Competing endogenous RNA: the key to posttranscriptional regulation. ScientificWorldJournal. 2014; 2014:896206.
9. Bachmayr-Heyda A, Reiner AT, Auer K, Sukhbaatar N, Aust S, Bachleitner-Hofmann T, Mesteri I, Grunt TW, Zeillinger R, Pils D. Correlation of circular RNA abundance with proliferation—exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015; 5:8057.
10. Yao T, Chen Q, Fu L, Guo J. Circular RNAs: Biogenesis, properties, roles, and their relationships with liver diseases. Hepatol Res. 2017; 47:497–504. doi: 10.1111/hepr.12871.
11. Caiment F, Gaj S, Claessen S, Kleinjans J. High-throughput data integration of RNA-miRNA-circRNA reveals novel insights into mechanisms of benzo[a]pyrene-induced carcinogenicity. Nucleic Acids Res. 2015; 43:2525–34.
12. Li P, Chen H, Chen S, Mo X, Li T, Xiao B, Yu R, Guo J. Circular RNA 0000096 affects cell growth and migration in gastric cancer. Br J Cancer. 2017; 116:626–633.
13. Zheng Q, Bao C, Guo W, Li S, Chen J, Chen B, Luo Y, Lyu D, Li Y, Shi G, Liang L, Gu J, He X, Huang S. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016; 7:11215.
14. Li F, Zhang L, Li W, Deng J, Zheng J, An M, Lu J, Zhou Y. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/β-catenin pathway. Oncotarget. 2015; 6:6001–13. doi: 10.18632/oncotarget.3469.
15. Yu L, Gong X, Sun L, Zhou Q, Lu B, Zhu L. The Circular RNA Cdr1as Act as an Oncogene in Hepatocellular Carcinoma through Targeting miR-7 Expression. PLoS One. 2016; 11:e0158347.
16. Li H, Hao X, Wang H, Liu Z, He Y, Pu M, Zhang H, Yu H, Duan J, Qu S. Circular RNA Expression Profile of Pancreatic Ductal Adenocarcinoma Revealed by Microarray. Cell Physiol Biochem. 2016; 40:1334–1344.
17. Fukuda Y, Kawasaki H, Taira K. Exploration of human miRNA target genes in neuronal differentiation. Nucleic Acids Symp Ser (Oxf). 2005; 341–342.
18. Jia M, Jiang L, Wang YD, Huang JZ, Yu M, Xue HZ. lincRNA-p21 inhibits invasion and metastasis of hepatocellular carcinoma through Notch signaling-induced epithelial-mesenchymal transition. Hepatol Res. 2016; 46:1137–1144.
19. Lu J, Xia Y, Chen K, Zheng Y, Wang J, Lu W, Yin Q, Wang F, Zhou Y, Guo C. Oncogenic role of the Notch pathway in primary liver cancer. Oncol Lett. 2016; 12:3–10.
20. Giovannini C, Minguzzi M, Genovese F, Baglioni M, Gualandi A, Ravaioli M, Milazzo M, Tavolari S, Bolondi L, Gramantieri L. Molecular and proteomic insight into Notch1 characterization in hepatocellular carcinoma. Oncotarget. 2016; 7:39609–26. doi: 10.18632/oncotarget.9203.
21. Sokolowski KM, Balamurugan M, Kunnimalaiyaan S, Wilson J, Gamblin TC, Kunnimalaiyaan M. Role of Akt inhibition on Notch1 expression in hepatocellular carcinoma: potential role for dual targeted therapy. Am J Surg. 2016; 211:755–760.
22. Yao M, Zhao J, Lu F. Alpha-fetoprotein still is a valuable diagnostic and prognosis predicting biomarker in hepatitis B virus infection-related hepatocellular carcinoma. Oncotarget. 2016; 7:3702–08. doi: 10.18632/oncotarget.6913.
23. Forner A. Hepatocellular carcinoma surveillance with miRNAs. Lancet Oncol. 2015; 16:743–745.
24. Zhao Y, Jia HL, Zhou HJ, Dong QZ, Fu LY, Yan ZW, Sun J, Ren N, Ye QH, Qin LX. Identification of metastasis-related microRNAs of hepatocellular carcinoma in hepatocellular carcinoma cell lines by quantitative real time PCR. Zhonghua Gan Zang Bing Za Zhi. 2009; 17:526–530.
25. Ebbesen KK, Kjems J, Hansen TB. Circular RNAs: Identification, biogenesis and function. Biochim Biophys Acta. 2016; 1859:163–168.
26. Glažar P, Papavasileiou P, Rajewsky N. circBase: a database for circular RNAs. RNA. 2014; 20:1666–1670.
27. Vicens Q, Westhof E. Biogenesis of Circular RNAs. Cell. 2014; 159:13–14.
28. Ebbesen KK, Hansen TB, Kjems J. Insights into circular RNA biology. RNA Biol. 2016; 1–11.
29. Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012; 7:e30733.
30. Guo JU, Agarwal V, Guo H, Bartel DP. Expanded identification and characterization of mammalian circular RNAs. Genome Biol. 2014; 15:409.
31. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015; 12:381–388.
32. Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO. Cell-type specific features of circular RNA expression. PLoS Genet. 2013; 9:e1003777.
33. Chen S, Li T, Zhao Q, Xiao B, Guo J. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin Chim Acta. 2017; 466:167–171.
34. Zhang YG, Yang HL, Long Y, Li WL. Circular RNA in blood corpuscles combined with plasma protein factor for early prediction of pre-eclampsia. BJOG. 2016; 123:2113–2118.
35. Talhouarne GJ, Gall JG. Lariat intronic RNAs in the cytoplasm of Xenopus tropicalis oocytes. RNA. 2014; 20:1476–1487.
36. Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z, Sharpless NE. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010; 6:e1001233.
37. Hansen TB, Kjems J, Damgaard CK. Circular RNA and miR-7 in cancer. Cancer Res. 2013; 73:5609–5612.
38. Xu H, Guo S, Li W, Yu P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep. 2015; 5:12453.
39. Du WW, Yang W, Liu E, Yang Z, Dhaliwal P, Yang BB. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016; 44:2846–2858.
40. Xia T, Chen S, Jiang Z, Shao Y, Jiang X, Li P, Xiao B, Guo J. Long noncoding RNA FER1L4 suppresses cancer cell growth by acting as a competing endogenous RNA and regulating PTEN expression. Sci Rep. 2015; 5:13445.
41. Zhao ZJ, Shen J. Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol. 2015; 1–8.
42. Li T, Mo X, Fu L, Xiao B, Guo J. Molecular mechanisms of long noncoding RNAs on gastric cancer. Oncotarget. 2016; 7:8601–12. doi: 10.18632/oncotarget.6926.
43. Wan L, Zhang L, Fan K, Cheng ZX, Sun QC, Wang JJ. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/β-Catenin Pathway. Biomed Res Int. 2016; 2016:1579490.
44. Huang G, Zhu H, Shi Y, Wu W, Cai H, Chen X. cir-ITCH plays an inhibitory role in colorectal cancer by regulating the Wnt/β-catenin pathway. PLoS One. 2015; 10:e0131225.
45. Han H, Li W, Shen H, Zhang J, Zhu Y, Li Y. microRNA-129-5p, a c-Myc negative target, affects hepatocellular carcinoma progression by blocking the Warburg effect. J Mol Cell Biol. 2016 Mar 21. pii: mjw010. [Epub ahead of print].
46. Ma N, Chen F, Shen SL, Chen W, Chen LZ, Su Q, Zhang LJ, Bi J, Zeng WT, Li W, Huang XH, Wang Q. MicroRNA-129-5p inhibits hepatocellular carcinoma cell metastasis and invasion via targeting ETS1. Biochem Biophys Res Commun. 2015; 461:618–623.
47. Barrett SP, Salzman J. Circular RNAs: analysis, expression and potential functions. Development. 2016; 143:1838–1847.
48. Vlachos IS, Zagganas K, Paraskevopoulou MD, Georgakilas G, Karagkouni D, Vergoulis T, Dalamagas T, Hatzigeorgiou AG. DIANA-miRPath v3.0: deciphering microRNA function with experimental support. Nucleic Acids Res. 2015; 43:W460–466.
49. Vlachos IS, Paraskevopoulou MD, Karagkouni D, Georgakilas G, Vergoulis T, Kanellos I, Anastasopoulos IL, Maniou S, Karathanou K, Kalfakakou D, Fevgas A, Dalamagas T, Hatzigeorgiou AG. DIANA-TarBase v7.0: indexing more than half a million experimentally supported miRNA:mRNA interactions. Nucleic Acids Res. 2015; 43:D153–159.

