Oncotarget: CRISPR/Cas9-directed epigenetic editing
2021-09-12
Oncotarget published "Diverse transcriptional regulation and functional effects revealed by CRISPR/Cas9-directed epigenetic editing" which reported that the adequate functioning of this process is indispensable for tissue homeostasis and cell fate determination. Here, the authors have taken advantage of CRISPR/dCas9 technology adapted for epigenetic editing through site-specific targeting of DNA methylation to characterize the transcriptional changes of the candidate gene and the functional effects on cell fate in different tumor settings.
Strikingly, this modification led to opposing expression profiles of the target gene in different cancer cell models and affected the expression of mesenchymal genes CDH1, VIM1, TGFB1 and apoptotic marker BCL2. Moreover, methylation-induced changes in expression profiles was also accompanied by a phenotypic switch in cell migration and cell morphology.
Dr. Miguel Vizoso and Dr. Jacco van Rheenen said, "Increasing number of studies report that proteins do not work in isolation but are part of a complex network of biomolecules, that may differ at various settings (e.g., different tumor types [1, 2] or stages of tumor progression."
Increasing number of studies report that proteins do not work in isolation but are part of a complex network of biomolecules, that may differ at various settings (e.g., different tumor types [1, 2] or stages of tumor progression.
Various examples of genes with opposing roles, e. DNA methylation may be one of the drivers of these opposing roles. For example, the same DNA methylation mark can lead to very opposite outcomes depending on which allele is tagged with this modification. Here they test whether the same epigenetic modification could also orchestrate molecular and phenotypic diversity in non-imprinted genes.
In this study, the authors focus on the insulin-like growth factor binding protein 2, a recently discovered multitasked gene regulated by DNA methylation which has also been reported to function both as a tumor-promoting and -suppressing gene. The diverse phenotypes upon single alterations in cancer driver genes may simply reflect the various roles of these genes in distinct signaling pathways, but it may also be caused by differential gene expression patterns of these genes.
DNA methylation comprises a significant mechanism involved in transcriptomic regulation based on the occupancy of CpG dinucleotides by 5′-methylcytosine chemical groups. Whether the consequence of DNA methylation on the expression of the target genes and cell fate is uniform regardless of the cell type or can drive to opposing phenotypes depending on the tumor cell context is still unknown, and it has remained elusive for many years due to the lack of technologies to target DNA methylation in-situ.
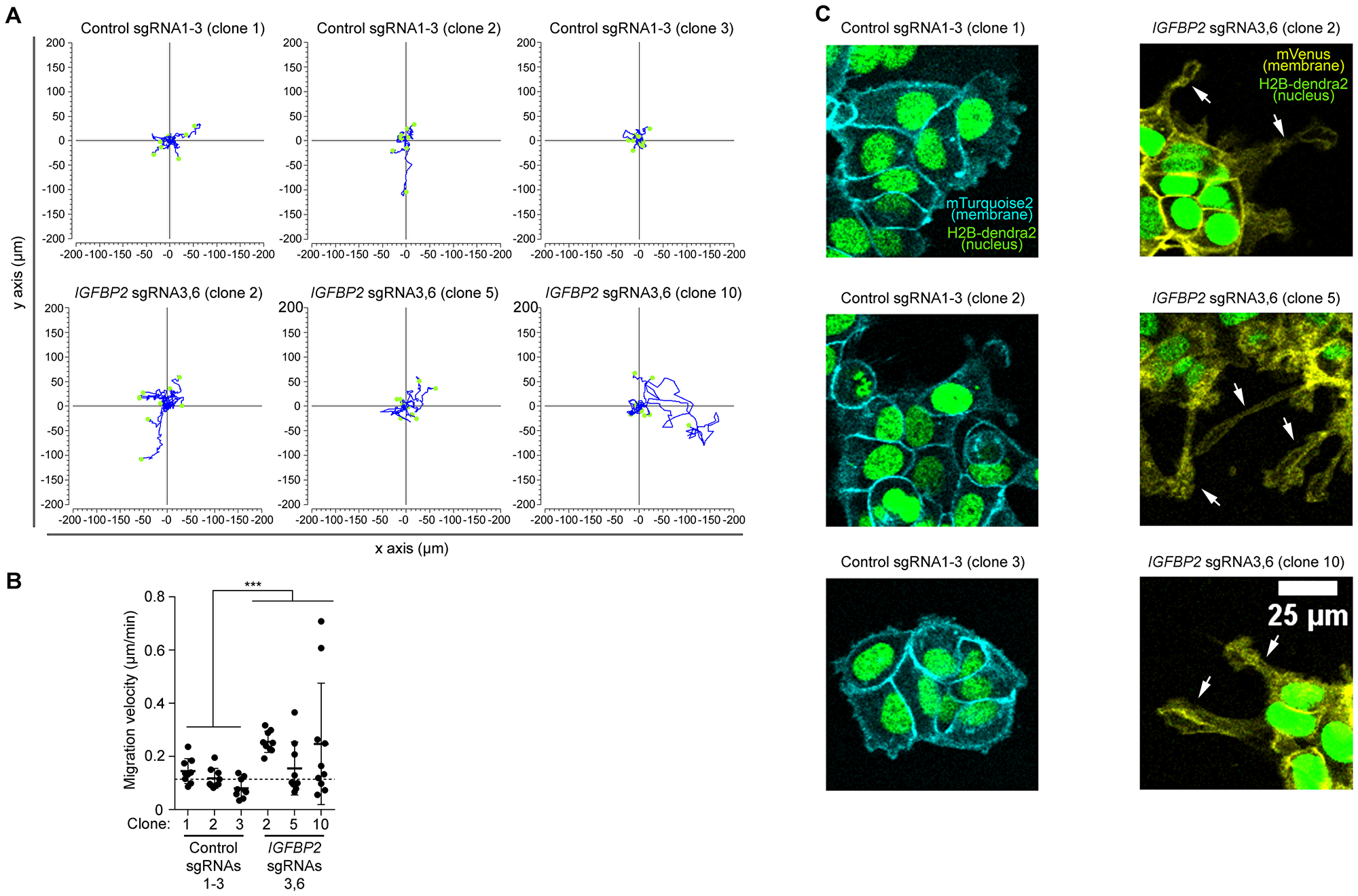
Figure 4: High resolution imaging reveal an increased migration speed of the epithelial cell line MCF7 clones upon CRISPR/dCas IGFBP2 epigenetic editing. (A) Display of 9 independent migratory tracks from 3 pooled control sgRNAs 1-3 clones and 3 pooled IGFBP2 sgRNAs 3 and 6 clones. (B) Distribution of migration velocity from 3 pooled control sgRNAs 1-3 clones and 3 pooled IGFBP2 sgRNAs 3 and 6 clones. Error bars represent the SD of n = 3–9 independent positions. Statistical analysis were performed using Mann-Whitney U test for speed and displacement and T-test for mitotic cell division analysis. p values ≤ 0.05, ≤ 0.01, or ≤ 0.001 are considered statistically significant and indicated by an asterisk (*, **, or ***, respectively). (C) Representative time-lapse images of 3 pooled control sgRNAs 1-3 clones and 3 pooled IGFBP2 sgRNAs 3 and 6 clones. White arrow indicates the appearance of cell protrusions (absent in the control conditions). Images show nuclear marker (H2B-dendra2) to follow cell divisions and migration and membrane markers (mTurquoise-2 and mVenus) to study cell morphology.
The Vizoso/van Rheenen Research Team concluded in their Oncotarget Research Output, "our study highlights the significance of exploring the effects of the epigenetic editing in different tumor settings by revealing the important consequences that this can have on transcriptomic regulation and tumor cell behavior."
Sign up for free Altmetric alerts about this article
DOI - https://doi.org/10.18632/oncotarget.28037
Full text - https://www.oncotarget.com/article/28037/text/
Correspondence to - Miguel Vizoso - orcid.org/0000-0002-9992-2851 and Jacco van Rheenen - [email protected]
Keywords - intestinal ischemia reperfusion (II/R), acute lung injury (ALI), prolyl oligopeptidase (POP), inflammation, angiogenesis
About Oncotarget
Oncotarget is a biweekly, peer-reviewed, open access biomedical journal covering research on all aspects of oncology.
To learn more about Oncotarget, please visit https://www.oncotarget.com or connect with:
SoundCloud - https://soundcloud.com/oncotarget
Facebook - https://www.facebook.com/Oncotarget/
Twitter - https://twitter.com/oncotarget
LinkedIn - https://www.linkedin.com/company/oncotarget
Pinterest - https://www.pinterest.com/oncotarget/
Reddit - https://www.reddit.com/user/Oncotarget/
Oncotarget is published by Impact Journals, LLC please visit https://www.ImpactJournals.com or connect with @ImpactJrnls
Media Contact
[email protected]
18009220957x105
Copyright © 2026 Rapamycin Press LLC dba Impact Journals
Oncotarget ® is a registered trademark of Rapamycin Press LLC
Impact Journals ® is a registered trademark of Rapamycin Press LLC
RAPAMYCIN PRESS ® is a registered trademark of Rapamycin Press LLC



