INTRODUCTION
Non-muscle invasive bladder cancers (NMIBC) have a 1- and 5-year disease-progression rate of up to 17% and 45%, respectively [1]. The current most important clinical prognosticators in NMIBC patients, according to the EORTC risk tables, are stage T1, high grade (HG) and presence of concomitant carcinoma in situ (CIS) [2]. Nevertheless, these prognostic factors do not have sufficient discriminative capacity to predict, at the patient level, the risk of tumour progression. Progressive patients deserve careful attention, particularly because therapeutic decisions change when patients are likely to progress. Many efforts have been made to develop biomarkers for assessing aggressiveness and for predicting the prognosis of NMIBC patients (reviewed in [3]), but to date none of the identified biomarkers is being used in the clinical setting due to their limited value. Thus, there is a clear need for accurate and reliable predictive biomarkers that can distinguish progressive from non-progressive NMIBC.
In recent years it has become apparent that gene expression is controlled post-transcriptionally by microRNAs (miRNAs), short (21–23 nt) non-coding RNAs which bind to specific mRNAs. Experimental work has shown that miRNAs are dysregulated in most cancer types, including bladder cancer (BC) [4–7], and have demonstrated significant diagnostic and prognostic values in different malignancies [8–11]. In the case of BC, the assessment of the levels of these aberrantly expressed miRNAs in urine samples appears to be a promising approach to identify diagnostic and prognostic biomarkers in a non-invasive way. Urine is a particular source of such biomarkers since tumour cells are in direct contact with it, it is collected noninvasively and miRNA have shown high stability and easy detectability in urine [12]. Using a case-control design, we have shown that urinary miRNA profiles vary significantly between urines from control and BC patients and between high and low grade tumours [13]. Since we have previously found a urinary miRNA signature representative of tumour aggressiveness, we decided to explore the hypothesis that urinary miRNAs may be useful as predictive biomarkers of progression in NMIBC patients. We thus compared the urinary miRNA profiles from individuals with NMIBC who progressed with those who did not progress. Subsequently, we developed a prognostic urinary miRNA signature that was further internally cross-validated.
RESULTS
Characteristics of the dataset
A total of 210 consecutive urinary samples from BC patients with NMIBC subjected to transurethral resection of the bladder (TURB) were included (Table 1). Overall, 22 (11%) patients developed tumour progression and 42 patients (20%) died, 10 of them (5%) due to BC, after a mean follow up of 47 months. The mean time to tumour progression and cancer-specific survival was 19 and 23 months, respectively.
Table 1: Clinicohistopathologic features of the bladder cancer patients
VARIABLES |
|
|---|---|
Gender_N (%) |
|
Male |
157 (75) |
Female |
53 (25) |
Age_years |
|
Mean |
71 |
Range |
37-94 |
Tumour stage and grade_N (%)1 |
|
Tis |
15 (7) |
Ta LG |
77 (37) |
Ta LG+CIS |
1 |
Ta HG |
22 (10) |
Ta HG+CIS |
3 (1) |
T1 LG |
1 |
T1 LG+CIS |
15 (7) |
T1 HG |
59 (28) |
T1 HG+CIS |
17 (8) |
Tumour focality_N (%) |
|
Unifocal |
108 (51) |
Multifocal |
102 (49) |
Tumour diameter (%) |
|
≤3 cm |
170 (81) |
>3 cm |
40 (19) |
Prior recurrence (%) |
|
Primary tumours |
104 (50) |
Recurrent tumours |
106 (50) |
BCG treatment (%) |
98 (47) |
TOTAL |
210 |
1The grade and stage of the tumours were determined according to WHO criteria [38] and TNM classification [39], respectively.
Abbreviations: CIS or Tis, Carcinoma in situ; LG, Low grade; HG, High grade; BCG, Bacillus Calmette-Guerin
The clinical classifiers for progression, stage, grade, presence of CIS, multiplicity and tumour size were analysed in both progressive and non-progressive patients. Stage T1 and HG tumours were identified in 44% and 54% of non-progressive and in 41% and 68% of progressive patients, respectively. Concomitant CIS was found in 18% of progressive and non-progressive patients. Multiplicity was found in 22% and 68% of non-progressive and progressive patients, respectively. A tumour diameter of > 3 cm was found in 34 out of 188 (18%) non-progressive and 6 out of 22 (27%) progressive patients. Recurrent tumours were identified in 59% of progressive and 49% of non-progressive patients.
Association of miRNA expression and clinical features with progression of NMIBC patients
Among the eight previously reported miRNA [13] tested in the present work, miR-187-3p and miR-25-3p were excluded from further analyses because expression data was not available in >30% of samples (insufficient cDNA was available to perform all qPCR reactions). Supplementary Table 1 shows the fold change differences of the miRNAs between the groups and in relation to the previous study on these miRNAs. Univariate Cox regression analysis including clinical covariates and urinary expression levels of miRNAs identified miR-140-5p, miR-142-3p, miR-18a-3p and miR-92a-3p as predictors of tumour progression (Table 2). Of note, none of the clinical covariates were significantly associated with progression in our cohort of NMIBC. After including the above-mentioned miRNAs in the multivariate stepwise regression analysis, urinary expression of miR-140-5p and miR-92a-3p were found as independent prognostic factors of tumour progression (Table 3).
Table 2: Univariate analysis of predictors of tumour progression
Variable |
Cut-off level/categories |
HR |
95% CI |
P value |
|---|---|---|---|---|
Tumour focality |
uni vs. multifocal |
2.25 |
0.92-5.52 |
0.0768 |
Tumour size |
≤3 cm vs. >3 cm |
1.69 |
0.66-4.33 |
0.2718 |
BCG treatment |
no vs yes |
1.29 |
0.56-2.98 |
0.5546 |
Stage1 |
Ta vs. T1 |
1.06 |
0.72-1.56 |
0.7738 |
Grade1 |
LG vs. HG |
1.85 |
0.75-4.54 |
0.1792 |
Concomitant CIS |
without CIS vs. CIS |
1 |
0.34-2.95 |
0.9974 |
miR-140-5p |
-4.4 |
2.89 |
1.13-7.39 |
0.0267* |
miR-142-3p |
1.01 |
3.09 |
1.26-7.6 |
0.0139* |
miR-18a-3p |
-7.02 |
4.40 |
1.72-11.29 |
0.0020* |
miR-92a-3p |
-0.555 |
5.35 |
1.81-15.53 |
0.0025* |
miR-125b-5p |
-1.74 |
0.60 |
0.23-1.53 |
0.2813 |
miR-204-5p |
-5.15 |
0.93 |
0.39-2.21 |
0.8612 |
Results of dichotomous clinical variables are expressed considering category placed in second term as the reference category
1The grade and stage of the tumours were determined according to WHO criteria [38] and TNM classification [39], respectively.
*Statistically significant
Abbreviations: CIS, Carcinoma in situ; LG, Low grade; HG, High grade; BCG, Bacillus Calmette-Guerin; HR, hazard ratio; 95%CI, confidence interval.
Table 3: Independent predictors of tumour progression resulting from multivariate stepwise Cox regression analysis
Variable |
HR |
95% CI |
P value |
|---|---|---|---|
miR-140-5p |
3.53 |
1.38-9.06 |
0.0086* |
miR-92a-3p |
6.21 |
2.09-18.45 |
0.0010* |
Abbreviations: HR, hazard ratio; 95%CI, confidence interval.
*Statistically significant
Development of a miRNA-based prognostic classifier for NMIBC patients
A 2-miRNA prognostic classifier was developed by combining miR-140-5p and miR-92a-3p expression values. A risk score (RS) for tumour progression was calculated for each patient according to a mathematical algorithm containing miR-140-5p and miR-92a-3p expression values. The median value of this RS was -0.206 (range -1.469 to 1.621). Thereafter, a receiver-operating characteristic (ROC) curve analysis of the model (Figure 1) allowed the selection of a cut-off value of 1.62 to classify patients into a high-risk group for tumour progression (20%) and low-risk group for tumour progression (80%) with an accuracy of 81% (55% sensitivity and 84% specificity). Figure 2 depicts Kaplan-Meier curves generated using the selected cut-off point. As shown, RS generated using miRNA expression values was able to discriminate between two groups of NMIBC patients with a highly significant different probability of tumour progression (HR, 5.21; 95%CI, 2.24-12.06; p=0.0001) (Figure 2A). It is important to point out that when patients were stratified according to BC risk groups, the RS continued to discriminate two subgroups of different probability of progression (non-high-risk NMIBC group: HR, 9.59; 95%CI, 1.84-50.09, p=0.007 and high-risk NMIBC group: HR, 3.92; 95%CI, 1.42-10.81, p=0.008) (Figure 3). Moreover, although it did not constitute a primary outcome, accuracy of the RS was also evaluated regarding the probability of being a predictor of cancer-specific survival. Using the same cut-off point, the algorithm was also able to distinguish between two groups with significantly different probabilities of cancer-specific survival (HR 3.88; 95%CI, 1.12-13.44, p=0.033) (Figure 2B).
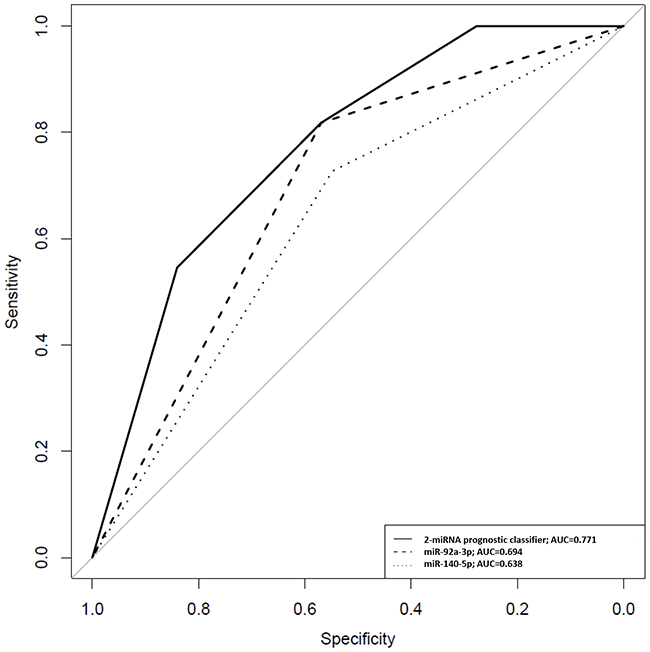
Figure 1: ROC curves for miR-140-5p, miR-92a-3p and the 2-miRNA prognostic classifier based on results obtained by quantitative PCR in urinary samples of NMIBC patients. A RS for tumour progression was calculated for each patient according to a mathematical algorithm containing miR-140-5p and miR-92a-3p expression values as described in Patients and methods section. In this equation, miR-140-5p and miR-92a-3p were introduced as dichotomous variables (miR-140-5p expression ≥-4.4=1; <-4.4=0 and miR-92a-3p expression ≥-0.555=1; <-0.555=0). At fixed sensitivity of 80%, specificity for miR-140-5p and miR-92a-3p individually and combined in the 2-miRNA classifier was 32%, 38% and 59%, respectively.
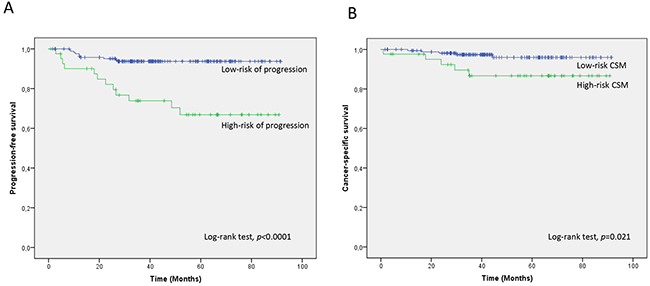
Figure 2: Kaplan-Meier curves for the two-miRNA prognostic classifier showing. A. time to progression and B. time to cancer-specific survival for low-risk (RS<1.62; n=168) and high-risk (RS>1.62; n=42) groups of NMIBC patients. CSM; cancer-specific mortality
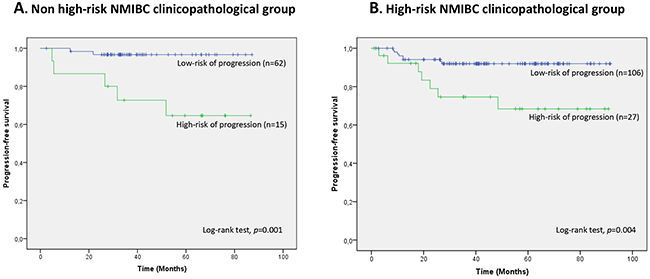
Figure 3: Kaplan-Meier curves for the two-miRNA prognostic classifier showing time to progression in clinicopathological groups of risk. A. non high-risk NMIBC (n=77) and B. high-risk NMIBC (n=133) patients. Patients were divided within each clinicopathological NMIBC risk group according to their RS (RS<1.62, low-risk of progression; RS>1.62, high-risk of progression).
Finally, robustness of the mathematical model was evaluated by bootstrapping with 1,000 resamples, obtaining a C-index after bootstrapping of 0.7306723.
Target prediction and functional enrichment of the two miRNA signature
The DIANA-miRPath miRNA analysis, by using the 2 miRNAs of the model, showed several statistically significant predicted KEGG terms related to cell cycle, TGF-β signalling pathway, MAPK signalling pathway, Wnt signalling pathway, p53 signalling pathway, and pathways in cancer among others. Consistently, when analysing the data by the gene intersection and union mode, with the two miRNAs targeting the same gene, common terms appear to be significantly enriched such as the p53 signalling pathway, TGF-β signalling pathway, cell cycle, and pathways in cancer (Figure 4 and Supplementary Figure 1).
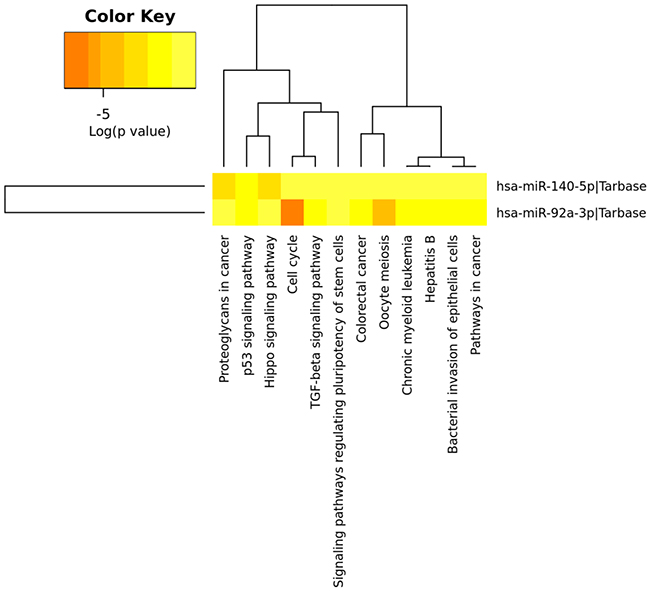
Figure 4: Heatmap of the KEGG pathways enriched in two miRNA target genes. Heatmap from intersection of targeted genes (genes targeted by the two miRNAs from the model) is shown. The two miRNAs are involved in multiple common pathways, especially in cancer-specific pathways (DIANA-miRpath computes log10 P-values).
DISCUSSION
Prediction of disease progression for patients diagnosed with NMIBC is an important clinical challenge. The most important clinical classifiers for progression currently used, stage T1, HG and presence of CIS [2] where unable to predict progression in our cohort of patients. Neither, the less distinctive clinical prognostic classifiers; recurrence, multiplicity and tumour size, could distinguish progressive from non-progressive patients. One explanation could be that the vast majority of our cohort corresponds to HG NMIBC tumours. This fact could minimize the impact of clinicopathological features on predicting progression, and precisely confirms the critical need for new objective methods to identify those NMIBC patients that are likely to progress. Here, we show that a 2-miRNA urinary signature is able to identify a subgroup of NMIBC patients with high-risk of progression, thus providing additional risk stratification beyond clinicopathological prognosticators.
We and others have previously provided evidence that urinary miRNAs can be used as non-invasive diagnostic biomarkers for BC patients [13–17]. Cell-free urine and serum have also been used as a source of non-invasive diagnostic and prognostic biomarkers of BC [18–20], but as far as we could ascertain, this is the first study that has evaluated urinary cell miRNAs as prognostic biomarkers of BC. The results of this study demonstrate that expression of miR-140-5p and miR-92a-3p in urinary cells independently predicts tumour progression in NMIBC patients. This miRNA signature provides an easy-to-use and reliable algorithm to identify a subgroup of patients with a higher probability of tumour progression and shorter cancer specific survival. An accurate estimation of the progression risk would help to identify the most appropriate therapy for each individual patient. For instance, patients at high risk of progression could benefit from re-TURB, adjuvant therapies, more aggressive treatments or more intensive surveillance. In addition, the 2-miRNA model maintains its prognostic ability when NMIBC patients are stratified according to their clinicopathological risk group, thus reinforcing the validity of our model. Of special interest is the predictive value of the model for progression in the subset of non-high risk NMIBC that are supposed to have low-intermediate risk of progression. Identification of those non-high risk NMIBC patients which are more likely to progress may help to change current treatment schedules for this particular subset of NMIBC patients, intensifying their surveillance schedule.
Regarding the genetic markers included in our algorithm, miR-140-5p has been previously suggested to inhibit the expression of TGF-β and MAPK/ERK signalling pathways in several malignancies [21–25]. In addition, a recent publication reveals that overexpression of miR-140-5p has an inhibitory effect on BC by downregulating the expression of the oncogenic isoform ΔNp63 of TP63 [26], a key gene localized in region 3q28 associated with BC risk [27]. In agreement with this data, we previously found a statistically significant downregulation of miR-140-5p in urinary cells from BC patients as well as in high grade BC tumours [13], although this statistically significant downregulation is not seen in the present cohort of NMIBC progressive patients. It is most likely that the different cell types and the variable proportion of tumour cells in the urine of each subject accounts for this lack of statistically significant differences.
Aberrations in miR-92a-3p expression have been previously reported in many cancers [28–30], but to our knowledge, this is the first time miR-92a-3p has been associated with BC. In recent studies, it was observed that overexpression of miR-92a-3p can disrupt PI3K/Akt/mTOR pathway by targeting two negative regulatory factors of this pathway, PTEN and PHLPP [29, 31]. Different studies have reported that overexpression of miR-92a-3p is involved in the development of metastasis and correlates with survival in different malignancies [32–34]. Our study shows that miR-92a-3p is significantly overexpressed in urinary cells of NMIBC progressing patients which is in concordance with its role as oncomiR.
KEGG pathway analysis corroborates that the 2-miRNA signature is biologically meaningful. It shows that the target genes of the two miRNA take part in many important signalling pathways such as cancer pathways, cell cycle, TGF-β pathway and p53 signalling pathway, among others. Moreover, it has been reported that several of the genes targeted by our miRNAs (e.g. TP63, PTEN) [35, 36] play a role in the genetic pathogenesis of BC.
The strengths of this study lie in the fact that it includes a cohort of patients attended in a single centre, homogeneously treated, with prospective data collection and long term follow-up. Furthermore, the use of urine samples to obtain the miRNA prognostic signature may allow the development of a non-invasive bladder cancer prognostic tool with an easy translation into clinical practice. However, some limitations should be acknowledged. First, banked urine samples were analysed, so we cannot calculate the rate of experiment failure due to low cellularity of the sample. Nonetheless, our experience is that the failure rate is low (~5%) in the case of tumour samples [37]. Second, the number of events in the cancer-specific survival analysis is limited. Thus, this part of the results must be taken with care until a validation in a cohort with larger number of events is performed. Next, because of our interest in magnifying the probability to identify reliable prognostic miRNAs, all available samples were included in the evaluation set, thus preventing an independent validation. To overcome this drawback, we strictly ascertained the robustness of the mathematical algorithm by bootstrapping, obtaining a high concordance index. However, the performance of this miRNA signature would be more reliable if validation is performed in an independent external data set. Finally, we have evaluated a limited number of candidate miRNAs previously included in our signatures. Other attractive candidate miRNAs with prognostic value in BC are reported in the literature. Thus, our study rather than establishing a definitive prognostic miRNA expression signature, may contribute to describe prognostic miRNAs which warrant further prospective evaluation in carefully and specifically designed studies.
In summary, our results showed a 2-miRNA urinary signature which significantly predicts progression and cancer-specific survival in NMIBC patients, indicating that it may be a novel potential biomarker for prognosis of NMIBC patients.
MATERIALS AND METHODS
Patients and clinical samples
Under Institutional Review Board approval and patients’ informed consent, we prospectively collected freshly voided urine samples from BC patients at Hospital Clínic of Barcelona, starting January 2008. Urines were centrifuged and separated into cellular pellet and supernatant prior to storage at -80ºC in a urinary biorepository.
For this study, consecutive available banked cellular pellets from urine samples collected between January 2008 and July 2013 at Hospital Clínic of Barcelona were used for analysis. These included 210 samples from urothelial cell carcinoma of the bladder patients subjected to TURB (Table 1) [38, 39]. The only exclusion criterion was the lack of follow-up after TURB. Tumours were classified according to their clinicopathological risk into two categories: high-risk NMIBC tumours [any of the following: T1 tumours, HG tumours, CIS, and multiple, recurrent, large (>3 cm) Ta low grade tumours (all conditions must present for the last point)] and non high-risk NMIBC tumours (all tumours not defined in the other category). None of the patients included had an upper urinary tract tumour. All patients were followed-up postoperatively following the EAU guidelines [1]. Tumours were considered to be in progression when pathological stage or histological grade increasing and/or distant metastasis or pathological nodes were developed during the follow-up.
RNA extraction
Around 50-100 ml of urine was collected from each BC patient the day before theTURB. Urine samples were processed as previously described [40], except that collected urines were stored at 4ºC and processed within the next 24 hours instead of ice cooled. RNAs from the urinary cell pellets were extracted using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions and quantified with a NanoDrop1000 (NanoDrop Technologies, Wilmington, DE, USA).
miRNA analysis
A total of 8 key miRNAs (miR-25-3p, miR-18a-3p, miR-92a-3p, miR-140-5p, miR-125b-5p, miR-142-3p, miR-204-5p and miR-187-3p), previously described by our group [13], were selected for examination in urinary samples from NMIBC patients by quantitative PCR by using miRCURY LNA Universal RT microRNA PCR kit (Exiqon, Vedbaek, Denmark), as previously described [13]. Briefly, total RNA (100 ng) containing miRNA was polyadenylated, and cDNA was synthesized using a poly(T) primer with a 3’ degenerate anchor and a 5’ universal tag. Then, cDNA was served as a template for miRNA RT-qPCR amplification with the specific locked nucleic acid (LNA) primers (Supplementary Table 2) and SYBR Green master mix. PCR reactions were carried out using a Light Cycler 480 instrument. The amplification profile was denatured at 95ºC for 10 min followed by 45 amplification cycles of 95ºC for 10s and 60ºC for 1 min. At the end of the PCR cycles, melting curve analyses were performed. miR-103-3p and miR-30c-5p were used as endogenous controls, as previously described [13, 41].
Statistical analysis
Univariate Cox regression analysis was performed on each covariate to examine its influence on tumour progression and cancer–specific survival. Thereafter, a multivariate forward stepwise Cox regression analysis was performed. After establishing the multivariate model, a RS for the miRNAs of the model was calculated for each patient as the linear combination of the model according to the following RS=β1× miR-140-5p +β2× miR-92a-3p. RS was subjected to a ROC analysis in order to choose the most appropriate threshold for predicting tumour progression. Then, Kaplan-Meier curves for tumour progression and cancer-specific survival were generated using the selected cut-off point and compared according to the log-rank test. Statistical significance was established at p-value of 0.05. R-software and SPSS v23.0 were used for calculations.
Finally, robustness of the mathematical algorithm resulting from the multivariate analysis was evaluated by bootstrapping with 1,000 resamples. For this purpose, validate function from RMS Package was used and the optimism-adjusted estimate of concordance index was computed (http://CRAN.R-project.org/package=rms).
Pathway enrichment analysis
DIANA-miRPath v3.0 software (http://www.microrna.gr/miRPathv3) [42] using experimentally validated miRNA interactions derived from DIANA-TarBase v7 was used to identify targets of the miRNAs composing the prognostic signature. Subsequent target enrichment analysis was performed in order to discover possible canonical altered pathways using DAVID (https://david.ncifcrf.gov).
Abbreviations
BC; bladder cancer
BCG; bacillus Calmette-Guerin
CI; confidence interval
CIS/Tis; carcinoma in situ
CSM; cancer-specific mortality
HG; high grade
HR; hazard ratio
KEGG; kyoto encyclopedia of genes and genomes
LG; low grade
LNA: locked nucleic acid
miRNA; microRNA
NMIBC; non-muscle invasive bladder cancer
ROC; receiver-operating characteristic
RS; risk score
TURB; transurethral resection of the bladder
ACKNOWLEDGMENTS
We thank all the patients who participated in this study and all the staff and nurses from the Urology Department of the Hospital Clínic for collaborating in collecting urine samples. We thank Helena Kruyer for the English correction of the manuscript.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
FUNDING
This work was supported by grants from the Spanish Urological Association (FIU 2009 to LM), Fondo de Investigaciones Sanitarias (PI10/01145 to AA), Fondo Europeo de Desarrollo Regional. Unión Europea. Una manera de hacer Europa and CERCA Programme/ Generalitat de Catalunya.
Author contributions
Study concepts and design: LM, MJR, AA, LI, JJL
Data acquisition: MIT, AC, MC, LG
Data analysis and interpretation: MIT, LM, JJL, MJR, AA, LI
Statistical analysis: JJL, LI
Manuscript preparation: LM
Manuscript review: ALL AUTHORS
REFERENCES
1. Babjuk M, Bohle A, Burger M, Capoun O, Cohen D, Comperat EM, Hernandez V, Kaasinen E, Palou J, Roupret M, van Rhijn BW, Shariat SF, Soukup V, et al. EAU Guidelines on Non-Muscle-invasive Urothelial Carcinoma of the Bladder: Update 2016. Eur Urol. 2016.
2. Sylvester RJ, van der MEIJDEN AP, Oosterlinck W, Witjes JA, Bouffioux C, Denis L, Newling DW, Kurth K. Predicting recurrence and progression in individual patients with stage Ta T1 bladder cancer using EORTC risk tables: a combined analysis of 2596 patients from seven EORTC trials. Eur Urol. 2006; 49: 466-5.
3. Mitra AP. Molecular substratification of bladder cancer: moving towards individualized patient management. Ther Adv Urol. 2016; 8: 215-233.
4. Catto JW, Alcaraz A, Bjartell AS, De Vere WR, Evans CP, Fussel S, Hamdy FC, Kallioniemi O, Mengual L, Schlomm T, Visakorpi T. MicroRNA in Prostate, Bladder, and Kidney Cancer: A Systematic Review. Eur Urol. 2011; 59: 671-681.
5. Ichimi T, Enokida H, Okuno Y, Kunimoto R, Chiyomaru T, Kawamoto K, Kawahara K, Toki K, Kawakami K, Nishiyama K, Tsujimoto G, Nakagawa M, Seki N. Identification of novel microRNA targets based on microRNA signatures in bladder cancer. Int J Cancer. 2009; 125: 345-352.
6. Veerla S, Lindgren D, Kvist A, Frigyesi A, Staaf J, Persson H, Liedberg F, Chebil G, Gudjonsson S, Borg A, Mansson W, Rovira C, Hoglund M. MiRNA expression in urothelial carcinomas: important roles of miR-10a, miR-222, miR-125b, miR-7 and miR-452 for tumor stage and metastasis, and frequent homozygous losses of miR-31. Int J Cancer. 2009; 124: 2236-2242.
7. Zabolotneva AA, Zhavoronkov A, Garazha AV, Roumiantsev SA, Buzdin AA. Characteristic patterns of microRNA expression in human bladder cancer. Front Genet. 2012; 3: 310.
8. Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, et al. MicroRNA expression profiles classify human cancers. Nature. 2005; 435: 834-838.
9. Dyrskjot L, Ostenfeld MS, Bramsen JB, Silahtaroglu AN, Lamy P, Ramanathan R, Fristrup N, Jensen JL, Andersen CL, Zieger K, Kauppinen S, Ulhoi BP, Kjems J, et al. Genomic profiling of microRNAs in bladder cancer: miR-129 is associated with poor outcome and promotes cell death in vitro. Cancer Res. 2009; 69: 4851-4860.
10. Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, Iorio MV, Visone R, Sever NI, Fabbri M, Iuliano R, Palumbo T, Pichiorri F, et al. A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med. 2005; 353: 1793-1801.
11. Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Calin GA, Liu CG, Croce CM, et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell. 2006; 9: 189-198.
12. Weber JA, Baxter DH, Zhang S, Huang DY, Huang KH, Lee MJ, Galas DJ, Wang K. The microRNA spectrum in 12 body fluids. Clin Chem. 2010; 56: 1733-1741.
13. Mengual L, Lozano JJ, Ingelmo-Torres M, Gazquez C, Ribal MJ, Alcaraz A. Using microRNA profiling in urine samples to develop a non-invasive test for bladder cancer. Int J Cancer. 2013; 133:2631-2641.
14. Hanke M, Hoefig K, Merz H, Feller AC, Kausch I, Jocham D, Warnecke JM, Sczakiel G. A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol Oncol. 2010; 28: 655-661.
15. Yamada Y, Enokida H, Kojima S, Kawakami K, Chiyomaru T, Tatarano S, Yoshino H, Kawahara K, Nishiyama K, Seki N, Nakagawa M. MiR-96 and miR-183 detection in urine serve as potential tumor markers of urothelial carcinoma: correlation with stage and grade, and comparison with urinary cytology. Cancer Sci. 2011; 102: 522-529.
16. Puerta-Gil P, Garcia-Baquero R, Jia AY, Ocana S, Alvarez-Mugica M, Alvarez-Ossorio JL, Cordon-Cardo C, Cava F, Sanchez-Carbayo M. miR-143, miR-222, and miR-452 Are Useful as Tumor Stratification and Noninvasive Diagnostic Biomarkers for Bladder Cancer. Am J Pathol. 2012; 180: 1808-1815.
17. Miah S, Dudziec E, Drayton RM, Zlotta AR, Morgan SL, Rosario DJ, Hamdy FC, Catto JW. An evaluation of urinary microRNA reveals a high sensitivity for bladder cancer. Br J Cancer. 2012; 107: 123-128.
18. Zhang DZ, Lau KM, Chan ES, Wang G, Szeto CC, Wong K, Choy RK, Ng CF. Cell-free urinary microRNA-99a and microRNA-125b are diagnostic markers for the non-invasive screening of bladder cancer. PLoS ONE. 2014; 9: e100793.
19. Kim SM, Kang HW, Kim WT, Kim YJ, Yun SJ, Lee SC, Kim WJ. Cell-Free microRNA-214 From Urine as a Biomarker for Non-Muscle-Invasive Bladder Cancer. Korean J Urol. 2013; 54: 791-796.
20. Jiang X, Du L, Duan W, Wang R, Yan K, Wang L, Li J, Zheng G, Zhang X, Yang Y, Wang C. Serum microRNA expression signatures as novel noninvasive biomarkers for prediction and prognosis of muscle-invasive bladder cancer. Oncotarget. 2016; 7: 36733-36742. doi: 10.18632/oncotarget.9166.
21. Yang H, Fang F, Chang R, Yang L. MicroRNA-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor beta receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013; 58: 205-217.
22. Zhai H, Fesler A, Ba Y, Wu S, Ju J. Inhibition of colorectal cancer stem cell survival and invasive potential by hsa-miR-140-5p mediated suppression of Smad2 and autophagy. Oncotarget. 2015; 6: 19735-19746. doi: 10.18632/oncotarget.3771.
23. Butz H, Liko I, Czirjak S, Igaz P, Korbonits M, Racz K, Patocs A. MicroRNA profile indicates downregulation of the TGFbeta pathway in sporadic non-functioning pituitary adenomas. Pituitary. 2011; 14: 112-124.
24. Pais H, Nicolas FE, Soond SM, Swingler TE, Clark IM, Chantry A, Moulton V, Dalmay T. Analyzing mRNA expression identifies Smad3 as a microRNA-140 target regulated only at protein level. RNA. 2010; 16: 489-494.
25. Smith AL, Robin TP, Ford HL. Molecular pathways: targeting the TGF-beta pathway for cancer therapy. Clin Cancer Res. 2012; 18: 4514-4521.
26. Wang M, Du M, Ma L, Chu H, Lv Q, Ye D, Guo J, Gu C, Xia G, Zhu Y, Ding Q, Yuan L, Fu G, et al. A functional variant in TP63 at 3q28 associated with bladder cancer risk by creating an miR-140-5p binding site. Int J Cancer. 2016; 139: 65-74.
27. Kiemeney LA, Thorlacius S, Sulem P, Geller F, Aben KK, Stacey SN, Gudmundsson J, Jakobsdottir M, Bergthorsson JT, Sigurdsson A, Blondal T, Witjes JA, Vermeulen SH, et al. Sequence variant on 8q24 confers susceptibility to urinary bladder cancer. Nat Genet. 2008; 40: 1307-1312.
28. Liu C, Zhang Y, Chen H, Jiang L, Xiao D. Function analysis of rs9589207 polymorphism in miR-92a in gastric cancer. Tumour Biol. 2016; 37: 4439-4444.
29. Ren P, Gong F, Zhang Y, Jiang J, Zhang H. MicroRNA-92a promotes growth, metastasis, and chemoresistance in non-small cell lung cancer cells by targeting PTEN. Tumour Biol. 2016; 37: 3215-3225.
30. Zhao JY, Wang F, Li Y, Zhang XB, Yang L, Wang W, Xu H, Liu DZ, Zhang LY. Five miRNAs Considered as Molecular Targets for Predicting Esophageal Cancer. Med Sci Monit. 2015; 21: 3222-3230.
31. Schee K, Boye K, Abrahamsen TW, Fodstad O, Flatmark K. Clinical relevance of microRNA miR-21, miR-31, miR-92a, miR-101, miR-106a and miR-145 in colorectal cancer. BMC Cancer. 2012; 12: 505.
32. Ahmadi S, Sharifi M, Salehi R. Locked nucleic acid inhibits miR-92a-3p in human colorectal cancer, induces apoptosis and inhibits cell proliferation. Cancer Gene Ther. 2016; 23: 199-205.
33. Wong N, Khwaja SS, Baker CM, Gay HA, Thorstad WL, Daly MD, Lewis JS, Jr., Wang X. Prognostic microRNA signatures derived from The Cancer Genome Atlas for head and neck squamous cell carcinomas. Cancer Med. 2016: 5:1619-1628.
34. Zavesky L, Jandakova E, Turyna R, Langmeierova L, Weinberger V, Zaveska DL, Hulkova M, Horinek A, Duskova D, Feyereisl J, Minar L, Kohoutova M. Evaluation of Cell-Free Urine microRNAs Expression for the Use in Diagnosis of Ovarian and Endometrial Cancers. A Pilot Study. Pathol Oncol Res. 2015; 21: 1027-1035.
35. Urist MJ, Di Como CJ, Lu ML, Charytonowicz E, Verbel D, Crum CP, Ince TA, McKeon FD, Cordon-Cardo C. Loss of p63 expression is associated with tumor progression in bladder cancer. Am J Pathol. 2002; 161: 1199-1206.
36. Sanguedolce F, Bufo P, Carrieri G, Cormio L. Predictive markers in bladder cancer: do we have molecular markers ready for clinical use? Crit Rev Clin Lab Sci. 2014; 51: 291-304.
37. Ribal MJ, Mengual L, Lozano JJ, Ingelmo-Torres M, Palou J, Rodriguez-Faba O, Witjes JA, Van der Heijden AG, Medina R, Conde JM, Marberger M, Schmidbauer J, Fernandez PL, et al. Gene expression test for the non-invasive diagnosis of bladder cancer: A prospective, blinded, international and multicenter validation study. Eur J Cancer. 2016; 54: 131-138.
38. Lopez-Beltran A, Sauter G, Gasser T, Hartmann A, Schmitz-Dräger BJ, Helpap B, Ayala AG, Tamboni P, Knowles MA, Sidransky D, Cordon-Cardo C, Jones PA, Cairns P, Simon R, Amin MB, Tyczynsky JE. Tumours of the Urinary System. In: Eble JN, Sauter G, Epstein JI, Sesterhenn IA. Pathology and Genetics of Tumours of the Urinary System and Male Genital Organs. World Health Organization Classification of Tumours. Lyon: IARC Press, 2004: p. 89-157.
39. Sobin LH, Wittekind CH, eds. TNM Classification of Malignant Tumours. International Union Against Cancer, 6th ed. New York: Jonh Wiley & Sons, 2002.
40. Mengual L, Burset M, Marin-Aguilera M, Ribal MJ, Alcaraz A. Multiplex preamplification of specific cDNA targets prior to gene expression analysis by TaqMan Arrays. BMC Res Notes. 2008; 1: 21.
41. Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002; 3: RESEARCH0034.
42. Vlachos IS, Zagganas K, Paraskevopoulou MD, Georgakilas G, Karagkouni D, Vergoulis T, Dalamagas T, Hatzigeorgiou AG. DIANA-miRPath v3.0: deciphering microRNA function with experimental support. Nucleic Acids Res. 2015; 43: W460-W466.

