Introduction
Colorectal cancer (CRC) is the 3rd most commonly diagnosed cancer in males and 2nd in females worldwide. It estimated that 1.4 million new cases and 693,900 deaths occurs in 2012 [1]. Numerous studies suggested that the first stage of metastasis occurs early and that more than 60% of patients have initiated the metastatic process by the time of diagnosis [2]. Based on the degree of metastasis, CRC is divided into localized stage, regional stage and stage with distant metastases. More seriously, compared to CRC patients at localized stage, the 5-year survival rates of patients with distant metastases drop from 90% to 10% [3, 4]. Over the past 3 dedicates, the 5-year survival rates of CRC patients at distant stage has not been improved. Besides, distant metastases lead to approximately 50% death of patients diagnosed with CRC [5]. Obviously, it is necessary to reveal the mechanisms underlying this process. The formation of metastases is a complex and multistep process of the dissemination of tumor cells from the primary tumor microenvironment to various distant organs and colonization of the secondary site [6]. Firstly, tumor cells escape from the primary tumor into the blood or lymphatic system. Secondly, some of them escaping the apoptosis survive and arrest at a secondary site. The third step is extravasation into the distant tissue and survival in the new microenvironment. Finally, they form metastatic colonization by proliferation in the distant location. All these steps are critical for us to understand the biological processes during metastasis. Therefore, angiogenesis, escape of apoptosis, proliferation, invasion and migration are essential for the colonization of metastatic outgrowth [7]. Although numerous genes have been identified as biomarkers of carcinogenesis, the emerging roles of long noncoding RNAs (lncRNAs) in the development of CRC metastasis are largely unknown.
With the advance of whole genome sequencing technology and high-resolution microarray, the massive amount of short RNA or long RNA without protein coding ability was revealed. These non-coding RNAs (ncRNAs) comprise of small interfering RNAs (siRNAs), microRNAs (miRNAs), PIWI-interacting RNAs (piRNAs), small nucleolar RNAs (snoRNAs) and long noncoding RNAs (lncRNAs). Long noncoding RNAs (lncRNAs) are non-coding transcripts of more than 200 nucleotides (nt) in length and majority of them are located in nuclear [8]. LncRNAs are generally transcribed by RNA polymerase II and subsequently are polyadenylated [9]. They have no open reading frame (ORF) resulting in the loss of translation capacity. Similar to protein coding genes, the transcriptional start sites (TSS) of lncRNAs are marked by histone 3 lysine 4 (H3K4me3) and the gene bodies are marked by histone 3 lysine 36 (H3K36me3) throughout. These imply that lncRNAs display epigenetic features [8]. Compared to messenger RNAs (mRNA), most lncRNAs only harbor two exons, resulting that they are shorter than mRNA [9]. Moreover, the expression levels of lncRNAs are significantly lower than mRNA expression and they show developmental and tissue specific expression patterns [10]. Based on the genetic point of view, five different types of lncRNAs are classified, including intergenic lncRNAs, intronic lncRNAs, bidirectional lncRNAs, sense lncRNAs and antisense lncRNAs [9]. These characteristics are essential for us to gain greater insights into the function of lncRNAs.
Recently, lncRNAs have caught more attention and accumulating studies suggested that lncRNAs play critical roles in tumourigenesis and cancer progression. They function as oncogenes and tumor suppressors in diverse biological processes, such as imprinting, epigenetic regulation, apoptosis, cell cycle, transcriptional and translational regulation, splicing, cell development and differentiation [11]. Besides, numerous studies have demonstrated that The metastatic pathways were highly associated with the aberrant expression of lncRNAs and lncRNAs played important roles in the development of metastases by activing or inhibiting the metastatic pathway in different cancers, such as hepatocellular carcinoma, gastric cancer and non-small cell lung cancer [12-14]. Due to the tumor or tissue specific expression patterns of lncRNAs, there is a critical need for amount of researches on the association between the dysregulation of lncRNAs and the metastatic pathways in CRC. However, the number of data is still limited. With the development of the therapeutic approaches based on lncRNAs, it is necessary to summarize the role of lncRNAs in the development of metastasis, especially in CRC.
In this review, we highlight the findings of roles for lncRNAs in CRC metastasis and review the association of lncRNAs and metastatic pathways in CRC, including escape of apoptosis, epithelia-mesenchymal transition (EMT), angiogenesis and proliferation, invasion and migration [7]. Furthermore, we also discuss the potential clinical application of lncRNAs in CRC.
LncRNAs involved in CRC metastasis
Recently, accumulating evidences have uncovered that lncRNAs directly regulate the metastatic pathways in CRC [15, 16]. By using Human lncRNA Assay, Han J et. al compared lncRNA expression profiles between metastatic lymph node (MLN), normal lymph node (NLN) and tumor tissues from three CRC patients. Five lncRNAs were down-regulated and fourteen lncRNAs were up-regulated in the MLN group compared with the NLN group and tumor tissue group. Besides, four gradually up-regulated lncRNAs and sixty-six down-regulated lncRNAs were identified from tumor tissue to MLN and NLN, respectively. All these lncRNAs were supposed to play important roles in the lymph node metastasis (LNM) of CRC [17]. Recently, three pairs of tumor tissues and MLNs were used to perform in another microarray analysis. A total of 390 aberrant expressed lncRNAs were observed in the tumor tissues compared with MLNs. Especially, ENST00000430471 exhibited the lowest expression in tumor tissues compared with MLNs. Further studies suggested that it promoted cell proliferation, invasion, migration and S-phase arrest and inhibited cell apoptosis [18]. To identified the role of lncRNAs in the progress of colorectal liver metastasis (CLM), Ye LC et. al profiled the lncRNAs expression in CRC tissues with synchronous, metachronous and non-liver metastasis. Three novel lncRNAs of forty differentially expressed lncRNAs in CLM tissues, were verified to be involved in CLM [19]. Besides, a genome-wide analysis comparing lncRNAs expression profiles between CRC tissues with or without liver metastasis identified 2636 differentially expressed lncRNAs. Six lncRNAs (POU6F2-AS1, RAB6C-AS1, DDP10-AS1, HOXA11-AS, LINC00944 and FEZF1-AS1) were verified to participate in the process of liver metastasis in CRC though further validation [20]. Interestingly, FEZF1-AS1 was subsequently reported to enhance CRC cells proliferation, invasion and migration partly through FEZF1 induction [21]. Recently, genome-wide lncRNA expression patterns were assessed by microarray analysis in metastatic lymph nodes and its paired normal lymph nodes of CRC patients. A new lncRNA named GAPLINC was found. It was demonstrated to be associated with tumor growth, tumor stage, node stage and overall survival of CRC. In vivo and vitro assays, it significantly promoted cell proliferation and invasion by interacting with PTB-associated splicing factor (PSF) and non-POU-domain-containing octamer-binding (NONO) [22]. All their results supposed that lncRNAs may play crucial roles in the metastasis of CRC.
Although the use of microarray makes to identified accumulate of lncRNAs, which are significantly associated with the process of CRC metastasis. Microarray analysis on colorectal tumor at multi-treatment center, of different subtypes and with large sample sizes were blank. Furthermore, to fully utilize the microarray data and better understand the intrinsic mechanisms, the specific signaling of these screened lncRNAs involved in influencing CRC metastasis should be investigated. Therefore, more studies in vivo and vitro are critical needed to continue.
LncRNAs and metastatic pathways in CRC
Multivariate evidences have demonstrated that aberrant expression of lncRNAs plays important roles in the metastatic process both in vitro and vivo in CRC (Table 1). Many metastatic pathway including escape of apoptosis, EMT, angiogenesis and invasion, migration and proliferation are crucial for the formation of metastases. Therefore, we discuss and describe the association of lncRNAs and these metastatic pathways in CRC in detail below.
Table 1: lncRNAs involved in metastasis CRC
lncRNA |
Chromosomal location |
Gene type |
Putative functions related to metastatic prognosis |
Signaling pathways |
Ref. |
CLMAT3 |
Chr14 |
Oncogene |
Liver metastasis |
G0/G1 cell-cycle arrest |
[19] |
FEZF1-AS1 |
7q31.32 |
Oncogene |
Proliferation, migration, invasion and metastasis |
||
GAPLINC |
Chr18 |
Oncogene |
LNM, proliferation and invasion |
GAPLINC/PSF/NONO |
|
PVT-1 |
8q24 |
Oncogene |
LNM, Proliferation, invasion and apoptosis |
[25] |
|
DQ786243 |
Chr1 |
Oncogene |
Cell cycle, apoptosis, metastasis proliferation and invasion, |
G2/M cell-cycle arrest |
[26] |
BANCR |
Chr9 |
Oncogene |
Proliferation, migration, EMT, cell cycle, and apoptosis |
EMT signaling pathway |
|
HOTTIP |
7q15.2 |
Oncogene |
Distance metastasis, proliferation and apoptosis |
G0/G1 cell-cycle arrest |
|
LincRNA-p21 |
6p21.2 |
Tumor suppressor |
Vascular invasion, proliferation and cell cycle |
Wnt/β-catenin signaling pathway |
|
ZFAS1 |
20q13.13 |
Oncogene |
Proliferation, apoptosis and cell cycle |
p53-dependent cell cycle |
[43] |
Loc554202 |
9p21.3 |
Tumor suppressor |
Proliferation, tumorigenesis and apoptosis |
caspase cleavage cascades |
[45] |
UCA1 |
19p13.12 |
Oncogene |
Proliferation, metastasis, cell cycle and apoptosis, , |
||
PRNCR1 |
8q24.21 |
Oncogene |
Proliferation and cell cycle |
G0/G1 cell-cycle arrest |
|
HOTAIR |
12q13.13 |
Oncogene |
Proliferation, invasion, EMT, LNM and Lung metastasis |
EMT signaling pathway |
[52] |
TUG1 |
22q12.2 |
Oncogene |
EMT, invasion and migration |
EMT signaling pathway |
[54] |
LncRNA-ATB |
Chr14 |
Oncogene |
LNM, EMT, tumorigenesis and invasion |
EMT signaling pathway |
|
H19 |
11p15.5 |
Oncogene |
Proliferation, EMT and cell cycle |
EMT signaling pathway β-catenin pathway |
|
CTD903 |
14q11.2 |
Oncogene |
Invasion and migration and EMT |
EMT signaling pathway Wnt/β-catenin pathway |
[65] |
GHET1 |
7q36.1 |
Oncogene |
Proliferation, invasion, migration and EMT |
EMT signaling pathway |
[66] |
CASC2 |
10q26.11 |
Tumor suppressor |
Proliferation and cell cycle |
G0/G1 cell-cycle arrest |
[109] |
LOC285194 |
3q13.31 |
Tumor suppressor |
Distant metastasis and proliferation |
||
FER1L4 |
20q11.22 |
Tumor suppressor |
LNM, cell cycle, proliferation, invasion, and migration |
[113] |
|
CCAL |
Chr3 |
Oncogene |
Proliferation, invasion , migration, apoptosis, cell cycle and LNM |
[118] |
|
CCAT1 |
8q24.21 |
Oncogene |
LNM, proliferation and invasion |
[120] |
|
CASC11 |
8q24 |
Oncogene |
Lymph metastasis and proliferation |
Wnt/β-catenin pathway |
[121] |
CCAT2 |
8q24.21 |
Oncogene |
Cell growth and metastasis |
[122] |
|
PCAT-1 |
8q24.21 |
Oncogene |
Distant metastasis |
[123] |
|
MALAT1 |
11q13.1 |
Oncogene |
Proliferation, migration, invasion and metastasis |
PI3K/Akt pathway Wnt/β-catenin pathway |
|
TINCR |
19q13.3 |
Tumor suppressor |
Proliferation and metastasis |
Wnt/β-catenin pathway |
[128] |
ncRAN |
17q25.1 |
Tumor suppressor |
Migration and invasion |
[129] |
|
RP11-462C24.1 |
4q25 |
Tumor suppressor |
Distant metastasis |
[130] |
|
91H |
7F5; 7.87.97cM |
Oncogene |
Distant metastasis, migration, invasion and proliferation |
[131] |
|
MEG3 |
14q.32 |
Tumor suppressor |
Proliferation, invasion and metastasis |
||
lncRNA-LET |
15q24.1 |
Tumor suppressor |
Invasion and metastasis |
[134] |
|
FTX |
Xq13.2 |
Oncogene |
Lymph vascular invasion, proliferation, invasion and migration |
[135] |
|
NEAT1 |
11q13.1 |
Oncogene |
Proliferation, invasion and metastasis |
Akt signaling pathway |
|
DANCR |
4q12 |
Oncogene |
LNM |
[138] |
|
AFAP1-AS1 |
4p16.1 |
Tumor suppressor |
Distant metastasis, proliferation and cell cycle |
[139] |
|
GAS5 |
1q25.1 |
Tumor suppressor |
Distant metastasis, proliferation and cell cycle |
[140] |
|
ANRIL |
9q21.3 |
Oncogene |
Proliferation, invasion and migration |
[141] |
|
ncRuPAR |
5q13.3 |
Tumor suppressor |
LNM and distant metastasis |
[142] |
|
CRNDE-h |
Chr16 |
Oncogene |
LNM and distant metastasis |
[143] |
LncRNAs and escape of apoptosis
The evasion of apoptosis and shear stress in the vasculature leading to arrest at a secondary site is an important step of metastatic process. Tumor cells with the phenotype of anti-apoptosis are more likely to metastasize [23]. Several researches have described the regulation of lncRNAs on cell cycle arrest and apoptosis [24]. The functional relationship of various lncRNAs in the escape of apoptotic cell death are reviewed in Figure 1.
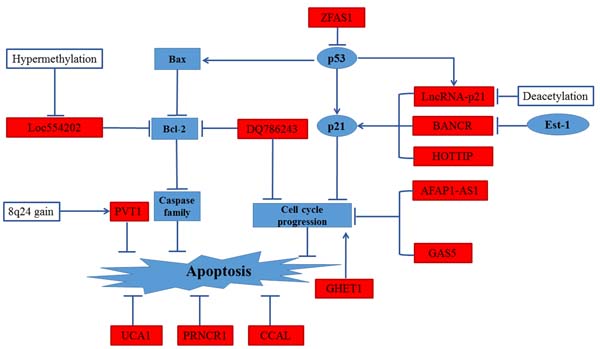
Figure 1: Regulation of apoptosis in CRC by lncRNAs. UCA1, PRNCR1 and CCAL could regulate the apoptosis pathway in CRC. High expression levels of PVT-1 response to 8q24 copy-number gain inhibited apoptosis pathway. Low expression levels of Loc554202 inhibited apoptosis pathway by down-regulation of Bcl-2. DQ786243 down-regulated Bcl-2 expression and led cell cycle arrest, leading to repressing apoptosis pathway. GHE1, GAS5 and AFAP1-AS1 influenced apoptosis by regulating cell cycle progression. HOTTIP modified apoptosis pathway and cell cycle progression by inducing expression of p21. BANCR, regulated by Est-1 mildly effected proliferation by promoting G1 arrest and causing p21 mediated- apoptosis. lincRNA-p21 regulates the G1/S the checkpoint and proliferation by promoting p53-dependent transcription of p21. ZFAS1 may influence cell cycle progress and inhibit apoptosis via destabilization of p53
PVT1, which maps to 8q24, is a novel promising biomarker in different solid cancers including CRC [25]. In CRC, high expression levels of PVT-1 in response to 8q24 copy-number gain showed greater lymph node metastasis (LNM). By gene expression microarray assays on CRC cell lines transfected with PVT-1 siRNA and NC group, it was demonstrated that apoptosis was induced by knockdown of PVT-1 in CRC cells [25]. More recently, another lncRNA DQ786243 was found to be differentially expressed between CRC tissues and adjacent normal tissues. In vitro, knockdown of DQ786243 was proved to inhibit cell proliferation, invasion and migration. In addition, DQ786243 is suggested to be involved in apoptosis and cell cycle progression [26].
Activation of the p53 signal pathway was proposed to play critical roles in both cell cycle arrest and apoptosis. p21, a key downstream effector of p53, was activated through the p53 dependent or independent pathway to inhibit cell proliferation by inducing G0/G1 arrest and apoptosis [27]. By examining BANCR levels in a cohort of 38 CRC patients, Shi et. al demonstrated that BANCR was low expressed in CRC and might be a promising biomarker for prognosis in CRC. In SW480 and HCT116 cells, BANCR mildly effected proliferation by promoting G1 arrest and causing p21 mediated- apoptosis [28]. As an oncogene, HOTTIP promotes cell proliferation, migration and inhibits cell apoptosis in different cancers [29-36]. In CRC, increased HOTTIP expression was supposed to be an unfavorable and independent prognostic factor for its association with clinical stage and distant metastasis [35]. In vitro assay suggested that HOTTIP inhibited G0/G1 arrest and promoted CRC cell growth partly by silencing of p21 expression [37]. Compared to healthy control, lincRNA-p21 was not only decreased in CRC tissues but also in plasma of CRC patients [38, 39]. Besides, numerous evidences revealed that lincRNA-p21 activated p53-mediated apoptosis pathway [40, 41]. To further explore the effects of lincRNA-p21 on the control of expression of p53 target genes, a conditional knockout mouse model was generated. Their results showed that lincRNA-p21 regulates the G1/S the checkpoint and proliferation by promoting p53-dependent transcription of p21 [42]. Recent studies demonstrated that ZFAS1 was identified as an oncogene in CRC. It may influence cell cycle progress and inhibit apoptosis via destabilization of p53 and through interaction with CDK1/cyclin B1 complex [43].
As we all know, caspase cleavage cascades act as the dominant regulator in the death signaling [44]. Due to the hypermethylation, Loc554202 was significantly decreased in human CRC tissues and CRC cell lines compared to controls. After transfection with pCDNA-Loc554202, the number of cells in the S-phase was obviously reduced and the proportions of apoptotic cells was significantly increased. Further studies indicated that activation of specific caspase cleavage cascades was partly responsible for Loc554202-induced CRC cell apoptosis [45]. Besides, UCA1 and PRNCR1 were both reported to inhibit cell cycle and apoptosis [46, 47]. However, UCA1 was pointed out to contribute to apoptosis by suppression of p53 expression in breast cancer cells and cardiomyocytes [48, 49], the exact mechanisms in CRC are still unclear.
LncRNAs and EMT
EMT is one of the underlying driving forces for primary tumor cells to acquire a migratory capacity and metastasis [50]. In addition, EMT property is also linked with a stem cell-like phenotype in invasive, de-differentiated cancer cells by analyzing gene expression patterns in CRC and their corresponding liver metastases [51]. Large numbers of lncRNAs have been reported to regulate EMT progress. Therefore, we summarized the regulation of EMT by lncRNAs in Figure 2.
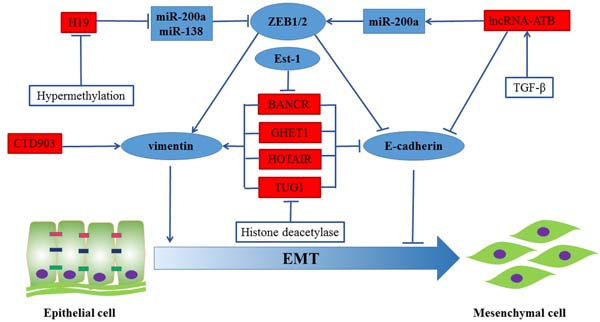
Figure 2: Regulation of EMT in CRC by lncRNAs. BANCR, GHET1, HOTAIR and TUG1 induced EMT phenotypes by repressing the expression of vimentin and promoting the expression of E-cadherin. The expression of long non-coding RNA-activated by TGF-β (lncRNA-ATB) mediated epithelial markers (E-cadherin, ZO-1) repression and increased the expression of mesenchymal markers ZEB1 and N-cadherin through sequestering miR-200a. Besides, H19 significantly promoted EMT progression by functioning as a ceRNA for miR-138 and miR-200a. Moreover, CTD903 induced EMT-like phenotypes.
Overexpression of zinc-finger-enhancer binding protein 1/2 (ZEB1/2), promotion of vimentin transcription and repression of E-cadherin transcription are accompanied by the activation of EMT. Recent study found out that HOTAIR, highly expressed in CRC stem cells, regulated the expression of EMT- associated molecules expression, including E-cadherin and vimentin/N-cadherin [52]. High-BANCR expression group was reported to be associated with more advanced LNM than the low- BANCR expression group. To further understand the internal mechanism, E-cadherin and vimentin were detected in HCT116 cells and Caco-2 cells. The results suggested that BANCR induced EMT phenotypes by repressing the expression of vimentin and promoting the expression of E-cadherin [53]. In vitro study pointed out that TUG1 played a critical role in CRC metastasis by activating EMT progress [54, 55]. The expression of long non-coding RNA-activated by TGF-β (lncRNA-ATB) was higher in metastatic cancer tissues [56]. Besides, lncRNA-ATB mediated epithelial markers (E-cadherin, ZO-1) repression and increased the expression of mesenchymal markers ZEB1 and N-cadherin through sequestering miR-200a [57, 58]. H19 was identified as a novel regulator of EMT in CRC. Multiple studies demonstrated that H19 may act as a competing endogenous RNA (ceRNA) [59-63]. In Liang’s study, it was reported that H19 was highly expressed and significantly promoted EMT progression. It mainly functioned as a ceRNA for miR-138 and miR-200a. to repress vimentin and ZEB2 expression, which were their endogenous targets and core mesenchymal markers [64]. Recently, the correlation of CTD903 expression and lymphatic and distant metastasis was observed in 115 pairs of CRC tissues. After treating with CTD903 siRNA, RKO and SW480 CRC cells exhibited the typical mesenchymal cell morphology, indicating that reduction of CTD903 induced EMT-like phenotypes. Further studies revealed that CTD903 repressed Wnt/β-catenin signal pathway and down-regulated the expression of Twist and Snail, whereas it had no effect on the expression of E-cadherin/N-cadherin and ZEB1 [65]. In addition, GHET1was also demonstrated to be involved in EMT prognosis in CRC. But, how it functions in CRC are still not reported [66]. Recently, SPRY4-IL1 and PANDAR both were proved to promote CRC metastasis via EMT pathway [67-69]. Besides, LINC01133 and SLC25A25-AS1 were tumor suppressors in CRC. Low levels of LINC01133 and SLC25A25-AS1 were suggested to promote EMT in CRC [70, 71]. Downregulation of SCL25A25-AS1 promoted EMT process dependent on Erk and p38 signaling. However, EMT process was regulated by LINC01133 dependent on the presence of SRSF6.
LncRNAs and angiogenesis
It has been demonstrated that tumor progresses are the development of tumor cells from prevascular phase to vascular phase. After the prevascular phage, capillaries were newly formed surrounding the tumor stroma, passed nutrients to it and allow tumor cells to enter into the circulation. Thus, compared to tumors in prevascular phase, vascularized tumors induced angiogenesis, are large in size and have a propensity to metastasize [72]. Angiogenesis is a complex process of new blood vessels formation. The first step is degradation of the extracellular matrix (ECM) and sprouting of the endothelial cells toward the gradient of vascular endothelial growth factor (VEGF). The next step is differentiation of the endothelial cells into tip, stalk and tube cells. Subsequently, it is the tube formation and maturation [73]. More and more experimental and clinical evidences suggest that angiogenesis is a hallmark of tumor metastasis and growth. It is significantly associated with advanced tumor growth and distant metastases in CRC [74]. Therefore, many anti-angiogenic agents were developed, including the VEGFA-targeted antibody bevacizumab [75]. Bevacizumab combined with capecitabine or irinotecan, fluorouracil and leucovorin chemotherapy can extend progression-free and overall-free survival in metastatic colorectal cancer (mCRC) [76-78].
Accumulating evidences proved that lncRNAs played critical roles in angiogenesis [79]. A lncRNA termed MVIH (lncRNAs associated with microvascular invasion in hepatocellular carinoma) was firstly reported to promote tumor growth and metastasis by activating angiogenesis. Using RNA pull-down (RIP) assays and enzyme-linked immunosorbent assays (ELISA) respectively, it was suggested that lncRNA MVIH activated angiogenesis via inhibiting the secretion of PGK1 (phosphoglycerate kinase 1), which is critical for angiogenesis [80]. Subsequently, MALAT1 was also proved to promote angiogenesis in vitro and vivo. The human umbilical vein endothelial cells (HUVECs) transfected with MALAT1 siRNA or GapmeRs directed against MALAT1 showed a significantly higher number of sprouts and more cell migration. Moreover, MALAT1 was observed to activate angiogenesis by regulating vessel density, vessel extension and blood flow recovery in MALAT1-/- mouse model [81, 82]. Recently, Microarray data of gene expression profiles suggested that MALAT1 promoted the secretion of fibroblast growth factor 2 (FGF2) in neuroblastoma cells under hypoxic conditions. It indicated that MALAT1 play a critical role in angiogenesis [83]. In osteoarthritis, MEG3 is found to be inversely associated with VEGF levels, which is involved in angiogenesis [84]. Therefore, we guess that MEG3 may regulate angiogenesis by influence the expression of VEGF. Moreover, HULC was proved to promote angiogenesis in gliomas and liver cancer [85, 86]. In glioma patients’ tissues, HULC was found positively associated with VEGF and microvessel density. Furthermore, the vitro assay results in U87MG and U251 cells showed that HULC silencing suppressed angiogenesis by inhibiting cell proliferation and invasion via PI3K/mTOR signaling pathway [85]. Besides, it was suggested to induce angiogenesis by upregulating sphingosine kinase 1 (SPHK1) [86]. Recently, accumulating lncRNAs, including H19, lincRNA-p21, TUG1 and HOTAIR, was proved to be involved in angiogenesis in different cancers [87-90]. Interestingly, all these lncRNAs have been proved to be involved in CRC metastasis. Therefore, they may affect CRC metastasis partly by regulating angiogenesis. However, there is no report about the functions of lncRNAs on angiogenesis in CRC. More importantly, by using next-generation ribonucleic acid sequencing and microarray assay, Fiedler J et. al found that LINC00313 and MIR503HG were significantly associated with angiogenesis. They also verified their potential clinical importance in an ex vivo model of human induced pluripotent stem cell-based engineered heart tissue [91]. These evidences suggested that lncRNA-based therapeutic strategies hold great promise to modulate tissue vascularization and be used to CRC treatment.
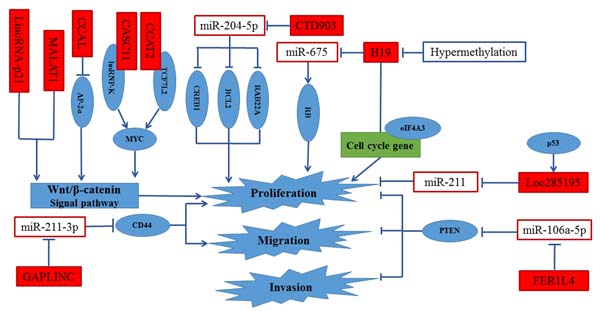
Figure 3: Regulation of proliferation, invasion and migration in CRC by lncRNAs. MALAT1 and lncRNA-p21 were supposed to promote cell proliferation, invasion and migration by activating Wnt/β-catenin signal pathway. CCAL enhanced cell proliferation by activated AP-2α-mediated Wnt/β-catenin signaling. CASC11 and CCAT2 activated Wnt/β-catenin signal pathway by directly targeting hnRNP-K and TCF7L2, respectively. CTD903, Loc285195 and H19 promoted proliferation through sequestering miR-204-5p, miR-211 and miR-675, respectively. Besides, the binding of eIF4A3 to H19 decreased the recruiting of eIF4A3 to the cell-cycle gene mRNA, resulting in the promotion of cell proliferation.
LncRNAs and proliferation, invasion and migration
Numerous studies show that microRNAs (miRNAs) play important roles in CRC metastasis. They are involved in various metastatic pathways of CRC [7]. LncRNAs, identified as the one kind of naturally competing endogenous RNAs (ceRNAs), inhibit the repression of miRNAs targets via competing for binding of the cellular miRNAs [92, 93]. Therefore, accumulating lncRNAs were found to serve as microRNAs sponges to regulate proliferation, invasion and migration in CRC. Urothelial carcinoma associated 1 (UCA1) was suggested to serve as a biomarker of several solid cancers including bladder cancer, breast cancer, gastric cancer and CRC. It was reported that UCA1 was a transcriptional target of C/EBPα, HIF-1α and Ets-2 [94-97] . It was demonstrated to regulate different pathways, including PI3K, AKT, KLF4-KRT6/13, mTOR-STAT3 and p27Kip1/CDK2 signaling [98-101]. Besides, it also was identified as a ceRNA [102, 103]. In CRC cells, UCA1 was proved to enhance cell proliferation by inhibiting the function of miR-204-5p via partly controlling its target genes [104]. By analyzing the expression of 45 pairs CRC tissue samples, a newly identified cancer-related lncRNA, lncRNA-uc002kmd.1(GAPLINC) was considered to be a key regulator of CRC. Their results also proved that lncRNA-uc002kmd.1 formed a molecular decoy for miR-211-3p, which targets CD44 for degradation. The lncRNA-uc002kmd.1-drived CD44-dependent cell growth by competing for miR-211-3p is respond for the cell proliferation and tumor growth in vitro and vivo [105]. LncRNAs are also identified as the precursor of miRNAs [106]. In CRC, H19 was shown to be the precursor of miR-675. H19 was associated with miR-675 expression in CRC cell lines and CRC tissues. Subsequently, the tumor suppressor retinoblastoma (RB) was proved to be a direct target of miR-675 by dual-luciferase reporter gene assay. In vitro assay, H19 promoted CRC cell proliferation by H19/miR-675/RB pathway [107].Besides, the binding of eIF4A3 to H19 decreased the recruiting of eIF4A3 to the cell-cycle gene mRNA, resulting in the promotion of cell proliferation. Interestingly, the results showed that H19 promoted cell proliferation by only regulating cell cycle progression but not apoptosis. [108]. Recently, evidences have suggested that Cancer Susceptibility Candidate 2 (CASC2) serves as an oncogene in CRC. Overexpression of CASC2 inhibits cell proliferation by extending G0/G1-S phase transition. Further studies indicated that CASC2, as a ceRNA by sponging miR-18a, activated the STAT3 signal pathway by modulating the level of the miR-18a targets [109]. Moreover, low expression of LOC285194 was correlated with larger tumor size and more distant metastasis [110]. In vitro and vivo assays, LOC285195, a transcriptional target of p53 was suggested to inhibit cell proliferation and tumor growth by repressing the expression of miR-211 [111]. FER1L4 is involved in various cancers including gastric cancer and colon cancer [112, 113]. In CRC, FER1L4 expression levels exhibited a negative associated with LNM, vascular invasion and depth of tumor invasion. Notably, 86.1% lymph node metastatic tissues showed lower expression of FER1L4 compared with primary cancer tissues. By gain-of- function assays, FER1L4 was revealed to inhibit cell proliferation, migration and invasion.
Wnt/β-catenin signal pathway plays a critical role in colorectal carcinogenesis. It controls not only EMT but also cell proliferation, invasion and migration in CRC [114]. As a tumor suppressor, LincRNA-p21 has been reported to inhibit the translation of β-catenin in Hela cells [115]. By detecting the expression of lincRNA-p21, β-catenin and Wnt/β-catenin target genes in 30 CRC tissues and its adjacent tissues, an inverse correlation between lincRNA-p21 and activity of β-catenin was found. Further studies demonstrated that lincRNA-p21 significantly repressed the Wnt/β-catenin signal pathway [116]. Subsequently, the results were verified in stem-like CRC tissues and cells [117]. By globally analyzing the protein-coding RNA and lncRNA expression profiles of normal colorectal tissues, colorectal adenoma and CRC tissues, a non-annotated lncRNA, CCAL, was emerged as oncogene. Using qRT-PCR analysis in 252 CRC tumor and paired non-tumor tissues and CRC cells, Ma et al revealed that CCAL overexpression was a frequent event no matter in CRC tissues and cells. In vitro and vivo assay, CCAL was suggested to be involved in several biological functions, including cell proliferation, invasion, migration, apoptosis and tumorigenesis. Further studies demonstrated that CCAL enhanced cell proliferation by activated AP-2α-mediated Wnt/β-catenin signaling [118]. Numerous studies have demonstrated that Multiple genes, located at 8q24, are often amplified and involved in metastasis in CRC [119]. Until now, six lncRNAs (PVT-1, PCAT-1, PRNCR1, CASC11, CCAT1 and CCAT2), all mapping to 8q24, were reported to overexpress in CRC [25, 47, 120-123]. By RNA-binding protein immunoprecipitation (RIP) experiments and western blot assays, CASC11 and CCAT2 activated Wnt/β-catenin signal pathway by directly targeting hnRNP-K and TCF7L2, respectively [121, 122]. Besides, MALAT1 was suggested to be a prognostic biomarker in in stage II/III CRC patients [124]. In LoVo cells, MALAT1 was supposed to promote cell proliferation, invasion and migration by increasing the nuclear localization of β-catenin, resulting in activating Wnt/β-catenin signal pathway [125]. More than Wnt/β-catenin signal pathway activation, it is reported that MALAT1 promotes cell proliferation, invasion and migration not only via PRKA kinase anchor protein 9 but also through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex [126, 127]. Recently, TINCR was proved to correlate with CRC proliferation and metastasis in vivo and vitro. The potential mechanisms was that loss of TINCR enhanced hydrolysis of EpCAM, subsequently, activated Wnt/β-catenin signal pathway [128]. Besides, several other lncRNAs were reported to participate in the prognosis of cell proliferation, invasion and migration in CRC [129-143]. However, the potential mechanisms are still unclear.
Future perspectives and clinical application
As discussed in above, lncRNAs are involved in different biological progressions in CRC. Moreover, similar to other nucleic acids, lncRNAs can also be detected in peripheral blood, such as serum, plasma, and peripheral blood mononuclear cells. Therefore, it likely that circulating lncRNAs may be new non-invasive molecular markers for tumor diagnosis [144, 145]. We list some lncRNAs with potential prognostic value for CRC diagnosis and therapy in Table 2. Recent studies showed that the combination of two exosomal mRNAs, KRTAP5-4 and MAGEA3 and one exosomal lncRNA, BCAR4 in serum could be potential candidates to detected CRC [146]. Moreover, by detecting 13 lncRNAs in serum samples from 71 CRC patients and 70 healthy individuals, it was suggested that a three- lncRNA signature (LOC285194, rp11-462C24.1 and Nbla12061) show potential as a diagnostic marker for CRC. More importantly, it showed much higher diagnostic ability than conventional blood biomarkers 0.793 (95% CI: 0.709 to 0.861), such as carcinoembryonic antigen (CEA), carbohydrate antigen 199 (CA199), carbohydrate antigen 125 (CA125) and carbohydrate antigen 724 (CA724) [147]. However, lncRNA-based tumor diagnostics has not been developed for use yet.
Table 2: LncRNA with potential biomarkers for CRC diagnosis and prognosis
LncRNAs |
Origin |
n |
AUC |
95%Cl |
Sensitivity(%) |
specificity(%) |
Ref. |
BANCR, NR_026817, NR_029373 and NR_034119 |
Serum |
120 |
0.881 |
0.833.919 |
89.17 |
75.83 |
[169] |
PRNCR1 |
Tissue |
63 |
0.799 |
80.4 |
70.0 |
[47] |
|
CCAT1&HOTAIR |
Plasma |
32 |
0.954 |
0.903-1.000 |
84.3 |
80.2 |
[38] |
LOC285194, RP11-462C24.1 and Nbla12061 |
Serum |
70 |
0.793 |
0.709-0.861 |
68.33 |
86.89 |
[170] |
MEG3 (diagnosed for CLM) |
Tissue |
51 |
0.62 |
0.48-0.74 |
[171] |
||
GAS5 ( diagnosed for CLM ) |
Tissue |
51 |
0.65 |
0.51-0.77 |
[171] |
||
H19( diagnosed for CLM ) |
Tissue |
51 |
0.56 |
0.43-0.69 |
[171] |
||
Yiya( diagnosed for CLM) |
Tissue |
51 |
0.70 |
0.56-0.81 |
[171] |
||
NEAT1_1 |
Blood |
30 |
0.732 |
0.724-0.842 |
56.7 |
83.3 |
[172] |
NEAT1_2 |
Blood |
30 |
0.845 |
0.816-0.914 |
86.6 |
83.3 |
[172] |
CRNDE-h |
Plasma |
15 |
0.888 |
87 |
93 |
[173] |
|
BCAR4 and 2 mRNA |
Exsome |
30 |
0.936 |
0.840-0.983 |
[146] |
Because of the critical roles of lncRNAs in cancer, modulation of lncRNAs expression seems to have vast potential in developing lncRNA-based cancer therapy. Based on viral and non-viral vectors, several efficient delivery systems to alter expression of lncRNA have been developed. Due to the serious side effects, non-viral inhibition of lncRNA is more practical in clinic. Non-viral inhibition of lncRNA is performed either with small molecule inhibitors or oligonucleotide -based therapeutics (antisense oligonucleotides and RNAi mediated gene silencing) [148]. Small molecule inhibitors are used to block the binding sites or change the secondary structure of the lncRNA, resulting in disrupting the interactions between lncRNAs with proteins or nucleic acids. However, there is still a huge challenge and more efforts are needed [149].
Currently, RNA interference (RNAi) and antisense oligonucleotides (ASOs) are another two applicable oligonucleotide based therapeutic approaches. RNAi is a specific, safe and cost-effective way to mimic the natural way of gene silencing by using synthetic siRNAs [150]. It has demonstrated that exogenous siRNAs are able to deplete lncRNA molecules within cytoplasm and nucleus [151]. Surprisingly, more evidences suggested that knockdown of some lncRNAs including HOTAIR and MALAT1 by siRNA induced significant anticancer effect in vivo and vitro [152, 153]. Until now, several formulation of siRNAs are undergoing different phase of clinical trials. However, all the siRNAs are targeting mRNAs not lncRNAs. Moreover, how to achieve successful siRNA delivery in order to make it stable in circulation, reaching the target tissue accurately, entering the cancer cells and load into the RISC complex successfully is still the main obstacle. Even through several strategies such as modifications of the siRNA molecules, the use of nanoparticle and lipid-based delivery tools can partly solve these issues, Obviously, the therapy using siRNAs targeting critical lncRNAs is still in the infancy stage and needs a long way to go. ASOs are short (13-25 nt) single stranded oligonucleotides complementary to the target RNA. They are used to down-regulate lncRNA in the nucleus by blocking lncRNAs and inducing lncRNAs degradation by nucleases [154]. Compared with RNAi, ASOs can not only knockdown the targeting lncRNAs effectively in vitro and vivo, but also show superiority over siRNAs in safety. Notoriously, lncRNAs can serve as endogenous microRNA sponge to reduce its activity. Similarly, evidences showed that microRNAs may naturally regulate lncRNAs expression vice versa [155].
Except RNAi, ASOs and microRNAs, hammerhead ribozyme (HanRz) and aptamer also show good inhibitory effect. However, some lncRNAs may be virtually undruggable, because of their low transcript abundance. Meanwhile, all these pharmaceutical technologies are aimed at the oncogenic lncRNAs. The important roles of other lncRNAs served as tumor suppressors in cancer should not be overlooked.
Conclusions
Evidence is accumulating that lncRNAs play critical roles in cancer progression and metastasis. In this review, we summarized the dysregulated lncRNAs associated with CRC metastasis. By microarray analysis, lncRNA expression profiles in primary tumors and metastatic tissues including MLN tissues and CLM tissues were compared [17-20, 22]. All these results supposed that lncRNAs were associated with the metastatic phenotype in CRC patients. However, the data are limited and microarray analysis on colorectal tumor at multi-treatment center, of different subtypes and with large sample size should be performed. Besides, we described the association of lncRNAs and metastatic pathways in CRC. Accumulating studies demonstrated that lncRNAs have a great influence on different biological progresses including apoptosis, cell cycle arrest, EMT, proliferation, invasion and migration. Except that, angiogenesis is essential for tumor growth and an important factor in the metastatic pathway [156, 157]. Accumulating lncRNAs were suggested to promote angiogenesis in liver cancer, non-small cell lung cancer, hepatocellular carcinoma and glioblastoma. However, the association between angiogenesis and lncRNAs in CRC are still not reported yet. Recently, more evidences indicated that angiogenesis may not be necessary for carcinoma metastases [158, 159], whereas co-opting host vessels is the essential choice for tumor cells survive [160]. Surprisingly, vessel co-option may respond for approximately 40% metastases in mCRC [161]. Moreover, evidences showed that the RAS/RAF/MAPK pathway and PI3K/AKT pathway suppresses invasion and promotes EMT, respectively [162, 163]. Therefore, the activation of these pathways might be associated with the phenotype of metastasis in CRC.
Recent studies have demonstrated that genomic variations in lncRNAs, especially in microRNA response elements (MREs) contributed to the risk and progress of CRC [164, 165]. These SNPs may also be associated with CRC metastasis. Because, they may lead to either complete or partial loss of miRNA mediated lncRNAs degradation, resulting in increased lncRNA expression. Moreover, lncRNAs in plasma were also proved to serve as prognostic factor [38, 166]. However, there is no report about the different expression of lncRNAs between early stage CRC patients and mCRC patients. These studies have shed lights on the future research direction on lncRNAs in CRC metastasis. In conclusion, current studies suggest the vital role of lncRNAs in the process of CRC metastasis. More studies with a large cohort of metastatic CRC patients should be performed in the future.
Acknowledgments and Funding
This study was funded by National Nature Science Foundation of China (81373489, 81300204, 81422052 and 81302850), National Science and Technology Major Project (2013ZX09509-107), Hunan Provincial Natural Science Foundation of China (13JJ1010) and Funds for Hunan education department program (12K006)
Conflicts of interest
All authors declare no conflict of interest. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
References
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65: 87-108. doi: 10.3322/caac.21262.
2. Schmidt-Kittler O, Ragg T, Daskalakis A, Granzow M, Ahr A, Blankenstein TJ, Kaufmann M, Diebold J, Arnholdt H, Muller P, Bischoff J, Harich D, Schlimok G, et al. From latent disseminated cells to overt metastasis: genetic analysis of systemic breast cancer progression. Proc Natl Acad Sci U S A. 2003; 100: 7737-42. doi: 10.1073/pnas.1331931100.
3. Stein U, Schlag PM. Clinical, biological, and molecular aspects of metastasis in colorectal cancer. Recent Results Cancer Res. 2007; 176: 61-80.
4. Fidler IJ. The pathogenesis of cancer metastasis: the ‘seed and soil’ hypothesis revisited. Nat Rev Cancer. 2003; 3: 453-8. doi: 10.1038/nrc1098.
5. Grothey A, Schmoll HJ. New chemotherapy approaches in colorectal cancer. Curr Opin Oncol. 2001; 13: 275-86.
6. Hunter KW, Crawford NP, Alsarraj J. Mechanisms of metastasis. Breast Cancer Res. 2008; 10 Suppl 1: S2. doi: 10.1186/bcr1988.
7. de Krijger I, Mekenkamp LJ, Punt CJ, Nagtegaal ID. MicroRNAs in colorectal cancer metastasis. J Pathol. 2011; 224: 438-47. doi: 10.1002/path.2922.
8. Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012; 22: 1775-89. doi: 10.1101/gr.132159.111.
9. Jia H, Osak M, Bogu GK, Stanton LW, Johnson R, Lipovich L. Genome-wide computational identification and manual annotation of human long noncoding RNA genes. Rna. 2010; 16: 1478-87. doi: 10.1261/rna.1951310.
10. Han D, Wang M, Ma N, Xu Y, Jiang Y, Gao X. Long noncoding RNAs: novel players in colorectal cancer. Cancer Lett. 2015; 361: 13-21. doi: 10.1016/j.canlet.2015.03.002.
11. Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014; 5: e944014. doi: 10.4161/21541272.2014.944014.
12. Yang ZG, Gao L, Guo XB, Shi YL. Roles of long non-coding RNAs in gastric cancer metastasis. World J Gastroenterol. 2015; 21: 5220-30. doi: 10.3748/wjg.v21.i17.5220.
13. Wu T, Yin X, Zhou Y, Wang Z, Shen S, Qiu Y, Sun R, Zhao Z. Roles of noncoding RNAs in metastasis of nonsmall cell lung cancer: A mini review. J Cancer Res Ther. 2015; 11 Suppl 1: C7-10. doi: 10.4103/0973-1482.163831.
14. Li G, Zhang H, Wan X, Yang X, Zhu C, Wang A, He L, Miao R, Chen S, Zhao H. Long noncoding RNA plays a key role in metastasis and prognosis of hepatocellular carcinoma. Biomed Res Int. 2014; 2014: 780521. doi: 10.1155/2014/780521.
15. Xie X, Tang B, Xiao YF, Xie R, Li BS, Dong H, Zhou JY, Yang SM. Long non-coding RNAs in colorectal cancer. Oncotarget. 2016; 7: 5226-39. doi: 10.18632/oncotarget.6446.
16. Shen XH, Qi P, Du X. Long non-coding RNAs in cancer invasion and metastasis. Mod Pathol. 2015; 28: 4-13. doi: 10.1038/modpathol.2014.75.
17. Han J, Rong LF, Shi CB, Dong XG, Wang J, Wang BL, Wen H, He ZY. Screening of lymph nodes metastasis associated lncRNAs in colorectal cancer patients. World J Gastroenterol. 2014; 20: 8139-50. doi: 10.3748/wjg.v20.i25.8139.
18. Yang P, Xu ZP, Chen T, He ZY. Long noncoding RNA expression profile analysis of colorectal cancer and metastatic lymph node based on microarray data. Onco Targets Ther. 2016; 9: 2465-78. doi: 10.2147/ott.s102348.
19. Ye LC, Ren L, Qiu JJ, Zhu DX, Chen T, Chang WJ, Lv SX, Xu J. Aberrant expression of long noncoding RNAs in colorectal cancer with liver metastasis. Tumour Biol. 2015; 36: 8747-54. doi: 10.1007/s13277-015-3627-4.
20. Chen D, Sun Q, Cheng X, Zhang L, Song W, Zhou D, Lin J, Wang W. Genome-wide analysis of long noncoding RNA (lncRNA) expression in colorectal cancer tissues from patients with liver metastasis. Cancer Med. 2016. doi: 10.1002/cam4.738.
21. Chen N, Guo D, Xu Q, Yang M, Wang D, Peng M, Ding Y, Wang S, Zhou J. Long non-coding RNA FEZF1-AS1 facilitates cell proliferation and migration in colorectal carcinoma. Oncotarget. 2016; 7: 11271-83. doi: 10.18632/oncotarget.7168.
22. Yang P, Chen T, Xu Z, Zhu H, Wang J, He Z. Long noncoding RNA GAPLINC promotes invasion in colorectal cancer by targeting SNAI2 through binding with PSF and NONO. Oncotarget. 2016; 7:42183-42194. doi: 10.18632/oncotarget.9741.
23. Celia-Terrassa T, Kang Y. Distinctive properties of metastasis-initiating cells. Genes Dev. 2016; 30: 892-908. doi: 10.1101/gad.277681.116.
24. Wapinski O, Chang HY. Long noncoding RNAs and human disease. Trends Cell Biol. 2011; 21: 354-61. doi: 10.1016/j.tcb.2011.04.001.
25. Takahashi Y, Sawada G, Kurashige J, Uchi R, Matsumura T, Ueo H, Takano Y, Eguchi H, Sudo T, Sugimachi K, Yamamoto H, Doki Y, Mori M, et al. Amplification of PVT-1 is involved in poor prognosis via apoptosis inhibition in colorectal cancers. Br J Cancer. 2014; 110: 164-71. doi: 10.1038/bjc.2013.698.
26. Sun L, Xue H, Jiang C, Zhou H, Gu L, Liu Y, Xu C, Xu Q. LncRNA DQ786243 contributes to proliferation and metastasis of colorectal cancer both in vitro and in vivo. Biosci Rep. 2016; 36. doi: 10.1042/bsr20160048.
27. Mirzayans R, Andrais B, Scott A, Murray D. New insights into p53 signaling and cancer cell response to DNA damage: implications for cancer therapy. J Biomed Biotechnol. 2012; 2012: 170325. doi: 10.1155/2012/170325.
28. Shi Y, Liu Y, Wang J, Jie D, Yun T, Li W, Yan L, Wang K, Feng J. Downregulated Long Noncoding RNA BANCR Promotes the Proliferation of Colorectal Cancer Cells via Downregualtion of p21 Expression. PLoS One. 2015; 10: e0122679. doi: 10.1371/journal.pone.0122679.
29. Cheng Y, Jutooru I, Chadalapaka G, Corton JC, Safe S. The long non-coding RNA HOTTIP enhances pancreatic cancer cell proliferation, survival and migration. Oncotarget. 2015; 6: 10840-52. doi: 10.18632/oncotarget.3450.
30. Zhang H, Zhao L, Wang YX, Xi M, Liu SL, Luo LL. Long non-coding RNA HOTTIP is correlated with progression and prognosis in tongue squamous cell carcinoma. Tumour Biol. 2015; 36: 8805-9. doi: 10.1007/s13277-015-3645-2.
31. Deng HP, Chen L, Fan T, Zhang B, Xu Y, Geng Q. Long non-coding RNA HOTTIP promotes tumor growth and inhibits cell apoptosis in lung cancer. Cell Mol Biol (Noisy-le-grand). 2015; 61: 34-40.
32. Li F, Cao L, Hang D, Wang F, Wang Q. Long non-coding RNA HOTTIP is up-regulated and associated with poor prognosis in patients with osteosarcoma. Int J Clin Exp Pathol. 2015; 8: 11414-20.
33. Zhang SR, Yang JK, Xie JK, Zhao LC. Long noncoding RNA HOTTIP contributes to the progression of prostate cancer by regulating HOXA13. Cell Mol Biol (Noisy-le-grand). 2016; 62: 84-8.
34. Ye H, Liu K, Qian K. Overexpression of long noncoding RNA HOTTIP promotes tumor invasion and predicts poor prognosis in gastric cancer. Onco Targets Ther. 2016; 9: 2081-8. doi: 10.2147/ott.s95414.
35. Ren YK, Xiao Y, Wan XB, Zhao YZ, Li J, Li Y, Han GS, Chen XB, Zou QY, Wang GC, Lu CM, Xu YC, Wang YC. Association of long non-coding RNA HOTTIP with progression and prognosis in colorectal cancer. Int J Clin Exp Pathol. 2015; 8: 11458-63.
36. Lian Y, Ding J, Zhang Z, Shi Y, Zhu Y, Li J, Peng P, Wang J, Fan Y, De W, Wang K. The long noncoding RNA HOXA transcript at the distal tip promotes colorectal cancer growth partially via silencing of p21 expression. Tumour Biol. 2016; 37: 7431-40. doi: 10.1007/s13277-015-4617-2.
37. Lian Y, Ding J, Zhang Z, Shi Y, Zhu Y, Li J, Peng P, Wang J, Fan Y, De W, Wang K. The long noncoding RNA HOXA transcript at the distal tip promotes colorectal cancer growth partially via silencing of p21 expression. Tumour Biol. 2015. doi: 10.1007/s13277-015-4617-2.
38. Zhao W, Song M, Zhang J, Kuerban M, Wang H. Combined identification of long non-coding RNA CCAT1 and HOTAIR in serum as an effective screening for colorectal carcinoma. Int J Clin Exp Pathol. 2015; 8: 14131-40.
39. Zhai H, Fesler A, Schee K, Fodstad O, Flatmark K, Ju J. Clinical significance of long intergenic noncoding RNA-p21 in colorectal cancer. Clin Colorectal Cancer. 2013; 12: 261-6. doi: 10.1016/j.clcc.2013.06.003.
40. Hall JR, Messenger ZJ, Tam HW, Phillips SL, Recio L, Smart RC. Long noncoding RNA lincRNA-p21 is the major mediator of UVB-induced and p53-dependent apoptosis in keratinocytes. Cell Death Dis. 2015; 6: e1700. doi: 10.1038/cddis.2015.67.
41. Wu G, Cai J, Han Y, Chen J, Huang ZP, Chen C, Cai Y, Huang H, Yang Y, Liu Y, Xu Z, He D, Zhang X, et al. LincRNA-p21 regulates neointima formation, vascular smooth muscle cell proliferation, apoptosis, and atherosclerosis by enhancing p53 activity. Circulation. 2014; 130: 1452-65. doi: 10.1161/circulationaha.114.011675.
42. Dimitrova N, Zamudio JR, Jong RM, Soukup D, Resnick R, Sarma K, Ward AJ, Raj A, Lee JT, Sharp PA, Jacks T. LincRNA-p21 activates p21 in cis to promote Polycomb target gene expression and to enforce the G1/S checkpoint. Mol Cell. 2014; 54: 777-90. doi: 10.1016/j.molcel.2014.04.025.
43. Thorenoor N, Faltejskova-Vychytilova P, Hombach S, Mlcochova J, Kretz M, Svoboda M, Slaby O. Long non-coding RNA ZFAS1 interacts with CDK1 and is involved in p53-dependent cell cycle control and apoptosis in colorectal cancer. Oncotarget. 2016; 7: 622-37. doi: 10.18632/oncotarget.5807.
44. Suzuki A, Tsutomi Y, Yamamoto N, Shibutani T, Akahane K. Mitochondrial regulation of cell death: mitochondria are essential for procaspase 3-p21 complex formation to resist Fas-mediated cell death. Mol Cell Biol. 1999; 19: 3842-7.
45. Ding J, Lu B, Wang J, Wang J, Shi Y, Lian Y, Zhu Y, Wang J, Fan Y, Wang Z, De W, Wang K. Long non-coding RNA Loc554202 induces apoptosis in colorectal cancer cells via the caspase cleavage cascades. J Exp Clin Cancer Res. 2015; 34: 100. doi: 10.1186/s13046-015-0217-7.
46. Han Y, Yang YN, Yuan HH, Zhang TT, Sui H, Wei XL, Liu L, Huang P, Zhang WJ, Bai YX. UCA1, a long non-coding RNA up-regulated in colorectal cancer influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014; 46: 396-401. doi: 10.1097/pat.0000000000000125.
47. Yang L, Qiu M, Xu Y, Wang J, Zheng Y, Li M, Xu L, Yin R. Upregulation of long non-coding RNA PRNCR1 in colorectal cancer promotes cell proliferation and cell cycle progression. Oncol Rep. 2016; 35: 318-24. doi: 10.3892/or.2015.4364.
48. Huang J, Zhou N, Watabe K, Lu Z, Wu F, Xu M, Mo YY. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014; 5: e1008. doi: 10.1038/cddis.2013.541.
49. Liu Y, Zhou D, Li G, Ming X, Tu Y, Tian J, Lu H, Yu B. Long non coding RNA-UCA1 contributes to cardiomyocyte apoptosis by suppression of p27 expression. Cell Physiol Biochem. 2015; 35: 1986-98. doi: 10.1159/000374006.
50. Cao H, Xu E, Liu H, Wan L, Lai M. Epithelial-mesenchymal transition in colorectal cancer metastasis: A system review. Pathol Res Pract. 2015; 211: 557-69. doi: 10.1016/j.prp.2015.05.010.
51. Brabletz T. To differentiate or not—routes towards metastasis. Nat Rev Cancer. 2012; 12: 425-36. doi: 10.1038/nrc3265.
52. Dou J, Ni Y, He X, Wu D, Li M, Wu S, Zhang R, Guo M, Zhao F. Decreasing lncRNA HOTAIR expression inhibits human colorectal cancer stem cells. Am J Transl Res. 2016; 8: 98-108.
53. Guo Q, Zhao Y, Chen J, Hu J, Wang S, Zhang D, Sun Y. BRAF-activated long non-coding RNA contributes to colorectal cancer migration by inducing epithelial-mesenchymal transition. Oncol Lett. 2014; 8: 869-75. doi: 10.3892/ol.2014.2154.
54. Sun J, Ding C, Yang Z, Liu T, Zhang X, Zhao C, Wang J. The long non-coding RNA TUG1 indicates a poor prognosis for colorectal cancer and promotes metastasis by affecting epithelial-mesenchymal transition. J Transl Med. 2016; 14: 42. doi: 10.1186/s12967-016-0786-z.
55. Wang L, Zhao Z, Feng W, Ye Z, Dai W, Zhang C, Peng J, Wu K. Long non-coding RNA TUG1 promotes colorectal cancer metastasis via EMT pathway. Oncotarget. 2016; 7:51713-51719. doi: 10.18632/oncotarget.10563.
56. Iguchi T, Uchi R, Nambara S, Saito T, Komatsu H, Hirata H, Ueda M, Sakimura S, Takano Y, Kurashige J, Shinden Y, Eguchi H, Sugimachi K, et al. A long noncoding RNA, lncRNA-ATB, is involved in the progression and prognosis of colorectal cancer. Anticancer Res. 2015; 35: 1385-8.
57. Yue B, Qiu S, Zhao S, Liu C, Zhang D, Yu F, Peng Z, Yan D. LncRNA-ATB mediated E-cadherin repression promotes the progression of colon cancer and predicts poor prognosis. J Gastroenterol Hepatol. 2016; 31: 595-603. doi: 10.1111/jgh.13206.
58. Li W, Kang Y. A new Lnc in metastasis: long noncoding RNA mediates the prometastatic functions of TGF-beta. Cancer Cell. 2014; 25: 557-9. doi: 10.1016/j.ccr.2014.04.014.
59. Li H, Yu B, Li J, Su L, Yan M, Zhu Z, Liu B. Overexpression of lncRNA H19 enhances carcinogenesis and metastasis of gastric cancer. Oncotarget. 2014; 5: 2318-29. doi: 10.18632/oncotarget.1913.
60. Zhu M, Chen Q, Liu X, Sun Q, Zhao X, Deng R, Wang Y, Huang J, Xu M, Yan J, Yu J. lncRNA H19/miR-675 axis represses prostate cancer metastasis by targeting TGFBI. Febs j. 2014; 281: 3766-75. doi: 10.1111/febs.12902.
61. Yan L, Zhou J, Gao Y, Ghazal S, Lu L, Bellone S, Yang Y, Liu N, Zhao X, Santin AD, Taylor H, Huang Y. Regulation of tumor cell migration and invasion by the H19/let-7 axis is antagonized by metformin-induced DNA methylation. Oncogene. 2015; 34: 3076-84. doi: 10.1038/onc.2014.236.
62. Liang WC, Fu WM, Wang YB, Sun YX, Xu LL, Wong CW, Chan KM, Li G, Waye MM, Zhang JF. H19 activates Wnt signaling and promotes osteoblast differentiation by functioning as a competing endogenous RNA. Sci Rep. 2016; 6: 20121. doi: 10.1038/srep20121.
63. Su Z, Zhi X, Zhang Q, Yang L, Xu H, Xu Z. LncRNA H19 functions as a competing endogenous RNA to regulate AQP3 expression by sponging miR-874 in the intestinal barrier. FEBS Lett. 2016; 590: 1354-64. doi: 10.1002/1873-3468.12171.
64. Liang WC, Fu WM, Wong CW, Wang Y, Wang WM, Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF, Waye MM. The lncRNA H19 promotes epithelial to mesenchymal transition by functioning as miRNA sponges in colorectal cancer. Oncotarget. 2015; 6: 22513-25. doi: 10.18632/oncotarget.4154.
65. Yuan Z, Yu X, Ni B, Chen D, Yang Z, Huang J, Wang J, Chen D, Wang L. Overexpression of long non-coding RNA-CTD903 inhibits colorectal cancer invasion and migration by repressing Wnt/beta-catenin signaling and predicts favorable prognosis. Int J Oncol. 2016; 48: 2675-85. doi: 10.3892/ijo.2016.3447.
66. Zhou J, Li X, Wu M, Lin C, Guo Y, Tian B. Knockdown of Long Noncoding RNA GHET1 Inhibits Cell Proliferation and Invasion of Colorectal Cancer. Oncol Res. 2016; 23: 303-9. doi: 10.3727/096504016x14567549091305.
67. Shen F, Cai WS, Feng Z, Chen JW, Feng JH, Liu QC, Fang YP, Li KP, Xiao HQ, Cao J, Xu B. Long non-coding RNA SPRY4-IT1 pormotes colorectal cancer metastasis by regulate epithelial-mesenchymal transition. Oncotarget. 2017; 8:14479-14486. doi: 10.18632/oncotarget.10407.
68. Cao D, Ding Q, Yu W, Gao M, Wang Y. Long noncoding RNA SPRY4-IT1 promotes malignant development of colorectal cancer by targeting epithelial-mesenchymal transition. Onco Targets Ther. 2016; 9: 5417-25. doi: 10.2147/ott.s111794.
69. Lu M, Liu Z, Li B, Wang G, Li D, Zhu Y. The high expression of long non-coding RNA PANDAR indicates a poor prognosis for colorectal cancer and promotes metastasis by EMT pathway. J Cancer Res Clin Oncol. 2016. doi: 10.1007/s00432-016-2252-y.
70. Kong J, Sun W, Li C, Wan L, Wang S, Wu Y, Xu E, Zhang H, Lai M. Long non-coding RNA LINC01133 inhibits epithelial-mesenchymal transition and metastasis in colorectal cancer by interacting with SRSF6. Cancer Lett. 2016; 380: 476-84. doi: 10.1016/j.canlet.2016.07.015.
71. Li Y, Huang S, Li Y, Zhang W, He K, Zhao M, Lin H, Li D, Zhang H, Zheng Z, Huang C. Decreased expression of LncRNA SLC25A25-AS1 promotes proliferation, chemoresistance, and EMT in colorectal cancer cells. Tumour Biol. 2016; 37: 14205-15. doi: 10.1007/s13277-016-5254-0.
72. Folkman J. The role of angiogenesis in tumor growth. Semin Cancer Biol. 1992; 3: 65-71.
73. Dimova I, Popivanov G, Djonov V. Angiogenesis in cancer - general pathways and their therapeutic implications. J buon. 2014; 19: 15-21.
74. Oklu R, Walker TG, Wicky S, Hesketh R. Angiogenesis and current antiangiogenic strategies for the treatment of cancer. J Vasc Interv Radiol. 2010; 21: 1791-805; quiz 806. doi: 10.1016/j.jvir.2010.08.009.
75. Ferrara N, Hillan KJ, Gerber HP, Novotny W. Discovery and development of bevacizumab, an anti-VEGF antibody for treating cancer. Nat Rev Drug Discov. 2004; 3: 391-400. doi: 10.1038/nrd1381.
76. Hurwitz H, Fehrenbacher L, Novotny W, Cartwright T, Hainsworth J, Heim W, Berlin J, Baron A, Griffing S, Holmgren E, Ferrara N, Fyfe G, Rogers B, et al. Bevacizumab plus irinotecan, fluorouracil, and leucovorin for metastatic colorectal cancer. N Engl J Med. 2004; 350: 2335-42. doi: 10.1056/NEJMoa032691.
77. Cunningham D, Lang I, Marcuello E, Lorusso V, Ocvirk J, Shin DB, Jonker D, Osborne S, Andre N, Waterkamp D, Saunders MP. Bevacizumab plus capecitabine versus capecitabine alone in elderly patients with previously untreated metastatic colorectal cancer (AVEX): an open-label, randomised phase 3 trial. Lancet Oncol. 2013; 14: 1077-85. doi: 10.1016/s1470-2045(13)70154-2.
78. Simkens LH, van Tinteren H, May A, ten Tije AJ, Creemers GJ, Loosveld OJ, de Jongh FE, Erdkamp FL, Erjavec Z, van der Torren AM, Tol J, Braun HJ, Nieboer P, et al. Maintenance treatment with capecitabine and bevacizumab in metastatic colorectal cancer (CAIRO3): a phase 3 randomised controlled trial of the Dutch Colorectal Cancer Group. Lancet. 2015; 385: 1843-52. doi: 10.1016/s0140-6736(14)62004-3.
79. Khorshidi A, Dhaliwal P, Yang BB. Noncoding RNAs in Tumor Angiogenesis. Adv Exp Med Biol. 2016; 927: 217-41. doi: 10.1007/978-981-10-1498-7_8.
80. Yuan SX, Yang F, Yang Y, Tao QF, Zhang J, Huang G, Yang Y, Wang RY, Yang S, Huo XS, Zhang L, Wang F, Sun SH, et al. Long noncoding RNA associated with microvascular invasion in hepatocellular carcinoma promotes angiogenesis and serves as a predictor for hepatocellular carcinoma patients’ poor recurrence-free survival after hepatectomy. Hepatology. 2012; 56: 2231-41. doi: 10.1002/hep.25895.
81. Michalik KM, You X, Manavski Y, Doddaballapur A, Zornig M, Braun T, John D, Ponomareva Y, Chen W, Uchida S, Boon RA, Dimmeler S. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ Res. 2014; 114: 1389-97. doi: 10.1161/circresaha.114.303265.
82. Thum T, Fiedler J. LINCing MALAT1 and angiogenesis. Circ Res. 2014; 114: 1366-8. doi: 10.1161/circresaha.114.303896.
83. Tee AE, Liu B, Song R, Li J, Pasquier E, Cheung BB, Jiang C, Marshall GM, Haber M, Norris MD, Fletcher JI, Dinger ME, Liu T. The long noncoding RNA MALAT1 promotes tumor-driven angiogenesis by up-regulating pro-angiogenic gene expression. Oncotarget. 2016; 7: 8663-75. doi: 10.18632/oncotarget.6675.
84. Su W, Xie W, Shang Q, Su B. The Long Noncoding RNA MEG3 Is Downregulated and Inversely Associated with VEGF Levels in Osteoarthritis. 2015; 2015: 356893. doi: 10.1155/2015/356893.
85. Zhu Y, Zhang X, Qi L, Cai Y, Yang P, Xuan G, Jiang Y. HULC long noncoding RNA silencing suppresses angiogenesis by regulating ESM-1 via the PI3K/Akt/mTOR signaling pathway in human gliomas. Oncotarget. 2016; 7: 14429-40. doi: 10.18632/oncotarget.7418.
86. Lu Z, Xiao Z, Liu F, Cui M, Li W, Yang Z, Li J, Ye L, Zhang X. Long non-coding RNA HULC promotes tumor angiogenesis in liver cancer by up-regulating sphingosine kinase 1 (SPHK1). Oncotarget. 2016; 7: 241-54. doi: 10.18632/oncotarget.6280.
87. Jiang X, Yan Y, Hu M, Chen X, Wang Y, Dai Y, Wu D, Wang Y, Zhuang Z, Xia H. Increased level of H19 long noncoding RNA promotes invasion, angiogenesis, and stemness of glioblastoma cells. J Neurosurg. 2016; 124: 129-36. doi: 10.3171/2014.12.jns1426.
88. Castellano JJ, Navarro A, Vinolas N, Marrades RM, Moises J, Cordeiro A, Saco A, Munoz C, Fuster D, Molins L, Ramirez J, Monzo M. LincRNA-p21 Impacts Prognosis in Resected Non-Small Cell Lung Cancer Patients through Angiogenesis Regulation. J Thorac Oncol. 2016. doi: 10.1016/j.jtho.2016.07.015.
89. Dong R, Liu GB, Liu BH, Chen G, Li K, Zheng S, Dong KR. Targeting long non-coding RNA-TUG1 inhibits tumor growth and angiogenesis in hepatoblastoma. Cell Death Dis. 2016; 7: e2278. doi: 10.1038/cddis.2016.143.
90. Fu WM, Lu YF, Hu BG, Liang WC, Zhu X, Yang HD, Li G, Zhang JF. Long noncoding RNA Hotair mediated angiogenesis in nasopharyngeal carcinoma by direct and indirect signaling pathways. Oncotarget. 2016; 7: 4712-23. doi: 10.18632/oncotarget.6731.
91. Fiedler J, Breckwoldt K, Remmele CW, Hartmann D, Dittrich M, Pfanne A, Just A, Xiao K, Kunz M, Muller T, Hansen A, Geffers R, Dandekar T, et al. Development of Long Noncoding RNA-Based Strategies to Modulate Tissue Vascularization. J Am Coll Cardiol. 2015; 66: 2005-15. doi: 10.1016/j.jacc.2015.07.081.
92. Ebert MS, Neilson JR, Sharp PA. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat Methods. 2007; 4: 721-6. doi: 10.1038/nmeth1079.
93. Tay Y, Rinn J, Pandolfi PP. The multilayered complexity of ceRNA crosstalk and competition. Nature. 2014; 505: 344-52. doi: 10.1038/nature12986.
94. Wu W, Zhang S, Li X, Xue M, Cao S, Chen W. Ets-2 regulates cell apoptosis via the Akt pathway, through the regulation of urothelial cancer associated 1, a long non-coding RNA, in bladder cancer cells. PLoS One. 2013; 8: e73920. doi: 10.1371/journal.pone.0073920.
95. Xue M, Li X, Wu W, Zhang S, Wu S, Li Z, Chen W. Upregulation of long non-coding RNA urothelial carcinoma associated 1 by CCAAT/enhancer binding protein alpha contributes to bladder cancer cell growth and reduced apoptosis. Oncol Rep. 2014; 31: 1993-2000. doi: 10.3892/or.2014.3092.
96. Xue M, Li X, Li Z, Chen W. Urothelial carcinoma associated 1 is a hypoxia-inducible factor-1alpha-targeted long noncoding RNA that enhances hypoxic bladder cancer cell proliferation, migration, and invasion. Tumour Biol. 2014; 35: 6901-12. doi: 10.1007/s13277-014-1925-x.
97. Hughes JM, Legnini I, Salvatori B, Masciarelli S, Marchioni M, Fazi F, Morlando M, Bozzoni I, Fatica A. C/EBPalpha-p30 protein induces expression of the oncogenic long non-coding RNA UCA1 in acute myeloid leukemia. Oncotarget. 2015; 6: 18534-44. doi: 10.18632/oncotarget.4069.
98. Yang C, Li X, Wang Y, Zhao L, Chen W. Long non-coding RNA UCA1 regulated cell cycle distribution via CREB through PI3-K dependent pathway in bladder carcinoma cells. Gene. 2012; 496: 8-16. doi: 10.1016/j.gene.2012.01.012.
99. Li Z, Li X, Wu S, Xue M, Chen W. Long non-coding RNA UCA1 promotes glycolysis by upregulating hexokinase 2 through the mTOR-STAT3/microRNA143 pathway. Cancer Sci. 2014; 105: 951-5. doi: 10.1111/cas.12461.
100. Na XY, Liu ZY, Ren PP, Yu R, Shang XS. Long non-coding RNA UCA1 contributes to the progression of prostate cancer and regulates proliferation through KLF4-KRT6/13 signaling pathway. Int J Clin Exp Med. 2015; 8: 12609-16.
101. Hu JJ, Song W, Zhang SD, Shen XH, Qiu XM, Wu HZ, Gong PH, Lu S, Zhao ZJ, He ML, Fan H. HBx-upregulated lncRNA UCA1 promotes cell growth and tumorigenesis by recruiting EZH2 and repressing p27Kip1/CDK2 signaling. Sci Rep. 2016; 6: 23521. doi: 10.1038/srep23521.
102. Xue M, Pang H, Li X, Li H, Pan J, Chen W. Long non-coding RNA urothelial cancer-associated 1 promotes bladder cancer cell migration and invasion by way of the hsa-miR-145-ZEB1/2-FSCN1 pathway. Cancer Sci. 2016; 107: 18-27. doi: 10.1111/cas.12844.
103. Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao X, Chen WS, Li B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016; 371: 99-106. doi: 10.1016/j.canlet.2015.11.024.
104. Bian Z, Jin L, Zhang J, Yin Y, Quan C, Hu Y, Feng Y, Liu H, Fei B, Mao Y, Zhou L, Qi X, Huang S, et al. LncRNA-UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci Rep. 2016; 6: 23892. doi: 10.1038/srep23892.
105. Wu X, He X, Li S, Xu X, Chen X, Zhu H. Long Non-Coding RNA ucoo2kmd.1 Regulates CD44-Dependent Cell Growth by Competing for miR-211-3p in Colorectal Cancer. PLoS One. 2016; 11: e0151287. doi: 10.1371/journal.pone.0151287.
106. Cai X, Cullen BR. The imprinted H19 noncoding RNA is a primary microRNA precursor. Rna. 2007; 13: 313-6. doi: 10.1261/rna.351707.
107. Tsang WP, Ng EK, Ng SS, Jin H, Yu J, Sung JJ, Kwok TT. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis. 2010; 31: 350-8. doi: 10.1093/carcin/bgp181.
108. Han D, Gao X, Wang M, Qiao Y, Xu Y, Yang J, Dong N, He J, Sun Q, Lv G, Xu C, Tao J, Ma N. Long noncoding RNA H19 indicates a poor prognosis of colorectal cancer and promotes tumor growth by recruiting and binding to eIF4A3. Oncotarget. 2016; 7:22159-22173. doi: 10.18632/oncotarget.8063.
109. Huang G, Wu X, Li S, Xu X, Zhu H, Chen X. The long noncoding RNA CASC2 functions as a competing endogenous RNA by sponging miR-18a in colorectal cancer. Sci Rep. 2016; 6: 26524. doi: 10.1038/srep26524.
110. Qi P, Xu MD, Ni SJ, Huang D, Wei P, Tan C, Zhou XY, Du X. Low expression of LOC285194 is associated with poor prognosis in colorectal cancer. J Transl Med. 2013; 11: 122. doi: 10.1186/1479-5876-11-122.
111. Liu Q, Huang J, Zhou N, Zhang Z, Zhang A, Lu Z, Wu F, Mo YY. LncRNA loc285194 is a p53-regulated tumor suppressor. Nucleic Acids Res. 2013; 41: 4976-87. doi: 10.1093/nar/gkt182.
112. Xia T, Chen S, Jiang Z, Shao Y, Jiang X, Li P, Xiao B, Guo J. Long noncoding RNA FER1L4 suppresses cancer cell growth by acting as a competing endogenous RNA and regulating PTEN expression. Sci Rep. 2015; 5: 13445. doi: 10.1038/srep13445.
113. Yue B, Sun B, Liu C, Zhao S, Zhang D, Yu F, Yan D. Long non-coding RNA Fer-1-like protein 4 suppresses oncogenesis and exhibits prognostic value by associating with miR-106a-5p in colon cancer. Cancer Sci. 2015; 106: 1323-32. doi: 10.1111/cas.12759.
114. Sebio A, Kahn M, Lenz HJ. The potential of targeting Wnt/beta-catenin in colon cancer. Expert Opin Ther Targets. 2014; 18: 611-5. doi: 10.1517/14728222.2014.906580.
115. Yoon JH, Abdelmohsen K, Srikantan S, Yang X, Martindale JL, De S, Huarte M, Zhan M, Becker KG, Gorospe M. LincRNA-p21 suppresses target mRNA translation. Mol Cell. 2012; 47: 648-55. doi: 10.1016/j.molcel.2012.06.027.
116. Wang G, Li Z, Zhao Q, Zhu Y, Zhao C, Li X, Ma Z, Li X, Zhang Y. LincRNA-p21 enhances the sensitivity of radiotherapy for human colorectal cancer by targeting the Wnt/beta-catenin signaling pathway. Oncol Rep. 2014; 31: 1839-45. doi: 10.3892/or.2014.3047.
117. Wang J, Lei ZJ, Guo Y, Wang T, Qin ZY, Xiao HL, Fan LL, Chen DF, Bian XW, Liu J, Wang B. miRNA-regulated delivery of lincRNA-p21 suppresses beta-catenin signaling and tumorigenicity of colorectal cancer stem cells. Oncotarget. 2015; 6: 37852-70. doi: 10.18632/oncotarget.5635.
118. Ma Y, Yang Y, Wang F, Moyer MP, Wei Q, Zhang P, Yang Z, Liu W, Zhang H, Chen N, Wang H, Wang H, Qin H. Long non-coding RNA CCAL regulates colorectal cancer progression by activating Wnt/beta-catenin signalling pathway via suppression of activator protein 2alpha. Gut. 2015. doi: 10.1136/gutjnl-2014-308392.
119. Buffart TE, Coffa J, Hermsen MA, Carvalho B, van der Sijp JR, Ylstra B, Pals G, Schouten JP, Meijer GA. DNA copy number changes at 8q11-24 in metastasized colorectal cancer. Cell Oncol. 2005; 27: 57-65.
120. He X, Tan X, Wang X, Jin H, Liu L, Ma L, Yu H, Fan Z. C-Myc-activated long noncoding RNA CCAT1 promotes colon cancer cell proliferation and invasion. Tumour Biol. 2014; 35: 12181-8. doi: 10.1007/s13277-014-2526-4.
121. Zhang Z, Zhou C, Chang Y, Zhang Z, Hu Y, Zhang F, Lu Y, Zheng L, Zhang W, Li X, Li X. Long non-coding RNA CASC11 interacts with hnRNP-K and activates the WNT/beta-catenin pathway to promote growth and metastasis in colorectal cancer. Cancer Lett. 2016; 376: 62-73. doi: 10.1016/j.canlet.2016.03.022.
122. Ling H, Spizzo R, Atlasi Y, Nicoloso M, Shimizu M, Redis RS, Nishida N, Gafa R, Song J, Guo Z, Ivan C, Barbarotto E, De Vries I, et al. CCAT2, a novel noncoding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res. 2013; 23: 1446-61. doi: 10.1101/gr.152942.112.
123. Ge X, Chen Y, Liao X, Liu D, Li F, Ruan H, Jia W. Overexpression of long noncoding RNA PCAT-1 is a novel biomarker of poor prognosis in patients with colorectal cancer. Med Oncol. 2013; 30: 588. doi: 10.1007/s12032-013-0588-6.
124. Zheng HT, Shi DB, Wang YW, Li XX, Xu Y, Tripathi P, Gu WL, Cai GX, Cai SJ. High expression of lncRNA MALAT1 suggests a biomarker of poor prognosis in colorectal cancer. Int J Clin Exp Pathol. 2014; 7: 3174-81.
125. Ji Q, Liu X, Fu X, Zhang L, Sui H, Zhou L, Sun J, Cai J, Qin J, Ren J, Li Q. Resveratrol inhibits invasion and metastasis of colorectal cancer cells via MALAT1 mediated Wnt/beta-catenin signal pathway. PLoS One. 2013; 8: e78700. doi: 10.1371/journal.pone.0078700.
126. Yang MH, Hu ZY, Xu C, Xie LY, Wang XY, Chen SY, Li ZG. MALAT1 promotes colorectal cancer cell proliferation/migration/invasion via PRKA kinase anchor protein 9. Biochim Biophys Acta. 2015; 1852: 166-74. doi: 10.1016/j.bbadis.2014.11.013.
127. Ji Q, Zhang L, Liu X, Zhou L, Wang W, Han Z, Sui H, Tang Y, Wang Y, Liu N, Ren J, Hou F, Li Q. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br J Cancer. 2014; 111: 736-48. doi: 10.1038/bjc.2014.383.
128. Zhang ZY, Lu YX, Zhang ZY, Chang YY, Zheng L, Yuan L, Zhang F, Hu YH, Zhang WJ, Li XN. Loss of TINCR expression promotes proliferation, metastasis through activating EpCAM cleavage in colorectal cancer. Oncotarget. 2016; 7:22639-49. doi: 10.18632/oncotarget.8141.
129. Qi P, Xu MD, Ni SJ, Shen XH, Wei P, Huang D, Tan C, Sheng WQ, Zhou XY, Du X. Down-regulation of ncRAN, a long non-coding RNA, contributes to colorectal cancer cell migration and invasion and predicts poor overall survival for colorectal cancer patients. Mol Carcinog. 2015; 54: 742-50. doi: 10.1002/mc.22137.
130. Shi D, Zheng H, Zhuo C, Peng J, Li D, Xu Y, Li X, Cai G, Cai S. Low expression of novel lncRNA RP11-462C24.1 suggests a biomarker of poor prognosis in colorectal cancer. Med Oncol. 2014; 31: 31. doi: 10.1007/s12032-014-0031-7.
131. Deng Q, He B, Gao T, Pan Y, Sun H, Xu Y, Li R, Ying H, Wang F, Liu X, Chen J, Wang S. Up-regulation of 91H promotes tumor metastasis and predicts poor prognosis for patients with colorectal cancer. PLoS One. 2014; 9: e103022. doi: 10.1371/journal.pone.0103022.
132. Cao X, Zhuang S, Hu Y, Xi L, Deng L, Sheng H, Shen W. Associations between polymorphisms of long non-coding RNA MEG3 and risk of colorectal cancer in Chinese. Oncotarget. 2016; 7:19054-9. doi: 10.18632/oncotarget.7764.
133. Yin DD, Liu ZJ, Zhang E, Kong R, Zhang ZH, Guo RH. Decreased expression of long noncoding RNA MEG3 affects cell proliferation and predicts a poor prognosis in patients with colorectal cancer. Tumour Biol. 2015; 36: 4851-9. doi: 10.1007/s13277-015-3139-2.
134. Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP, Wang F, Sun SH. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell. 2013; 49: 1083-96. doi: 10.1016/j.molcel.2013.01.010.
135. Guo XB, Hua Z, Li C, Peng LP, Wang JS, Wang B, Zhi QM. Biological significance of long non-coding RNA FTX expression in human colorectal cancer. Int J Clin Exp Med. 2015; 8: 15591-600.
136. Wu Y, Yang L, Zhao J, Li C, Nie J, Liu F, Zhuo C, Zheng Y, Li B, Wang Z, Xu Y. Nuclear-enriched abundant transcript 1 as a diagnostic and prognostic biomarker in colorectal cancer. Mol Cancer. 2015; 14: 191. doi: 10.1186/s12943-015-0455-5.
137. Li Y, Li Y, Chen W, He F, Tan Z, Zheng J, Wang W, Zhao Q, Li J. NEAT expression is associated with tumor recurrence and unfavorable prognosis in colorectal cancer. Oncotarget. 2015; 6: 27641-50. doi: 10.18632/oncotarget.4737.
138. Liu Y, Zhang M, Liang L, Li J, Chen YX. Over-expression of lncRNA DANCR is associated with advanced tumor progression and poor prognosis in patients with colorectal cancer. Int J Clin Exp Pathol. 2015; 8: 11480-4.
139. Wang F, Ni H, Sun F, Li M, Chen L. Overexpression of lncRNA AFAP1-AS1 correlates with poor prognosis and promotes tumorigenesis in colorectal cancer. Biomed Pharmacother. 2016; 81: 152-9. doi: 10.1016/j.biopha.2016.04.009.
140. Yin D, He X, Zhang E, Kong R, De W, Zhang Z. Long noncoding RNA GAS5 affects cell proliferation and predicts a poor prognosis in patients with colorectal cancer. Med Oncol. 2014; 31: 253. doi: 10.1007/s12032-014-0253-8.
141. Sun Y, Zheng ZP, Li H, Zhang HQ, Ma FQ. ANRIL is associated with the survival rate of patients with colorectal cancer, and affects cell migration and invasion in vitro. Mol Med Rep. 2016. doi: 10.3892/mmr.2016.5409.
142. Yan B, Gu W, Yang Z, Gu Z, Yue X, Gu Q, Liu L. Downregulation of a long noncoding RNA-ncRuPAR contributes to tumor inhibition in colorectal cancer. Tumour Biol. 2014; 35: 11329-35. doi: 10.1007/s13277-014-2465-0.
143. Liu T, Zhang X, Yang YM, Du LT, Wang CX. Increased expression of the long noncoding RNA CRNDE-h indicates a poor prognosis in colorectal cancer, and is positively correlated with IRX5 mRNA expression. Onco Targets Ther. 2016; 9: 1437-48. doi: 10.2147/ott.s98268.
144. Slaby O. Non-coding RNAs as Biomarkers for Colorectal Cancer Screening and Early Detection. Adv Exp Med Biol. 2016; 937: 153-70. doi: 10.1007/978-3-319-42059-2_8.
145. Huang X, Yuan T, Tschannen M, Sun Z, Jacob H, Du M, Liang M, Dittmar RL, Liu Y, Liang M, Kohli M, Thibodeau SN, Boardman L, et al. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genomics. 2013; 14: 319. doi: 10.1186/1471-2164-14-319.
146. Dong L, Lin W, Qi P, Xu MD, Wu X, Ni S, Huang D, Weng WW, Tan C, Sheng W, Zhou X, Du X. Circulating Long RNAs in Serum Extracellular Vesicles: Their Characterization and Potential Application as Biomarkers for Diagnosis of Colorectal Cancer. Cancer Epidemiol Biomarkers Prev. 2016; 25: 1158-66. doi: 10.1158/1055-9965.epi-16-0006.
147. Wang C, Yu J, Han Y, Li L, Li J, Li T, Qi P. Long non-coding RNAs LOC285194, RP11-462C24.1 and Nbla12061 in serum provide a new approach for distinguishing patients with colorectal cancer from healthy controls. Oncotarget. 2016; 7:70769-70778. doi: 10.18632/oncotarget.12220.
148. Li CH, Chen Y. Targeting long non-coding RNAs in cancers: progress and prospects. Int J Biochem Cell Biol. 2013; 45: 1895-910. doi: 10.1016/j.biocel.2013.05.030.
149. Pedram Fatemi R, Salah-Uddin S, Modarresi F, Khoury N, Wahlestedt C, Faghihi MA. Screening for Small-Molecule Modulators of Long Noncoding RNA-Protein Interactions Using AlphaScreen. J Biomol Screen. 2015; 20: 1132-41. doi: 10.1177/1087057115594187.
150. Hannon GJ. RNA interference. Nature. 2002; 418: 244-51. doi: 10.1038/418244a.
151. Robb GB, Brown KM, Khurana J, Rana TM. Specific and potent RNAi in the nucleus of human cells. Nat Struct Mol Biol. 2005; 12: 133-7. doi: 10.1038/nsmb886.
152. Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010; 464: 1071-6. doi: 10.1038/nature08975.
153. Gutschner T, Hammerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M, Zornig M, MacLeod AR, Spector DL, et al. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013; 73: 1180-9. doi: 10.1158/0008-5472.can-12-2850.
154. Bennett CF, Swayze EE. RNA targeting therapeutics: molecular mechanisms of antisense oligonucleotides as a therapeutic platform. Annu Rev Pharmacol Toxicol. 2010; 50: 259-93. doi: 10.1146/annurev.pharmtox.010909.105654.
155. Jalali S, Bhartiya D, Lalwani MK, Sivasubbu S, Scaria V. Systematic transcriptome wide analysis of lncRNA-miRNA interactions. PLoS One. 2013; 8: e53823. doi: 10.1371/journal.pone.0053823.
156. Lee SH, Jeong D, Han YS, Baek MJ. Pivotal role of vascular endothelial growth factor pathway in tumor angiogenesis. Ann Surg Treat Res. 2015; 89: 1-8. doi: 10.4174/astr.2015.89.1.1.
157. Hicklin DJ, Ellis LM. Role of the vascular endothelial growth factor pathway in tumor growth and angiogenesis. J Clin Oncol. 2005; 23: 1011-27. doi: 10.1200/jco.2005.06.081.
158. Naresh KN, Nerurkar AY, Borges AM. Angiogenesis is redundant for tumour growth in lymph node metastases. Histopathology. 2001; 38: 466-70.
159. Edel M, Robbins P, D’Antuono M, Harvey J, Papadimitrion J, Mitchell C, Dawkins H. Assessment of endothelial cell proliferation in primary breast carcinoma and its association with axillary lymph node status. Breast. 2000; 9: 28-34. doi: 10.1054/brst.1999.0082.
160. Zhao C, Yang H, Shi H, Wang X, Chen X, Yuan Y, Lin S, Wei Y. Distinct contributions of angiogenesis and vascular co-option during the initiation of primary microtumors and micrometastases. Carcinogenesis. 2011; 32: 1143-50. doi: 10.1093/carcin/bgr076.
161. Frentzas S, Simoneau E, Bridgeman VL, Vermeulen PB, Foo S, Kostaras E, Nathan MR, Wotherspoon A, Gao ZH, Shi Y, Van den Eynden G, Daley F, Peckitt C, et al. Vessel co-option mediates resistance to anti-angiogenic therapy in liver metastases. 2016; 22:1294-1302. doi: 10.1038/nm.4197.
162. Blasi F, Sidenius N. The urokinase receptor: focused cell surface proteolysis, cell adhesion and signaling. FEBS Lett. 2010; 584: 1923-30. doi: 10.1016/j.febslet.2009.12.039.
163. Xu W, Yang Z, Lu N. A new role for the PI3K/Akt signaling pathway in the epithelial-mesenchymal transition. Cell Adh Migr. 2015; 9: 317-24. doi: 10.1080/19336918.2015.1016686.
164. Xu C, Yang M, Tian J, Wang X, Li Z. MALAT-1: a long non-coding RNA and its important 3’ end functional motif in colorectal cancer metastasis. Int J Oncol. 2011; 39: 169-75. doi: 10.3892/ijo.2011.1007.
165. Gong J, Tian J, Lou J, Ke J, Li L, Li J, Yang Y, Gong Y, Zhu Y, Zhang Y, Zhong R, Chang J, Miao X. A functional polymorphism in lnc-LAMC2-1:1 confers risk of colorectal cancer by affecting miRNA binding. Carcinogenesis. 2016; 37: 443-51. doi: 10.1093/carcin/bgw024.
166. Shi J, Li X, Zhang F, Zhang C, Guan Q, Cao X, Zhu W, Zhang X, Cheng Y, Ou K, Chen Q, Hu S. Circulating lncRNAs associated with occurrence of colorectal cancer progression. Am J Cancer Res. 2015; 5: 2258-65.
167. Li AX, Xin WQ, Ma CG. Fentanyl inhibits the invasion and migration of colorectal cancer cells via inhibiting the negative regulation of Ets-1 on BANCR. Biochem Biophys Res Commun. 2015; 465: 594-600. doi: 10.1016/j.bbrc.2015.08.068.
168. Li L, Sun R, Liang Y, Pan X, Li Z, Bai P, Zeng X, Zhang D, Zhang L, Gao L. Association between polymorphisms in long non-coding RNA PRNCR1 in 8q24 and risk of colorectal cancer. J Exp Clin Cancer Res. 2013; 32: 104. doi: 10.1186/1756-9966-32-104.
169. Wang R, Du L, Yang X, Jiang X, Duan W, Yan S, Xie Y, Zhu Y, Wang Q, Wang L, Yang Y, Wang C. Identification of long noncoding RNAs as potential novel diagnosis and prognosis biomarkers in colorectal cancer. J Cancer Res Clin Oncol. 2016; 142: 2291-301. doi: 10.1007/s00432-016-2238-9.
170. Wang C, Yu J, Han Y, Li L, Li J, Li T, Qi P. Long non-coding RNAs LOC285194, RP11-462C24.1 and Nbla12061 in serum provide a new approach for distinguishing patients with colorectal cancer from healthy controls. Oncotarget. 2016; 7: 70769-78. doi: 10.18632/oncotarget.12220.
171. Kong H, Wu Y, Zhu M, Zhai C, Qian J, Gao X, Wang S, Hou Y, Lu S, Zhu H. Long non-coding RNAs: novel prognostic biomarkers for liver metastases in patients with early stage colorectal cancer. Oncotarget. 2016; 7: 50428-36. doi: 10.18632/oncotarget.10416.
172. Wu Y, Yang L, Zhao J, Li C, Nie J, Liu F, Zhuo C, Zheng Y, Li B, Wang Z, Xu Y. Nuclear-enriched abundant transcript 1 as a diagnostic and prognostic biomarker in colorectal cancer. Mol Cancer. 2015; 14: 191. doi: 10.1186/s12943-015-0455-5.
173. Liu T, Zhang X, Gao S, Jing F, Yang Y, Du L, Zheng G, Li P, Li C, Wang C. Exosomal long noncoding RNA CRNDE-h as a novel serum-based biomarker for diagnosis and prognosis of colorectal cancer. Oncotarget. 2016; 7: 85551-63. doi: 10.18632/oncotarget.13465.

