INTRODUCTION
Non-protein-coding genes have accounted for the vast majority of the human genome, while protein coding potential was observed in only ~2% of human genes. These abundant transcripts have been referred to as non-coding RNAs (ncRNAs). Long ncRNAs (lncRNAs) were one class of transcripts defined as transcribed RNA molecules greater than 200 nucleotides in length, without open reading frame of a significant length [1]. There is considerable evidence to suggest that lncRNAs were involved in a wide range of biological processes. They have emerged as critical factors in cancer progression [2–4]. Many studies have shown that lncRNAs played the role of oncogenes or cancer suppressors in tumor development [5–6]. Several lncRNAs, such as HOTAIR, MALAT1, and GAS5, were reported to be dys-regulated in cancer and were closely related to tumorigenesis, metastasis, and prognosis [7–9]. Hence, lncRNAs have opened up a new avenue towards to the researches on cancer initiation, progression, and metastasis. lncRNAs have showed their application potential in cancer diagnosis, prognosis and therapeutic target.
In recent years, a newly identified lncRNA, human urothelial carcinoma associated 1 (UCA1), has attracted much attention. This lncRNA was originally reported to play a role and its expression would be elevated in the oncogenesis of urinary bladder cancer [10]. UCA1 was found to promote tumorigenicity, with invasive potential in bladder cancer. In addition, the motility, invasion and drug resistance was enhanced by the ectopic expression of UCA1 in BLS-211 cells [11]. A number of other studies also revealed that UCA1 played a crucial role in carcinogenesis and that its expression would be elevated in a variety of malignancies [12–14]. Furthermore, the expression level of UCA1 was found to be associated with tumor clinicopathological features and patient prognosis. Therefore, in present study, relevant publications were collected and a meta-analysis was performed to investigate the relationship between UCA1 expression and clinical outcome. It aimed to further determine whether the over-expression of UCA1 could be applied as a potential biomarker for predicting the patient prognosis.
RESULTS
Study characteristics
The process of literature retrieval was shown in detail (Figure 1). A total of 15 eligible articles (16 studies) were ultimately identified [15–29]. A total of 1,240 cancer patients were included in present meta-analysis, and the mean patient sample size was 82.7 (ranging from 20 to 117). The included 16 studies were all conducted in China. Seven different solid tumor types were evaluated in our study, with five colorectal cancers (CRC), one esophageal squamous cell carcinomas (ESCC), one prostate cancer (PC), two hepatocellular carcinomas (HCC), two non-small cell lung cancers (NSCLC), two gastric cancers (GC) and two ovarian cancers (OC). All cancerous specimens were well preserved before RNA extraction. Diagnoses were all made based on pathology. The main characteristics were summarized (Table 1).
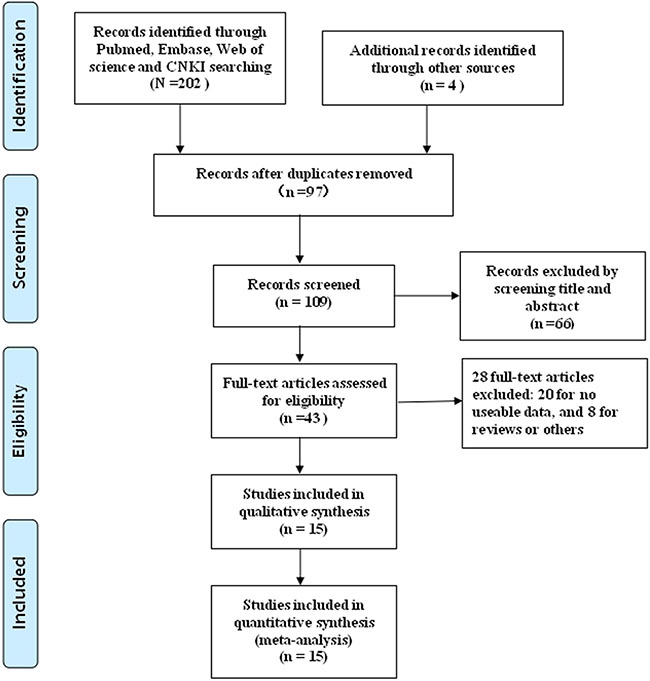
Figure 1: Flowchart presenting the steps of literature search and selection.
Table 1: Main characteristics of all included studies
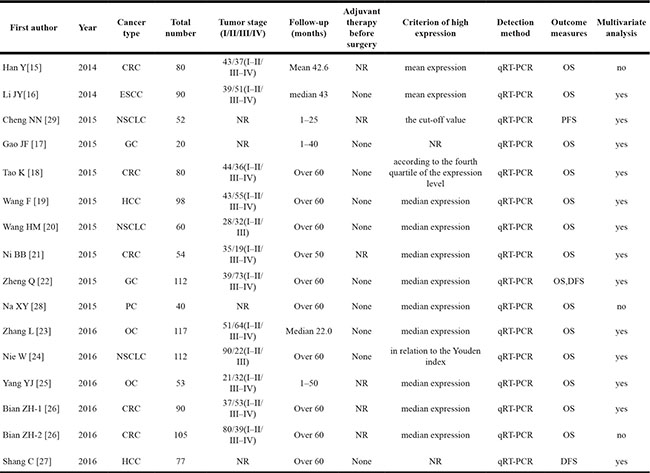
Abbreviations: CRC: colorectal cancer; ESCC: esophageal squamous cell carcinoma; PC: prostate cancer; HCC: hepatocellular carcinoma; NSCLC: non-small cell lung cancer; GC: gastric cancer; OC: ovarian cancer; OS: overall survival; DFS: disease-free survival; PFS: progression free survival; qRT-PCR: quantitative real-time-polymerase chain reaction; NR: not reported.
Increased UCA1 expression and OS
Among the 15 eligible articles, the OS according to UCA1 expression were reported in 13 articles (14 studies; 1,111 cancer patients). The fixed-effects model was adopted to estimate the pooled hazard ratios (HRs), as well as corresponding 95% confidence interval (CI). The result showed no heterogeneity across-studies (I2 = 0%, Ph = 0.883). The HRs, expressed as the high UCA1 expression group versus the low UCA1 expression group, was 1.71 (95% CI: 1.43–1.99, P = 0.000) (Figure 2). The result indicated that there was a significant difference in the OS between the two groups. A significantly shorter OS was observed in the patients with high UCA1 expression level, compared to those with low UCA1 expression level. Thus, it concluded that the expression of UCA1 at high levels was associated with poor OS.
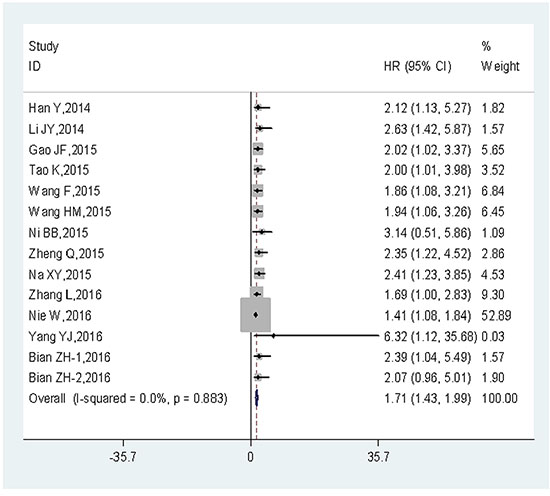
Figure 2: Forest plot of HR for the relationship between high UCA1 expression and OS.
Additionally, pooled HRs for OS based on different types of tumor were shown (Figure 3). The negative effect of increased UCA1 expression on OS was demonstrated in patients with colorectal cancer (HR = 2.22; 95% CI = 1.34–3.11; P = 0.000), gastric cancer (HR= 2.13; 95% CI = 1.17–3.09; P = 0.000), non-small cell lung cancer (HR = 1.47; 95% CI = 1.10–1.83; P = 0.000) ovarian cancer (HR = 1.70; 95% CI = 0.79–2.61; P = 0.000) and other cancers (HR = 2.14; 95% CI = 1.37–2.92; P = 0.000). When all cancer types were roughly grouped into 2 categories (Digestive system cancers and others), a similar result was observed in digestive system cancers (HR = 2.12; 95% CI = 1.59–2.66; P = 0.000) (Table 2).
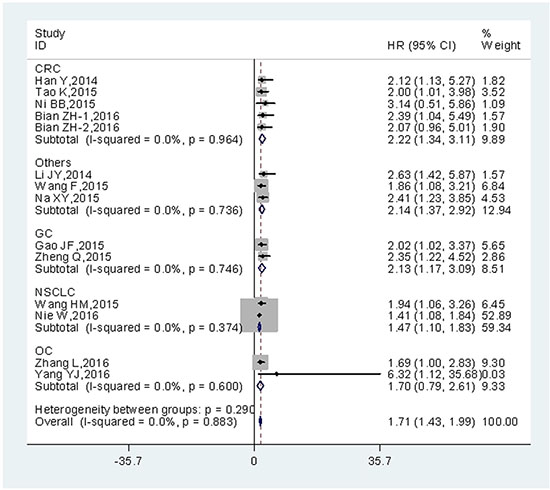
Figure 3: Forest plot of HR for the relationship between high UCA1 expression and OS in patients with various cancers.
Table 2: Pooled HR for OS according to subgroup analysis
Categories |
Studies (n) |
Number of patients |
Fixed-effects model |
Heterogeneity |
||
|---|---|---|---|---|---|---|
HR (95% CI) for OS |
P-value |
I2 (%) |
Ph |
|||
[1] OS |
14 |
1111 |
1.71 (1.43–1.99) |
0.000 |
0 |
0.883 |
[2] Cancer type |
|
|
|
|
|
|
1) Digestive system cancers |
9 |
729 |
2.12 (1.59–2.66) |
0.000 |
0 |
0.997 |
Others |
5 |
382 |
1.55 (1.23–1.88) |
0.000 |
0 |
0.553 |
2) CRC |
5 |
409 |
2.22 (1.34–3.11) |
0.000 |
0 |
0.964 |
GC |
2 |
132 |
2.13 (1.17–3.09) |
0.000 |
0 |
0.764 |
NSCLC |
2 |
172 |
1.47 (1.10–1.83) |
0.000 |
0 |
0.374 |
OC |
2 |
170 |
1.70 (0.79–2.61) |
0.000 |
0 |
0.600 |
Others |
3 |
228 |
2.14 (1.37–2.92) |
0.000 |
0 |
0.736 |
[3] Cutoff value |
|
|
|
|
|
|
Median |
9 |
729 |
2.02 (1.55–2.49) |
0.000 |
0 |
0.977 |
Mean |
2 |
170 |
2.35 (0.84–3.87) |
0.002 |
0 |
0.744 |
Others |
3 |
212 |
1.50 (1.14–1.85) |
0.000 |
0 |
0.497 |
[4] Analysis type |
|
|
|
|
|
|
Multivariate |
11 |
886 |
1.66 (1.37–1.95) |
0.000 |
0 |
0.828 |
Survival curves |
3 |
225 |
2.27 (1.30–3.24) |
0.000 |
0 |
0.951 |
[5] Sample size |
|
|
|
|
|
|
≥ 100 |
4 |
446 |
1.51 (1.17–1.85) |
0.000 |
0 |
0.636 |
< 100 |
10 |
665 |
2.11 (1.63–2.60) |
0.000 |
0 |
0.996 |
Moreover, for OS, the pooled HR values > 1 were consistently calculated in subgroup meta-analysis stratified by the cut-off value, analysis type and sample size, which was also statistically significant (Table 2, the Figures were presented in Supplementary Information).
Increased UCA1 expression and DFS
Only two studies, comprising a total of 189 patients, provided appropriate data for DFS analysis. No severe statistical heterogeneity was observed across-studies (I2 = 0%; Ph = 0.996), the fixed-effects model was applied to analyze the pooled hazard ratios (HRs) with corresponding 95% confidence interval (CI). The overall result indicated a significantly positive association between high expression level of UCA1 and poor DFS (HR = 2.54; 95% CI = 1.09–4.00; P = 0.001) (Figure 4).
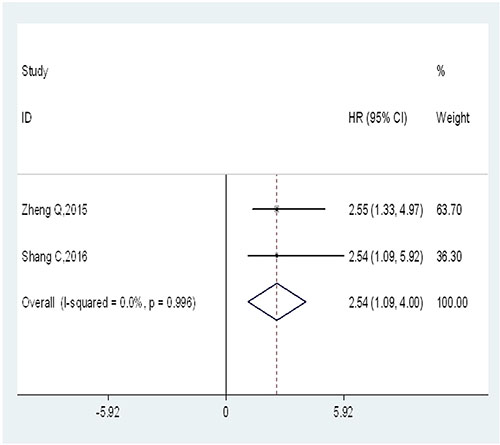
Figure 4: Forest plot of HR for the relationship between high UCA1 expression and DFS.
Associations between lncRNA-UCA1 expression and clinicopathological parameters
From the pooled results (Table 3), it found that increased UCA1 was significantly associated with lymph node metastasis (OR = 2.98, 95% CI: 2.06–4.30, P = 0.000), distant metastasis (OR = 3.14, 95% CI: 1.77–5.58, P = 0.000), and poor clinical stage (OR = 2.76, 95% CI: 2.08–3.68, P = 0.000). However, no significant correlation was observed between the increased UCA1 expression with the age, sex, tumor differentiation, lymphatic invasion and tumor size (All the figures were presented in Supplementary Information). Because of the insufficient data, we failed to detect the relationship between the over-expression of UCA1 and some other clinicopathological parameters.
Table 3: Meta-analysis results of the associations of increased UCA1 expression with clinicopathological parameters
Clinicopathological parameter |
Studies (n) |
Number of patients |
OR (95% CI) |
P-value |
Heterogeneity |
||
|---|---|---|---|---|---|---|---|
I2 (%) |
Ph |
Model |
|||||
Age (≥ 60 vs. < 60) |
7 |
635 |
0.92 (0.67––1.28) |
0.62 |
0 |
0.66 |
Fixed effects |
Sex (Male vs. female) |
9 |
776 |
0.77 (0.57–1.05) |
0.09 |
0 |
0.45 |
Fixed effects |
Lymphatic invasion (Present vs. Absent) |
4 |
336 |
1.26 (0.80–1.99) |
0.32 |
0 |
0.55 |
Fixed effects |
Tumor size (≥ 5 vs. < 5) |
5 |
460 |
1.41 (0.61–3.23) |
0.42 |
78 |
0.001 |
Random effects |
Tumor differentiation (Poorly/others vs. Well/moderately) |
7 |
572 |
1.12 (0.78–1.59) |
0.54 |
42 |
0.11 |
Fixed effects |
Lymph node metastasis (Yes vs. No) |
7 |
566 |
2.98 (2.06–4.30) |
0.000 |
0 |
0.58 |
Fixed effects |
Distant metastasis (Yes vs. No) |
4 |
322 |
3.14 (1.77–5.58) |
0.000 |
41 |
0.17 |
Fixed effects |
TNM stage (III–IV vs. I–II) |
10 |
884 |
2.76 (2.08–3.68) |
0.000 |
6 |
0.39 |
Fixed effects |
Sensitivity analysis
For meta-analysis of the association between UCA1 expression level and OS, the sensitivity analysis was performed by removing each study in turn from the pooled analysis. It aimed to assess the influence of the removed data set on the overall HRs. The result was not significantly influenced after the exclusion of any study, indicating the robustness of the results.
Publication bias
For meta-analysis of the association between UCA1 expression levels and OS, the funnel plot was asymmetric (Figure 5), and the trim and fill method was also adopted to test for publication bias. The results showed no severe publication bias was observed between the included studies.
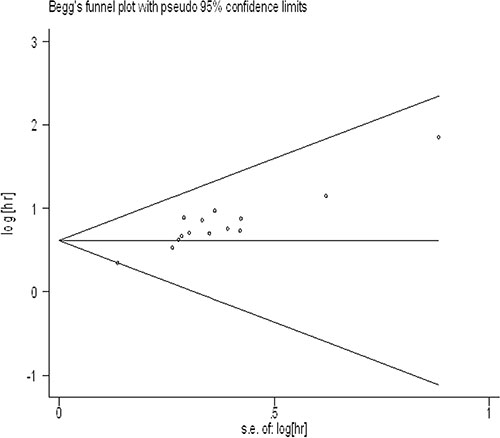
Figure 5: Funnel plot analysis of potential publication bias.
DISCUSSION
UCA1 was a novel ncRNA gene, located in chromosome 19p13.12, which contained three exons and two introns. UCA1 was a long intergenic ncRNA that first discovered in bladder cancer in 2006 [10]. As an oncogenic lncRNA, it was reported to be abundant in many cancer cells and tumor tissues. UCA1 was involved in cancer progression, and its aberrant expression was associated with a broad range of cellular processes, including cell cycle distribution, apoptosis and proliferation [28, 30–31]. Importantly, UCA1 has been considered as a promising diagnostic marker and potential therapeutic target for human cancers [32–34]. Furthermore, some studies have shown that UCA1 was related to drug resistance in some malignancies, including ovarian cancer, bladder cancer and EGFR-mutant NSCLC, suggesting that UCA1 could be applied as a biomarker for monitoring the efficacy of chemotherapy [29, 35–36].
A growing number of studies have focused on the relationship between UCA1 and carcinoma, as well as interpreting its function in the progression of human cancer. However, its underlying molecular mechanism remained to be largely unclear. Numerous studies have shown that UCA1 made effects as an oncogenic lncRNA by inhibiting known tumor suppressors, such as p27 and miR-143 in breast cancer [37, 38] and BRG1 in bladder cancer [39]. It also acted as a molecular sponge for certain microRNAs (miRs), such as miR-193a-3p in NSCLC [24].
The mechanism on regulatory activity of UCA1 in cancer invasion and metastasis has been explored in several cancer types. In a study by Xue et al. [40], cell invasion of bladder cancer cells was promoted by UCA1 via the hsa-miR-145-ZEB1⁄2-FSCN1 pathway. Another study investigated on the function of UCA1 in hypoxic bladder cancer, revealing that UCA1 could function as a HIF-1α-targeted lncRNA to enhance bladder cancer cell invasion [41]. Chen et al. [42] found that the invasiveness of breast cancer cells could be promoted by the macrophage infiltration in the setting of up-regulated UCA1. Furthermore, a recent study confirmed a connection between UCA1 and miR-485-5p/MMP14 [25]. Regarding the role of UCA1 in HCC tumorigenesis, Wang et al. [19] found that UCA1 could facilitate HCC cell growth and metastasis through the inhibition of miR-216b and activation of the FGFR1/ERK signaling pathway. On the other hand, some studies have shown that aerobic glycolysis could be promoted by UCA1 in bladder cancer cells by targeting miR-16 and up-regulating hexokinase-2 [43–44], thus affecting their malignant potential.
Our meta-analysis provided evidence that high UCA1 expression was significantly correlated with a poor clinical prognosis in patients with various cancer types. Firstly, the combined results indicated that increased UCA1 expression was associated with a shorter OS in solid tumor patients. A shorter overall survival time was observed in the patients with high expression level of UCA1, compared to those with low UCA1 expression. It suggested that UCA1 could act as an independent prognostic factor for OS in cancer patients. In subgroup analysis for OS, UCA1 at high level in cancerous tissues may be a reliable prognostic marker specifically for digestive system cancers.
Secondly, this meta-analysis showed that patients with elevated UCA1 may suffer from a significantly poorer DFS. Only one study reported the PFS, thus we couldn’t do further analysis. The result of both univariate and multivariate analysis indicated that the UCA1 expression level was an independent prognostic factor for PFS in patients with EGFR-TKI-sensitive NSCLC [29].
Thirdly, the clinicopathological significance of over-expressed UCA1 was also demonstrated in present meta-analysis. From the pooled results, we could find that increased UCA1 expression was positively correlated with advanced clinical stage, and cancer patients with high UCA1 expression may develop with an increased risk of LNM and DM.
However, there would be some limitations in our meta-analysis. For instance, the total sample size was relatively small, and patients included in the meta-analysis were all Asians from China. Additionally, publication bias may exist, despite the fact that no significant publication bias was observed based on the trim and fill method, as well as stable results revealed in sensitivity analysis. Finally, the cut-off definition for high UCA1 expression was not consistent. Therefore, larger-size, multi-center and higher-quality studies with a unified criterion for determining UCA1 expression are necessary to validate the results in this study.
MATERIALS AND METHODS
Literature search
For obtaining potentially eligible studies, comprehensive literature retrieval was conducted in various databases: PubMed, Web of Science, Embase, and CNKI, with a cut-off date of April 11, 2016. The keywords for the search were as follows: “urothelial cancer associated 1”, “UCA1”, “lncRNA UCA1”, “cancer”, “carcinoma”, “neoplasm” and “tumor”. In addition, other relevant articles were also manually viewed from the references lists.
Inclusion and exclusion criteria
Inclusion criteria for the articles were as follows: (1) the role of lncRNA UCA1 in the development of human cancer was investigated, (2) associations of lncRNA UCA1 expression with prognosis or clinicopathological features were described, (3) the expression level of lncRNA UCA1 in primary cancerous tissue was determined by RT-qPCR and (4) patients were divided into high and low expression groups according to the expression level of lncRNA UCA1.
Exclusion criteria for the articles were as follows: (1) duplicate publications; (2) studies without valuable data; and (3) reviews, letters, case reports and expert opinions.
Date extraction and quality assessment
The data and information from all included studies were independently extracted by two investigators (ZY and QC). The following information were collected from each study: first author name, publication year, the study country, cancer type, total patients number, tumor stage, follow-up period, outcome measures, the criteria for high UCA1 expression, determination method, HR and corresponding 95% CI. Besides, the data of clinicopathological parameters were also extracted from the eligible studies. For studies that provided both the results of univariate and multivariate analysis, only the latter was selected because of its increased precision on interpreting confounding factors. If a study reported only Kaplan-Meier curves, Engauge Digitizer version 4.1 was applied to extract the survival data. In the event of a disagreement, a consensus was reached by a third investigator (ZPQ). The Newcastle-Ottawa Scale (NOS) was applied to assess the quality of all included studies. The NOS scores ranged from 0 to 9, and the study with an NOS score ≥ 6 was considered to be of high quality. The quality of all studies included in this meta-analysis was varied from 4 to 9, with a mean value of 6.5.
Statistical methods
The current meta-analysis was performed with RevMan5.3 software and Stata SE12.0. The heterogeneity between studies was determined with the Chi square-based Q test and I2 statistics. A P value less than 0.05 for the Q test and I2 value above 50% were considered to be significantly heterogeneous. The fixed effects model was applied for studies with no obvious heterogeneity (Ph > 0.05, I2 < 50%); otherwise, the random effects model was adopted (Ph ≤ 0.05, I2 ≥ 50%). Potential publication bias was assessed with a funnel plot. The sensitivity analysis was also performed to assess the stability of the results. A P value less than 0.05 was considered statistically significant.
ACKNOWLEDGMENTS AND FUNDING
We thank for the financial support of Graduate Innovation Fund of Nanchang University (No.cx2015168).
CONFLICTS OF INTEREST
The authors declared that there are no conflicts of interest regarding the publication of this paper.
REFERENCES
1. Tsai MC, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011; 71:3–7.
2. Dey BK, Mueller AC, Dutta A. Long non-coding RNAs as emerging regulators of differentiation, development, and disease. Transcription. 2014; 5:e944014.
3. Dhamija S, Diederichs S. From junk to master regulators of invasion: lncRNA functions in migration, EMT and metastasis. Int J Cancer. 2016; 139:269–280.
4. Yamashita A, Shichino Y, Yamamoto M. The long non-coding RNA world in yeasts. Biochim Biophys Acta. 2016; 1859:147–154.
5. Colombo T, Farina L, Macino G, Paci P. PVT1: a rising star among oncogenic long noncoding RNAs. Biomed Res Int. 2015; 2015:304208.
6. Zhou Y, Zhang X, Klibanski A. MEG3 noncoding RNA: a tumor suppressor. J Mol Endocrinol. 2012; 48:R45–53.
7. Hajjari M, Salavaty A. HOTAIR: an oncogenic long non-coding RNA in different cancers. Cancer Biol Med. 2015; 12:1–9.
8. Gutschner T, Hämmerle M, Diederichs S. MALAT1-a paradigm for long noncoding RNA function in cancer. J Mol Med (Berl). 2013; 91:791–801.
9. Pickard MR, Williams GT. Molecular and Cellular Mechanisms of Action of Tumour Suppressor GAS5 LncRNA. Genes (Basel). 2015; 6:484–499.
10. Wang XS, Zhang Z, Wang HC, Cai JL, Xu QW, Li MQ, Chen YC, Qian XP, Lu TJ, Yu LZ, Zhang Y, Xin DQ, Na YQ, et al. Rapid identification of UCA1 as a very sensitive and specific unique marker for human bladder carcinoma. Clin Cancer Res. 2006; 12:4851–4858.
11. Wang F, Li X, Xie X, Zhao L, Chen W. UCA1, a non-protein-coding RNA up-regulated in bladder carcinoma and embryo, influencing cell growth and promoting invasion. FEBS Lett. 2008; 582:1919–1927.
12. Fang Z, Wu L, Wang L, Yang Y, Meng Y, Yang H. Increased expression of the long non-coding RNA UCA1 in tongue squamous cell carcinomas: a possible correlation with cancer metastasis. Oral Surg Oral Med Oral Pathol Oral Radiol. 2014; 117:89–95.
13. Tian Y, Zhang X, Hao Y, Fang Z, He Y. Potential roles of abnormally expressed long noncoding RNA UCA1 and Malat-1 in metastasis of melanoma. Melanoma Res. 2014; 24:335–341.
14. Li Y, Wang T, Li Y, Chen D, Yu Z, Jin L, Ni L, Yang S, Mao X, Gui Y, Lai Y. Identification of long-non coding RNA UCA1 as an oncogene in renal cell carcinoma. Mol Med Rep. 2016; 13:3326–3334.
15. Han Y, Yang YN, Yuan HH, Zhang TT, Sui H, Wei XL, Liu L, Huang P, Zhang WJ, Bai YX. UCA1, a long non-coding RNA up-regulated in colorectal cancerb influences cell proliferation, apoptosis and cell cycle distribution. Pathology. 2014; 46:396–401.
16. Li JY, Ma X, Zhang CB. Overexpression of long non-coding RNA UCA1 predicts a poor prognosis in patients with esophageal squamous cell carcinoma. Int J Clin Exp Pathol. 2014; 7:7938–7944.
17. Gao J, Cao R, Mu H. Long non-coding RNA UCA1 may be a novel diagnostic and predictive biomarker in plasma for early gastric cancer. Int J Clin Exp Pathol. 2015; 8:12936–12942.
18. Tao K, Yang J, Hu Y, Sun Y, Tan Z, Duan J, Zhang F, Yan H, Deng A. Clinical significance of urothelial carcinoma associated 1 in colon cancer. Int J Clin Exp Med. 2015; 8:21854–21860.
19. Wang F, Ying HQ, He BS, Pan YQ, Deng QW, Sun HL, Chen J, Liu X, Wang SK. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway. Oncotarget. 2015; 6:7899–7917. doi: 10.18632/oncotarget.3219.
20. Wang HM, Lu JH, Chen WY, Gu AQ. Upregulated lncRNA-UCA1 contributes to progression of lung cancer and is closely related to clinical diagnosis as a predictive biomarker in plasma. Int J Clin Exp Med. 2015; 8:11824–11830.
21. Ni B, Yu X, Guo X, Fan X, Yang Z, Wu P, Yuan Z, Deng Y, Wang J, Chen D, Wang L. Increased urothelial cancer associated 1 is associated with tumor proliferation and metastasis and predictsnpoor prognosis in colorectal cancer. Int J Oncol. 2015; 47:1329–1338.
22. Zheng Q, Wu F, Dai WY, Zheng DC, Zheng C, Ye H, Zhou B, Chen JJ, Chen P. Aberrant expression of UCA1 in gastric cancer and its clinical significance. Clin Transl Oncol. 2015; 17:640–646.
23. Zhang L, Cao X, Zhang L, Zhang X, Sheng H, Tao K. UCA1 overexpression predicts clinical outcome of patients with ovarian cancer receiving adjuvant chemotherapy. Cancer Chemother Pharmacol. 2016; 77:629–634.
24. Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao X, Chen WS, Li B. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett. 2016; 371:99–106.
25. Yang Y, Jiang Y, Wan Y, Zhang L, Qiu J, Zhou S, Cheng W. UCA1 functions as a competing endogenous RNA to suppress epithelial ovarian cancer metastasis. Tumour Biol. 2016.
26. Bian Z, Jin L, Zhang J, Yin Y, Quan C, Hu Y, Feng Y, Liu H, Fei B, Mao Y, Zhou L, Qi X, Huang S, et al. LncRNA-UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci Rep. 2016; 6:23892.
27. Shang C, Guo Y, Zhang J, Huang B. Silence of long noncoding RNA UCA1 inhibits malignant proliferation and chemotherapy resistance to adriamycin in gastric cancer. Cancer Chemother Pharmacol. 2016; 77:1061–1067.
28. Na XY, Liu ZY, Ren PP, Yu R, Shang XS. Long non-coding RNA UCA1 contributes to the progression of prostate cancer and regulates proliferation through KLF4-KRT6/13 signaling pathway. Int J Clin Exp Med. 2015; 8:12609–12616.
29. Cheng N, Cai W, Ren S, Li X, Wang Q, Pan H, Zhao M, Li J, Zhang Y, Zhao C, Chen X, Fei K, Zhou C, et al. Long non-coding RNA UCA1 induces non-T790M acquired resistance to EGFR-TKIs by activating the AKT/mTOR pathway in EGFR-mutant non-small cell lung cancer. Oncotarget. 2015; 6:23582–23593. doi: 10.18632/oncotarget.4361.
30. Yang C, Li X, Wang Y, Zhao L, Chen W. Long non-coding RNA UCA1 regulated cell cycle distribution via CREB through PI3-K dependent pathway in bladder carcinoma cells. Gene. 2012; 496:8–16.
31. Wu W, Zhang S, Li X, Xue M, Cao S, Chen W. Ets-2 regulates cell apoptosis via the Akt pathway, through the regulation of urothelial cancer associated 1, a long non-coding RNA, in bladder cancer cells. PLoS One. 2013; 8:e73920.
32. Srivastava AK, Singh PK, Rath SK, Dalela D, Goel MM, Bhatt ML. Appraisal of diagnostic ability of UCA1 as a biomarker of carcinoma of the urinary bladder. Tumour Biol. 2014; 35:11435–11442.
33. Kamel MM, Matboli M, Sallam M, Montasser IF, Saad AS, El-Tawdi AH. Investigation of long noncoding RNAs expression profile as potential serum biomarkers in patients with hepatocellular carcinoma. Transl Res. 2016; 168:134–145.
34. Eissa S, Matboli M, Essawy NO, Shehta M, Kotb YM. Rapid detection of urinary long non-coding RNA urothelial carcinoma associated one using a PCR-free nanoparticle-based assay. Biomarkers. 2015; 20:212–217.
35. Fan Y, Shen B, Tan M, Mu X, Qin Y, Zhang F, Liu Y. Long non-coding RNA UCA1 increases chemoresistance of bladder cancer cells by regulating Wnt signaling. FEBS J. 2014; 281:1750–1758.
36. Wang F, Zhou J, Xie X, Hu J, Chen L, Hu Q, Guo H, Yu C. Involvement of SRPK1 in cisplatin resistance related to long non-coding RNA UCA1 in human ovarian cancer cells. Neoplasma. 2015; 62:432–438.
37. Huang J, Zhou N, Watabe K, Lu Z, Wu F, Xu M, Mo YY. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014; 5:e1008.
38. Tuo YL, Li XM, Luo J. Long noncoding RNA UCA1 modulates breast cancer cell growth and apoptosis through decreasing tumor suppressive miR-143. Eur Rev Med Pharmacol Sci. 2015; 19:3403–3411.
39. Wang X, Gong Y, Jin B, Wu C, Yang J, Wang L, Zhang Z, Mao Z. Long non-coding RNA urothelial carcinoma associated 1 induces cell replication by inhibiting BRG1 in 5637 cells. Oncol Rep. 2014; 32:1281–1290.
40. Xue M, Pang H, Li X, Li H, Pan J, Chen W. Long non-coding RNA urothelial cancer-associated 1 promotes bladder cancer cell migration and invasion by way of the hsa-miR-145-ZEB1/2-FSCN1 pathway. Cancer Sci. 2016; 107:18–27.
41. Xue M, Li X, Li Z, Chen W. Urothelial carcinoma associated 1 is a hypoxia-inducible factor-1α-targeted long noncoding RNA that enhances hypoxic bladder cancer cell proliferation, migration, and invasion. Tumour Biol. 2014; 35:6901–6912.
42. Chen S, Shao C, Xu M, Ji J1, Xie Y, Lei Y, Wang X. Macrophage infiltration promotes invasiveness of breast cancer cells via activating long non-coding RNA UCA1. Int J Clin Exp Pathol. 2015; 8:9052–9061.
43. Li HJ, Li X, Pang H, Pan JJ, Xie XJ, Chen W. Long non-coding RNA UCA1 promotes glutamine metabolism by targeting miR-16 in human bladder cancer. Jpn J Clin Oncol. 2015; 45:1055–1063.
44. Li Z, Li X, Wu S, Xue M, Chen W. Long non-coding RNA UCA1 promotes glycolysis by upregulating hexokinase-2 through the mTOR-STAT3/microRNA143 pathway. Cancer Sci. 2014; 105:951–955.

