INTRODUCTION
Colorectal cancer (CRC) is a leading cause of cancer and cancer deaths in the Western countries [1]. In many developing countries in Asia, the incidence of CRC is rising [2] as diet and lifestyle change.
CRC, as other complex diseases, is caused by both genetic and environmental factors. Twin studies have shown that inherited genetic factors contribute approximately 35% of the disease etiology [3, 4]. For the past decades, linkage studies on multi-case families have identified a number of rare mutations in highly penetrant genes (e.g. FAP, HNPCC, DNA mismatch repair (MMR) gene). Mutations in those genes are responsible for less than 5% of cases in the pathogenesis of CRC [5]; While, mildly or moderately penetrant genes could explain about 8.3% of etiology in cases with familial aggregation [6, 7]. It is expected that low-risk variants likely explain the remaining proportion of inherited susceptibility [5]. The candidate gene screening at some susceptibility loci such as (MTHFR) [8, 9] and cyclin D1 [10, 11], as well as the genome-wide expression studies [12, 13] were reported to identify associations with colorectal tumorigenesis. However, the replication and functional studies are needed to support the findings.
Recently association studies, directly comparing the frequencies of genetic variants between large series of affected cases and unrelated controls, are now considered to be more appropriate than linkage studies for the identification of susceptibility loci for complex diseases including CRC [14]. Over the past decade, with the resources of human genome and high-throughput platforms, Genome-wide association studies (GWAS) provide a powerful new approach allowing the scanning of the entire genome for association with disease to identify common, low-penetrance susceptibility loci without prior knowledge of biological function. GWAS have identified a number of loci that increase the risk of developing CRC. Early GWAS conducted in populations of European ancestry living in the United Kingdom (UK) and Canada have been published for the identification of 11 well-replicated disease loci, of which most were not previously suspected to be related to CRC. These variants map to 8q24.21 (rs6983267), 8q23.3 (rs16892766, EIF3H), 9q24 (rs719725), 10p14 (rs10795668), 11q23 (rs3802842), 14q22.2 (rs4444235, BMP4), 15q13.3 (rs4779584), 16q22.1 (rs9929218, CDH1), 18q21.1 (rs4939827, SMAD7), 19q13.1 (rs10411210, RHPN2) and 20p12.3 (rs961253) [15, 16]. The proportions of the familial and population risks explained by the published loci are small, thus a great deal of work is needed to understand the biological mechanisms underlying these associations. However, individual GWAS require large sample sizes to account for the inflated Type I error resulting from the very high number of case-control comparisons and to detect effect sizes that are expected to be small. Therefore, pooled data analysis can enhance the statistical power of the study. It enables us to summarize available evidence from larger sample sizes.
To date, over 40 loci have been reported to be associated with CRC risk, among which 19 Single nucleotide polymorphisms (SNPs) (Table 1) were reported at least twice by 2 or more study groups. Five SNPs at loci 10p14, 14q22.2, 15q13.3, 18q21.1 and 8q24.21 were replicated at least 3 times in publications [16-30]. However, the results from past studies on these SNPs are either controversial or inconclusive, such as rs10795668 and rs4779584 with some studies supporting a significant association, whereas others finding no association or an association in opposite direction. Thus, we performed an up to date meta-analysis to more precisely characterize the association between these 5 SNPs and CRC on a larger sample sizes.
Table 1: SNPs frequently reported to be associated with CRC risk
SNPs* |
Chr/nearest Gene |
Position |
Reference |
|---|---|---|---|
rs10411210 |
19q13.1/RHPN2 |
38224140 |
17,18 |
rs10774214 |
12p13.32/CCND2 |
4238613 |
10,22 |
rs10795668 |
10p14 |
8741225 |
15,18,20 |
rs12953717 |
18q21.1/SMAD7 |
44707927 |
13,16 |
rs16892766 |
8q23.3/EIF3H |
117699995 |
15,18 |
rs1862748 |
16q22/CDH1 |
67390444 |
17,21 |
rs2423279 |
20p12.3/HAO1 |
7760350 |
10,22 |
rs3802842 |
11q23 |
110676919 |
16,18 |
rs4444235 |
14q22.2/BMP4 |
53480669 |
17,18,24 |
rs4779584 |
15q13.3/GREM1 |
30782048 |
15,18,21 |
rs4925386 |
20q13.33/LAMA5 |
60354439 |
12,18 |
rs4939827 |
18q21.1/SMAD7 |
44707461 |
13,15,16,18 |
rs5934683 |
Xp22.2/SHROOM2 |
9711474 |
10,25 |
rs647161 |
5q31.1/PITX1 |
134526991 |
10,22 |
rs6983267 |
8q24.21/POU5F1, MYC |
128482487 |
11,14,15,18,19,24 |
rs7136702 |
12q13.13/LARP4 |
49166483 |
12,25 |
rs7229639 |
18q21.1/SMAD7 |
44704974 |
22,23 |
rs961253 |
20p12.3/BMP2 |
6352281 |
17,18 |
rs9929218 |
16q22.1/CDH1 |
67378447 |
17,18 |
*5 SNPs in bold were selected for further analysis.
RESULTS AND DISCUSSION
Five most frequently reported SNPs were selected
Thirty relevant articles of CRC GWAS were identified after searching GWAS Central and PubMed. There were 149 individual SNPs reported to be associated with CRC risk. 19 of them were reported at least twice by different studies, 5 SNPs among which were most frequently reported: rs6983267, rs4939827, rs10795668, rs4444235 and rs4779584 (Table 1).
Characteristics of included studies
As shown in Figure 1, 32 potentially relevant articles were identified. 17 of these 32 articles met the inclusion criteria. However, one study including patients with Lynch syndrome and one study containing overlapping participants with other included studies were excluded after further examination. Finally, as a publication has more than one cohort in the study while another 3 have more than one population, a total of 15 publications with 24 data sets comprising 26,322 cases and 25,157 controls were included in the meta-analysis (Table 2). The allele frequencies of rs6983267 in controls conformed to Hardy-Weinberg equilibrium for all included studies (Supplementary Table S1).
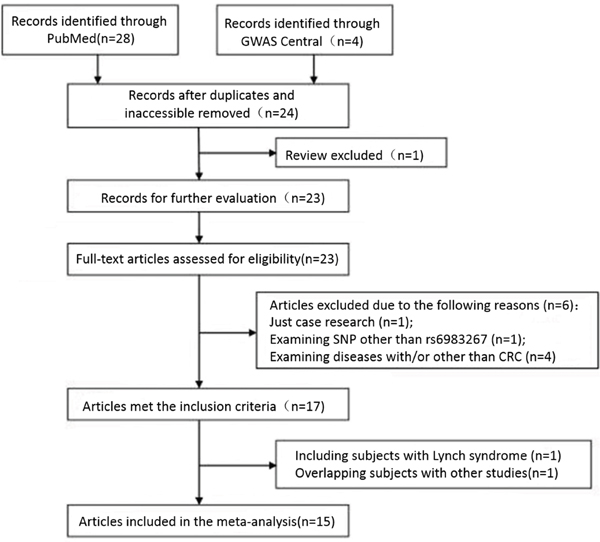
Figure 1: Flow chart of study selection on rs6983267.
Table 2: Characteristics of studies on the association between rs6938267 and CRC risk included in the meta-analysis
First author |
Year |
Ethnicity (Population) |
Number |
|
|---|---|---|---|---|
Case |
Control |
|||
Real LM |
2014 |
Spanish |
500 |
801 |
Haerian MS |
2014 |
Iranian |
380 |
335 |
Li FX |
2012 |
Southern Chinese |
229 |
267 |
Thean LF |
2012 |
Chinese |
991 |
993 |
Tuupanen S |
2008 |
Finnish |
996 |
1012 |
Daraei A |
2012 |
Iranian |
115 |
120 |
Cui R |
2011 |
Japanese |
6167 |
4494 |
He J |
2011 |
European American |
1171 |
1534 |
He J |
2011 |
African American |
382 |
510 |
He J |
2011 |
Native Hawaiian |
323 |
472 |
He J |
2011 |
Japanese American |
1042 |
1426 |
He J |
2011 |
Latino |
393 |
524 |
Kupfer SS |
2010 |
European American |
399 |
367 |
Kupfer SS |
2010 |
African American |
795 |
985 |
von Holst S |
2010 |
Swedish |
1786 |
1749 |
Matsuo k |
2009 |
Japanese |
481 |
962 |
Middeldorp A |
2009 |
Dutch |
995 |
1340 |
Kupfer SS |
2009 |
European Americans |
288 |
202 |
Kupfer SS |
2009 |
African Americans |
281 |
237 |
Curtin K |
2009 |
UK |
654 |
621 |
Tomlinson I (A) |
2007 |
UK |
620 |
960 |
Tomlinson I (B) |
2007 |
UK |
4361 |
3752 |
Tomlinson I (C) |
2007 |
UK |
1901 |
1079 |
Tomlinson I (D) |
2007 |
UK |
1072 |
415 |
The studies were included for the other 4 SNPs (Supplementary Table S2-S5) based on the same criteria showing in material and methods. The numbers of studies included for all 5 SNPs were listed in Table 3, and their locations on chromosomes are shown in Figure 2.
Table 3: Search results of studies on 5SNPs included in meta-analysis
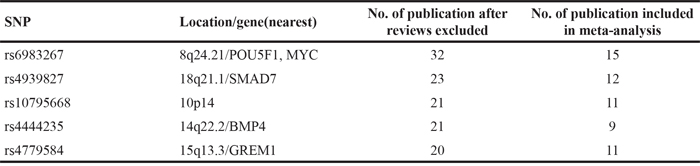
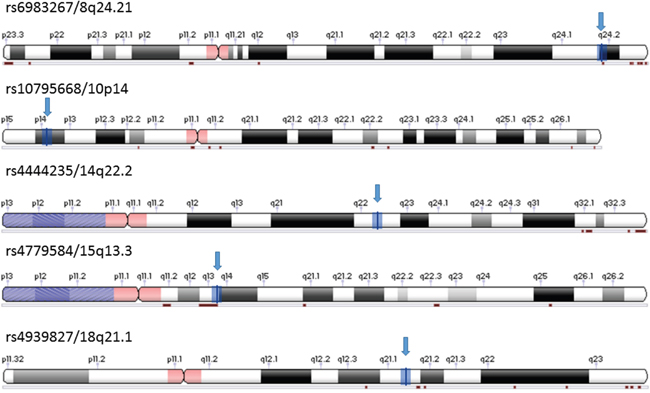
Figure 2: Location of five SNPs selected for meta-analysis. Adopted from NCBI Variation Viewer 1.5.
The meta-analysis of 5 SNPs in associated with CRC risk
The meta-analysis were performed based on two types of the data extracted from the included studies. One type was data with genotype counts (Supplementary Table S1), and the other type was that with OR and 95%CI values generated under various genetic models. Different genetic models were used in the meta-analysis. After pooling all of the included studies into the meta-analyses for each selected SNP, we found that all 5 SNPs were significantly associated with an increased risk of CRC (Table 4). Forest plots in the homogeneous genetic models are shown in Figure 3A-3E.
Table 4: Results of Meta-analysis for the selected 5 SNPs with CRC risk
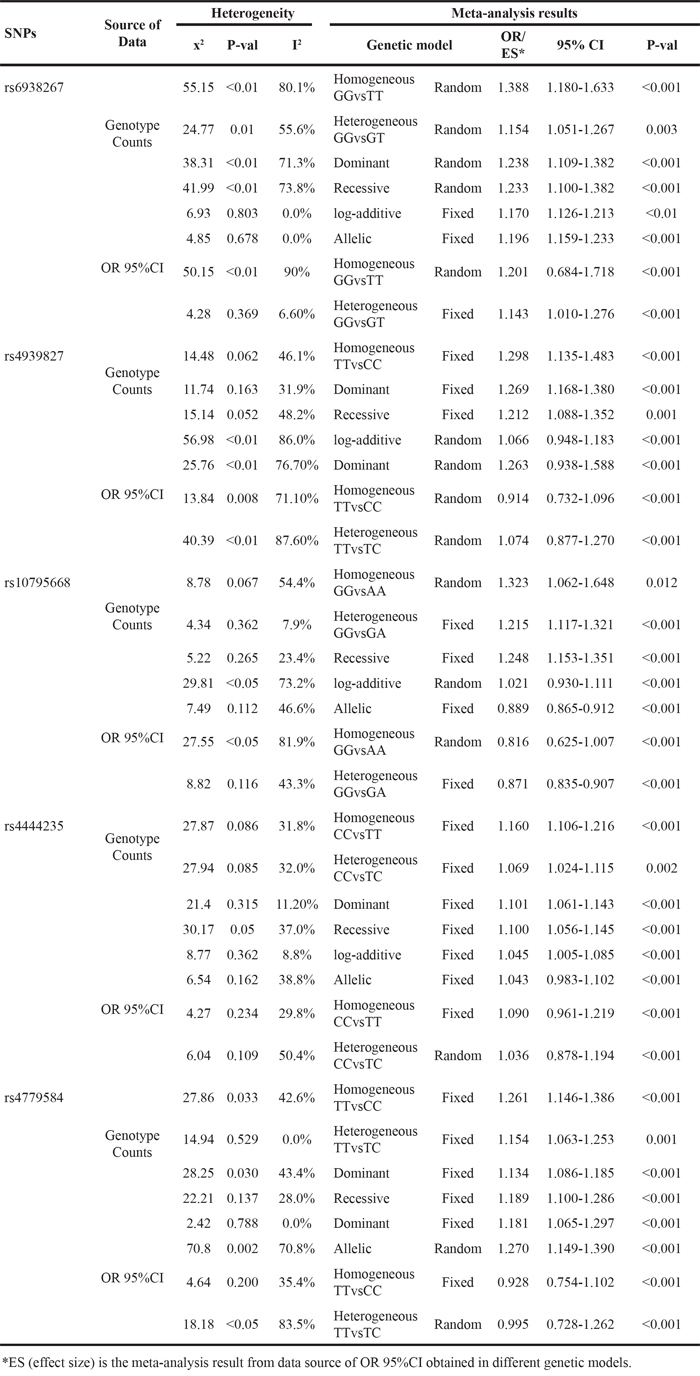
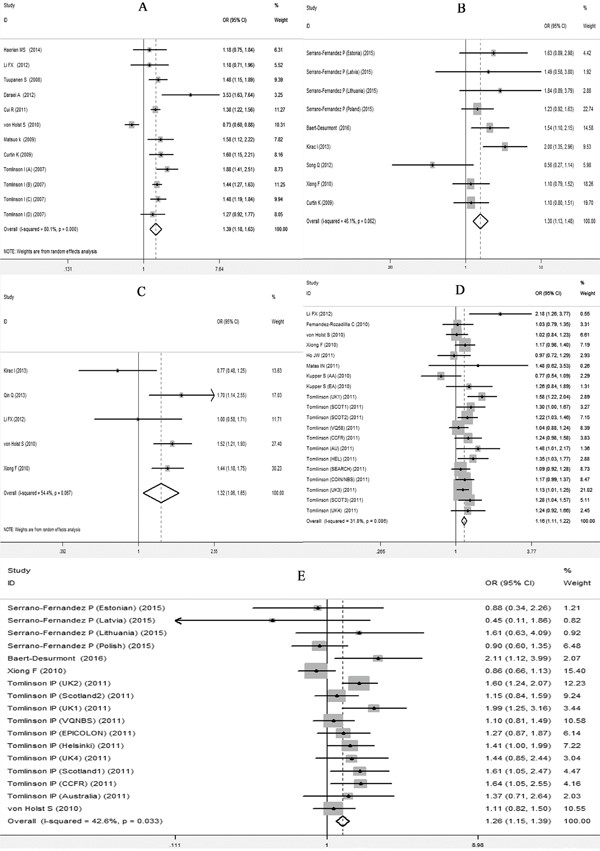
Figure 3: Forest plots for the 5 SNPs and risk of CRC in the homogenous model. A. rs6938267, B. rs4939827, C. rs10795668, D. rs4444235, E. rs4779584.
rs6983267
The analysis of rs6983267 G allele carriers showed a strong association with CRC, with an OR of 1.388 (95% CI 1.180-1.8633; P<0.001) for the homogeneous model, an OR of 1.238 (95% CI 1.109-1.382; P<0.001) for the dominant model and an OR of 1.233 (95% CI 1.100-1.382; P<0.001) for the recessive model. These results were consistent with the risk estimates analyzed using OR values (95% CI, under homogenous model) collected from the included studies, with an effect size (ES) of 1.201 (ES>0; 95% CI 0.684-1.718; P<0.001).
The rs6983267 polymorphism is a G/T single-nucleotide variation on human chromosome 8p24. It has been associated with both prostate and CRC, and it was also the first SNP published to be associated with CRC risk [21]. Our meta-analysis showed a similar result to the previous study (OR=1.47 for GG homozygous). The SNP rs6983267 is located far away from any coding sequences. The nearest genes are the MYC oncogene and the putative pseudogene POU5F1P1, a transcription factor expressed in colon cancer cell lines [21, 31]. Berndt et al. discussed the potential contribution of MYC which is located > 300 KB distant to rs6983267 [32]. While, Tuupanen et al. didn’t find clear association between rs6983267 genotype and MYC expression [33]. Thus, further studies are required as there still remains controversy between MYC and rs6983267.
The present meta-analysis was performed based on larger sample size and various source of data including 26,322 cases and 25,157 controls. The sensitivity analysis results showed that the significance of the pooled ORs was not influenced by any single study. However, an individual study von Holst S (2010) increased the heterogeneity (I2=71.3%, P=0.000) in homogeneous model across the studies included. The value of I2 in the same model reduced greatly (I2=14.3%, p=0.308) in the same genetic model after this study had been removed (Supplementary Figure S1A, S1B). The heterogeneity within the white subgroup also existed when stratified analysis was performed. The difference within the ethnicity may be due to the tumor location (proximal or distal distance) or/and stage. We couldn’t have further analysis and discussion because the clinical and histopathologic characteristics were not provided by all studies.
rs4939827
The rs4939827 T allele had a significant association with CRC with an OR of 1.298 (95% CI 1.135-1.483, P<0.001) for the homogeneous model, an OR of 1.269 (95% CI 1.168-1.380; P<0.001) and an OR of 1.212 (95% CI 1.088-1.352; P<0.001) for the recessive model. The analysis on ORs (95% CI, under dominant model) in our meta-analysis showed a similar results with an ES of 1.263 (95% CI 0.938-1.588; P<0.001).
Variant rs4939827 is a C/T single-nucleotide variation located within intron 3 of SMAD7 on chromosome 18q21. It has been associated with CRC. SMAD7 functions as an important pathway regulation step in the signaling of the TGF-β superfamily. The experiment demonstrates that some of the risk-associated alleles correlate with increased expression of SMAD7 in normal colon tissues, potentially perturb TGF-β negative feedback loop in TGF-β/BMP signaling pathways [34], thus make the progression of cell malignancy.
Broderick et al firstly identified rs4939827 in a GWA set of 620 cases and 960 controls and 3 replication sets of 7,377 cases and 5,867 controls [20], but in an opposite direction with an OR of 0.73 for CC homozygous. Multiple independent studies have replicated the association. The allele frequencies of this SNP in study von Holst S (2010) was found inconsistent with HWE. So it was removed from the present analysis for rs4939827 (data source of genotype counts). The results of present meta-analysis including 39,537 cases and 39,117 controls confirmed that the rs4939827 polymorphism is significantly associated with the increased CRC risk. Our results were similar to other independent studies [23, 35-37]. In addition, it is worth mentioning that our stratified analyses of ethnicities and regions indicate that the heterogeneity among the European descendants is higher than that among the Asians (Supplementary Figure S2).
rs10795668
In our study we found that rs10795668 A allele was significantly associated with CRC, having an OR of 1.323 (95% CI 1.062-1.648; P=0.012) for the homogeneous model, an OR of 1.248 (95% CI 1.153-1.351; P<0.001) for the recessive model. The analysis on ORs (95% CI, under homogeneous model) in our meta-analysis supported the risk estimate with an ES of 0.816 (ES>0; 95% CI 0.625-1.007; P<0.001).
The SNP rs10795668 with an A/G single-nucleotide variation maps to an 82-kb LD block (8.73–8.81 Mb) within 10p14 [22]. At present little is known about its functions. The SNP is located outside the coding regions of genes, and within 400 kb area around the SNP there are no known genes coding proteins. Loo LW et al. performed cis-expression quantitative trait loci analyses to investigate possible regulatory functions on the expression of neighboring genes [34]. They observed a significant association between the low CRC risk A allele for rs10795668 at 10p14 and increased expression of ATP5C1, which is a mitochondrial protein involved in cellular metabolism. Their findings suggested putative functional activities for the CRC GWAS identified risk loci as regulating the expression of neighboring genes.
The result of present meta-analysis including 37,294 cases and 41,037 controls also verified a significant association between the rs10795668 polymorphism and CRC risk, but in an opposite direction to the first report [22]. rs10795668 has been associated with CRC, yet with most conflicting results reported by multiple independent studies [5, 22, 24, 35, 36, 38-44]. The inconsistency in independent studies may be due to the limited sample size of some populations. In addition, rs10795668 exhibited a population difference among the European, African American and Asian populations [45], suggesting genetic heterogeneity across different ethnic groups or possible gene-environment interaction. Our stratified analysis also exhibited a population difference among different ethnicities. Therefore, further functional investigation is necessary to verify the study findings, which will help understand the mechanisms of the variant in the CRC development and tumor progression.
rs4444235
For SNP rs4444235, a significant association with CRC risk was confirmed with an OR equal to 1.160 (95% CI 1.106-1.216; P<0.001) for homogeneous model, 1.101 (95%CI 1.061-1.14; P<0.001) for the dominant and 1.100 (95% CI 1.056-1.145; P<0.001) for the recessive model. The effect size analyzed on homogeneous ORs source also showed similar effect (ES 1.090>0; 95%CI 0.961-1.219; P<0.001) to support the results.
SNP rs4444235 located 9.4 kb from the transcription start site of the gene encoding bone morphogenetic protein 4 preproprotein (BMP4), is a C/T single-nucleotide variation on human chromosome 14q22.2.
GWAS have identified rs4444235 as a new CRC and colorectal adenoma (CRA) susceptibility locus in populations of European descent. This hypothesis has been variously examined by a number of studies but with contradictory results [46-54]. A recent meta-analysis was performed with 54,631 CRC cases, 3,995 CRA cases and 88,098 controls from 15 studies [55]. In the stratified analysis on ethnicity, significantly increased risks were found in East Asians (OR = 1.07, 95% CI 1.01–1.12, P = 0.01) and Caucasians (OR = 1.07, 95% CI: 1.05–1.10, P < 10−5); while no significant associations were found among African Americans and other ethnic populations in all genetic models. The results implicated that rs4444235/BMP4 is a risk factor associated with increased CRC and CRA susceptibility, but these associations are vary in different ethnic groups.
The present meta-analysis including 33,024 CRC cases (according to inclusion criteria, CRA cases were excluded) and 33,348 controls showed that the rs4444235 polymorphism was significantly associated with increased risk of CRC under different genetic models.
rs4779584
Our analysis verified that rs4779584 was significantly associated with increased risk of CRC with an OR of 1.261 (95% CI 1.146-1.386; P<0.001) for the homogeneous model, an OR of 1.134 (1.086-1.185) and 1.189 (1.100-1.286) for the dominant and recessive model, respectively (P<0.001). The analysis on ORs (95% CI, under dominant model) in our meta-analysis showed a similar results with an ES of 1.181 (95% CI 1.065-1.297; P<0.001).
The rs4779584 polymorphism is a C/T single-nucleotide variation in the human genome on 15q13.3. It resides between GREM1 and SCG5. GREM1 is an important signaling molecule in the TGF-ß pathway involved in tumor invasion and metastasis [56] acting as a bone morphogenetic protein (BMP) antagonist. Secretogranin V, encoded by SCG5 is an important neuroendocrine signaling molecule that appears to influence cellular proliferation in the large bowel based on nutrient availability or systemic hormonal effects. Several studies have reported association between this SNP and the risk of CRC. However, the results from the studies are inconclusive. Yang H et al. [57] conducted a meta-analysis of 12 independent case-control studies including 11,769 CRC cases and 14,328 healthy controls. The result showed that the rs4779584 polymorphism may increase the risk of developing CRC in Caucasian population.
The meta-analysis of this study included 33,008 cases of CRC and 35,715 controls. The analyses in various genetic models showed significant association between rs4779584 and increased risk of CRC. However, there was no significant difference among different ethnic populations.
The meta-analysis of this study was performed on 5 SNPs frequently reported by different research groups. The overall results showed that the selected 5 SNPs were strongly associated with CRC. The stratified analyses such as ethnicity and country or region suggested a difference among ethnicities, some even within the ethnicity. Cui R et al. [18] identified a novel susceptible locus at rs7758229/SLC22A3 on 6q26-q27 region that was significantly associated with distal colon cancer (OR 1.28; P=7.92× 10-9) in Asian population. This result suggests an ethnic diversity between Asians and Caucasians in the colorectal pathogenesis.
Publication bias
The Funnel plot and the Egger’s test were used to assess publication bias. As showing in Figure 4a-4e, the shape of the funnel plots appeared to be symmetrical. The Egger’s test did not detect any publication bias (data not shown).
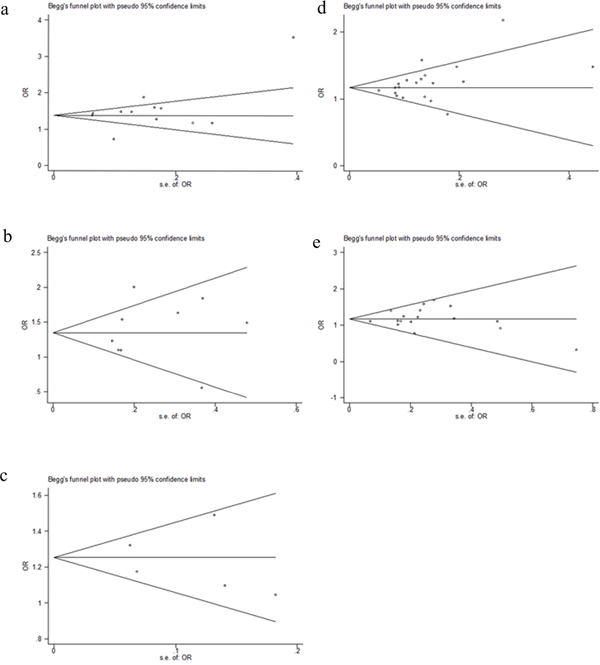
Figure 4: Begg’s Funnel plot of 5 SNPs for Publication bias in the homogenous model. a. rs6938267, b. rs4939827, c. rs10795668, d. rs4444235, e. rs4779584.
Sensitivity analyses
Sensitivity analyses were performed after sequentially removing each eligible study. This approach is regarded as an indispensable step for analyzing multiple criteria. The significance of the pooled ORs was not influenced by any single study under dominant genetic model (Figure 5a-5e), indicating that our results were statistically robust and stable.
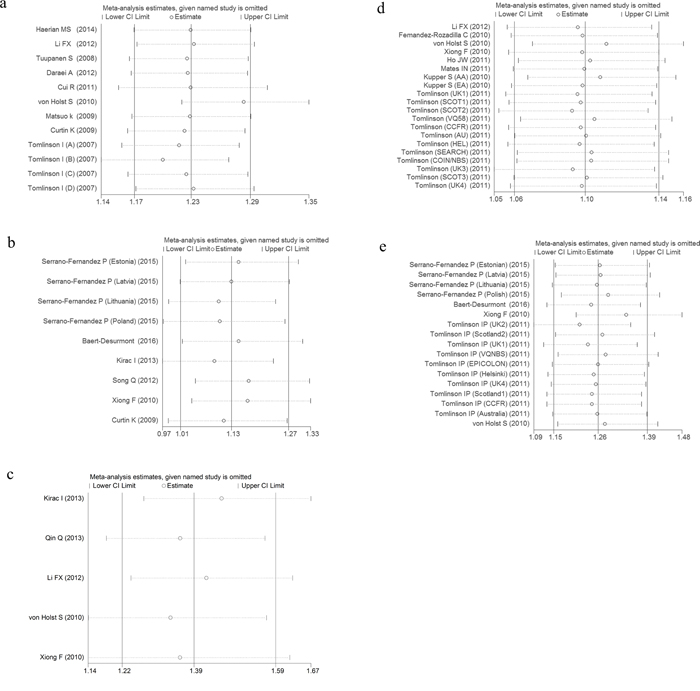
Figure 5: Sensitivity analysis of the odds ratio coefficients under dominant (a, c) and homogeneous (b, d, e) genetic models. a. rs6938267, b. rs4939827, c. rs10795668, d. rs4444235, e. rs4779584.
CONCLUSION
In summary, this meta-analysis included a greater number of studies after SNP screening and publication inclusion. Therefore, a larger sample size (average 33,000 CRC cases and 34,000 controls per SNP) and increased statistical power were obtained. The analyses results of the selected 5 SNPs including rs6983267/8q24.21, rs4939827/18q21.1, rs10795668/10p14, rs44444235/ 14q22.2 and rs4779584/15q13.3 verified their associations with CRC risk. Moreover, the sensitivity analysis suggested no publication bias. The results are stable and reliable.
Early GWAS and replication studies mostly focused on the European and American white population. In comparison, the number of the studies on the Asians and African Americans is relatively small. However, the CRC incidence of these two populations are increasing. Different populations have common variants. While, the specific susceptibility loci might exist as well. The heterogeneity between different population groups was observed in the studies including our analysis. Therefore, further association studies on the Asian and African American population are needed to identify and verify common and unique CRC susceptibility loci in different populations. That will improve the understanding of the pathogenesis of CRC, lead to prognosis and therapeutic strategies and further help design potential CRC risk assessment model.
MATERIALS AND METHODS
Candidate SNPs screening
A literature searching was performed in GWAS central and PubMed without language restriction up to the end of 2014. The search strategy was based on combinations of the terms “GWAS, SNP” and “colorectal cancer or CRC”.
Literature search
The most frequently reported SNPs were further searched in PubMed to end of June 2015. In addition, references in retrieved articles were scanned.
Studies were included if they met the all of the following criteria: (1) original study; (2) assessment of the association between the selected 5 SNPs and CRC risk; (3) case-control or cohort study design; (4) providing genotype numbers, odds ratio (OR) with corresponding confidence 95% interval (95%CI) or sufficient data to calculate them; (5) studies of humans; (6) the genotypes of the SNPs in controls are in HWE. We didn’t included investigations in subjects with Lynch syndrome. Case reports, comments, reviews and editorials were also excluded. If the studies had overlapping subjects, only the study with the largest population was finally included. If more than one geographic or ethnic population were included in one report, each population was considered separately. All data were extracted independently by two reviewers and any disagreement was adjudicated by a third author.
Data extraction
The following information was extracted from each study: first author’s name, publication year, study country, ethnicity of study population, study type, genotyping method, numbers of cases and controls, control source, genotype numbers of the cases and controls or odds ratio (OR) and 95% confidence interval (CI).
Statistical analyses
Hardy–Weinberg equilibrium (HWE) was tested by a goodness-of-fit χ2 test to compare the observed genotype frequencies to the expected genotype frequencies in controls (P<0.05).
To evaluate associations between SNP polymorphisms and risk of CRC, the pooled OR and associated 95% CI where calculated. We used the following models to calculate different ORs: the homogeneous and heterogeneous genetic models (AA vs. aa; Aa vs aa), the dominant genetic model (AA+Aa vs. aa), and the recessive genetic model (AA vs. Aa+aa).
Statistical heterogeneity across studies included was assessed by Cochran’s Q-statistic and considered significant at P<0.05 [58]. The I2 statistic was then utilized to estimate heterogeneity quantitatively (I2= 0–25%, no heterogeneity; I2= 25–50%, moderate heterogeneity; I2= 50–75%, large heterogeneity; I2= 75–100%, extreme heterogeneity) [59]. The fixed effects model, by Mentel-Haenszel (M-H) method, was used to calculate the pooled estimate when I2<50; otherwise, the random-effects model, by DerSimonian-Laird (D+L) method, was applied when I2≥50 [60]. Sensitivity analysis was performed to assess the influence of each study on overall estimate by sequential removing of each study [61]. Publication bias was estimated by Begg’s funnel plot and Eegger’s test [62, 63]. All analyses were performed with the STATA Version 11.0 software (Stata Corp, College Station, TX). All P values in this study were two-tailed tested with a significant level at 0.05.
ACKNOWLEDGMENTS
This work was supported by the Scientific Research Foundation and Academic & Technology Leaders Introduction Project, and 211 Project of Anhui University (10117700023, 02303203-32030081), and The Student Research Training Program of Anhui University (J18520131), and Natural Science Foundation Project of Anhui Province (1508085MH189, 1508085QC63) as well as The Education Revitalization Project of Anhui Province: Stem Cell and Translational Medicine (Y05201374). We acknowledge and are grateful to all participants for their contributions to this study. The authors thank Mr. Simpkins for manuscript reviewing.
CONFLICTS OF INTEREST
The authors declare that they have no conflicts of interest.
REFERENCES
1. Siegel R, Desantis C, Jemal A. Colorectal cancer statistics, 2014. CA Cancer J Clin. 2014; 64:104-117.
2. Ferlay J, Steliarova-Foucher E, Lortet-Tieulent J, Rosso S, Coebergh JW, Comber H, Forman D, Bray F. Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. Eur J Cancer. 2013; 49:1374-1403.
3. Lichtenstein P, Holm NV, Verkasalo PK, Iliadou A, Kaprio J, Koskenvuo M, Pukkala E, Skytthe A, Hemminki K. Environmental and heritable factors in the causation of cancer--analyses of cohorts of twins from Sweden, Denmark, and Finland. N Engl J Med. 2000; 343:78-85.
4. Hemminki K, Chen B. Familial risk for colorectal cancers are mainly due to heritable causes. Cancer Epidemiol Biomarkers Prev. 2004; 13:1253-1256.
5. Mates IN, Jinga V, Csiki IE, Mates D, Dinu D, Constantin A, Jinga M. Single nucleotide polymorphisms in colorectal cancer: associations with tumor site and TNM stage. J Gastrointestin Liver Dis. 2012; 21:45-52.
6. Lindgren G, Liljegren A, Jaramillo E, Rubio C, Lindblom A. Adenoma prevalence and cancer risk in familial non-polyposis colorectal cancer. Gut. 2002; 50:228-234.
7. Liljegren A, Lindblom A, Rotstein S, Nilsson B, Rubio C, Jaramillo E. Prevalence and incidence of hyperplastic polyps and adenomas in familial colorectal cancer: correlation between the two types of colon polyps. Gut. 2003; 52:1140-1147.
8. Pande M, Chen J, Amos CI, Lynch PM, Broaddus R, Frazier ML. Influence of methylenetetrahydrofolate reductase gene polymorphisms C677T and A1298C on age-associated risk for colorectal cancer in a caucasian lynch syndrome population. Cancer Epidemiol Biomarkers Prev. 2007; 16:1753-1759.
9. Lima CS, Nascimento H, Bonadia LC, Teori MT, Coy CS, Goes JR, Costa FF, Bertuzzo CS. Polymorphisms in methylenetetrahydrofolate reductase gene (MTHFR) and the age of onset of sporadic colorectal adenocarcinoma. Int J Colorectal Dis. 2007; 22:757-763.
10. Lewis RC, Bostick RM, Xie D, Deng Z, Wargovich MJ, Fina MF, Roufail WM, Geisinger KR. Polymorphism of the cyclin D1 gene, CCND1, and risk for incident sporadic colorectal adenomas. Cancer Res. 2003; 63:8549-8553.
11. Hong Y, Eu KW, Seow-Choen F, Fook-Chong S, Cheah PY. GG genotype of cyclin D1 G870A polymorphism is associated with increased risk and advanced colorectal cancer in patients in Singapore. Eur J Cancer. 2005; 41:1037-1044.
12. Hong Y, Ho KS, Eu KW, Cheah PY. A susceptibility gene set for early onset colorectal cancer that integrates diverse signaling pathways: implication for tumorigenesis. Clin Cancer Res. 2007; 13:1107-1114.
13. Wang Y, Jatkoe T, Zhang Y, Mutch MG, Talantov D, Jiang J, McLeod HL, Atkins D. Gene expression profiles and molecular markers to predict recurrence of Dukes' B colon cancer. Journal of clinical oncology. 2004; 22:1564-1571.
14. Risch NJ. Searching for genetic determinants in the new millennium. Nature. 2000; 405:847-856.
15. Zanke BW, Greenwood CM, Rangrej J, Kustra R, Tenesa A, Farrington SM, Prendergast J, Olschwang S, Chiang T, Crowdy E, Ferretti V, Laflamme P, Sundararajan S, et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on chromosome 8q24. Nat Genet. 2007; 39:989-994.
16. Houlston RS, Webb E, Broderick P, Pittman AM, Di Bernardo MC, Lubbe S, Chandler I, Vijayakrishnan J, Sullivan K, Penegar S, Carvajal-Carmona L, Howarth K, Jaeger E, et al. Meta-analysis of genome-wide association data identifies four new susceptibility loci for colorectal cancer. Nat Genet. 2008; 40:1426-1435.
17. Jia WH, Zhang B, Matsuo K, Shin A, Xiang YB, Jee SH, Kim DH, Ren Z, Cai Q, Long J, Shi J, Wen W, Yang G, et al. Genome-wide association analyses in East Asians identify new susceptibility loci for colorectal cancer. Nat Genet. 2013; 45:191-196.
18. Cui R, Okada Y, Jang SG, Ku JL, Park JG, Kamatani Y, Hosono N, Tsunoda T, Kumar V, Tanikawa C, Kamatani N, Yamada R, Kubo M, et al. Common variant in 6q26-q27 is associated with distal colon cancer in an Asian population. Gut. 2011; 60:799-805.
19. Houlston RS, Cheadle J, Dobbins SE, Tenesa A, Jones AM, Howarth K, Spain SL, Broderick P, Domingo E, Farrington S, Prendergast JG, Pittman AM, Theodoratou E, et al. Meta-analysis of three genome-wide association studies identifies susceptibility loci for colorectal cancer at 1q41, 3q26.2, 12q13.13 and 20q13.33. Nat Genet. 2010; 42:973-977.
20. Broderick P, Carvajal-Carmona L, Pittman AM, Webb E, Howarth K, Rowan A, Lubbe S, Spain S, Sullivan K, Fielding S, Jaeger E, Vijayakrishnan J, Kemp Z, et al. A genome-wide association study shows that common alleles of SMAD7 influence colorectal cancer risk. Nat Genet. 2007; 39:1315-1317.
21. Tomlinson I, Webb E, Carvajal-Carmona L, Broderick P, Kemp Z, Spain S, Penegar S, Chandler I, Gorman M, Wood W, Barclay E, Lubbe S, Martin L, et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat Genet. 2007; 39:984-988.
22. Tomlinson IP, Webb E, Carvajal-Carmona L, Broderick P, Howarth K, Pittman AM, Spain S, Lubbe S, Walther A, Sullivan K, Jaeger E, Fielding S, Rowan A, et al. A genome-wide association study identifies colorectal cancer susceptibility loci on chromosomes 10p14 and 8q23.3. Nat Genet. 2008; 40:623-630.
23. Tenesa A, Farrington SM, Prendergast JG, Porteous ME, Walker M, Haq N, Barnetson RA, Theodoratou E, Cetnarskyj R, Cartwright N, Semple C, Clark AJ, Reid FJ, et al. Genome-wide association scan identifies a colorectal cancer susceptibility locus on 11q23 and replicates risk loci at 8q24 and 18q21. Nat Genet. 2008; 40:631-637.
24. Peters U, Hutter CM, Hsu L, Schumacher FR, Conti DV, Carlson CS, Edlund CK, Haile RW, Gallinger S, Zanke BW, Lemire M, Rangrej J, Vijayaraghavan R, et al. Meta-analysis of new genome-wide association studies of colorectal cancer risk. Hum Genet. 2012; 131:217-234.
25. Thean LF, Li HH, Teo YY, Koh WP, Yuan JM, Teoh ML, Koh PK, Tang CL, Cheah PY. Association of Caucasian-identified variants with colorectal cancer risk in Singapore Chinese. PLoS One. 2012; 7:e42407.
26. Yang CY, Lu RH, Lin CH, Jen CH, Tung CY, Yang SH, Lin JK, Jiang JK, Lin CH. Single nucleotide polymorphisms associated with colorectal cancer susceptibility and loss of heterozygosity in a Taiwanese population. PLoS One. 2014; 9:e100060.
27. Kupfer SS, Skol AD, Hong E, Ludvik A, Kittles RA, Keku TO, Sandler RS, Ellis NA. Shared and independent colorectal cancer risk alleles in TGFbeta-related genes in African and European Americans. Carcinogenesis. 2014; 35:2025-2030.
28. Zhang B, Jia WH, Matsuda K, Kweon SS, Matsuo K, Xiang YB, Shin A, Jee SH, Kim DH, Cai Q, Long J, Shi J, Wen W, et al. Large-scale genetic study in East Asians identifies six new loci associated with colorectal cancer risk. Nat Genet. 2014; 46:533-542.
29. Zhang B, Jia WH, Matsuo K, Shin A, Xiang YB, Matsuda K, Jee SH, Kim DH, Cheah PY, Ren Z, Cai Q, Long J, Shi J, et al. Genome-wide association study identifies a new SMAD7 risk variant associated with colorectal cancer risk in East Asians. Int J Cancer. 2014; 135:948-955.
30. Closa A, Cordero D, Sanz-Pamplona R, Sole X, Crous-Bou M, Pare-Brunet L, Berenguer A, Guino E, Lopez-Doriga A, Guardiola J, Biondo S, Salazar R, Moreno V. Identification of candidate susceptibility genes for colorectal cancer through eQTL analysis. Carcinogenesis. 2014; 35:2039-2046.
31. Berndt SI, Potter JD, Hazra A, Yeager M, Thomas G, Makar KW, Welch R, Cross AJ, Huang WY, Schoen RE, Giovannucci E, Chan AT, Chanock SJ, et al. Pooled analysis of genetic variation at chromosome 8q24 and colorectal neoplasia risk. Hum Mol Genet. 2008; 17:2665-2672.
32. Suo G, Han J, Wang X, Zhang J, Zhao Y, Zhao Y, Dai J. Oct4 pseudogenes are transcribed in cancers. Biochem Biophys Res Commun. 2005; 337:1047-1051.
33. Tuupanen S, Yan J, Turunen M, Gylfe AE, Kaasinen E, Li L, Eng C, Culver DA, Kalady MF, Pennison MJ, Pasche B, Manne U, de la Chapelle A, et al. Characterization of the colorectal cancer-associated enhancer MYC-335 at 8q24: the role of rs67491583. Cancer Genet. 2012; 205:25-33.
34. Loo LW, Cheng I, Tiirikainen M, Lum-Jones A, Seifried A, Dunklee LM, Church JM, Gryfe R, Weisenberger DJ, Haile RW, Gallinger S, Duggan DJ, Thibodeau SN, et al. cis-Expression QTL analysis of established colorectal cancer risk variants in colon tumors and adjacent normal tissue. PLoS One. 2012; 7:e30477.
35. Xiong F, Wu C, Bi X, Yu D, Huang L, Xu J, Zhang T, Zhai K, Chang J, Tan W, Cai J, Lin D. Risk of genome-wide association study-identified genetic variants for colorectal cancer in a Chinese population. Cancer Epidemiol Biomarkers Prev. 2010; 19:1855-1861.
36. He J, Wilkens LR, Stram DO, Kolonel LN, Henderson BE, Wu AH, Le Marchand L, Haiman CA. Generalizability and epidemiologic characterization of eleven colorectal cancer GWAS hits in multiple populations. Cancer Epidemiol Biomarkers Prev. 2011; 20:70-81.
37. Song Q, Zhu B, Hu W, Cheng L, Gong H, Xu B, Zheng X, Zou L, Zhong R, Duan S, Chen W, Rui R, Wu J, et al. A common SMAD7 variant is associated with risk of colorectal cancer: evidence from a case-control study and a meta-analysis. PLoS One. 2012; 7:e33318.
38. Ho JW, Choi SC, Lee YF, Hui TC, Cherny SS, Garcia-Barcelo MM, Carvajal-Carmona L, Liu R, To SH, Yau TK, Chung CC, Yau CC, Hui SM, et al. Replication study of SNP associations for colorectal cancer in Hong Kong Chinese. Br J Cancer. 2011; 104:369-375.
39. von Holst S, Picelli S, Edler D, Lenander C, Dalen J, Hjern F, Lundqvist N, Lindforss U, Pahlman L, Smedh K, Tornqvist A, Holm J, Janson M, et al. Association studies on 11 published colorectal cancer risk loci. Br J Cancer. 2010; 103:575-580.
40. Kupfer SS, Anderson JR, Hooker S, Skol A, Kittles RA, Keku TO, Sandler RS, Ellis NA. Genetic heterogeneity in colorectal cancer associations between African and European americans. Gastroenterology. 2010; 139:1677-1685, 1685 e1671-1678.
41. Li FX, Yang XX, Hu NY, Du HY, Ma Q, Li M. Single-nucleotide polymorphism associations for colorectal cancer in southern chinese population. Chin J Cancer Res. 2012; 24:29-35.
42. Middeldorp A, Jagmohan-Changur S, van Eijk R, Tops C, Devilee P, Vasen HF, Hes FJ, Houlston R, Tomlinson I, Houwing-Duistermaat JJ, Wijnen JT, Morreau H, van Wezel T. Enrichment of low penetrance susceptibility loci in a Dutch familial colorectal cancer cohort. Cancer Epidemiol Biomarkers Prev. 2009; 18:3062-3067.
43. Talseth-Palmer BA, Brenne IS, Ashton KA, Evans TJ, McPhillips M, Groombridge C, Suchy J, Kurzawski G, Spigelman A, Lubinski J, Scott RJ. Colorectal cancer susceptibility loci on chromosome 8q23.3 and 11q23.1 as modifiers for disease expression in Lynch syndrome. J Med Genet. 2011; 48:279-284.
44. Wijnen JT, Brohet RM, van Eijk R, Jagmohan-Changur S, Middeldorp A, Tops CM, van Puijenbroek M, Ausems MG, Gomez Garcia E, Hes FJ, Hoogerbrugge N, Menko FH, van Os TA, et al. Chromosome 8q23.3 and 11q23.1 variants modify colorectal cancer risk in Lynch syndrome. Gastroenterology. 2009; 136:131-137.
45. Qin Q, Liu L, Zhong R, Zou L, Yin J, Zhu B, Cao B, Chen W, Chen J, Li X, Li T, Lu X, Lou J, et al. The genetic variant on chromosome 10p14 is associated with risk of colorectal cancer: results from a case-control study and a meta-analysis. PLoS One. 2013; 8:e64310.
46. Abuli A, Lozano JJ, Rodriguez-Soler M, Jover R, Bessa X, Munoz J, Esteban-Jurado C, Fernandez-Rozadilla C, Carracedo A, Ruiz-Ponte C, Cubiella J, Balaguer F, Bujanda L, et al. Genetic susceptibility variants associated with colorectal cancer prognosis. Carcinogenesis. 2013; 34:2286-2291.
47. Dai J, Gu J, Huang M, Eng C, Kopetz ES, Ellis LM, Hawk E, Wu X. GWAS-identified colorectal cancer susceptibility loci associated with clinical outcomes. Carcinogenesis. 2012; 33:1327-1331.
48. Hoskins JM, Ong PS, Keku TO, Galanko JA, Martin CF, Coleman CA, Wolfe M, Sandler RS, McLeod HL. Association of eleven common, low-penetrance colorectal cancer susceptibility genetic variants at six risk loci with clinical outcome. PLoS One. 2012; 7:e41954.
49. Phipps AI, Newcomb PA, Garcia-Albeniz X, Hutter CM, White E, Fuchs CS, Hazra A, Ogino S, Nan H, Ma J, Campbell PT, Figueiredo JC, Peters U, et al. Association between colorectal cancer susceptibility loci and survival time after diagnosis with colorectal cancer. Gastroenterology. 2012; 143:51-54 e54.
50. Tenesa A, Theodoratou E, Din FV, Farrington SM, Cetnarskyj R, Barnetson RA, Porteous ME, Campbell H, Dunlop MG. Ten common genetic variants associated with colorectal cancer risk are not associated with survival after diagnosis. Clin Cancer Res. 2010; 16:3754-3759.
51. Xing J, Myers RE, He X, Qu F, Zhou F, Ma X, Hyslop T, Bao G, Wan S, Yang H, Chen Z. GWAS-identified colorectal cancer susceptibility locus associates with disease prognosis. Eur J Cancer. 2011; 47:1699-1707.
52. Cicek MS, Slager SL, Achenbach SJ, French AJ, Blair HE, Fink SR, Foster NR, Kabat BF, Halling KC, Cunningham JM, Cerhan JR, Jenkins RB, Boardman LA, et al. Functional and clinical significance of variants localized to 8q24 in colon cancer. Cancer Epidemiol Biomarkers Prev. 2009; 18:2492-2500.
53. Garcia-Albeniz X, Nan H, Valeri L, Morikawa T, Kuchiba A, Phipps AI, Hutter CM, Peters U, Newcomb PA, Fuchs CS, Giovannucci EL, Ogino S, Chan AT. Phenotypic and tumor molecular characterization of colorectal cancer in relation to a susceptibility SMAD7 variant associated with survival. Carcinogenesis. 2013; 34:292-298.
54. Morris EJ, Penegar S, Whiffin N, Broderick P, Bishop DT, Northwood E, Quirke P, Finan P, Houlston RS. A retrospective observational study of the relationship between single nucleotide polymorphisms associated with the risk of developing colorectal cancer and survival. PLoS One. 2015; 10:e0117816.
55. Zhou L, Xie J, Gu EL, Huang Y, Qu Y, Xu AP, Zhu Y, Wang H. Common genetic variant on BMP4 contributes to colorectal adenoma and cancer: A meta-analysis based on 15 studies. Cytokine. 2015; 72:154-159.
56. Derynck R, Akhurst RJ, Balmain A. TGF-beta signaling in tumor suppression and cancer progression. Nat Genet. 2001; 29:117-129.
57. Yang H, Gao Y, Feng T, Jin TB, Kang LL, Chen C. Meta-analysis of the rs4779584 polymorphism and colorectal cancer risk. PLoS One. 2014; 9:e89736.
58. Lau J, Ioannidis JP, Schmid CH. Quantitative synthesis in systematic reviews. Ann Intern Med. 1997; 127:820-826.
59. Higgins JP, Thompson SG, Deeks JJ and Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003; 327:557-560.
60. DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986; 7:177-188.
61. Thakkinstian A, McElduff P, D'Este C, Duffy D, Attia J. A method for meta-analysis of molecular association studies. Stat Med. 2005; 24:1291-1306.
62. Begg CB, Mazumdar M. Operating characteristics of a rank correlation test for publication bias. Biometrics. 1994; 50:1088-1101.
63. Egger M, Davey Smith G, Schneider M, Minder C. Bias in meta-analysis detected by a simple, graphical test. BMJ. 1997; 315:629-634.

