Introduction
The importance of the ligand-receptor axis involving high mobility group protein box1 (HMGB1) and the receptor for advanced glycation end products (RAGE) in inflammation and tumorigenesis has been increasingly recognized by many experimental and clinical investigators [1-4]. HMGB1 is a nuclear DNA-binding factor that exerts an important regulatory role in chromatin architecture and transcriptional regulation [5]. The binding of HMGB1 to RAGE can stimulate the expression of Ras, PI3K, and Rho, which are major downstream signaling molecules responsible for the NF-κB activation [6, 7]. Studies in growing numbers have reported the concordant over-expression of HMGB1 and RAGE in a variety of cancer types, including hepatocellular carcinoma [8-11]. Several lines of clinical evidence suggested that HMGB1 and RAGE possessed the potentials to serve as powerful prognostic and therapeutic targets for hepatocellular carcinoma [12-14]. The genes coding for HMGB1 and RAGE mapped on different chromosomes carry many polymorphic loci. There is compelling evidence that circulating variation of HMGB1 and RAGE is predominantly under genetic control [15-17]. One therefore could hypothesize that HMGB1 and RAGE genes are two potential candidate susceptibility genes for hepatocellular carcinoma.
To test this hypothesis, we designed a multicenter case-control study and genotyped seven widely-studied polymorphisms from HMGB1 and RAGE genes among 540 hepatocellular carcinoma patients and 540 age- and gender-matched controls.
Results
Basic characteristics
The summarization and comparison of basic characteristics between 540 patients and 540 matched controls are shown in Table 1. Patients were matched with controls on age and gender. The mean level of BMI was comparable between patients and controls (P = 0.372). By contrast, the percentages of current or ever cigarette smokers and alcohol drinkers were significantly higher in patients than in controls (P < 0.001).
Table 1: The summarization and comparison of basic characteristics between 540 patients and 540 age- and gender-matched controls
Characteristics |
Patients |
Controls |
P* |
Age (years) |
51.5 (6.7) |
50.4 (6.8) |
0.165 |
Male gender |
61.7% |
61.3% |
0.900 |
BMI (kg/m2) |
25.4 ± 2.2 |
25.7 ± 2.0 |
0.372 |
Current/ever smokers |
58.9% |
33.9% |
<0.001 |
Current/ever alcohol drinkers |
63.9% |
34.4% |
<0.001 |
BMI: body mass index. Quantitative data are listed as the mean ± standard deviation, and categorical data are listed as the percentage. Quantitative and categorical data are compared between patients and controls with the t test and Chi-squared test, respectively.
Single polymorphism analysis
The genotypes and alleles of seven polymorphisms under study are compared in Table 2. There was no significant deviation from the Hardy-Weinberg equilibrium for all seven polymorphisms in control subjects (all P > 0.05), and their linkage patterns within each gene are illustrated in Figure 1. The mutant alleles of two polymorphisms, rs1045411 in HMGB1 gene and rs2070600 in RAGE gene, had significantly higher frequencies in hepatocellular carcinoma patients than in controls (both P < 0.001), and this significance was attained even after the Bonferroni correction (P < 0.05/7). Further, the power to detect the significance was 99.6% and 99.9% respectively for allele comparisons of rs1045411 and rs2070600 between patients and controls. In addition, the genotypes of rs1800625 differed significantly between the two groups (P = 0.008), while its allele distributions were only marginally significant (P = 0.034) that did not survive the Bonferroni correction. The other polymorphisms showed no statistical significance in susceptibility to hepatocellular carcinoma.
Table 2: The comparison of the genotypes and alleles of seven polymorphisms from HMGB1 and RAGE genes between 540 patients and 540 age- and gender-matched controls
Polymorphisms |
w/m |
Group |
Genotype |
Allele |
Pgenotype |
Pallele |
||
ww |
wm |
mm |
m |
|||||
rs1045411 |
C/T |
Patients |
349 |
158 |
33 |
20.82% |
<0.001 |
<0.001 |
Controls |
405 |
127 |
8 |
13.26% |
||||
rs2249825 |
C/G |
Patients |
349 |
168 |
23 |
19.84% |
0.521 |
0.513 |
Controls |
354 |
170 |
16 |
18.75% |
||||
rs1415125 |
T/C |
Patients |
273 |
216 |
51 |
29.38% |
0.588 |
0.272 |
Controls |
290 |
205 |
45 |
27.27% |
||||
rs1800625 |
T/C |
Patients |
403 |
107 |
30 |
15.37% |
0.008 |
0.034 |
Controls |
417 |
113 |
10 |
12.31% |
||||
rs1800624 |
T/A |
Patients |
374 |
147 |
19 |
17.12% |
0.610 |
0.999 |
Controls |
370 |
155 |
15 |
17.05% |
||||
rs2070600 |
G/A |
Patients |
254 |
202 |
84 |
34.24% |
<0.001 |
<0.001 |
Controls |
321 |
190 |
29 |
22.92% |
||||
rs184003 |
G/T |
Patients |
277 |
221 |
42 |
28.21% |
0.194 |
0.064 |
Controls |
303 |
207 |
30 |
24.81% |
||||
ww: homozygous wild genotype; wm: heterozygous genotype; mm: homozygous mutant genotype; m: mutant allele. Genotypes and alleles were compared between patients and controls with the Chi-squared test.
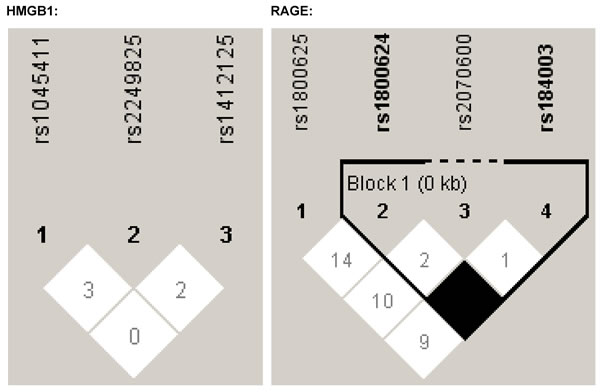
Figure 1: The linkage patterns of polymorphisms in HMGB1 and RAGE genes
The risk prediction of each polymorphism under study for hepatocellular carcinoma is shown in Table 3. After adjusting for body mass index (BMI), smoking and drinking, only rs1045411 and rs2070600 polymorphisms were significantly associated with an increased risk of hepatocellular carcinoma, particularly for rs2070600 under both genetic models. In detail, the mutant allele carriers of rs2070600 had a 1.77-fold (95% CI: 1.34-2.32; P < 0.001) and 1.75-fold (95% CI: 1.23-2.50; P = 0.002) increased risk after adjustment under the additive and dominant models, respectively. There was no statistical significance for the other five polymorphisms under study.
Table 3: The risk prediction of seven polymorphisms from HMGB1 and RAGE genes for hepatocellular carcinoma before and after adjusting for confounders under the additive and dominant models
Polymorphisms |
Additive model |
Dominant model |
||
Unadjusted |
Adjusted |
Unadjusted |
Adjusted |
|
rs1045411 |
1.67; 1.21-2.31; 0.002 |
1.64; 1.18-2.29; 0.003 |
1.64; 1.13-2.40; 0.010 |
1.62; 1.10-2.39; 0.014 |
rs2249825 |
1.07; 0.79-1.46; 0.654 |
1.00; 0.73-1.37; 0.997 |
1.04; 0.73-1.49; 0.822 |
0.95; 0.66-1.38; 0.784 |
rs1415125 |
1.10; 0.85-1.44; 0.460 |
1.12; 0.85-1.47; 0.414 |
1.14; 0.81-1.60; 0.464 |
1.13; 0.80-1.61; 0.482 |
rs1800625 |
1.25; 0.90-1.74; 0.183 |
1.20; 0.85-1.68; 0.295 |
1.15; 0.77-1.72; 0.493 |
1.08; 0.71-1.63; 0.718 |
rs1800624 |
1.01; 0.73-1.39; 0.974 |
1.02; 0.73-1.41; 0.919 |
0.97; 0.67-1.40; 0.863 |
0.96; 0.66-1.40; 0.831 |
rs2070600 |
1.67; 1.28-2.17; <0.001 |
1.77; 1.34-2.32; <0.001 |
1.65; 1.17-2.33; 0.005 |
1.75; 1.23-2.50; 0.002 |
rs184003 |
1.19; 0.90-1.58; 0.211 |
1.20; 0.90-1.59; 0.215 |
1.21; 0.86-1.71; 0.282 |
1.21; 0.85-1.72; 0.295 |
OR: odds ratio; 95% CI: 95% confidence interval. *P was calculated after adjusting for body mass index, smoking and drinking under the conditional binary Logistic regression model. Additive model was defined as homozygous wild genotype vs. heterozygous genotype vs. homozygous mutant genotype. Dominant model was defined as homozygous wild genotype vs. heterozygous genotype plus homozygous mutant genotype.
Haplotype analysis
Haplotype analysis was separately conducted in HMGB1 and RAGE genes in view of different chromosomes they mapped on, and the corresponding results are shown in Table 4. As the low-penetrance haplotypes usually carry a high risk of producing chance false-positive findings, haplotype analysis was only restricted to the common haplotypes, which had an estimated frequency of at least 3% in both patients and controls.
In HMGB1 gene, the frequency of T-C-T haplotype (rs1045411-rs2249825-rs1415125, similarly hereinafter) was significantly higher in patients than in controls (12.59% vs. 5.56%, P = 0.003) with the study power reaching 99.9%, and this haplotype was associated with a 2.47-fold (95% CI: 1.41-4.34; P = 0.002) increased risk of hepatocellular carcinoma after adjusting for BMI, smoking and drinking when compared with the commonest haplotype C-C-T.
In RAGE gene, the commonest T-T-G-G haplotype (rs1800625-rs1800624-rs2070600-rs184003, similarly hereinafter) was frequently seen in controls compared with patients, and relative to this haplotype, T-T-A-G (OR; 95% CI; P: 1.75; 1.02-3.03; 0.045) and T-T-A-T (OR; 95% CI; P: 1.95; 1.01-3.76; 0.048) haplotypes were associated with a marginally increased risk of hepatocellular carcinoma after adjusting for confounders mentioned above, with the study power of 28.0% and 71.9%, respectively.
Table 4: The comparison of derived common haplotypes based on seven polymorphisms from HMGB1 and RAGE genes between 540 patients and 540 age- and gender-matched controls, and their risk prediction for hepatocellular carcinoma
Haplotypes |
Patients |
Controls |
P |
Crude OR; 95% CI; P |
Adjusted OR; 95% CI; P* |
HMGB1 gene (rs1045411-rs2249825-rs1415125) |
|||||
C-C-T |
45.01% |
53.84% |
0.013 |
Reference haplotype |
|
C-C-C |
17.18% |
17.79% |
0.632 |
1.14; 0.77-1.69; 0.516 |
1.18; 0.79-1.76; 0.428 |
T-C-T |
12.59% |
5.56% |
0.003 |
2.55; 1.44-4.5; 0.001 |
2.47; 1.41-4.34; 0.002 |
C-G-T |
10.66% |
9.71% |
0.946 |
1.26; 0.76-2.11; 0.371 |
1.18; 0.7-1.98; 0.544 |
C-G-C |
6.33% |
5.42% |
0.466 |
1.39; 0.72-2.66; 0.321 |
1.27; 0.66-2.47; 0.476 |
T-C-C |
5.38% |
4.07% |
0.087 |
1.48; 0.75-2.95; 0.262 |
1.46; 0.73-2.92; 0.278 |
RAGE gene (rs1800625-rs1800624-rs2070600-rs184003) |
|||||
T-T-G-G |
30.43% |
39.43% |
0.002 |
Reference haplotype |
|
T-T-G-T |
16.02% |
18.29% |
0.482 |
1.16; 0.75-1.78; 0.502 |
1.22; 0.78-1.9; 0.382 |
T-T-A-G |
13.98% |
11.99% |
0.118 |
1.53; 0.9-2.58; 0.116 |
1.75; 1.02-3.03; 0.045 |
T-A-G-G |
10.64% |
10.11% |
0.615 |
1.36; 0.81-2.29; 0.243 |
1.44; 0.84-2.46; 0.182 |
T-T-A-T |
7.44% |
4.82% |
0.032 |
1.88; 0.99-3.6; 0.056 |
1.95; 1.01-3.76; 0.048 |
C-T-G-G |
5.93% |
4.82% |
0.785 |
1.43; 0.73-2.8; 0.293 |
1.41; 0.71-2.79; 0.326 |
OR: odds ratio; 95% CI: 95% confidence interval. *P was calculated after adjusting for body mass index, smoking and drinking.
Discussion
In this multicenter matched case-control study, we set out to examine the hypothesis that HMGB1 and RAGE genes are two potential candidate susceptibility genes for hepatocellular carcinoma. Through extensive analyses, our findings support this hypothesis by identifying two risk associated polymorphisms, rs1045411 and rs2070600 from the HMGB1/RAGE axis, and more importantly a joint impact of seven polymorphisms in susceptibility to hepatocellular carcinoma.
Hepatocellular carcinoma is one of the common solid malignancies in the world, and it is particularly prevalent in China [18, 19]. Many epidemiological studies have suggested that a strong genetic component underlies the hepatocarcinogenesis [20, 21]. Although genome-wide association studies are widely utilized to characterize the genetic profiles of hepatocellular carcinoma, the results seem far from promising [22, 23]. Given the ubiquity of genetic heterogeneity, it is of special importance to develop a catalogue of cancer-susceptibility genes or polymorphisms within individual ethnic groups. Bearing this in mind, we selected seven well-characterized polymorphisms from HMGB1 and RAGE genes to assess their genetic susceptibility to hepatocellular carcinoma in a large Han Chinese population.
First, it is interesting to notice a leading role played by rs1045411 and rs2070600 polymorphisms that conferred a significant increased risk for hepatocellular carcinoma, and the two polymorphisms were respectively resided in HMGB1 and RAGE genes. On one hand, the polymorphism rs1045411, which is located in the 3’-untranslated region of HMGB1, might be biologically functional as the mutant allele of this polymorphism was recently reported to be significantly associated with higher positive blood culture rates and elevated cytokine levels, particularly in patients with chronic lung diseases or diabetes as co-morbidities [24]. So far the physiological significance of rs1045411 is uncertain, and the location of this polymorphism could suggest a role in mRNA stability as microRNAs can bind the 3’-untranslated regions of mRNA transcripts and inhibit gene expression at the posttranscriptional level. Moreover, in the present study, the mutant allele of rs1045411 was associated with a significantly increased risk of hepatocellualr carcinoma, while another study in a Taiwan population found that carriers of this mutant allele had a lower risk [25], likely due to differences in lifestyles, diets or study power. On the other hand, the polymorphism rs2070600 is non-synonymous (Gly82Ser) in the exon 3 of RAGE gene, and the association of this polymorphism was significant with a wide range of cancer types, including lung cancer [26], breast cancer [27], ovarian cancer [28] and colorectal cancer [29]. However, a recent study by Su et al failed to support this significant association in Taiwanese, and instead they found that another promoter polymorphism rs1800625 in RAGE gene was linked to the increased risk for hepatocellular carcinoma [30]. Besides genetic profiles, the pathogenesis of hepatocellular carcinoma is also closely related to environmental, dietary, life style factors, which are sharply different between northern Han Chinese in the present study and Taiwanese. It remains possible to speculate that if involved, the rs2070600 mutation might account for circulating variation of soluble RAGE, which was demonstrated to play a protective role in the development of cancer by a meta-analysis [16].
Second, it is widely recognized that single locus may not by itself have a significant association with a disease because its effect may be small and dependent on the nearby loci that compensate for variation in the locus under study [31]. To address this claim, we performed the haplotype analysis, and it is worth noting for a joint impact of multiple polymorphisms from the HMGB1/RAGE axis on the hepatocarcinogenesis. This finding is understandable in view of the fact that the role of any given locus in the course of hepatocellular carcinoma is likely to be small on average. Nevertheless, this study provides preliminary evidence that potentially unfavorable combinations of polymorphisms from the HMGB1/RAGE axis could synergistically affect the onset and progression of hepatocellular carcinoma.
Finally, there are several limitations to this study. Firstly, this is a multicenter case-control study, and the probability of population stratification and admixture might be high. However, the conformity of seven polymorphisms to the Hardy-Weinberg equilibrium can somewhat reduce this probability. Secondly, the total sample size of the present study (N = 1080) is not large enough to justify the impact of low-penetrance alleles or genotypes. However, in case of significant findings in the present study, the statistical power to detect a significant allele difference between patients and controls was over 99%, indicating the somewhat validity of our association analyses. Thirdly, because this study was carried out in a Han Chinese population, we are reticent in extrapolating our chief conclusions to the other races or ethnicities, as it is widely recognized that strong genetic heterogeneity underlies the development of hepatocellular carcinoma [32]. Fourthly, although patients and controls were matched on age and gender at enrolment, other lifestyle factors (such as smoking and drinking habits) different remarkably between the two groups, and clinical markers are not available for us to analyze, leaving some room for further explorations especially from the aspect of gene-to-environment interaction. We agree that a large-scale well-designed study is warranted to overcome these limitations and produce convincing evidence.
In summary, we identified two risk-associated polymorphisms (rs1045411 and rs2070600) and more importantly a joint impact of seven polymorphisms from the HMGB1/RAGE axis in susceptibility to hepatocellular carcinoma. From a broader clinical perspective, a greater understanding of this axis is critical to furthering our understanding of the genetic underpinnings of hepatocellular carcinoma or related solid tumors.
materials and Methods
Study subjects
All study subjects were gathered from three local hospitals (Yuhuangding Hospital, Yantai Affiliated Hospital of Binzhou Medical University and Shandong Provincial Hospital) in Shandong province, China between February 2014 and September 2016, and they gave informed, written consent prior to participation. The protocols of this study were approved by the ethics committees at all participating hospitals.
Basic characteristics
The clinical diagnosis of hepatocellular carcinoma was confirmed by histopathological examinations. Hepatocellular carcinoma patients (n = 540) were matched on age (within 2-year strata) and gender with 540 cancer-free controls gathered from the same hospital that had no self-reported family history of any malignancies excluding non-melanoma skin cancer within three generations. For patients, age was recorded for the initial onset age of hepatocellular carcinoma. Besides baseline age and gender, we also recorded body height, body weight, tobacco smoking and alcohol drinking. Body height and weight were measured in light clothing, and body mass index (BMI) was calculated as weight in kilometer divided by height in meters-squared. Tobacco smokers refer to the current or ever smokers (at least ten cigarettes per week) who smoked more than 3 months. Alcohol drinkers refer to the current or ever drinkers (at least one time a week) who drank more than three months.
At the time of enrolment, each study subject provided a fasted peripheral venous blood sample, which was centrifuged within the same day at 3000×g for 5 min at room temperature, and then was kept at minus 40ºC temperature until batch assayed.
Polymorphism selection
From HMGB1 gene, three polymorphisms including rs1045411 (3’-untranslated region), rs2249825 (promoter region) and rs1415125 (promoter region) were selected. From RAGE gene, four polymorphisms including rs1800625 (promoter region), rs1800624 (promoter region), rs2070600 (exon 3) and rs184003 (intron 7) were selected. These polymorphisms were widely studied in predisposition to a variety of cancer types, including breast cancer [33], gastric cancer [34, 35], colorectal cancer [29, 36], lung cancer [37, 38] and so on.
Genotype determination
Genomic DNA was prepared from white blood cells, and was extracted by either a phenol-chloroform method or a salting-out method. Genotypes of seven polymorphisms under study were determined by the popular ligase detection reaction method, as previously reported by Niederhauser and coworker [39]. The accuracy of this genotyping method was validated by re-genotyping 10% (N = 108) of all DNA samples through a random selection manner. The genotyping results of these 108 duplicated samples fitted perfectly.
Statistical analysis
Quantitative data were summarized as the mean ± standard deviation, and categorical data were summarized as the percentage. The Hardy-Weinberg equilibrium refers to the consistency of genotype frequencies with the two alleles being independently sampled from a population of alleles, and it was tested by the Chi-squared test in controls to avoid gross genotyping misclassifications. Genetic inheritance pattern of each polymorphism was assessed to be either additive (wild homozygote vs. heterozygote vs. mutant homozygote) or dominant (wild homozygote vs. combined heterozygote and mutant homozygote). A multivariate conditional binary Logistic regression analysis was conducted to model individual polymorphisms after considering the confounding impact of BMI, smoking and drinking. Further haplotype analysis was performed to examine possible joint impact of individual polymorphisms within the same gene after adjustment. In theory, a haplotype is defined as a combination of two or more alleles on a single chromosome. The frequencies of derived haplotypes were compared between patients and controls, and their risk prediction for hepatocellular carcinoma was calculated after adjusting for BMI, smoking and drinking. All tests are two-sided, and P < 0.05 was considered statistically significant unless otherwise indicated. Above calculations were performed with the R software (v3.3.1), which can be freely downloaded from the website https://www.r-project.org/. Haplotype analyses were performed with the HAPLO.STATS program (v1.7.1) realized under the R environment (v3.3.1). The HAPLOTYPE.STATS program provides different methods to testing the association of haplotypes with traits after adjustment for effects from other confounders, which has a statistical advantage to further the understanding of the roles of different haplotypes [40, 41]. Linkage disequilibrium patterns between polymorphisms within the same gene was assessed by calculating D’/LOD (LOD is the log of the likelihood odds ratio, a measure of confidence in the value of D’) with the Haploview (v4.2). Study power was estimated with the CaTS software (http://www.sph.umich.edu/csg/abecasis/CaTS/index.html).
Funding
This work was conducted under the supports from the Taishan Scholars Construction Engineering, the National Natural Science Foundation of China (81400771 and 31671139), the Shandong Provincial Natural Science Foundation (ZR2014HL033), the Shandong Provincial Science & Technology Project (J14LE01 and J15LK03), the Yantai Science & Technology Project (2015ZH083) and the Binzhou Medical University Scientific Research Funds (BY2013KYQD17 and BY2013KYQD18).
Conflicts of interests
There are no competing interests.
References
1. Hiwatashi K, Ueno S, Abeyama K, Kubo F, Sakoda M, Maruyama I, Hamanoue M, Natsugoe S, Aikou T. A novel function of the receptor for advanced glycation end-products (RAGE) in association with tumorigenesis and tumor differentiation of HCC. Ann Surg Oncol. 2008; 15l:923-933.
2. DiNorcia J, Moroziewicz DN, Ippagunta N, Lee MK, Foster M, Rotterdam HZ, Bao F, Zhou YS, Yan SF, Emond J, Schmidt AM, Allendorf JD. RAGE signaling significantly impacts tumorigenesis and hepatic tumor growth in murine models of colorectal carcinoma. J Gastrointest Surg. 2010; 14l:1680-1690.
3. Pusterla T, Nemeth J, Stein I, Wiechert L, Knigin D, Marhenke S, Longerich T, Kumar V, Arnold B, Vogel A, Bierhaus A, Pikarsky E, Hess J, et al. Receptor for advanced glycation endproducts (RAGE) is a key regulator of oval cell activation and inflammation-associated liver carcinogenesis in mice. Hepatology. 2013; 58l:363-373.
4. Sundar IK, Javed F, Romanos GE, Rahman I. E-cigarettes and flavorings induce inflammatory and pro-senescence responses in oral epithelial cells and periodontal fibroblasts. Oncotarget. 2016; 7:77196-77204. doi: 10.18632/oncotarget.12857.
5. Lamore SD, Cabello CM, Wondrak GT. HMGB1-directed drug discovery targeting cutaneous inflammatory dysregulation. Curr Drug Metab. 2010; 11l:250-265.
6. Lee SA, Kwak MS, Kim S, Shin JS. The role of high mobility group box 1 in innate immunity. Yonsei Med J. 2014; 55l:1165-1176.
7. Reed KR, Song F, Young MA, Hassan N, Antoine DJ, Gemici NB, Clarke AR, Jenkins JR. Secreted HMGB1 from Wnt activated intestinal cells is required to maintain a crypt progenitor phenotype. Oncotarget. 2016; 7:51665-51673. doi: 10.18632/oncotarget.10076.
8. Cheng BQ, Jia CQ, Liu CT, Lu XF, Zhong N, Zhang ZL, Fan W, Li YQ. Serum high mobility group box chromosomal protein 1 is associated with clinicopathologic features in patients with hepatocellular carcinoma. Dig Liver Dis. 2008; 40l:446-452.
9. Chung HW, Lee SG, Kim H, Hong DJ, Chung JB, Stroncek D, Lim JB. Serum high mobility group box-1 (HMGB1) is closely associated with the clinical and pathologic features of gastric cancer. J Transl Med. 2009; 7l:38.
10. Chen RC, Yi PP, Zhou RR, Xiao MF, Huang ZB, Tang DL, Huang Y, Fan XG. The role of HMGB1-RAGE axis in migration and invasion of hepatocellular carcinoma cell lines. Mol Cell Biochem. 2014; 390l:271-280.
11. Wu T, Zhang W, Yang G, Li H, Chen Q, Song R, Zhao L. HMGB1 overexpression as a prognostic factor for survival in cancer: a meta-analysis and systematic review. Oncotarget. 2016; 7:50417-50427. doi: 10.18632/oncotarget.10413.
12. Ito R, Ishii Y, Wakiyama S, Shiba H, Fujioka S, Misawa T, Ishida Y, Hano H, Yanaga K. Prognostic significance of receptor for advanced glycation end products expression in hepatocellular carcinoma after hepatectomy. J Surg Res. 2014; 192l:503-508.
13. Kohles N, Nagel D, Jungst D, Stieber P, Holdenrieder S. Predictive value of immunogenic cell death biomarkers HMGB1, sRAGE, and DNase in liver cancer patients receiving transarterial chemoembolization therapy. Tumour Biol. 2012; 33l:2401-2409.
14. Moy KA, Jiao L, Freedman ND, Weinstein SJ, Sinha R, Virtamo J, Albanes D, Stolzenberg-Solomon RZ. Soluble receptor for advanced glycation end products and risk of liver cancer. Hepatology. 2013; 57l:2338-2345.
15. Wang H, Li Y, Yu W, Ma L, Ji X, Xiao W. Expression of the receptor for advanced glycation end-products and frequency of polymorphism in lung cancer. Oncol Lett. 2015; 10l:51-60.
16. He L, Bao H, Xue J, Zheng L, Zhang Q, Sun L, Pan H. Circulating soluble advanced glycation end product is inversely associated with the significant risk of developing cancer: evidence from a meta-analysis. Tumour Biol. 2014; 35l:8749-8755.
17. Tesarova P, Kalousova M, Jachymova M, Mestek O, Petruzelka L, Zima T. Receptor for advanced glycation end products (RAGE)--soluble form (sRAGE) and gene polymorphisms in patients with breast cancer. Cancer Invest. 2007; 25l:720-725.
18. Tanaka M, Katayama F, Kato H, Tanaka H, Wang J, Qiao YL, Inoue M. Hepatitis B and C virus infection and hepatocellular carcinoma in China: a review of epidemiology and control measures. J Epidemiol. 2011; 21l:401-416.
19. Zhu Q, Li N, Zeng X, Han Q, Li F, Yang C, Lv Y, Zhou Z, Liu Z. Hepatocellular carcinoma in a large medical center of China over a 10-year period: evolving therapeutic option and improving survival. Oncotarget. 2015; 6l:4440-4450. doi: 10.18632/oncotarget.2913.
20. Buendia MA. Genetics of hepatocellular carcinoma. Semin Cancer Biol. 2000; 10l:185-200.
21. Knudsen ES, Gopal P, Singal AG. The changing landscape of hepatocellular carcinoma: etiology, genetics, and therapy. Am J Pathol. 2014; 184l:574-583.
22. Kumar V, Kato N, Urabe Y, Takahashi A, Muroyama R, Hosono N, Otsuka M, Tateishi R, Omata M, Nakagawa H, Koike K, Kamatani N, Kubo M, et al. Genome-wide association study identifies a susceptibility locus for HCV-induced hepatocellular carcinoma. Nat Genet. 2011; 43l:455-458.
23. Zhang H, Zhai Y, Hu Z, Wu C, Qian J, Jia W, Ma F, Huang W, Yu L, Yue W, Wang Z, Li P, Zhang Y, et al. Genome-wide association study identifies 1p36.22 as a new susceptibility locus for hepatocellular carcinoma in chronic hepatitis B virus carriers. Nat Genet. 2010; 42l:755-758.
24. Lee K, Chang Y, Song K, Park YY, Huh JW, Hong SB, Lim CM, Koh Y. Associations between Single Nucleotide Polymorphisms of High Mobility Group Box 1 Protein and Clinical Outcomes in Korean Sepsis Patients. Yonsei Med J. 2016; 57l:111-117.
25. Wang B, Yeh CB, Lein MY, Su CM, Yang SF, Liu YF, Tang CH. Effects of HMGB1 Polymorphisms on the Susceptibility and Progression of Hepatocellular Carcinoma. Int J Med Sci. 2016; 13l:304-309.
26. Wang X, Cui E, Zeng H, Hua F, Wang B, Mao W, Feng X. RAGE genetic polymorphisms are associated with risk, chemotherapy response and prognosis in patients with advanced NSCLC. PLoS One. 2012; 7l:e43734.
27. Archontogeorgis K, Nena E, Papanas N, Steiropoulos P. Biomarkers to improve diagnosis and monitoring of obstructive sleep apnea syndrome: current status and future perspectives. Pulm Med. 2014; 2014l:930535.
28. Zhang S, Hou X, Zi S, Wang Y, Chen L, Kong B. Polymorphisms of receptor for advanced glycation end products and risk of epithelial ovarian cancer in Chinese patients. Cell Physiol Biochem. 2013; 31l:525-531.
29. Qian F, Sun BL, Zhang WY, Ke J, Zhu J. Gly82Ser polymorphism of the receptor for advanced glycation end-product (RAGE) potential high risk in patients with colorectal cancer. Tumour Biol. 2014; 35l:3171-3175.
30. Su SC, Hsieh MJ, Chou YE, Fan WL, Yeh CB, Yang SF. Effects of RAGE Gene Polymorphisms on the Risk and Progression of Hepatocellular Carcinoma. Medicine (Baltimore). 2015; 94l:e1396.
31. Papaemmanuil E, Carvajal-Carmona L, Sellick GS, Kemp Z, Webb E, Spain S, Sullivan K, Barclay E, Lubbe S, Jaeger E, Vijayakrishnan J, Broderick P, Gorman M, et al. Deciphering the genetics of hereditary non-syndromic colorectal cancer. Eur J Hum Genet. 2008; 16l:1477-1486.
32. Lin ZH, Xin YN, Dong QJ, Wang Q, Jiang XJ, Zhan SH, Sun Y, Xuan SY. Association between HLA-DRB1 alleles polymorphism and hepatocellular carcinoma: a meta-analysis. BMC Gastroenterol. 2010; 10l:145.
33. Yue L, Zhang Q, He L, Zhang M, Dong J, Zhao D, Ma H, Pan H, Zheng L. Genetic predisposition of six well-defined polymorphisms in HMGB1/RAGE pathway to breast cancer in a large Han Chinese population. J Cell Mol Med. 2016; 20l:1966-1973.
34. Gu H, Yang L, Sun Q, Zhou B, Tang N, Cong R, Zeng Y, Wang B. Gly82Ser polymorphism of the receptor for advanced glycation end products is associated with an increased risk of gastric cancer in a Chinese population. Clin Cancer Res. 2008; 14l:3627-3632.
35. Bao G, Qu F, He L, Zhao H, Wang N, Ji G, He X. Prognostic Significance of Tag SNP rs1045411 in HMGB1 of the Aggressive Gastric Cancer in a Chinese Population. PLoS One. 2016; 11l:e0154378.
36. Wang JX, Yu HL, Bei SS, Cui ZH, Li ZW, Liu ZJ, Lv YF. Association of HMGB1 Gene Polymorphisms with Risk of Colorectal Cancer in a Chinese Population. Med Sci Monit. 2016; 22l:3419-3425.
37. Wang Y, Li XP, Yin JY, Zhang Y, He H, Qian CY, Chen J, Zheng Y, Smieszkol K, Fu YL, Chen ZY, Zhou HH, Liu ZQ. Association of HMGB1 and HMGB2 genetic polymorphisms with lung cancer chemotherapy response. Clin Exp Pharmacol Physiol. 2014; 41l:408-415.
38. Pan H, Niu W, He L, Wang B, Cao J, Zhao F, Liu Y, Li S, Wu H. Contributory role of five common polymorphisms of RAGE and APE1 genes in lung cancer among Han Chinese. PLoS One. 2013; 8l:e69018.
39. Niederhauser C, Kaempf L, Heinzer I. Use of the ligase detection reaction-polymerase chain reaction to identify point mutations in extended-spectrum beta-lactamases. Eur J Clin Microbiol Infect Dis. 2000; 19l:477-480.
40. Schaid DJ, Rowland CM, Tines DE, Jacobson RM, Poland GA. Score tests for association between traits and haplotypes when linkage phase is ambiguous. Am J Hum Genet. 2002; 70l:425-434.
41. Wang Y, Li B, Zhao W, Liu P, Zhao Q, Chen S, Li H, Gu D. Association study of G protein-coupled receptor kinase 4 gene variants with essential hypertension in northern Han Chinese. Ann Hum Genet. 2006; 70l:778-783.

