Introduction
Neurodegenerative diseases are a group of disorders with progressive neuronal loss in particular regions of brain, including Alzheimer’s disease (AD), Parkinson’s disease (PD), progressive supranuclear palsy (PSP), corticobasal degeneration (CBD), frontotemporal dementia (FTD), amyotrophic lateral sclerosis (ALS) and many others. The etiology of neurodegenerative diseases is complicated and multifactorial, mainly including genetic variants and environmental exposure. However, epidemiologic evidence for the association between the environmental exposure and neurodegenerative diseases is not conclusive [1]. Genetic variant is a crucial factor in etiology and pathogenic mechanisms of neurodegenerative diseases [2]. Hundreds of genetic variants have been confirmed significantly associated with neurodegenerative diseases, but the majority of these genes do not overlap across diseases [2]. Only several susceptibility genes relate to diverse neurodegenerative diseases. Among them, microtubule-associated protein tau (MAPT) gene is compelling [3]. Mutations in MAPT have been reported to participate in AD [4-6], PD [7-9], PSP [10, 11], CBD [12, 13], FTD [14, 15] and ALS [16, 17].
MAPT is located on chromosome 17q21.3, and encodes six tau isoforms ranging from 352 to 441 amino acids in length expressed in neurons [18]. To date, nearly 60 mutations in MAPT have been proved pathogenic to neurodegenerative diseases [3]. Most MAPT variants probably cause abnormal structure and function of hyperphosphorylated, insoluble and aggregated tau. These pathological changes are crucial for the pathogenesis of tau-related neurodegenerative diseases, which are called tauopathies [19]. As known to all, there are two main haplotypes in MAPT, H1 and H2. Several single-nucleotide polymorphisms (SNPs) throughout MAPT gene are in complete linkage disequilibrium (LD) and largely tags the H1 and H2 haplotypes, called haplotype tagging SNPs (htSNPs) [20]. Six htSNPs were identified, including 5 htSNPs that represent the intra-H1 variation (rs1467967, rs242557, rs3785883, rs2471738, and rs7521) and a 238-bp insertion/deletion polymorphism within intron 9 (del-In9) (Haplotype H1c: rs1467967 = A, rs242557 = A, rs3785883 = G, rs2471738 = T, del-In9 = ins and rs7521 = G) [20, 21]. The insertion of the del-In9 tags H1 haplotype and the deletion tags H2 haplotype. So, the available data of del-In9 were included into the H2 group to be further analyzed. Two htSNPs were the promoter polymorphisms (rs1467967 and rs242557) and three were intronic (rs3785883, rs2471738, and rs7521) [22]. Additionally, the Q7R (rs62063857) polymorphism of the Saitohin gene (STH), nested in intron 9 of the MAPT, is in complete LD with the extended H1/H2 haplotype [23, 24]. The Q and R alleles were in LD with H1 and H2 haplotypes, respectively [23, 24]. Unfortunately, some studies showed obvious association of MAPT variants with neurodegenerative diseases, while others showed none. The results of the related studies were inconsistent.
Therefore, we performed a meta-analysis by pooling the whole related data from previously studies to reach a more precise estimate of the relationship between variants in MAPT and their risk on neurodegenerative diseases.
Results
5357 studies were retrieved through the literature search, and 1226 papers of these were identified as potentially meeting the inclusion criteria after reviewing the titles and abstracts. Besides, we included 148 studies by manual searching. After further reviewing the full text, 82 studies were identified eligible (Figure 1). The results of meta-analysis on the association between variants in MAPT and neurodegenerative diseases were shown in Figure 2-5.
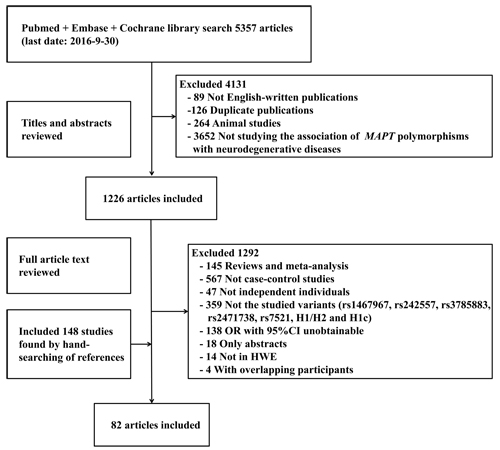
Figure 1: Flow diagram of the study selection process. Abbreviations: OR, odds ratio; CI, confidence interval; HWE, Hardy-Weinberg equilibrium.
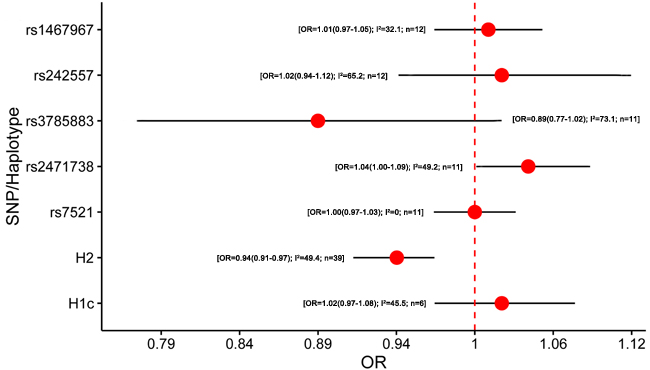
Figure 2: Results of the meta-analysis for five htSNPs, H2 haplotype and H1c subhaplotype in Alzheimer’s disease. Abbreviations: OR, odds ratio; SNP, single nucleotide polymorphism; I2, the heterogeneity calculated by the Cochran Q test; n, the number of included studies.
Alzheimer’s disease
A total of 37 studies were included in the meta-analysis of MAPT polymorphism in AD [4-6, 20, 25-57] (Supplementary Table 2). Among them, 34 studies were conducted in Caucasian populations, and 3 studies were performed in Asian populations. Two studies did not state the diagnostic criteria of AD, and 15 studies did not state whether the polymorphisms were in Hardy-Weinberg equilibrium (HWE). AD cases were diagnosed mainly using the criteria of the National Institute of Neurological and Communication Disorders and Stroke-Alzheimer Disease and Related Disorders Association (NINCDS-ADRDA) [58], or confirmed by autopsy. Notably, our meta-analysis showed that the minor allele (T allele) within rs2471738 was mildly associated with an increased risk of AD (odds ratio (OR) = 1.04, 95% confidence interval (CI) = 1.00 - 1.09) and H2 haplotype might be a protective factor for AD (OR = 0.94, 95% CI = 0.91 - 0.97) (Figure 2; Supplementary Table 8). No associations were found in rs1467967, rs242557, rs3785883, rs7521 and H1c for AD (OR = 1.01, 95% CI = 0.97 -1.05; OR = 1.02, 95% CI = 0.94 -1.12; OR = 0.89, 95% CI = 0.77 -1.02; OR = 1.00, 95% CI = 0.97 -1.03; OR = 1.02, 95% CI = 0.97 -1.08; respectively) (Figure 2; Supplementary Table 8). Due to the ethnic heterogeneity, we performed the subgroup analyses by ethnicity. Interestingly, the minor allele (A allele) within rs3785883 was a protective factor for AD risk in Caucasian (OR = 0.87, 95% CI = 0.76 -1.00) (Supplementary Table 9). This result was calculated by the random-effects model because the heterogeneity existed in the included studies of rs3785883 in Caucasian (I2 = 75.2). The heterogeneity was reduced to 56.2% and the pooled effect was changed into negative (OR = 0.94, 95% CI = 0.84 - 1.05) when one single study was excluded [53]. We speculated that the influence of this study maybe result from the origin of the Spanish which was different from other Caucasian populations. Publication bias was assessed by Egger test, and only the subgroup of rs3785883 in Caucasian that showed little publication bias (P Egger = 0.0499) (Supplementary Table 9). Using the trim and fill method to account for the bias did not influence the summary estimate for rs3785883 in Caucasian [59].
Parkinson’s disease
A total of 32 studies were included in the meta-analysis of MAPT polymorphism in PD [7, 8, 36, 39, 43, 53, 60-85] (Supplementary Table 3). Among them, 27 studies were conducted in Caucasian populations, 3 studies were performed in Asian populations, and one study was conducted in African populations. Besides, one study included both Caucasian and Asian populations [66]. Six studies lacked diagnostic criteria for PD. Seven studies did not state whether the polymorphisms were within the range of HWE. Most included studies were based on the UK Parkinson’s Disease Society Brain Bank clinical diagnostic criteria (UKPDBB) or the extended [86-88], or autopsy. Remarkable, our meta-analysis found that H2 haplotype might be a protective factor for PD (OR = 0.76, 95% CI = 0.74 - 0.79) (Figure 3; Supplementary Table 11). Then we performed the subgroup analyses by ethnicity. Similarly, H2 haplotype showed protective effect on PD in Caucasian subgroup (OR = 0.76, 95% CI = 0.74 - 0.79) (Supplementary Table 12). Additionally, the minor allele within rs242557 (A allele) and rs7521 (A allele) were mildly associated with an increased risk of PD in Caucasians (OR = 1.06, 95% CI = 1.01 - 1.12; OR = 1.11, 95% CI = 1.00 - 1.23, respectively) (Supplementary Table 12). No heterogeneity was existed in each subgroup. The data above were calculated by the fixed-effects model. In addition, based on the results from the Egger test, there was no publication bias.
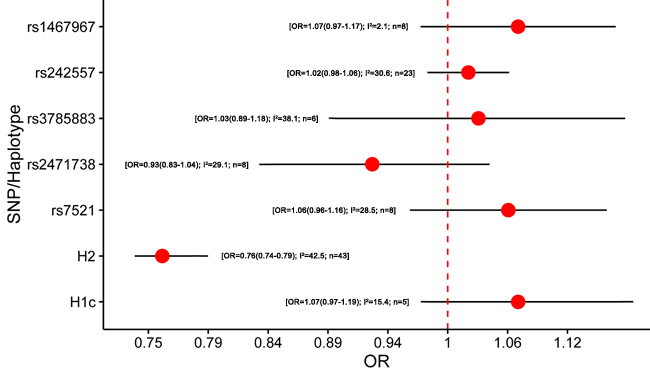
Figure 3: Results of the meta-analysis for five htSNPs, H2 haplotype and H1c subhaplotype in Parkinson’s disease. Abbreviations: OR, odds ratio; SNP, single nucleotide polymorphism; I2, the heterogeneity calculated by the Cochran Q test; n, the number of included studies.
Progressive supranuclear palsy
Twelves studies were included in the meta-analysis of MAPT polymorphism in PSP [11, 12, 15, 21, 23, 26, 45, 85, 89-92] (Supplementary Table 4). All of the studies were according to standard diagnostic criteria [93, 94]. Five studies did not perform the HWE test. All studies were performed in Caucasians. Notably, our results showed that the minor allele within rs242557 (A allele) and rs2471738 (T allele) may be risk factors for PSP (OR = 1. 96, 95% CI = 1. 71 - 2.25; OR = 1. 85, 95% CI = 1. 48 - 2.31; respectively) (Figure 4; Supplementary Table 14). H2 haplotype showed significantly protective effect on PSP risk (OR = 0.20, 95% CI = 0.18 - 0.23). Additionally, H1c significantly increased the risk of PSP (OR = 2.33, 95% CI = 1.28 - 4.25). The pooled ORs and 95%CIs of three subgroups were calculated by the random-effects model because the heterogeneity evidently existed. For rs242557 subgroup sensitivity analysis, the heterogeneity was reduced to 41.1% and the pooled effect was mildly increased (OR from 1.96 to 2.09) when the stage 2 of the study of Hoglinger et al. was excluded [11]. The majority of cases in the stage 2 of this study were clinically diagnosed and this diagnostic misclassification rate as 12% [11]. Hence, we inferred that the highly diagnostic misclassification rate might affect the result. In addition, for H1c subgroup sensitivity analysis, the heterogeneity was reduced to 28.4% and the pooled effect was mildly increased (OR from 2.33 to 3.17) when one single study conducted in the Spanish was excluded [12]. However, the heterogeneity cannot be further explained for the rs3785883 subgroup, maybe due to too few studies included. In conclusion, heterogeneity and sensitivity analyses showed that the exclusion of the related study did not influence the statistical significances of factors. In addition, no publication bias was found.
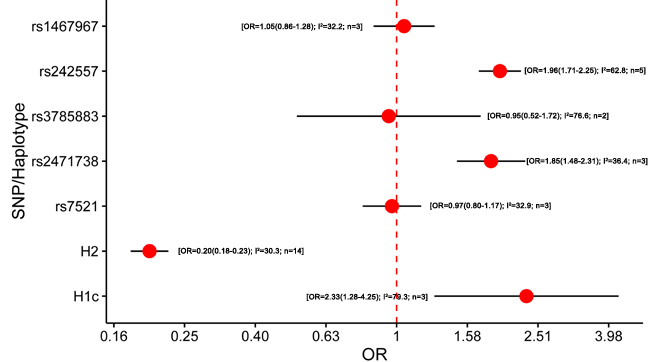
Figure 4: Results of the meta-analysis for five htSNPs, H2 haplotype and H1c subhaplotype in progressive supranuclear palsy. Abbreviations: OR, odds ratio; SNP, single nucleotide polymorphism; I2, the heterogeneity calculated by the Cochran Q test; n, the number of included studies.
Corticobasal degeneration
Six studies were included in the meta-analysis of MAPT polymorphism in CBD [12, 13, 15, 21, 85, 95] (Supplementary Table 5). Only one study lacked the diagnostic criteria for CBD [15], while two studies did not analyze the HWE [13, 85]. All of the included studies were conducted in Caucasian populations. Remarkably, our meta-analysis showed that the minor allele within rs242557 (A allele) and rs2471738 (T allele) may increase the risk of CBD (OR = 2.51, 95%CI = 1. 66 -3.78; OR = 2.07, 95%CI = 1. 32 -3.23; respectively). Notably, H2 haplotype showed protective effect on CBD (OR = 0.30, 95% CI = 0.23 - 0.41), whereas H1c may be risk factor for CBD (OR = 2.57, 95%CI = 1.51 - 4.40) (Figure 5; Supplementary Table 15). The heterogeneity among included studies only existed in rs7521 subgroup. For rs7521 subgroup sensitivity analysis, the heterogeneity was reduced to 0% when one single study which did not report the HWE test was excluded [13]. However, heterogeneity and sensitivity analyses showed that the exclusion of the related study did not influence the statistical significances.
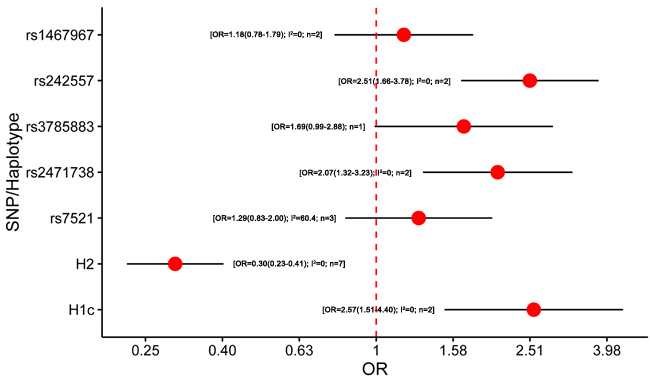
Figure 5: Results of the meta-analysis for five htSNPs, H2 haplotype and H1c subhaplotype in corticobasal degeneration. Abbreviations: OR, odds ratio; SNP, single nucleotide polymorphism; I2, the heterogeneity calculated by the Cochran Q test; n, the number of included studies.
Frontotemporal dementia
Seven studies were included in the meta-analysis of MAPT polymorphism in FTD [14-16, 20, 35, 43, 96, 97] (Supplementary Table 6). All studies were conducted in Caucasians. All of the studies were according to the Lund-Manchester criteria or the extended [98, 99]. Three studies did not perform the HWE test [14, 16, 20]. All studies included were only explored the association between MAPT haplotypes and FTD. However, the results showed that H2 haplotype was not associated with FTD risk (OR = 1.02, 95% CI = 0.78 - 1.32) (Figure 6; Supplementary Table 16). Mild heterogeneity was existed (I2 = 55.5) and reduced to 42.5% when the three studies which did not report the HWE test was excluded. But the exclusion of the related study did not influence the statistical significances.
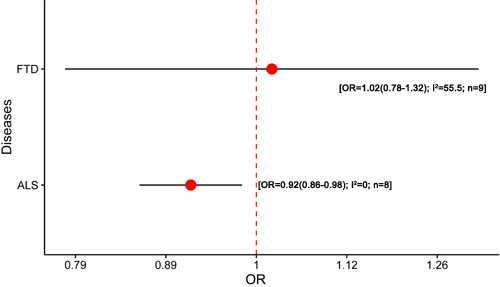
Figure 6: Results of the meta-analysis for H2 haplotype in FTD and ALS. Abbreviations: OR, odds ratio; FTD, frontotemporal dementia; ALS, amyotrophic lateral sclerosis; I2, the heterogeneity calculated by the Cochran Q test; n, the number of included studies.
Amyotrophic lateral sclerosis
Two studies were included in the meta-analysis of MAPT polymorphism in ALS [16, 17] (Supplementary Table 7). All studies were conducted in Caucasians and all cases were diagnosed according to El Escarol criteria for ALS. One study had not performed the HWE test [16]. Like FTD, the studies included were only restricted to the association between MAPT haplotypes and ALS. Notably, the H2 haplotype showed mildly protective effect on ALS risk (OR = 0.92, 95% CI = 0.86 - 0.98, I2 = 0) (Figure 6; Supplementary Table 17). No heterogeneity was found (I2 = 0).
Discussion
Our findings provide confirmatory evidence that multiple variants in distinct regions in MAPT are associated with different neurodegenerative diseases. The minor alleles within rs1467967 and rs242557 are not all the same in different populations (Supplementary Table 1). For uniformity, we all choose the same allele (the minor allele in Caucasians) in subgroups divided by populations for accuracy.
AD is histopathologically characterized by two neuropathological hallmarks: neuritic plaques primarily composed of extracellular Amyloid-β (Aβ) deposits and intracellular neurofibrillary tangles (NFTs) composed of hyper-phosphorylated tau protein [100]. Obviously, tau protein, the MAPT gene encoded, plays an essential role in the pathogenesis of AD. It participates in the forming of NFTs. However, the genetic association between MAPT variants and AD risk has been inconsistent. Our results showed that rs2471738, rs3785883 and H2 haplotype were associated with AD. Several studies found that the pathogenesis of AD conferred by MAPT variants might be increasing levels of total or/and 4-repeat (4R) tau [6, 48, 101]. The T allele at rs2471738 has been associated with increased AD risk. But this SNP lies within a region of intron 9, and does not appear to interrupt the splice sites. This suggests that perhaps rs2471738 is not functional, but is in LD with other functional variants [4, 50]. Notably, the A allele at rs3785883 was found with a decreased risk of AD only in Caucasians. However, after the heterogeneity and sensitivity analysis, the association changed into negative when one study was excluded [53]. Hence, the protective effect of the A allele at rs3785883 on AD development in Caucasians is unreliable and awaits further investigation. Remarkably, both the two included studies showed that the A allele at rs242557 was associated with a decreased risk of AD in Asian [55, 56]. When pooling together, our result showed no association. So further studies are needed to verify this association since our result was only based on two studies.
PD is the second most common neurodegenerative disease following AD. Recently, some genome-wide association studies (GWAS) have provided robust evidence for the genetic association of MAPT with PD [7, 8, 73]. Likewise, our pooled analysis showed a robust association between MAPT and PD. We found that rs242557, rs7521 and H2 haplotype were associated with PD in Caucasian. Besides, MAPT variants influence the susceptibility of PD risk mainly by affecting the levels of total or 4R tau [8]. Then the tau would interact with α-synuclein, which forms the Lewy body in PD [102]. This process can promote the fibrillization of both tau and α-synuclein, and drive the formation of pathological inclusions in PD [103]. Rs242557 in the promoter of MAPT has been identified as a functional variant, which could affect the expression or splicing of MAPT, acting as cis-factors [83, 104]. Therefore, we speculate that rs242557-A allele can increase the brain tau levels in PD patients and thus raise the risk of PD susceptibility in Caucasians. For rs7521, the A allele could lower the age at onset (AAO) of PD and in reverse the G allele delays the AAO [70]. This finding is identical to our result that the A allele at rs7521 could increase the risk of PD in Caucasians. The mechanism may be that rs7521 is in LD with other risky SNPs [70]. We confirmed that H2 haplotype could decrease the risk of PD in Caucasians. However, only one study in Asian and one in African were included, and so the association of MAPT haplotype with PD in non-Caucasians awaits further investigation.
PSP and CBD are two parkinsonian syndromes characterized by deposits of neurofibrillary tangles in the brain, which are mainly composed of 4R MAPT protein isoforms [105]. In healthy adult brain, the levels of 3R-tau and 4R-tau are approximately equal. 4R-tau were nearly three times faster than 3R-tau for the rates of promoting to microtubule assemble to form the NTFs [3]. Probably, the two disorders are genetically related diseases and share a similar cause that involves tau dysfunction [12, 95]. However, PSP and CBD are usually considered sporadic disorders, so the genetics of the two disorders has been seldom researched [89, 95]. Hence, our meta-analysis is essential for identifying the genetics of PSP and CBD by pooling the limited studies. We confirmed that rs242557, rs2471738 and H1c subhaplotype could increase the risk of PSP and CBD, while H2 haplotype decrease. What’s more, these results were further confirmed that PSP and CBD might share a similar genetic background. Similarly, the mechanisms for the associations of rs242557, rs2471738, H2 and H1c with PSP and CBD may be due to influence the levels of tau, especially the 4R tau. However, only two studies were included in the subgroup of rs242557, rs2471738 and H1c in CBD. So, these results should be treated with cautious and more related studies are needed in the further.
FTD is a form of presenile dementia characterized clinically by behavioral and personality changes, mutism and decline of memory later in the disease [96], which neuropathologically affecting the frontal and/or temporal lobes. Many studies confirmed that mutations in the MAPT gene had been detected in autosomal dominant FTD with parkinsonism (FTDP). In addition, the accumulation of aberrant tau protein was found in FTD. These findings suggested that MAPT may be a genetic risk factor for FTD. However, our meta-analysis was failed to find the MAPT haplotypes had been associated with FTD. We suggest that more studies on other variants in MAPT are needed to probe the association and also in other populations.
ALS is a rarely progressive neurodegenerative disorder, but is the most common motor neuron disease in adult-onset. ALS is characterized by the simultaneous deficits of upper and lower motor neurons, which lead to muscle atrophy and paralysis [106]. The hyperphosphorylated and aggregated tau have been found in ALS, while the association was uncertain. We pooled that presently limited studies and found that H2 haplotype may be a mildly protective factor for ALS.
Our meta-analyses provide confirmatory evidence that multiple variants in MAPT are associated with neurodegenerative diseases risk. But there are several limits in our study. One limitation of our study is that the present studies were only limited to Caucasian populations in PSP, CBD, FTD and ALS, to our knowledge. What is more, in FTD and ALS, the studies were only limited to H1/H2 haplotype. Therefore, more studies are needed in the future, particularly those which performed in different populations and SNPs. Another limitation is different diagnostic criteria across studies. The accuracy of clinical diagnostic criteria is lower than pathological diagnostic criteria. For example, we assumed that about 6% of the patients had PSP but might be clinically diagnosed as PD by mistake [88]. Furthermore, we divided the data into subgroups by different criteria. But there was no difference across the pathological and clinical diagnostic subgroups.
In conclusion, this is the first comprehensive meta-analysis and systematic review which focused on the association between MAPT polymorphisms and neurodegenerative diseases so far. Our results robustly confirm the susceptibility role of MAPT in neurodegenerative diseases in a large meta-analysis, using highly informative htSNPs which capture 95% of MAPT haplotype diversity. Here we analyze these variants in MAPT and their association with neurodegenerative diseases as a step toward determining the precise mechanisms of genetic susceptibility for these group diseases. Significantly, this will have important implications for identifying potential therapeutic targets.
materials and Methods
Identification and selection of relevant studies
In order to identify the association between the MAPT polymorphism with neurodegenerative diseases, we conducted a literature search in MEDLINE, EMBASE and the Cochrane library up to September 2016. We used the key search terms including “microtubule-associated protein tau”, “MAPT”, “saitohin”, “STH”, “polymorphism”, “Alzheimer’s disease”, “Parkinson’s disease”, “progressive supranuclear palsy”, “corticobasal degeneration”, “frontotemporal dementia”, “amyotrophic lateral sclerosis” and “neurodegenerative disease”, combined with Boolean operators as appropriate. Additional studies were obtained from the reference lists of relevant original studies or review articles.
We assessed all studies if they meet the following criteria: (1) Evaluation of an association between MAPT polymorphisms and neurodegenerative diseases; (2) Case-control studies design; (3) Available allelic frequency for MAPT rs1467967, rs242557, rs3785883, rs2471738, rs7521, H2 and H1c haplotypes; Data of allelic frequency for estimating the OR with corresponding 95% CI were available in the report or could be calculated. We exclude the study, which the genotype frequency in the control group was not in HWE. Additionally, when two or more studies had overlapping participants, only the one with larger sample size was included.
Data extraction and quality assessment
Two reviewers independently read the studies and extracted data according to predefined criteria. Data of allelic frequency and its OR with corresponding 95% CI were extracted. Besides, the following information was extracted: author, publication year, state, ethnicity, diagnostic criteria, type of neurodegenerative diseases, sample size, mean age, the percentage of female, minor allele frequency, and the HWE test in each control group. The quality of studies was assessed with the Newcastle-Ottawa quality Scale (NOS): 1) the Selection; 2) the Comparability; 3) the Exposure [107]. Studies with a score of at least seven points were considered to be high quality.
Statistical analysis
To determine the strength of associations between individual MAPT polymorphism and neurodegenerative disease, we calculated a pooled OR and 95 % CI using R software. The OR and 95 % CI were evaluated by comparison in minor and major allele frequency using the Pearson χ2 test or Fisher’s exact test. We used the random-effects and fixed-effects model to calculate the subgroup and pooled ORs with 95% CIs in the presence and absence of heterogeneity, respectively. We assessed the heterogeneity by the Cochran Q and I2 (P > 0.10 or I2 < 50%, mean a lack of heterogeneity). Publication bias was detected by Egger’s test (P > 0.05, mean no publication bias) [108]. When k ≥ 10, we calculate the P value of Egger’s test using R software (k, means the number of studies). We used the trim and fill method to adjust the statistically significant publication bias [59]. In addition, as ethnic-specific susceptibility variants exist, we reanalyzed the data according to different races (Caucasian and Asian).
Abbreviations
MAPT = microtubule-associated protein tau; AD = Alzheimer’s disease; PD = Parkinson’s disease; PSP = progressive supranuclear palsy; CBD = corticobasal degeneration; FTD = frontotemporal dementia; ALS = amyotrophic lateral sclerosis; SNPs = single-nucleotide polymorphisms; htSNPs = haplotype tagging SNPs; LD = complete linkage disequilibrium; del-In9 = a 238-bp insertion/deletion polymorphism within intron 9; STH = Saitohin gene; HWE = Hardy-Weinberg equilibrium; NINCDS-ADRDA = the diagnostic criteria of the National Institute of Neurological and Communication Disorders and Stroke-Alzheimer Disease and Related Disorders Association; OR = odds ratio; CI = confidence interval; UKPDBB = the UK Parkinson’s Disease Society Brain Bank clinical diagnostic criteria; Aβ = Amyloid-β; 4R = 4-repeat; GWAS = genome-wide association studies; AAO = age at onset; FTDP = FTD with parkinsonism; NOS = Newcastle-Ottawa quality Scale.
Author contributions
Lan Tan and Jin-Tai Yu participated in the design of the study; Jun-Xia Zhu, Yu Wan and Lin Tan selected reports and extracted the data; Hui-Fu Wang and Cheng-Cheng Zhang analyzed data; Cheng-Cheng Zhang wrote the manuscript; All authors reviewed and approved the final version of the manuscript.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (81471309, 81371406, 81571245, and 81501103), the Shandong Provincial Outstanding Medical Academic Professional Program, Qingdao Key Health Discipline Development Fund, Qingdao Outstanding Health Professional Development Fund, and Shandong Provincial Collaborative Innovation Center for Neurodegenerative Disorders.
Conflicts of interest
The authors disclose no conflicts of interest.
Funding
This work was supported by grants from the National Natural Science Foundation of China (81471309, 81371406, 81571245, and 81501103), the Shandong Provincial Outstanding Medical Academic Professional Program, Qingdao Key Health Discipline Development Fund, Qingdao Outstanding Health Professional Development Fund, and Shandong Provincial Collaborative Innovation Center for Neurodegenerative Disorders.
References
1. Brown RC, Lockwood AH, Sonawane BR. Neurodegenerative diseases: an overview of environmental risk factors. Environ Health Perspect. 2005; 113:1250–56.
2. Chen Y, Li S, Su L, Sheng J, Lv W, Chen G, Xu Z. Association of progranulin polymorphism rs5848 with neurodegenerative diseases: a meta-analysis. J Neurol. 2015; 262:814–22.
3. Zhang CC, Xing A, Tan MS, Tan L, Yu JT. The Role of MAPT in Neurodegenerative Diseases: Genetics, Mechanisms and Therapy. Mol Neurobiol. 2015.
4. Myers AJ, Kaleem M, Marlowe L, Pittman AM, Lees AJ, Fung HC, Duckworth J, Leung D, Gibson A, Morris CM, de Silva R, Hardy J. The H1c haplotype at the MAPT locus is associated with Alzheimer’s disease. Hum Mol Genet. 2005; 14:2399–404.
5. Abraham R, Sims R, Carroll L, Hollingworth P, O’Donovan MC, Williams J, Owen MJ. An association study of common variation at the MAPT locus with late-onset Alzheimer’s disease. American journal of medical genetics Part B, Neuropsychiatric genetics. 2009; 150B:1152-1155.
6. Allen M, Kachadoorian M, Quicksall Z, Zou F, Chai HS, Younkin C, Crook JE, Pankratz VS, Carrasquillo MM, Krishnan S, Nguyen T, Ma L, Malphrus K, et al. Association of MAPT haplotypes with Alzheimer’s disease risk and MAPT brain gene expression levels. Alzheimers Res Ther. 2014; 6:39.
7. Satake W, Nakabayashi Y, Mizuta I, Hirota Y, Ito C, Kubo M, Kawaguchi T, Tsunoda T, Watanabe M, Takeda A, Tomiyama H, Nakashima K, Hasegawa K, et al. Genome-wide association study identifies common variants at four loci as genetic risk factors for Parkinson’s disease. Nat Genet. 2009; 41:1303–07.
8. Simón-Sánchez J, Schulte C, Bras JM, Sharma M, Gibbs JR, Berg D, Paisan-Ruiz C, Lichtner P, Scholz SW, Hernandez DG, Krüger R, Federoff M, Klein C, et al. Genome-wide association study reveals genetic risk underlying Parkinson’s disease. Nat Genet. 2009; 41:1308–12.
9. Lill CM, Roehr JT, McQueen MB, Kavvoura FK, Bagade S, Schjeide BM, Schjeide LM, Meissner E, Zauft U, Allen NC, Liu T, Schilling M, Anderson KJ, et al. Comprehensive research synopsis and systematic meta-analyses in Parkinson’s disease genetics: the PDGene database. PLoS Genet. 2012; 8:e1002548.
10. Higgins JJ, Golbe LI, De Biase A, Jankovic J, Factor SA, Adler RL. An extended 5′-tau susceptibility haplotype in progressive supranuclear palsy. Neurology. 2000; 55:1364–67.
11. Höglinger GU, Melhem NM, Dickson DW, Sleiman PM, Wang LS, Klei L, Rademakers R, de Silva R, Litvan I, Riley DE, van Swieten JC, Heutink P, Wszolek ZK, et al. Identification of common variants influencing risk of the tauopathy progressive supranuclear palsy. Nat Genet. 2011; 43:699–705.
12. Cruchaga C, Vidal-Taboada JM, Ezquerra M, Lorenzo E, Martinez-Lage P, Blazquez M, Tolosa E, Pastor P, and Iberian Atypical Parkinsonism Study Group Researchers. 5′-Upstream variants of CRHR1 and MAPT genes associated with age at onset in progressive supranuclear palsy and cortical basal degeneration. Neurobiol Dis. 2009; 33:164–70.
13. Kouri N, Carlomagno Y, Baker M, Liesinger AM, Caselli RJ, Wszolek ZK, Petrucelli L, Boeve BF, Parisi JE, Josephs KA, Uitti RJ, Ross OA, Graff-Radford NR, et al. Novel mutation in MAPT exon 13 (p.N410H) causes corticobasal degeneration. Acta Neuropathol. 2014; 127:271–82.
14. Sobrido MJ, Miller BL, Havlioglu N, Zhukareva V, Jiang Z, Nasreddine ZS, Lee VM, Chow TW, Wilhelmsen KC, Cummings JL, Wu JY, Geschwind DH. Novel tau polymorphisms, tau haplotypes, and splicing in familial and sporadic frontotemporal dementia. Arch Neurol. 2003; 60:698–702.
15. Webb A, Miller B, Bonasera S, Boxer A, Karydas A, Wilhelmsen KC. Role of the tau gene region chromosome inversion in progressive supranuclear palsy, corticobasal degeneration, and related disorders. Arch Neurol. 2008; 65:1473–78.
16. Hughes A, Mann D, Pickering-Brown S. Tau haplotype frequency in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Exp Neurol. 2003; 181:12–16.
17. Taes I, Goris A, Lemmens R, van Es MA, van den Berg LH, Chio A, Traynor BJ, Birve A, Andersen P, Slowik A, Tomik B, Brown RH Jr, Shaw CE, et al. Tau levels do not influence human ALS or motor neuron degeneration in the SOD1G93A mouse. Neurology. 2010; 74:1687–93.
18. Goedert M, Jakes R. Expression of separate isoforms of human tau protein: correlation with the tau pattern in brain and effects on tubulin polymerization. EMBO J. 1990; 9:4225–30.
19. Spillantini MG, Goedert M. Tau pathology and neurodegeneration. Lancet Neurol. 2013; 12:609–22.
20. Kaivorinne AL, Krüger J, Kuivaniemi K, Tuominen H, Moilanen V, Majamaa K, Remes AM. Role of MAPT mutations and haplotype in frontotemporal lobar degeneration in Northern Finland. BMC Neurol. 2008; 8:48.
21. Pittman AM, Myers AJ, Abou-Sleiman P, Fung HC, Kaleem M, Marlowe L, Duckworth J, Leung D, Williams D, Kilford L, Thomas N, Morris CM, Dickson D, et al. Linkage disequilibrium fine mapping and haplotype association analysis of the tau gene in progressive supranuclear palsy and corticobasal degeneration. J Med Genet. 2005; 42:837–46.
22. Pittman AM, Myers AJ, Duckworth J, Bryden L, Hanson M, Abou-Sleiman P, Wood NW, Hardy J, Lees A, de Silva R. The structure of the tau haplotype in controls and in progressive supranuclear palsy. Hum Mol Genet. 2004; 13:1267–74.
23. de Silva R, Hope A, Pittman A, Weale ME, Morris HR, Wood NW, Lees AJ. Strong association of the Saitohin gene Q7 variant with progressive supranuclear palsy. Neurology. 2003; 61:407–09.
24. Tobin JE, Latourelle JC, Lew MF, Klein C, Suchowersky O, Shill HA, Golbe LI, Mark MH, Growdon JH, Wooten GF, Racette BA, Perlmutter JS, Watts R, et al. Haplotypes and gene expression implicate the MAPT region for Parkinson disease: the GenePD Study. Neurology. 2008; 71:28–34.
25. Crawford F, Freeman M, Town T, Fallin D, Gold M, Duara R, Mullan M. No genetic association between polymorphisms in the Tau gene and Alzheimer’s disease in clinic or population based samples. Neurosci Lett. 1999; 266:193–96.
26. Ezquerra M, Pastor P, Valldeoriola F, Molinuevo JL, Blesa R, Tolosa E, Oliva R. Identification of a novel polymorphism in the promoter region of the tau gene highly associated to progressive supranuclear palsy in humans. Neurosci Lett. 1999; 275:183–86.
27. Lilius L, Froelich Fabre S, Basun H, Forsell C, Axelman K, Mattila K, Andreadis A, Viitanen M, Winblad B, Fratiglioni L, Lannfelt L. Tau gene polymorphisms and apolipoprotein E epsilon4 may interact to increase risk for Alzheimer’s disease. Neurosci Lett. 1999; 277:29–32.
28. Roks G, Dermaut B, Heutink P, Julliams A, Backhovens H, Van de Broeck M, Serneels S, Hofman A, Van Broeckhoven C, van Duijn CM, Cruts M. Mutation screening of the tau gene in patients with early-onset Alzheimer’s disease. Neurosci Lett. 1999; 277:137–39.
29. Baker M, Graff-Radford D, Wavrant DeVrièze F, Graff-Radford N, Petersen RC, Kokmen E, Boeve B, Myllykangas L, Polvikoski T, Sulkava R, Verkoniemmi A, Tienari P, Haltia M, et al. No association between TAU haplotype and Alzheimer’s disease in population or clinic based series or in familial disease. Neurosci Lett. 2000; 285:147–49.
30. Bullido MJ, Aldudo J, Frank A, Coria F, Avila J, Valdivieso F. A polymorphism in the tau gene associated with risk for Alzheimer’s disease. Neurosci Lett. 2000; 278:49–52.
31. Kwon JM, Nowotny P, Shah PK, Chakraverty S, Norton J, Morris JC, Goate AM. Tau polymorphisms are not associated with Alzheimer’s disease. Neurosci Lett. 2000; 284:77–80.
32. Russ C, Powell JF, Zhao J, Baker M, Hutton M, Crawford F, Mullan M, Roks G, Cruts M, Lovestone S. The microtubule associated protein Tau gene and Alzheimer’s disease—an association study and meta-analysis. Neurosci Lett. 2001; 314:92–96.
33. Conrad C, Vianna C, Freeman M, Davies P. A polymorphic gene nested within an intron of the tau gene: implications for Alzheimer’s disease. Proc Natl Acad Sci USA. 2002; 99:7751–56.
34. Cook L, Brayne CE, Easton D, Evans JG, Xuereb J, Cairns NJ, Rubinsztein DC. No evidence for an association between Saitohin Q7R polymorphism and Alzheimer’s disease. Ann Neurol. 2002; 52:690–91.
35. Verpillat P, Ricard S, Hannequin D, Dubois B, Bou J, Camuzat A, Pradier L, Frebourg T, Brice A, Clerget-Darpoux F, Deleuze JF, Campion D, and French Study Group on Alzheimer’s Disease and Frontotemporal Dementia. Is the saitohin gene involved in neurodegenerative diseases? Ann Neurol. 2002; 52:829–32.
36. Clark LN, Levy G, Tang MX, Mejia-Santana H, Ciappa A, Tycko B, Cote LJ, Louis ED, Mayeux R, Marder K. The Saitohin ‘Q7R’ polymorphism and tau haplotype in multi-ethnic Alzheimer disease and Parkinson’s disease cohorts. Neurosci Lett. 2003; 347:17–20.
37. Combarros O, Rodero L, Infante J, Palacio E, Llorca J, Fernández-Viadero C, Peña N, Berciano J. Age-dependent association between the Q7R polymorphism in the Saitohin gene and sporadic Alzheimer’s disease. Dement Geriatr Cogn Disord. 2003; 16:132–35.
38. Oliveira SA, Martin ER, Scott WK, Nicodemus KK, Small GW, Schmechel DE, Doraiswamy PM, Roses AD, Saunders AM, Gilbert JR, Haines JL, Vance JM, Pericak-Vance MA. The Q7R Saitohin gene polymorphism is not associated with Alzheimer disease. Neurosci Lett. 2003; 347:143–46.
39. Pepłońska B, Zekanowski C, Religa D, Czyzewski K, Styczyńska M, Pfeffer A, Gabryelewicz T, Gołebiowski M, Luczywek E, Wasiak B, Barczak A, Chodakowska M, Barcikowska M, Kuźnicki J. Strong association between Saitohin gene polymorphism and tau haplotype in the Polish population. Neurosci Lett. 2003; 348:163–66.
40. Streffer JR, Papassotiropoulos A, Kurosinski P, Signorell A, Wollmer MA, Tsolaki M, Iakovidou V, Hörndli F, Bosset J, Götz J, Nitsch RM, Hock C. Saitohin gene is not associated with Alzheimer’s disease. J Neurol Neurosurg Psychiatry. 2003; 74:362–63.
41. Conrad C, Vianna C, Schultz C, Thal DR, Ghebremedhin E, Lenz J, Braak H, Davies P. Molecular evolution and genetics of the Saitohin gene and tau haplotype in Alzheimer’s disease and argyrophilic grain disease. J Neurochem. 2004; 89:179–88.
42. Seripa D, Matera MG, D’Andrea RP, Gravina C, Masullo C, Daniele A, Bizzarro A, Rinaldi M, Antuono P, Wekstein DR, Dal Forno G, Fazio VM. Alzheimer disease risk associated with APOE4 is modified by STH gene polymorphism. Neurology. 2004; 62:1631–33.
43. Johansson A, Zetterberg H, Håkansson A, Nissbrandt H, Blennow K. TAU haplotype and the Saitohin Q7R gene polymorphism do not influence CSF Tau in Alzheimer’s disease and are not associated with frontotemporal dementia or Parkinson’s disease. Neurodegener Dis. 2005; 2:28–35.
44. Zuo L, van Dyck CH, Luo X, Kranzler HR, Yang BZ, Gelernter J. Variation at APOE and STH loci and Alzheimer’s disease. Behav Brain Funct. 2006; 2:13.
45. Ezquerra M, Gaig C, Ascaso C, Muñoz E, Tolosa E. Tau and saitohin gene expression pattern in progressive supranuclear palsy. Brain Res. 2007; 1145:168–76.
46. Laws SM, Friedrich P, Diehl-Schmid J, Müller J, Eisele T, Bäuml J, Förstl H, Kurz A, Riemenschneider M. Fine mapping of the MAPT locus using quantitative trait analysis identifies possible causal variants in Alzheimer’s disease. Mol Psychiatry. 2007; 12:510–17.
47. Mukherjee O, Kauwe JS, Mayo K, Morris JC, Goate AM. Haplotype-based association analysis of the MAPT locus in late onset Alzheimer’s disease. BMC Genet. 2007; 8:3.
48. Myers AJ, Pittman AM, Zhao AS, Rohrer K, Kaleem M, Marlowe L, Lees A, Leung D, McKeith IG, Perry RH, Morris CM, Trojanowski JQ, Clark C, et al. The MAPT H1c risk haplotype is associated with increased expression of tau and especially of 4 repeat containing transcripts. Neurobiol Dis. 2007; 25:561–70.
49. Lin K, Tang M, Guo Y, Han H, Lin Y, Ma C. The Q7R polymorphism in the saitohin gene is rare in a southern Chinese population. Neurological sciences. 2008; 29:431-434.
50. Mateo I, Sánchez-Juan P, Rodríguez-Rodríguez E, Infante J, Fernández-Viadero C, Peña N, Berciano J, Combarros O. 14-3-3 zeta and tau genes interactively decrease Alzheimer’s disease risk. Dement Geriatr Cogn Disord. 2008; 25:317–20.
51. Feulner TM, Laws SM, Friedrich P, Wagenpfeil S, Wurst SH, Riehle C, Kuhn KA, Krawczak M, Schreiber S, Nikolaus S, Förstl H, Kurz A, Riemenschneider M. Examination of the current top candidate genes for AD in a genome-wide association study. Mol Psychiatry. 2010; 15:756–66.
52. Cousin E, Macé S, Rocher C, Dib C, Muzard G, Hannequin D, Pradier L, Deleuze JF, Génin E, Brice A, Campion D. No replication of genetic association between candidate polymorphisms and Alzheimer’s disease. Neurobiol Aging. 2011; 32:1443–51.
53. Setó-Salvia N, Clarimón J, Pagonabarraga J, Pascual-Sedano B, Campolongo A, Combarros O, Mateo JI, Regaña D, Martínez-Corral M, Marquié M, Alcolea D, Suárez-Calvet M, Molina-Porcel L, et al. Dementia risk in Parkinson disease: disentangling the role of MAPT haplotypes. Arch Neurol. 2011; 68:359–64.
54. Elias-Sonnenschein LS, Helisalmi S, Natunen T, Hall A, Paajanen T, Herukka SK, Laitinen M, Remes AM, Koivisto AM, Mattila KM, Lehtimäki T, Verhey FR, Visser PJ, et al. Genetic loci associated with Alzheimer’s disease and cerebrospinal fluid biomarkers in a Finnish case-control cohort. PLoS One. 2013; 8:e59676.
55. Liu QY, Yu JT, Miao D, Ma XY, Wang HF, Wang W, Tan L. An exploratory study on STX6, MOBP, MAPT, and EIF2AK3 and late-onset Alzheimer’s disease. Neurobiol Aging. 2013; 34:1519 e13-7.
56. Chang CW, Hsu WC, Pittman A, Wu YR, Hardy J, Fung HC. Structural study of the microtubule-associated protein tau locus of Alzheimer’s disease in Taiwan. Biomed J. 2014; 37:127–32.
57. Pastor P, Moreno F, Clarimon J, Ruiz A, Combarros O, Calero M, Lopez de Munain A, Bullido MJ, de Pancorbo MM, Carro E, Antonell A, Coto E, Ortega-Cubero S, et al. MAPT H1 Haplotype is Associated with Late-Onset Alzheimer’s Disease Risk in APOEvarepsilon4 Noncarriers: Results from the Dementia Genetics Spanish Consortium. Journal of Alzheimer’s disease. JAD. 2015; 49:343–52.
58. McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology. 1984; 34:939–44.
59. Duval S, Tweedie R. Trim and fill: A simple funnel-plot-based method of testing and adjusting for publication bias in meta-analysis. Biometrics. 2000; 56:455–63.
60. Maraganore DM, Hernandez DG, Singleton AB, Farrer MJ, McDonnell SK, Hutton ML, Hardy JA, Rocca WA. Case-Control study of the extended tau gene haplotype in Parkinson’s disease. Ann Neurol. 2001; 50:658–61.
61. Farrer M, Skipper L, Berg M, Bisceglio G, Hanson M, Hardy J, Adam A, Gwinn-Hardy K, Aasly J. The tau H1 haplotype is associated with Parkinson’s disease in the Norwegian population. Neurosci Lett. 2002; 322:83–86.
62. Kwok JB, Teber ET, Loy C, Hallupp M, Nicholson G, Mellick GD, Buchanan DD, Silburn PA, Schofield PR. Tau haplotypes regulate transcription and are associated with Parkinson’s disease. Ann Neurol. 2004; 55:329–34.
63. Levecque C, Elbaz A, Clavel J, Vidal JS, Amouyel P, Alpérovitch A, Tzourio C, Chartier-Harlin MC. Association of polymorphisms in the Tau and Saitohin genes with Parkinson’s disease. J Neurol Neurosurg Psychiatry. 2004; 75:478–80.
64. Skipper L, Wilkes K, Toft M, Baker M, Lincoln S, Hulihan M, Ross OA, Hutton M, Aasly J, Farrer M. Linkage disequilibrium and association of MAPT H1 in Parkinson disease. Am J Hum Genet. 2004; 75:669–77.
65. Fidani L, Kalinderi K, Bostantjopoulou S, Clarimon J, Goulas A, Katsarou Z, Hardy J and Kotsis A. Association of the Tau haplotype with Parkinson’s disease in the Greek population. Movement disorders : official journal of the Movement Disorder Society. 2006; 21:1036-1039.
66. Fung HC, Xiromerisiou G, Gibbs JR, Wu YR, Eerola J, Gourbali V, Hellström O, Chen CM, Duckworth J, Papadimitriou A, Tienari PJ, Hadjigeorgiou GM, Hardy J, Singleton AB. Association of tau haplotype-tagging polymorphisms with Parkinson’s disease in diverse ethnic Parkinson’s disease cohorts. Neurodegener Dis. 2006; 3:327–33.
67. Winkler S, König IR, Lohmann-Hedrich K, Vieregge P, Kostic V, Klein C. Role of ethnicity on the association of MAPT H1 haplotypes and subhaplotypes in Parkinson’s disease. Eur J Hum Genet. 2007; 15:1163–68.
68. Zabetian CP, Hutter CM, Factor SA, Nutt JG, Higgins DS, Griffith A, Roberts JW, Leis BC, Kay DM, Yearout D, Montimurro JS, Edwards KL, Samii A, Payami H. Association analysis of MAPT H1 haplotype and subhaplotypes in Parkinson’s disease. Ann Neurol. 2007; 62:137–44.
69. Camuzat A, Romana M, Durr A, Feingold J, Brice A, Ruberg M, Lannuzel A. The PSP-associated MAPT H1 subhaplotype in Guadeloupean atypical parkinsonism. Movement disorders. 2008; 23:2384-2391.
70. Das G, Misra AK, Das SK, Ray K, Ray J. Microtubule-associated protein tau (MAPT) influences the risk of Parkinson’s disease among Indians. Neurosci Lett. 2009; 460:16–20.
71. Refenes N, Bolbrinker J, Tagaris G, Orlacchio A, Drakoulis N, Kreutz R. Role of the H1 haplotype of microtubule-associated protein tau (MAPT) gene in Greek patients with Parkinson’s disease. BMC Neurol. 2009; 9:26.
72. Vandrovcova J, Pittman AM, Malzer E, Abou-Sleiman PM, Lees AJ, Wood NW, de Silva R. Association of MAPT haplotype-tagging SNPs with sporadic Parkinson’s disease. Neurobiol Aging. 2009; 30:1477–82.
73. Edwards TL, Scott WK, Almonte C, Burt A, Powell EH, Beecham GW, Wang L, Züchner S, Konidari I, Wang G, Singer C, Nahab F, Scott B, et al. Genome-wide association study confirms SNPs in SNCA and the MAPT region as common risk factors for Parkinson disease. Ann Hum Genet. 2010; 74:97–109.
74. Wider C, Vilariño-Güell C, Jasinska-Myga B, Heckman MG, Soto-Ortolaza AI, Cobb SA, Aasly JO, Gibson JM, Lynch T, Uitti RJ, Wszolek ZK, Farrer MJ, Ross OA. Association of the MAPT locus with Parkinson’s disease. Eur J Neurol. 2010; 17:483–86.
75. Spencer CC, Plagnol V, Strange A, Gardner M, Paisan-Ruiz C, Band G, Barker RA, Bellenguez C, Bhatia K, Blackburn H, Blackwell JM, Bramon E, Brown MA, et al. Dissection of the genetics of Parkinson’s disease identifies an additional association 5′ of SNCA and multiple associated haplotypes at 17q21. Hum Mol Genet. 2011; 20:345–53.
76. Elbaz A, Ross OA, Ioannidis JP, Soto-Ortolaza AI, Moisan F, Aasly J, Annesi G, Bozi M, Brighina L, Chartier-Harlin MC, Destée A, Ferrarese C, Ferraris A, et al. Independent and joint effects of the MAPT and SNCA genes in Parkinson disease. Ann Neurol. 2011; 69:778–92.
77. Ezquerra M, Pastor P, Gaig C, Vidal-Taboada JM, Cruchaga C, Munoz E, Marti MJ, Valldeoriola F, Aguilar M, Calopa M, Hernandez-Vara J, Tolosa E. Different MAPT haplotypes are associated with Parkinson’s disease and progressive supranuclear palsy. Neurobiology of aging. 2011; 32:547 e511-546.
78. Kalinderi K, Fidani L, Katsarou Z, Clarimon J, Bostantjopoulou S, Kotsis A. GSK3beta polymorphisms, MAPT H1 haplotype and Parkinson’s disease in a Greek cohort. Neurobiology of aging. 2011; 32:546 e541-545.
79. Mata IF, Yearout D, Alvarez V, Coto E, de Mena L, Ribacoba R, Lorenzo-Betancor O, Samaranch L, Pastor P, Cervantes S, Infante J, Garcia-Gorostiaga I, Sierra M, et al. Replication of MAPT and SNCA, but not PARK16-18, as susceptibility genes for Parkinson’s disease. Movement disorders. 2011; 26:819-823.
80. Simón-Sánchez J, van Hilten JJ, van de Warrenburg B, Post B, Berendse HW, Arepalli S, Hernandez DG, de Bie RM, Velseboer D, Scheffer H, Bloem B, van Dijk KD, Rivadeneira F, et al. Genome-wide association study confirms extant PD risk loci among the Dutch. Eur J Hum Genet. 2011; 19:655–61.
81. Trotta L, Guella I, Soldà G, Sironi F, Tesei S, Canesi M, Pezzoli G, Goldwurm S, Duga S, Asselta R. SNCA and MAPT genes: independent and joint effects in Parkinson disease in the Italian population. Parkinsonism Relat Disord. 2012; 18:257–62.
82. Emelyanov A, Andoskin P, Yakimovskii A, Usenko T, Nuzhnyi E, Nikolaev M, Pchelina S. SNCA, LRRK2, MAPT polymorphisms and Parkinson’s disease in Russia. Parkinsonism Relat Disord. 2013; 19:1064–65.
83. Chen Y, Cao B, Ou R, Chen X, Zhao B, Wei Q, Wu Y, Shang HF. Association analysis of the GRN rs5848 and MAPT rs242557 polymorphisms in Parkinson’s disease and multiple system atrophy: a large-scale population-based study and meta-analysis. Int J Neurosci. 2015;126:947-54
84. Sonmez E, Ozel MD, Islek EE, Sazci A, Idrisoglu HA. Association of rs62063857 variant of the saitohin gene with Parkinson’s disease. Cell Mol Neurobiol. 2015; 35:115–21.
85. Cervera-Carles L, Pagonabarraga J, Pascual-Sedano B, Pastor P, Campolongo A, Fortea J, Blesa R, Alcolea D, Morenas-Rodriguez E, Sala I, Lleo A, Kulisevsky J, Clarimon J. Copy number variation analysis of the 17q21.31 region and its role in neurodegenerative diseases. Am J Med Genet B Neuropsychiatr Genet. 2016; 171:175-180.
86. Bower JH, Maraganore DM, McDonnell SK, Rocca WA. Incidence and distribution of parkinsonism in Olmsted County, Minnesota, 1976-1990. Neurology. 1999; 52:1214–20.
87. Gelb DJ, Oliver E, Gilman S. Diagnostic criteria for Parkinson disease. Arch Neurol. 1999; 56:33–39.
88. Hughes AJ, Daniel SE, Kilford L, Lees AJ. Accuracy of clinical diagnosis of idiopathic Parkinson’s disease: a clinico-pathological study of 100 cases. J Neurol Neurosurg Psychiatry. 1992; 55:181–84.
89. Baker M, Litvan I, Houlden H, Adamson J, Dickson D, Perez-Tur J, Hardy J, Lynch T, Bigio E, Hutton M. Association of an extended haplotype in the tau gene with progressive supranuclear palsy. Hum Mol Genet. 1999; 8:711–15.
90. de Silva R, Weiler M, Morris HR, Martin ER, Wood NW, Lees AJ. Strong association of a novel Tau promoter haplotype in progressive supranuclear palsy. Neurosci Lett. 2001; 311:145–48.
91. Ezquerra M, Campdelacreu J, Muñoz E, Oliva R, Tolosa E. Sequence analysis of tau 3’untranslated region and saitohin gene in sporadic progressive supranuclear palsy. J Neurol Neurosurg Psychiatry. 2004; 75:155–57.
92. Rademakers R, Melquist S, Cruts M, Theuns J, Del-Favero J, Poorkaj P, Baker M, Sleegers K, Crook R, De Pooter T, Bel Kacem S, Adamson J, Van den Bossche D, et al. High-density SNP haplotyping suggests altered regulation of tau gene expression in progressive supranuclear palsy. Hum Mol Genet. 2005; 14:3281–92.
93. Tolosa E, Valldeoriola F, Marti MJ. Clinical diagnosis and diagnostic criteria of progressive supranuclear palsy (Steele-Richardson-Olszewski syndrome). J Neural Transm Suppl. 1994; 42:15–31.
94. Litvan I, Agid Y, Calne D, Campbell G, Dubois B, Duvoisin RC, Goetz CG, Golbe LI, Grafman J, Growdon JH, Hallett M, Jankovic J, Quinn NP, et al. Clinical research criteria for the diagnosis of progressive supranuclear palsy (Steele-Richardson-Olszewski syndrome): report of the NINDS-SPSP international workshop. Neurology. 1996; 47:1–9.
95. Houlden H, Baker M, Morris HR, MacDonald N, Pickering-Brown S, Adamson J, Lees AJ, Rossor MN, Quinn NP, Kertesz A, Khan MN, Hardy J, Lantos PL, et al. Corticobasal degeneration and progressive supranuclear palsy share a common tau haplotype. Neurology. 2001; 56:1702–06.
96. Ghidoni R, Signorini S, Barbiero L, Sina E, Cominelli P, Villa A, Benussi L, Binetti G. The H2 MAPT haplotype is associated with familial frontotemporal dementia. Neurobiol Dis. 2006; 22:357–62.
97. Verpillat P, Camuzat A, Hannequin D, Thomas-Anterion C, Puel M, Belliard S, Dubois B, Didic M, Michel BF, Lacomblez L, Moreaud O, Sellal F, Golfier V, et al. Association between the extended tau haplotype and frontotemporal dementia. Arch Neurol. 2002; 59:935–39.
98. Miller BL, Ikonte C, Ponton M, Levy M, Boone K, Darby A, Berman N, Mena I, Cummings JL. A study of the Lund-Manchester research criteria for frontotemporal dementia: clinical and single-photon emission CT correlations. Neurology. 1997; 48:937–42.
99. Neary D, Snowden JS, Gustafson L, Passant U, Stuss D, Black S, Freedman M, Kertesz A, Robert PH, Albert M, Boone K, Miller BL, Cummings J, Benson DF. Frontotemporal lobar degeneration: a consensus on clinical diagnostic criteria. Neurology. 1998; 51:1546–54.
100. Yu JT, Tan L, Hardy J. Apolipoprotein E in Alzheimer’s disease: an update. Annu Rev Neurosci. 2014; 37:79–100.
101. Zou F, Chai HS, Younkin CS, Allen M, Crook J, Pankratz VS, Carrasquillo MM, Rowley CN, Nair AA, Middha S, Maharjan S, Nguyen T, Ma L, et al. Brain expression genome-wide association study (eGWAS) identifies human disease-associated variants. PLoS Genet. 2012; 8:e1002707.
102. Spillantini MG, Schmidt ML, Lee VM, Trojanowski JQ, Jakes R, Goedert M. Alpha-synuclein in Lewy bodies. Nature. 1997; 388:839–40.
103. Giasson BI, Forman MS, Higuchi M, Golbe LI, Graves CL, Kotzbauer PT, Trojanowski JQ, Lee VM. Initiation and synergistic fibrillization of tau and alpha-synuclein. Science. 2003; 300:636–40.
104. Myers AJ, Gibbs JR, Webster JA, Rohrer K, Zhao A, Marlowe L, Kaleem M, Leung D, Bryden L, Nath P, Zismann VL, Joshipura K, Huentelman MJ, et al. A survey of genetic human cortical gene expression. Nat Genet. 2007; 39:1494–99.
105. Spillantini MG, Goedert M. Tau protein pathology in neurodegenerative diseases. Trends Neurosci. 1998; 21:428–33.
106. Vercruysse P, Sinniger J, El Oussini H, Scekic-Zahirovic J, Dieterlé S, Dengler R, Meyer T, Zierz S, Kassubek J, Fischer W, Dreyhaupt J, Grehl T, Hermann A, et al. Alterations in the hypothalamic melanocortin pathway in amyotrophic lateral sclerosis. Brain. 2016; 139:1106–22.
107. Stang A. Critical evaluation of the Newcastle-Ottawa scale for the assessment of the quality of nonrandomized studies in meta-analyses. Eur J Epidemiol. 2010; 25:603–05.
108. Hayashino Y, Noguchi Y, Fukui T. Systematic evaluation and comparison of statistical tests for publication bias. Journal of epidemiology/Japan Epidemiological Association. 2005; 15:235-243.
